Synthesis of Oligonucleotides Containing 2′-N-alkylaminocarbonyl-2′-amino-LNA (2′-urea-LNA) Moieties Using Post-Synthetic Modification Strategy
Abstract
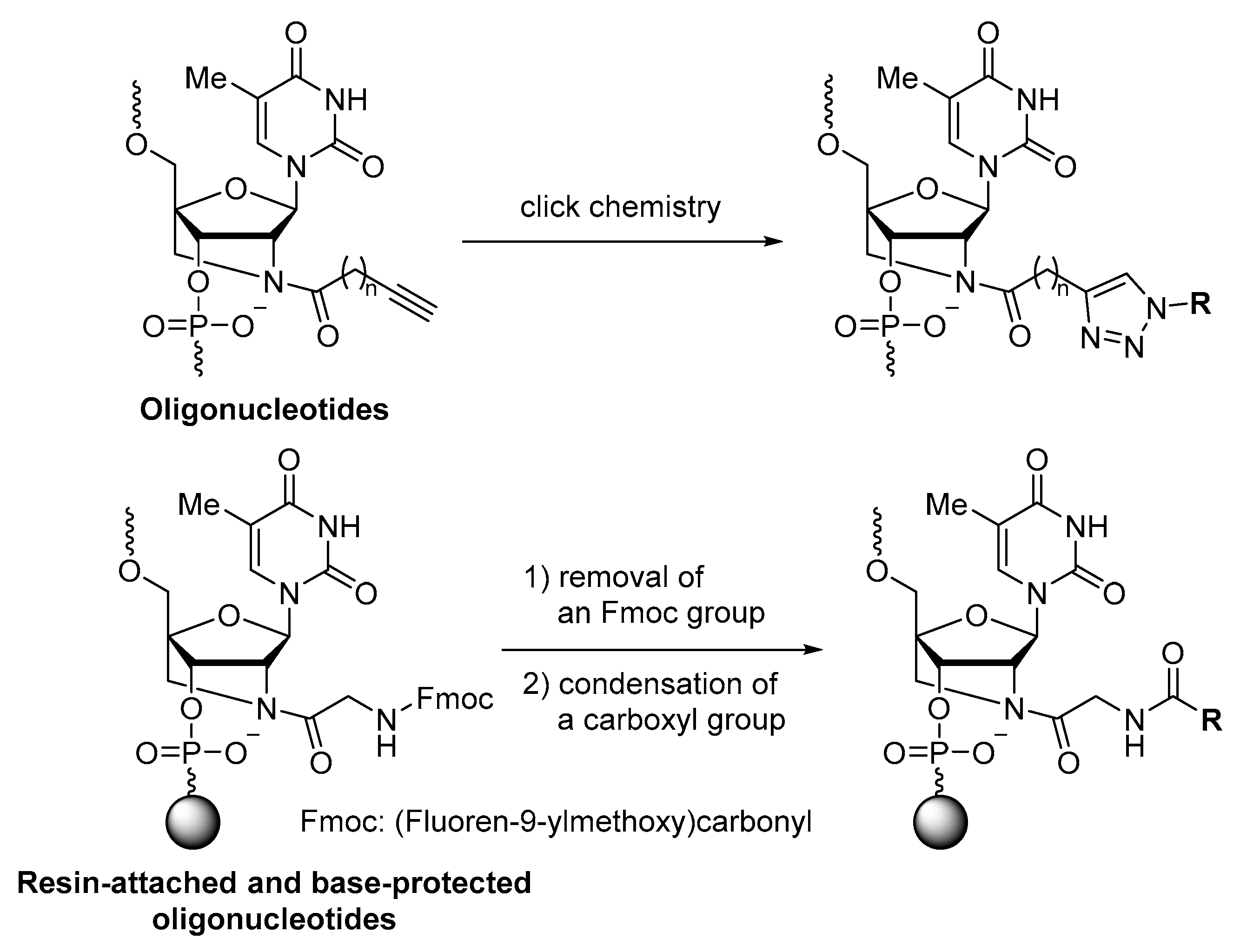
1. Introduction
2. Results and Discussion
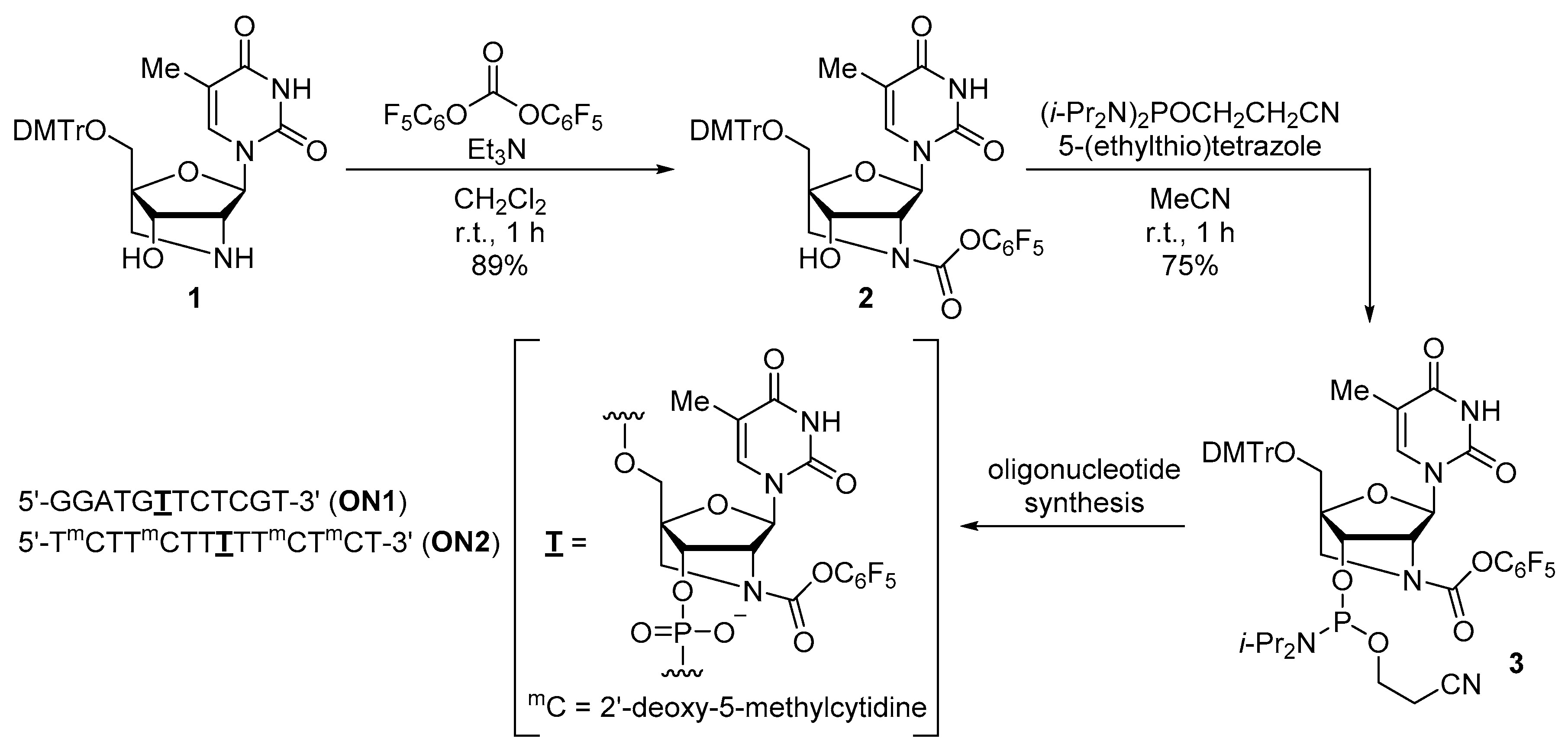
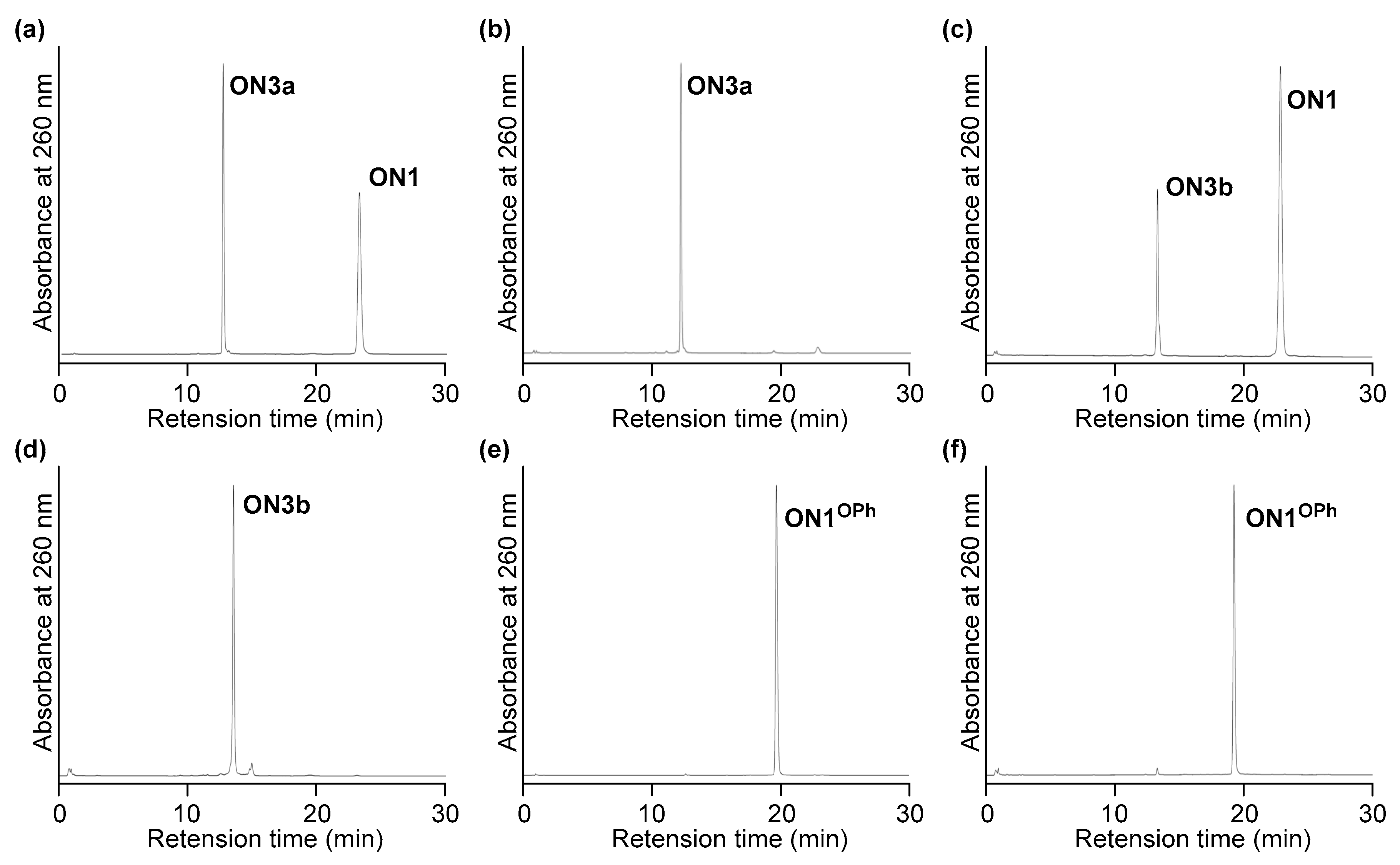
2.1. Synthesis
2.2. Evaluation
3. Materials and Methods
3.1. General
3.2. Synthesis
3.3. UV-Melting Experiment
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Goodchild, J. Conjugations of oligonucleotides and modified oligonucleotides: A review of their synthesis and properties. Bioconj. Chem. 1990, 1, 165–187. [Google Scholar] [CrossRef]
- Verma, S.; Eckstein, F. Modified oligonucleotides: Synthesis and strategy for users. Annu. Rev. Biochem. 1998, 67, 99–134. [Google Scholar] [CrossRef]
- Gramlich, P.M.E.; Wirges, C.T.; Manetto, A.; Carell, T. Postsynthetic DNA modification through the copper-catalyzed azide-alkyne cycloaddition reaction. Angew. Chem. Int. Ed. 2008, 47, 8350–8358. [Google Scholar] [CrossRef] [PubMed]
- Singh, Y.; Murat, P.; Defrancq, E. Recent developments in oligonucleotide conjugation. Chem. Soc. Rev. 2010, 39, 2054–2070. [Google Scholar] [CrossRef] [PubMed]
- El-Sagheer, A.H.; Brown, T. Click chemistry with DNA. Chem. Soc. Rev. 2010, 39, 1388–1405. [Google Scholar] [CrossRef]
- Shaughnessy, K.H. Palladium-catalyzed modification of unprotected nucleosides, nucleotides, and oligonucleotides. Molecules 2015, 20, 9419–9454. [Google Scholar] [CrossRef]
- Defrancq, E.; Messaoudi, S. Palladium-mediated labeling of nucleic acids. ChemBioChem 2017, 18, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Hari, Y. Site-specific modification of nucleobases in oligonucleotides. In Synthesis of Therapeutic Oligonucleotides; Obika, S., Sekine, M., Eds.; Springer: Singapore, 2018; pp. 131–145. [Google Scholar]
- Obika, S.; Rahman, S.M.A.; Fujisaka, A.; Kawada, Y.; Baba, T.; Imanishi, T. Bridged nucleic acids: Development, synthesis and properties. Heterocycles 2010, 81, 1347–1392. [Google Scholar] [CrossRef]
- Yamamoto, T.; Nakatani, M.; Narukawa, K.; Obika, S. Antisense drug discovery and development. Future Med. Chem. 2011, 3, 339–365. [Google Scholar] [CrossRef] [PubMed]
- Zhou, C.; Chattopadhyaya, J. Intramolecular free-radical cyclization reactions on pentose sugars for the synthesis of carba-LNA and carba-ENA and the application of their modified oligonucleotides as potential RNA targeted therapeutics. Chem. Rev. 2012, 112, 3808–3832. [Google Scholar] [CrossRef]
- Wan, W.B.; Seth, P.P. The medicinal chemistry of therapeutic oligonucleotides. J. Med. Chem. 2016, 59, 9645–9667. [Google Scholar] [CrossRef] [PubMed]
- Hari, Y. Bridged nucleosides as building blocks of oligonucleotides: Synthesis and properties. Heterocycles 2020. [Google Scholar] [CrossRef]
- Singh, S.K.; Kumar, R.; Wengel, J. Synthesis of 2′-amino-LNA: A novel conformationally restricted high-affinity oligonucleotide analogue with a handle. J. Org. Chem. 1998, 63, 10035–10039. [Google Scholar] [CrossRef]
- Astakhova, I.K.; Wengel, J. Scaffolding along nucleic acid duplexes using 2′-amino-locked nucleic acids. Acc. Chem. Res. 2014, 47, 1768–1777. [Google Scholar] [CrossRef] [PubMed]
- Lou, C.; Vester, B.; Wengel, J. Oligonucleotides containing a piperazino-modified 2′-amino-LNA monomer exhibit very high duplex stability and remarkable nuclease resistance. Chem. Commun. 2015, 51, 4024–4027. [Google Scholar] [CrossRef] [PubMed]
- Ries, A.; Kumar, R.; Lou, C.; Kosbar, T.; Vengut-Climent, E.; Jørgensen, P.T.; Morales, J.C.; Wengel, J. Synthesis and biophysical investigations of oligonucleotides containing galactose-modified DNA, LNA, and 2′-amino-LNA monomers. J. Org. Chem. 2016, 81, 10845–10856. [Google Scholar] [CrossRef]
- Lou, C.; Samuelsen, S.V.; Christensen, N.J.; Vester, B.; Wengel, J. Oligonucleotides containing aminated 2′-amino-LNA nucleotides: Synthesis and strong binding to complementary DNA and RNA. Bioconj. Chem. 2017, 28, 1214–1220. [Google Scholar] [CrossRef]
- Hvam, M.L.; Cai, Y.; Dagnæs-Hansen, F.; Nielsen, J.S.; Wengel, J.; Kjems, J.; Howard, K.A. Fatty acid-modified gapmer antisense oligonucleotide and serum albumin constructs for pharmacokinetic modulation. Mol. Ther. 2017, 25, 1710–1717. [Google Scholar] [CrossRef]
- Kumar, R.; Ries, A.; Wengel, J. Synthesis and excellent duplex stability of oligonucleotides containing 2′-amino-LNA functionalized with galactose units. Molecules 2017, 22, 852. [Google Scholar] [CrossRef]
- Ejlersen, M.; Christensen, N.J.; Sørensen, K.K.; Jensen, K.J.; Wengel, J.; Lou, C. Synergy of two highly specific biomolecular recognition events: Aligning an AT-hook peptide in DNA minor grooves via covalent conjugation to 2′-amino-LNA. Bioconj. Chem. 2018, 29, 1025–1029. [Google Scholar] [CrossRef]
- Osawa, T.; Yamashita, S.; Nakanishi, A.; Ito, Y.; Hari, Y. Synthesis and hybridization properties of oligonucleotides including 2′-N-alkoxycarbonyl-2′-amino-LNA derivatives. Heterocycles 2019, 99, 502–520. [Google Scholar]
- Jørgensen, A.S.; Gupta, P.; Wengel, J.; Astakhova, I.K. “Clickable” LNA/DNA probes for fluorescence sensing of nucleic acids and autoimmune antibodies. Chem. Commun. 2013, 49, 10751–10753. [Google Scholar] [CrossRef] [PubMed]
- Johannsen, M.W.; Crispino, L.; Wamberg, M.C.; Kalra, N.; Wengel, J. Amino acids attached to 2′-amino-LNA: Synthesis and excellent duplex stability. Org. Biomol. Chem. 2011, 9, 243–252. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not available. |
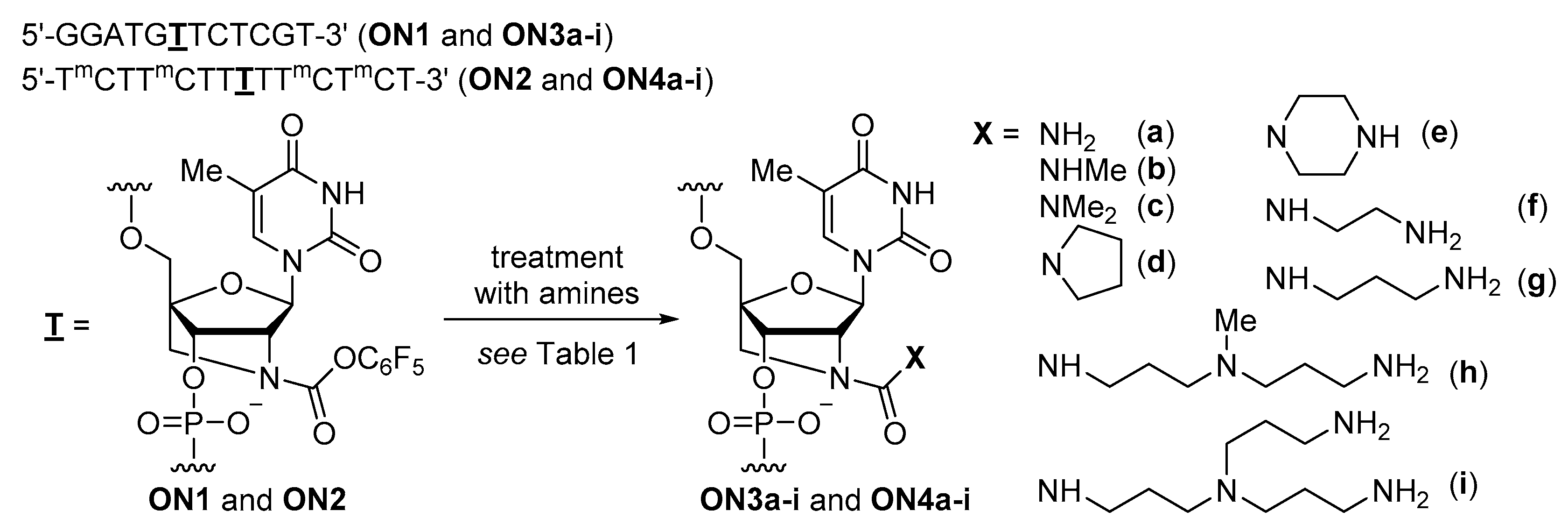
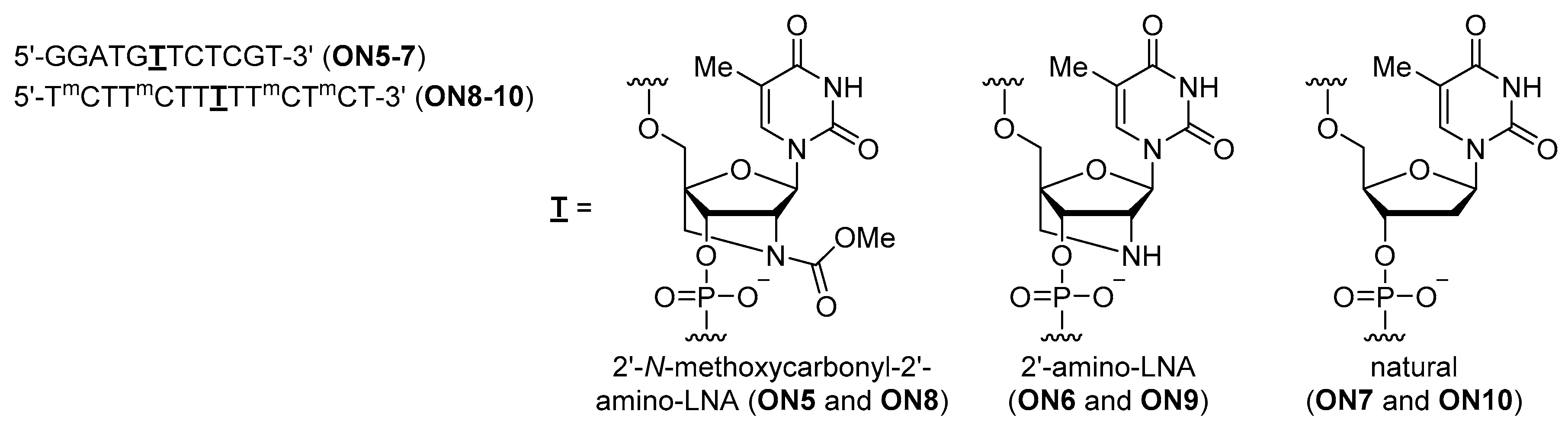
| Substrates | Products | Isolated Yield | Substrates | Products | Isolated Yield |
|---|---|---|---|---|---|
| ON11 | ON3a | 76% | ON21 | ON4a | 67% |
| ON12 | ON3b | 86% | ON22 | ON4b | 69% |
| ON12 | ON3c | 68% | ON22 | ON4c | 46% |
| ON13 | ON3d | 84% | ON23 | ON4d | 82% |
| ON13 | ON3e | 80% | ON23 | ON4e | 79% |
| ON13 | ON3f | 95% | ON23 | ON4f | 70% |
| ON13 | ON3g | 94% | ON23 | ON4g | 77% |
| ON13 | ON3h | 94% | ON23 | ON4h | 76% |
| ON13 | ON3i | 83% | ON23 | ON4i | 73% |
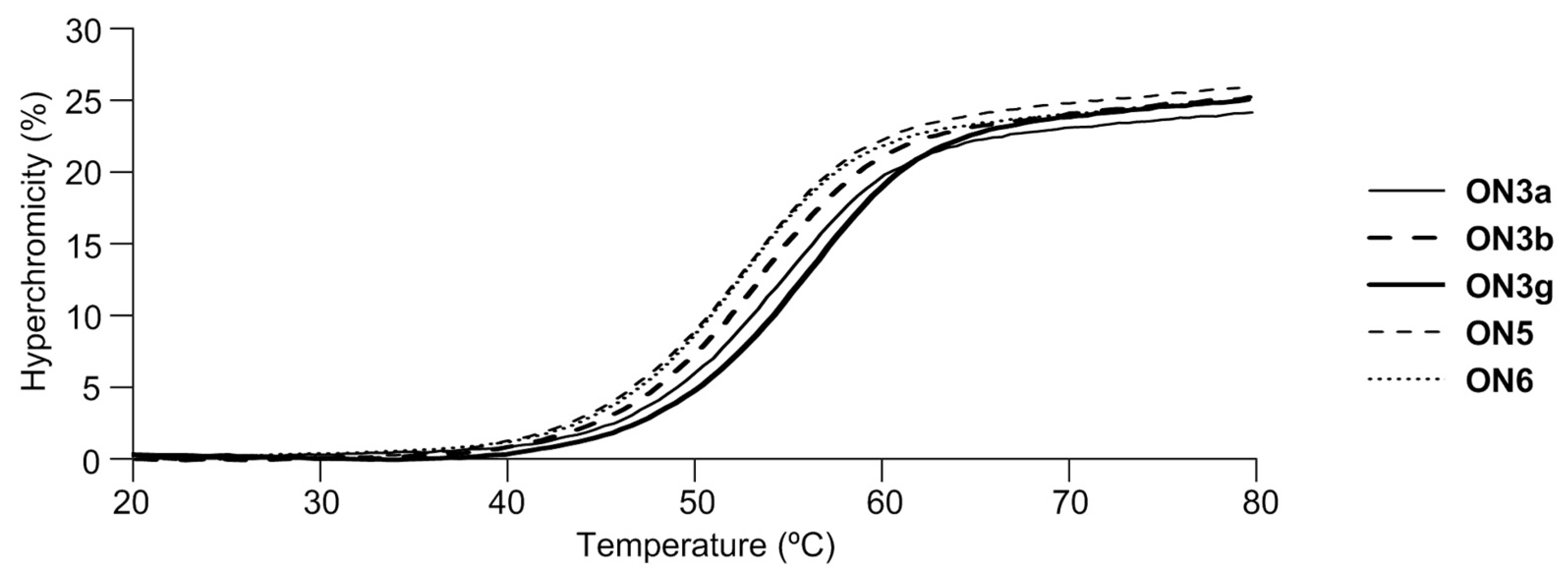
| Oligonucleotides 2 | Tm (ΔTm 3) with ssDNA | Tm (ΔTm 3) with ssRNA | |
|---|---|---|---|
| ON3a | X = NH2 | 54 °C (+3 °C) | 58 °C (+6 °C) |
| ON3b | X = NHMe | 53 °C (+2 °C) | 58 °C (+6 °C) |
| ON3c | X = NMe2 | 52 °C (+1 °C) | 57 °C (+5 °C) |
| ON3d | X = pyrrolidin-1-yl | 52 °C (+1 °C) | 57 °C (+5 °C) |
| ON3e | X = piperazin-1-yl | 54 °C (+3 °C) | 57 °C (+5 °C) |
| ON3f | X = NH(CH2)2NH2 | 55 °C (+4 °C) | 57 °C (+5 °C) |
| ON3g | X = NH(CH2)3NH2 | 55 °C (+4 °C) | 58 °C (+6 °C) |
| ON3h | X = NH(CH2)3NH(Me)(CH2)3NH2 | 54 °C (+3 °C) | 57 °C (+5 °C) |
| ON3i | X = NH(CH2)3N[(CH2)3NH2]2 | 55 °C (+4 °C) | 55 °C (+3 °C) |
| ON5 | X = OMe | 52 °C (+1 °C) | 58 °C (+6 °C) |
| ON6 | 2′-amino-LNA | 52 °C (+1 °C) | 58 °C (+6 °C) |
| ON7 | natural | 51 °C | 52 °C |
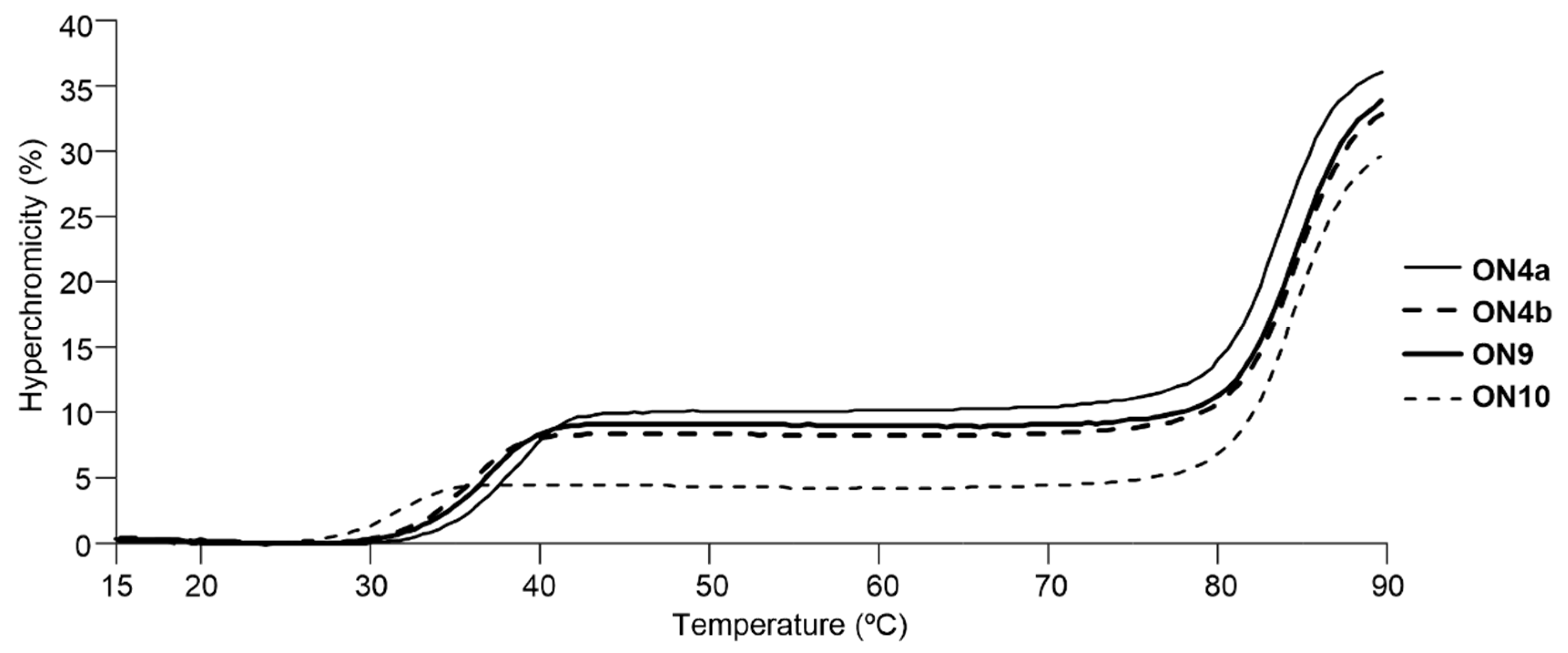
| Oligonucleotides 2 | Tm (ΔTm 3) | |
|---|---|---|
| ON4a | X = NH2 | 38 °C (+7 °C) |
| ON4b | X = NHMe | 36 °C (+5 °C) |
| ON4c | X = NMe2 | 38 °C (+7 °C) |
| ON4d | X = pyrrolidin-1-yl | 31 °C (0 °C) |
| ON4e | X = piperazin-1-yl | 34 °C (+3 °C) |
| ON4f | X = NH(CH2)2NH2 | 35 °C (+4 °C) |
| ON4g | X = NH(CH2)3NH2 | 35 °C (+4 °C) |
| ON4h | X = NH(CH2)3NH(Me)(CH2)3NH2 | 35 °C (+4 °C) |
| ON4i | X = NH(CH2)3N[(CH2)3NH2]2 | 35 °C (+4 °C) |
| ON8 | X = OMe | 36 °C (+5 °C) |
| ON9 | 2′-amino-LNA | 37 °C (+6 °C) |
| ON10 | natural | 31 °C |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yamashita, S.; Nishida, K.; Osawa, T.; Nakanishi, A.; Ito, Y.; Hari, Y. Synthesis of Oligonucleotides Containing 2′-N-alkylaminocarbonyl-2′-amino-LNA (2′-urea-LNA) Moieties Using Post-Synthetic Modification Strategy. Molecules 2020, 25, 346. https://doi.org/10.3390/molecules25020346
Yamashita S, Nishida K, Osawa T, Nakanishi A, Ito Y, Hari Y. Synthesis of Oligonucleotides Containing 2′-N-alkylaminocarbonyl-2′-amino-LNA (2′-urea-LNA) Moieties Using Post-Synthetic Modification Strategy. Molecules. 2020; 25(2):346. https://doi.org/10.3390/molecules25020346
Chicago/Turabian StyleYamashita, Shoko, Kodai Nishida, Takashi Osawa, Ayumi Nakanishi, Yuta Ito, and Yoshiyuki Hari. 2020. "Synthesis of Oligonucleotides Containing 2′-N-alkylaminocarbonyl-2′-amino-LNA (2′-urea-LNA) Moieties Using Post-Synthetic Modification Strategy" Molecules 25, no. 2: 346. https://doi.org/10.3390/molecules25020346
APA StyleYamashita, S., Nishida, K., Osawa, T., Nakanishi, A., Ito, Y., & Hari, Y. (2020). Synthesis of Oligonucleotides Containing 2′-N-alkylaminocarbonyl-2′-amino-LNA (2′-urea-LNA) Moieties Using Post-Synthetic Modification Strategy. Molecules, 25(2), 346. https://doi.org/10.3390/molecules25020346






