Characterization of a Flavonoid 3’/5’/7-O-Methyltransferase from Citrus reticulata and Evaluation of the In Vitro Cytotoxicity of Its Methylated Products
Abstract
1. Introduction
2. Results
2.1. CrOMT2 Isolation and Bioinformatic Analysis
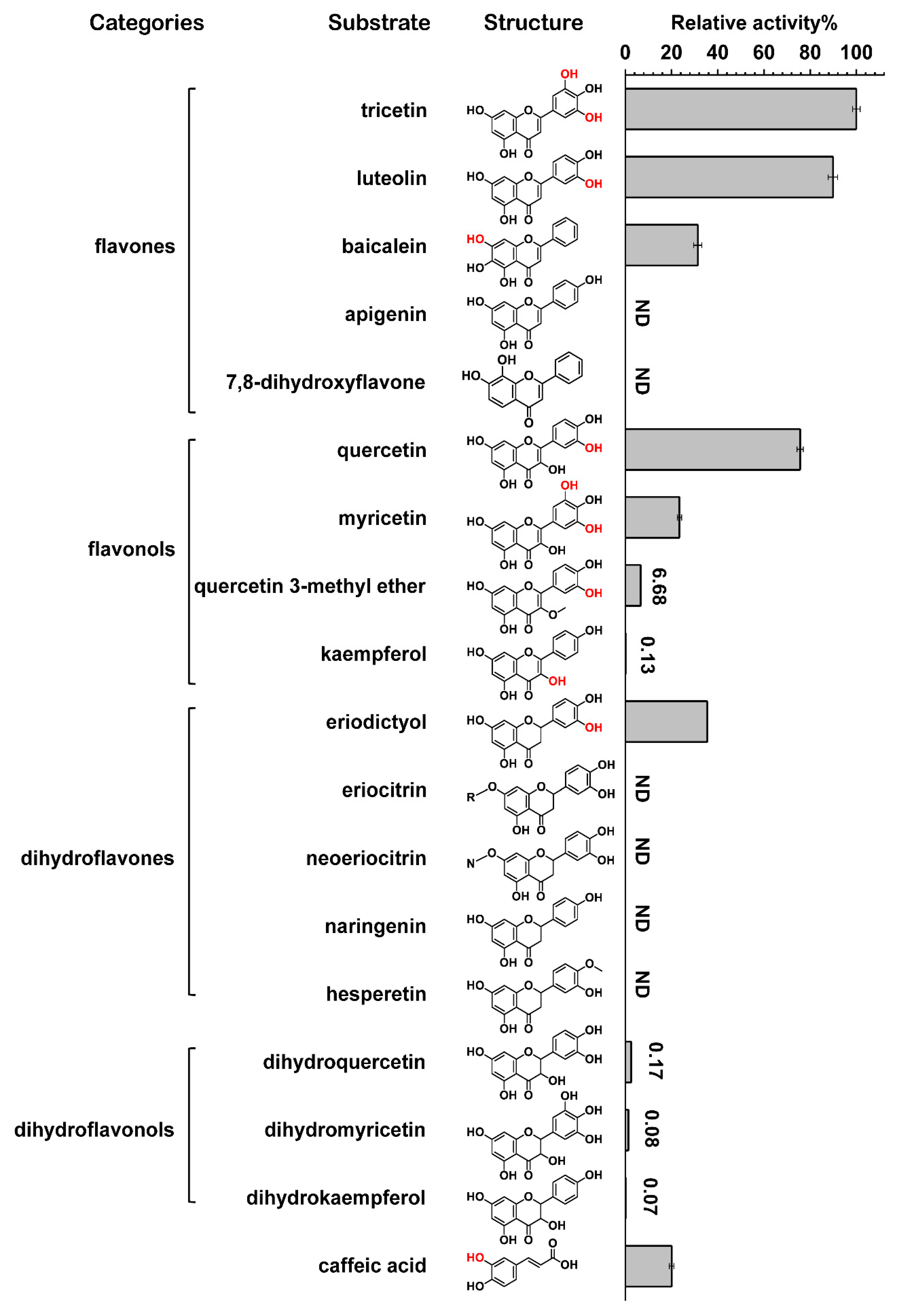
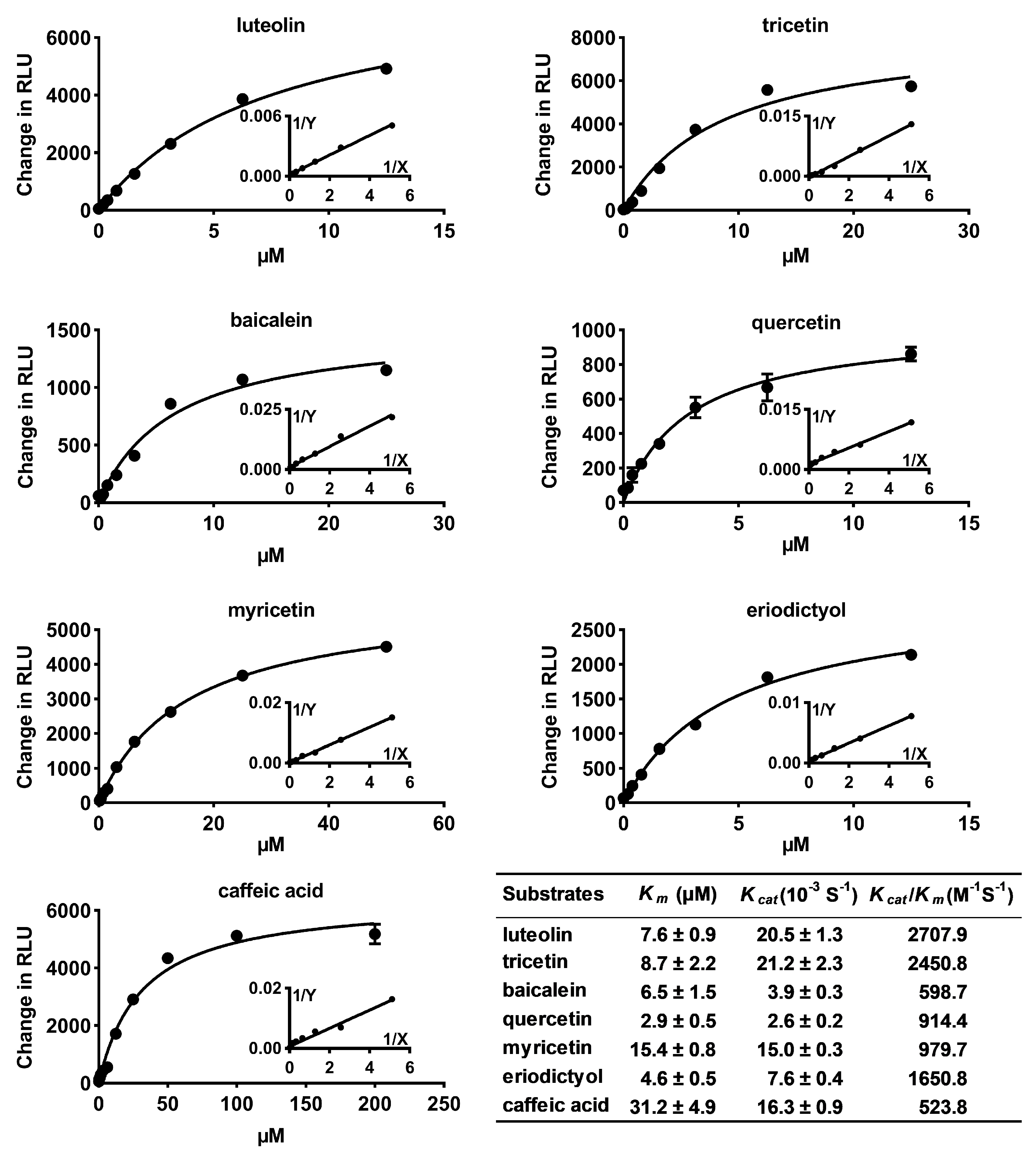
2.2. Substrate Specificity and Kinetic Parameters of CrOMT2
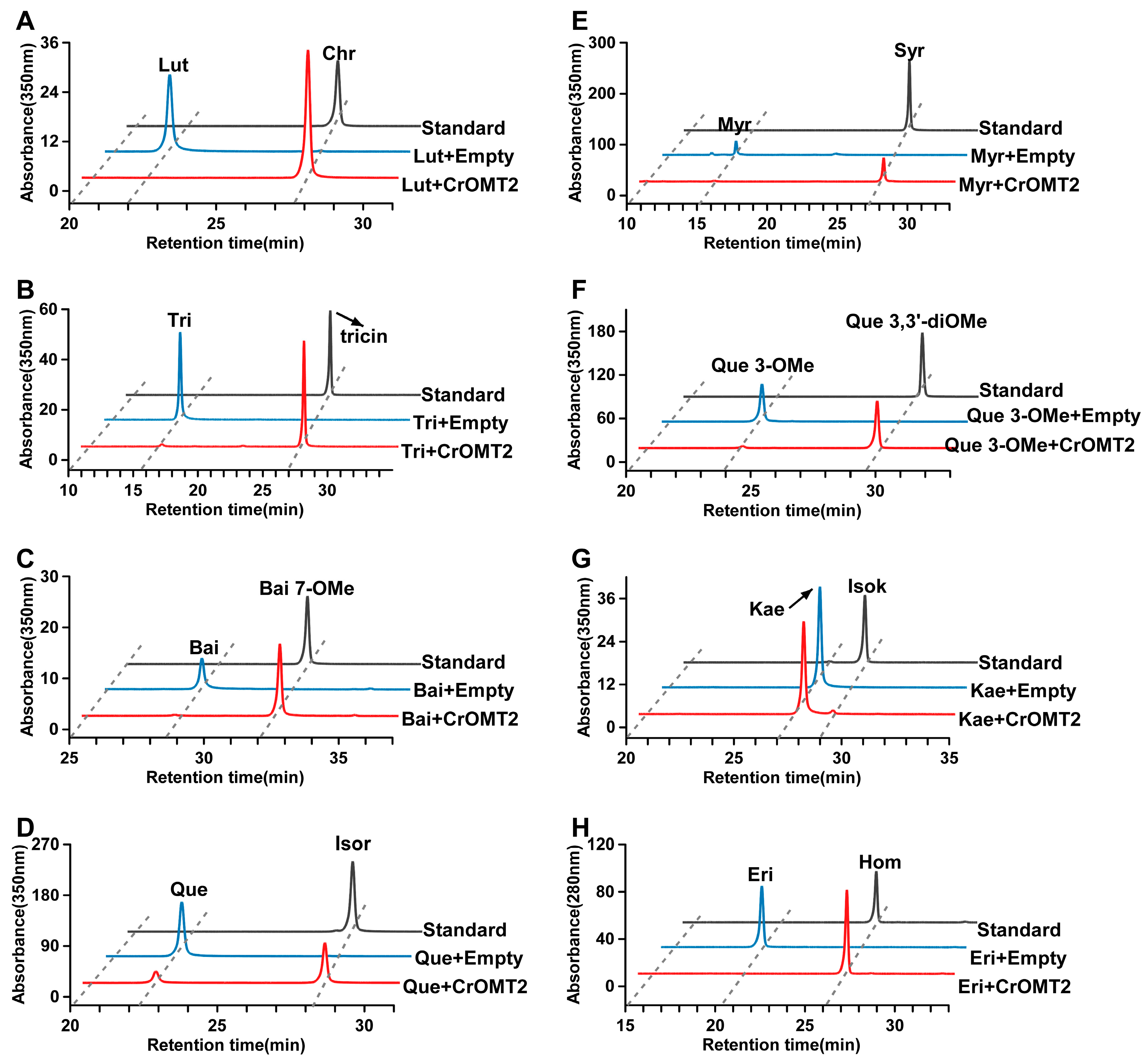
2.3. Identification of Methylated Products
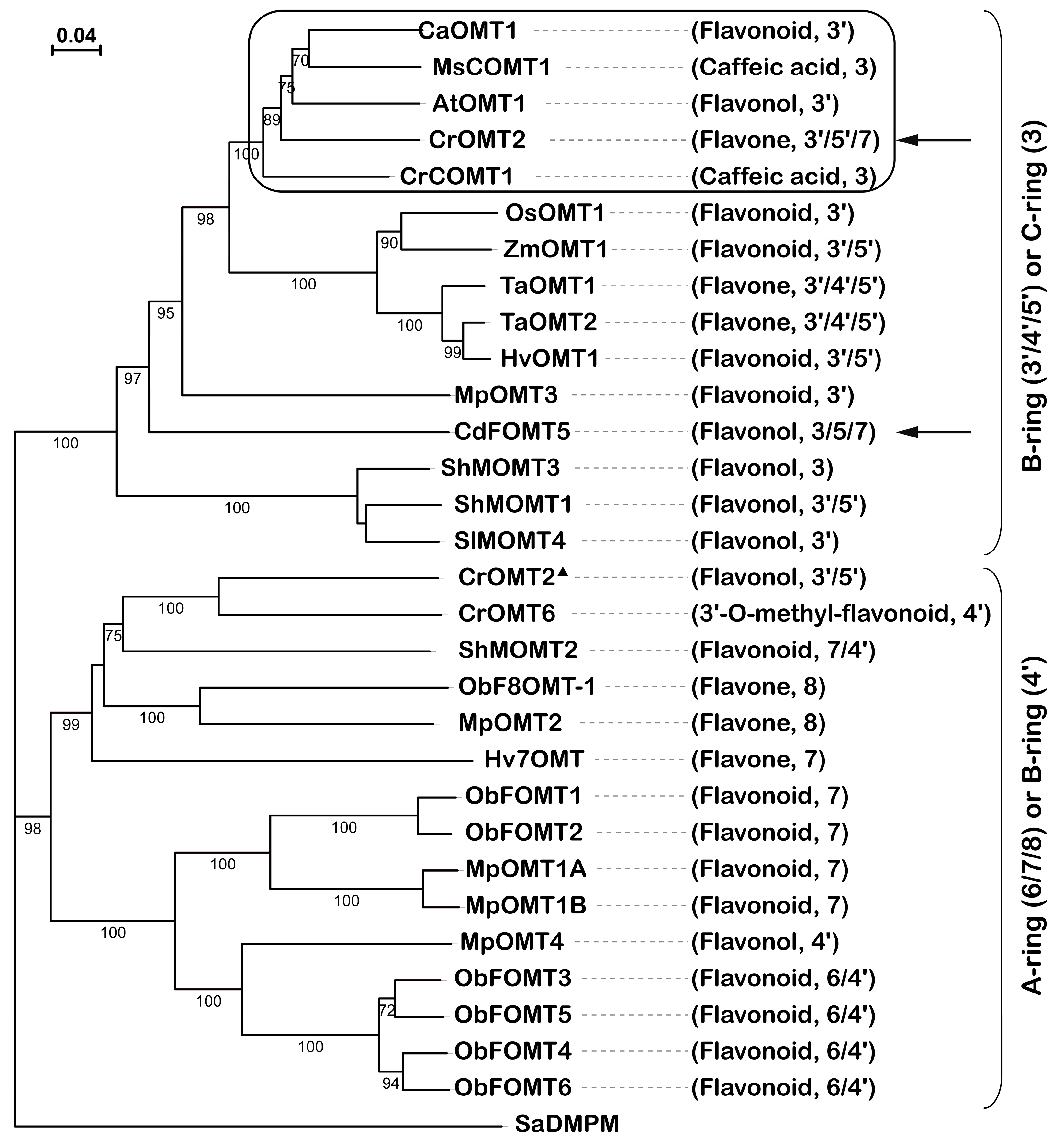
2.4. Phylogenetic Relationship of CrOMT2 to other COMTs
2.5. In Vitro Cytotoxicity of Flavonoids on Human Gastric Cancer Cells
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Chemicals and Reagents
4.3. Isolation, Cloning, and Heterologous Expression of CrOMT2
4.4. Phylogenetic Tree and Protein Alignments
4.5. Enzyme Activity Measurements and Kinetic Analysis
4.6. Cell Culture and Cell Viability Assay
4.7. HPLC and MS/MS Analysis
4.8. Statistical Analyses
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Samanta, A.; Das, G.; Das, S.K. Roles of flavonoids in plants. Int. J. Pharm. Sci. Nanotechnol. 2011, 100, 12–35. [Google Scholar]
- Winkel-Shirley, B. Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol. 2001, 126, 485–493. [Google Scholar] [CrossRef]
- Wink, M. Plant secondary metabolism: Diversity, function and its evolution. Nat. Prod. Commun. 2008, 3, 1205–1216. [Google Scholar] [CrossRef]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J Nutr Sci. 2016, 5, E47. [Google Scholar] [CrossRef] [PubMed]
- Tohge, T.; Fernie, A.R. An overview of compounds derived from the shikimate and phenylpro-panoid pathways and their medicinal importance. Mini-Rev. Med. Chem. 2017, 17, 1013–1027. [Google Scholar] [CrossRef] [PubMed]
- Nabavi, S.M.; Samec, D.; Tomczyk, M.; Milella, L.; Russo, D.; Habtemariam, S.; Suntar, I.; Rastrelli, L.; Daglia, M.; Xiao, J.; et al. Flavonoid biosynthetic pathways in plants: Versatile targets for metabolic engineering. Biotechnol.. Adv. 2020, 38, 107316. [Google Scholar] [CrossRef] [PubMed]
- Ghasemzadeh, A.; Ghasemzadeh, N. Flavonoids and phenolic acids: Role and biochemical activity in plants and human. J Med Plants Res. 2011, 5, 6697–6703. [Google Scholar] [CrossRef]
- D’Amelia, V.; Aversano, R.; Chiaiese, P.; Carputo, D. The antioxidant properties of plant flavonoids: Their exploitation by molecular plant breeding. Phytochem. Rev. 2018, 17, 611–625. [Google Scholar] [CrossRef]
- Wang, Y.; Zang, W.J.; Ji, S.Y.; Cao, J.P.; Sun, C.D. Three polymethoxyflavones purified from Ougan (Citrus reticulata Cv. Suavissima) inhibited LPS-induced NO elevation in the neuroglia BV-2 cell line via the JAK2/STAT3 pathway. Nutrients 2019, 11, 791. [Google Scholar] [CrossRef]
- Wang, Y.; Ji, S.Y.; Zang, W.J.; Wang, N.C.; Cao, J.P.; Li, X.; Sun, C.D. Identification of phenolic compounds from a unique citrus species, finger lime (Citrus australasica) and their inhibition of LPS-induced NO-releasing in BV-2 cell line. Food Chem. Toxicol. 2019, 129, 54–63. [Google Scholar] [CrossRef]
- Wang, Y.; Zhang, X.N.; Xie, W.H.; Zheng, Y.X.; Cao, J.P.; Cao, P.R.; Chen, Q.J.; Li, X.; Sun, C.D. The growth of SGC-7901 tumor xenografts was suppressed by Chinese bayberry anthocyanin extract through upregulating KLF6 gene expression. Nutrients 2016, 8, 599. [Google Scholar] [CrossRef]
- Kim, B.G.; Sung, H.S.; Chong, Y.; Lim, Y.; Ahn, J.H. Plant flavonoid O-methyltransferases: Substrate specificity and application. J. Plant Biol. 2010, 53, 321–329. [Google Scholar] [CrossRef]
- Berim, A.; Gang, D.R. Methoxylated flavones: Occurrence, importance, biosynthesis. Phytochem. Rev. 2016, 15, 363–390. [Google Scholar] [CrossRef]
- Joshi, C.P.; Chiang, V.L. Conserved sequence motifs in plant S-adenosyl-L-methionine-dependent methyltransferases. Plant Mol. Biol. 1998, 37, 663–674. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.G.; Lee, Y.; Hur, H.G.; Lim, Y.; Ahn, J.H. Flavonoid 3’-O-methyltransferase from rice: cDNA cloning, characterization and functional expression. Phytochemistry 2006, 67, 387–394. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Matsuba, Y.; Ning, J.; Schilmiller, A.L.; Hammar, D.; Jones, A.D.; Pichersky, E.; Last, R.L. Analysis of natural and induced variation in tomato glandular trichome flavonoids identifies a gene not present in the reference genome. Plant Cell 2014, 26, 3272–3285. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.M.; Fukushi, Y.; Wollenweber, E.; Ibrahim, R.K. Characterization of two O-methyltransferase-like genes in barley and maize. Pharm. Biol. 2008, 46, 26–34. [Google Scholar] [CrossRef]
- Schmidt, A.; Li, C.; Shi, F.; Jones, A.D.; Pichersky, E. Polymethylated myricetin in trichomes of the wild tomato species Solanum habrochaites and characterization of trichome-specific 3’/5’- and 7/4’-myricetin O-methyltransferases. Plant Physiol. 2011, 155, 1999–2009. [Google Scholar] [CrossRef]
- Cacace, S.; Schroder, G.; Wehinger, E.; Strack, D.; Schmidt, J.; Schroder, J. A flavonol O-methyltransferase from Catharanthus roseus performing two sequential methylations. Phytochemistry 2003, 62, 127–137. [Google Scholar] [CrossRef]
- Cho, M.H.; Park, H.L.; Park, J.H.; Lee, S.W.; Bhoo, S.H.; Hahn, T.R. Characterization of regiospecific flavonoid 3’/5’-O-methyltransferase from tomato and its application in flavonoid biotransformation. J Korean Soc Appl Bi. 2012, 55, 749–755. [Google Scholar] [CrossRef]
- Zhou, J.M.; Seo, Y.W.; Ibrahim, R.K. Biochemical characterization of a putative wheat caffeic acid O-methyltransferase. Plant Physiol. Biochem. 2009, 47, 322–326. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.M.; Gold, N.D.; Martin, V.J.J.; Wollenweber, E.; Ibrahim, R.K. Sequential O-methylation of tricetin by a single gene product in wheat. Bba-Gen Subjects. 2006, 1760, 1115–1124. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Qian, J.; Cao, J.P.; Wang, D.L.; Liu, C.R.; Yang, R.X.; Li, X.; Sun, C.D. Antioxidant capacity, anticancer ability and flavonoids composition of 35 citrus (Citrus reticulata Blanco) varieties. Molecules 2017, 22, 1114. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.G.; Luo, Y.; Wu, H.K.; Xi, W.P.; Yu, J.; Zhang, Q.Y.; Zhou, Z.Q. Systematic analysis of O-methyltransferase gene family and identification of potential members involved in the formation of O-methylated flavonoids in Citrus. Gene 2016, 575, 458–472. [Google Scholar] [CrossRef]
- Itoh, N.; Iwata, C.; Toda, H. Molecular cloning and characterization of a flavonoid-O-methyltransferase with broad substrate specificity and regioselectivity from Citrus depressa. BMC Plant Biol. 2016, 16, 180. [Google Scholar] [CrossRef]
- Brunet, G.; Saleh, N.A.; Ibrahim, R.K. Stepwise methylation of quercetin by cell-free extracts of Citrus tissues. Zeitschrift für Naturforschung C 1978, 33, 786–788. [Google Scholar] [CrossRef]
- Brunet, G.; Ibrahim, R.K. O-methylation of flavonoids by cell-free extracts of calamondin orange. Phytochemistry 1980, 19, 741–746. [Google Scholar] [CrossRef]
- Muzac, I.; Wang, J.; Anzellotti, D.; Zhang, H.; Ibrahim, R.K. Functional expression of an Arabidopsis cDNA clone encoding a flavonol 3’-O-methyltransferase and characterization of the gene product. Arch. Biochem. Biophys. 2000, 375, 385–388. [Google Scholar] [CrossRef]
- Zubieta, C.; Kota, P.; Ferrer, J.L.; Dixon, R.A.; Noel, J.P. Structural basis for the modulation of lignin monomer methylation by caffeic acid/5-hydroxyferulic acid 3/5-O-methyltransferase. Plant Cell 2002, 14, 1265–1277. [Google Scholar] [CrossRef] [PubMed]
- Schroder, G.; Wehinger, E.; Lukacin, R.; Wellmann, F.; Seefelder, W.; Schwab, W.; Schroder, J. Flavonoid methylation: A novel 4’-O-methyltransferase from Catharanthus roseus, and evidence that partially methylated flavanones are substrates of four different flavonoid dioxygenases. Phytochemistry 2004, 65, 1085–1094. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.; Kim, B.G.; Chong, Y.; Lim, Y.; Ahn, J.-H. Cation dependent O-methyltransferases from rice. Planta 2008, 227, 641–647. [Google Scholar] [CrossRef] [PubMed]
- Tsukasa, I. Flavonoid properties of five families newly incorporated into the order Caryophyllales (review). Bull. Natl. Mus. Nat. Sci. 2013, 39, 25–51. [Google Scholar]
- Boerjan, W.; Ralph, J.; Baucher, M. Lignin biosynthesis. Annu. Rev. Plant. Biol. 2003, 54, 519–546. [Google Scholar] [CrossRef] [PubMed]
- Shimizu, T.; Lin, F.Q.; Hasegawa, M.; Okada, K.; Nojiri, H.; Yamane, H. Purification and identification of naringenin 7-O-methyltransferase, a key enzyme in biosynthesis of flavonoid phytoalexin sakuranetin in rice. J. Biol. Chem. 2012, 287, 19315–19325. [Google Scholar] [CrossRef] [PubMed]
- Willits, M.G.; Giovanni, M.; Prata, R.T.N.; Kramer, C.M.; De Luca, V.; Steffens, J.C.; Graser, G. Bio-fermentation of modified flavonoids: An example of in vivo diversification of secondary metabolites. Phytochemistry 2004, 65, 31–41. [Google Scholar] [CrossRef]
- Tohge, T.; Yonekura-Sakakibara, K.; Niida, R.; Watanabe-Takahashi, A.; Saito, K. Phytochemical genomics in Arabidopsis thaliana: A case study for functional identification of flavonoid biosynthesis genes. Pure Appl. Chem. 2007, 79, 811–823. [Google Scholar] [CrossRef]
- Goujon, T.; Sibout, R.; Pollet, B.; Maba, B.; Nussaume, L.; Bechtold, N.; Lu, F.C.; Ralph, J.; Mila, I.; Barriere, Y.; et al. A new Arabidopsis thaliana mutant deficient in the expression of O-methyltransferase impacts lignins and sinapoyl esters. Plant. Mol. Biol. 2003, 51, 973–989. [Google Scholar] [CrossRef]
- Guo, D.; Chen, F.; Inoue, K.; Blount, J.W.; Dixon, R.A. Downregulation of caffeic acid 3-O-methyltransferase and caffeoyl CoA 3-O-methyltransferase in transgenic alfalfa: Impacts on lignin structure and implications for the biosynthesis of G and S lignin. Plant Cell 2001, 13, 73–88. [Google Scholar] [CrossRef]
- Schroder, G.; Wehinger, E.; Schroder, J. Predicting the substrates of cloned plant O-methyltransferases. Phytochemistry 2002, 59, 1–8. [Google Scholar] [CrossRef]
- Berim, A.; Hyatt, D.C.; Gang, D.R. A set of regioselective O-methyltransferases gives rise to the complex pattern of methoxylated flavones in sweet basil. Plant Physiol. 2012, 160, 1052–1069. [Google Scholar] [CrossRef]
- Christensen, A.B.; Gregersen, P.L.; Olsen, C.E.; Collinge, D.B. A flavonoid 7-O-methyltransferase is expressed in barley leaves in response to pathogen attack. Plant. Mol. Biol. 1998, 36, 219–227. [Google Scholar] [CrossRef] [PubMed]
- Ibrahim, R.K. A forty-year journey in plant research: Original contributions to flavonoid biochemistry. Can. J. Botany-Revue Canadienne De Botanique 2005, 83, 433–450. [Google Scholar] [CrossRef]
- Kiefer, E.; Heller, W.; Ernst, D. A simple and efficient protocol for isolation of functional RNA from plant tissues rich in secondary metabolites. Plant. Mol. Biol. Report. 2000, 18, 33–39. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Notredame, C.; Higgins, D.G.; Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 2000, 302, 205–217. [Google Scholar] [CrossRef]
- Robert, X.; Gouet, P. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Res. 2014, 42, W320–W324. [Google Scholar] [CrossRef]
- Zhuo, W.; Liu, Y.; Li, S.; Guo, D.; Sun, Q.; Jin, J.; Rao, X.; Li, M.; Sun, M.; Jiang, M.; et al. Long noncoding RNA GMAN, up-regulated in gastric cancer tissues, is associated with metastasis in patients and promotes translation of ephrin A1 by competitively binding GMAN-AS. Gastroenterology 2019, 156, 676–691. [Google Scholar] [CrossRef]
Sample Availability: All samples are available from the authors. |


| Substrate | IC50 (μM) | Product | IC50 (μM) | |
|---|---|---|---|---|
| SCG-7910 | tricetin | 78.8 | tricin | \ |
| luteolin | 15.1 | chrysoeriol | 13.3 | |
| baicalein | 37.5 | baicalein 7-methyl ether | 40.2 | |
| quercetin | 37.7 | isorhamnetin | 10.7 | |
| myricetin | \ | syringetin | \ | |
| eriodictyol | 66.4 | homoeriodictyol | 166 | |
| BGC-823 | tricetin | \ | tricin | \ |
| luteolin | 41.2 | chrysoeriol | 24.2 | |
| baicalein | 86.9 | baicalein 7-methyl ether | \ | |
| quercetin | 322 | isorhamnetin | 48.4 | |
| myricetin | \ | syringetin | \ | |
| eriodictyol | 188 | homoeriodictyol | \ |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liu, X.; Wang, Y.; Chen, Y.; Xu, S.; Gong, Q.; Zhao, C.; Cao, J.; Sun, C. Characterization of a Flavonoid 3’/5’/7-O-Methyltransferase from Citrus reticulata and Evaluation of the In Vitro Cytotoxicity of Its Methylated Products. Molecules 2020, 25, 858. https://doi.org/10.3390/molecules25040858
Liu X, Wang Y, Chen Y, Xu S, Gong Q, Zhao C, Cao J, Sun C. Characterization of a Flavonoid 3’/5’/7-O-Methyltransferase from Citrus reticulata and Evaluation of the In Vitro Cytotoxicity of Its Methylated Products. Molecules. 2020; 25(4):858. https://doi.org/10.3390/molecules25040858
Chicago/Turabian StyleLiu, Xiaojuan, Yue Wang, Yezhi Chen, Shuting Xu, Qin Gong, Chenning Zhao, Jinping Cao, and Chongde Sun. 2020. "Characterization of a Flavonoid 3’/5’/7-O-Methyltransferase from Citrus reticulata and Evaluation of the In Vitro Cytotoxicity of Its Methylated Products" Molecules 25, no. 4: 858. https://doi.org/10.3390/molecules25040858
APA StyleLiu, X., Wang, Y., Chen, Y., Xu, S., Gong, Q., Zhao, C., Cao, J., & Sun, C. (2020). Characterization of a Flavonoid 3’/5’/7-O-Methyltransferase from Citrus reticulata and Evaluation of the In Vitro Cytotoxicity of Its Methylated Products. Molecules, 25(4), 858. https://doi.org/10.3390/molecules25040858









