Peptides with Dual Antimicrobial–Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides
Abstract
1. Introduction
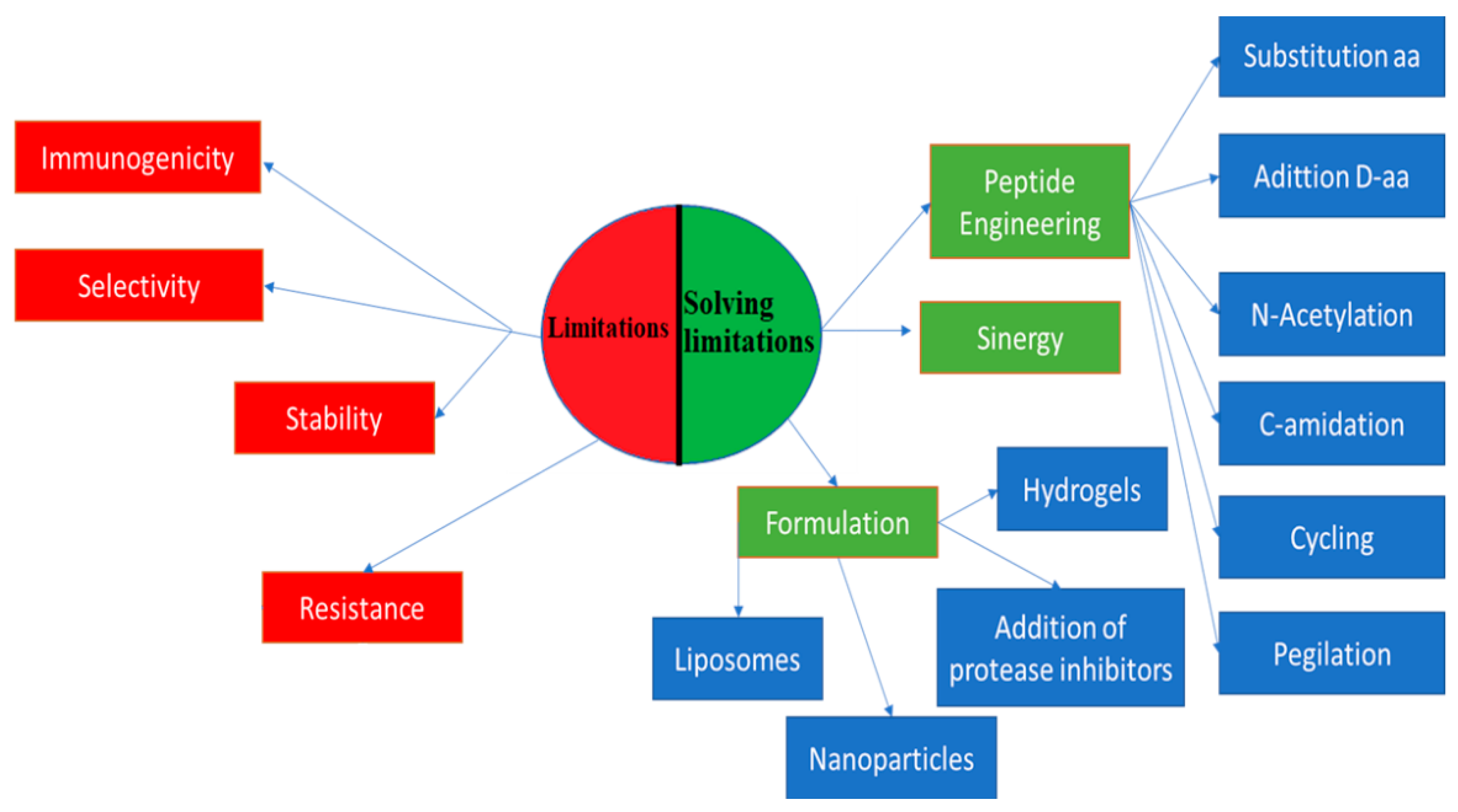
2. Efforts to Overcome Peptide Limitations
3. Peptides with Dual Antimicrobial–Anticancer Activity
4. Toward Rational Peptide Design
5. Physicochemical Methods of Rational Design of Anticancer Peptides
6. Sequence Template Methods of Rational Design of Anticancer Peptides
7. Automated Computational Methods for Anticancer Peptide Prediction
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ACTH | Adrenocorticotropic hormone |
| POPS | Phosphatidylserine |
| GnRH | Gonadotropin releasing hormone |
| TUNEL | Terminal deoxynucleotidyl transferase dUTP nick end labeling |
| LDH | Lactate dehydrogenase |
| ROS | Reactive oxygen intermediates |
| Pgp | P-glycoprotein |
| MTT | 3-(4,5-dimethylthiazolyl-2)-2,5-diphenyltetrazolium bromide |
| DAPI | 4′,6-diamidine-2-phenylindol |
| FITC | Fluorescein isothiocyanate |
| ML | Machine learning |
| HKP | Hunter–killer peptide |
| MMP | Matrix metalloproteinase |
| HOP | Hsp organizing protein |
| SVM | Support vector machine |
| DL | Deep learning |
| AAC | Amino acid composition |
| DPC | Dipeptide composition |
References
- Boopathi, V.; Subramaniyam, S.; Malik, A.; Lee, G.; Manavalan, B.; Yang, D.C. MACPpred: A support vector machine-based meta-predictor for identification of anticancer peptides. Int. J. Mol. Sci. 2019, 20, 1964. [Google Scholar] [CrossRef] [PubMed]
- Kakde, D.; Jain, D.; Shrivastava, V.; Kakde, R.; Patil, A.T. Cancer therapeutics—Opportunities, challenges and advances in drug delivery. J. Appl. Pharm. Sci. 2011, 1, 1–10. [Google Scholar]
- Felício, M.R.; Silva, O.N.; Gonçalves, S.; Santos, N.C.; Franco, O.L. Peptides with dual antimicrobial and anticancer activities. Front. Chem. 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Zompra, A.A.; Galanis, A.; Werbitzky, O.; Albericio, F. Manufacturing peptides as active pharmaceutical ingredients. Future Med. Chem. 2009, 1, 361–377. [Google Scholar] [CrossRef]
- Uhlig, T.; Kyprianou, T.; Martinelli, F.G.; Oppici, C.A.; Heiligers, D.; Hills, D.; Calvo, X.R.; Verhaert, P. The emergence of peptides in the pharmaceutical business: From exploration to exploitation. EuPA Open Proteom. 2014, 4, 58–69. [Google Scholar] [CrossRef]
- Lau, J.L.; Dunn, M.K. Therapeutic peptides: Historical perspectives, current development trends, and future directions. Bioorg. Med. Chem. 2018, 26, 2700–2707. [Google Scholar] [CrossRef]
- Craik, D.J.; Fairlie, D.P.; Liras, S.; Price, D. The future of peptide-based drugs: Peptides in drug development. Chem. Biol. Drug Des. 2013, 81, 136–147. [Google Scholar] [CrossRef]
- Eckert, R. Road to clinical efficacy: Challenges and novel strategies for antimicrobial peptide development. Future Microbiol. 2011, 6, 635–651. [Google Scholar] [CrossRef]
- Otvos, L.; Wade, J.D. Current challenges in peptide-based drug discovery. Front. Chem. 2014, 2. [Google Scholar] [CrossRef]
- Koutsopoulos, S. Peptide Applications in Biomedicine, Biotechnology and Bioengineering; Elsevier: Amsterdam, The Netherlands, 2018. [Google Scholar]
- Correa, W.; Heinbockel, L.; Martinez-de-Tejada, G.; Sánchez, S.; Garidel, P.; Schürholz, T.; Mier, W.; Dupont, A.; Hornef, M.; Gutsmann, T.; et al. Synthetic anti-lipopolysaccharide peptides (SALPs) as effective inhibitors of pathogen-associated molecular patterns (PAMPs). In Antimicrobial Peptides: Basics for Clinical Application; Matsuzaki, K., Ed.; Springer: Singapore, 2019; pp. 111–129. [Google Scholar]
- Marqus, S.; Pirogova, E.; Piva, T.J. Evaluation of the use of therapeutic peptides for cancer treatment. J. Biomed. Sci. 2017, 24, 21. [Google Scholar] [CrossRef]
- Albericio, F.; Kruger, H.G. Therapeutic peptides. Future Med. Chem. 2012, 4, 1527–1531. [Google Scholar] [CrossRef] [PubMed]
- Mahlapuu, M.; Håkansson, J.; Ringstad, L.; Björn, C. Antimicrobial peptides: An emerging category of therapeutic agents. Front. Cell. Infect. Microbiol. 2016, 6. [Google Scholar] [CrossRef] [PubMed]
- Riedl, S.; Zweytick, D.; Lohner, K. Membrane-active host defense peptides—Challenges and perspectives for the development of novel anticancer drugs. Chem. Phys. Lipids 2011, 164, 766–781. [Google Scholar] [CrossRef] [PubMed]
- Gaspar, D.; Veiga, A.S.; Castanho, M.A.R.B. From antimicrobial to anticancer peptides. A review. Front. Microbiol. 2013, 4. [Google Scholar] [CrossRef]
- Gabernet, G.; Gautschi, D.; Müller, A.T.; Neuhaus, C.S.; Armbrecht, L.; Dittrich, P.S.; Hiss, J.A.; Schneider, G. In silico design and optimization of selective membranolytic anticancer peptides. Sci. Rep. 2012, 9, 11282. [Google Scholar] [CrossRef] [PubMed]
- Hoskin, D.W.; Ramamoorthy, A. Studies on anticancer activities of antimicrobial peptides. Biochim. Biophys. Acta (BBA) Biomembr. 2008, 1778, 357–375. [Google Scholar] [CrossRef]
- Lee, D.; Hahm, K.-S.; Park, Y.; Kim, H.-Y.; Lee, W.; Lim, S.-C.; Seo, Y.-K.; Choi, C.-H. Functional and structural characteristics of anticancer peptide Pep27 analogues. Cancer Cell Int. 2005, 5, 21. [Google Scholar] [CrossRef][Green Version]
- Shoombuatong, W.; Schaduangrat, N.; Nantasenamat, C. Unraveling the bioactivity of anticancer peptides as deduced from machine learning. EXCLI J. 2018, 17, 734–752. [Google Scholar]
- Ellerby, H.M.; Arap, W.; Ellerby, L.M.; Kain, R.; Andrusiak, R.; Rio, G.D.; Krajewski, S.; Lombardo, C.R.; Rao, R.; Ruoslahti, E.; et al. Anti-cancer activity of targeted pro-apoptotic peptides. Nat. Med. 1999, 5, 1032–1038. [Google Scholar] [CrossRef]
- Agyei, D.; Tan, K.-X.; Pan, S.; Udenigwe, C.C.; Danquah, M.K. Peptides for biopharmaceutical applications. In Peptide Applications in Biomedicine, Biotechnology and Bioengineering; Koutsopoulos, S., Ed.; Woodhead Publishing: Sawston, UK, 2018; pp. 231–251. [Google Scholar]
- Drayton, M.; Kizhakkedathu, J.N.; Straus, S.K. Towards robust delivery of antimicrobial peptides to combat bacterial resistance. Molecules 2020, 25, 3048. [Google Scholar] [CrossRef]
- Kowalczyk, R.; Harris, P.W.R.; Williams, G.M.; Yang, S.-H.; Brimble, M.A. Peptide lipidation—A synthetic strategy to afford peptide based therapeutics. In Peptides and Peptide-Based Biomaterials and Their Biomedical Applications; Sunna, A., Care, A., Bergquist, P.L., Eds.; Springer International Publishing: Cham, Switzerland, 2017; Volume 1030, pp. 185–227. [Google Scholar]
- Jacques, P. Surfactin and other lipopeptides from Bacillus spp. In Microbiology Monographs; Springer Science and Business Media: Berlin/Heidelberg, Germany, 2010; Volume 20, pp. 57–91. [Google Scholar]
- Kampshoff, F.; Willcox, M.D.P.; Dutta, D. A pilot study of the synergy between two antimicrobial peptides and two common antibiotics. Antibiotics 2019, 8, 60. [Google Scholar] [CrossRef] [PubMed]
- Kumar, P.; Kizhakkedathu, J.; Straus, S. Antimicrobial peptides: Diversity, mechanism of action and strategies to improve the activity and biocompatibility in vivo. Biomolecules 2018, 8, 4. [Google Scholar] [CrossRef] [PubMed]
- Jonker, A.M.; Löwik, D.W.P.M.; van Hest, J.C.M. Peptide- and protein-based hydrogels. Chem. Mater. 2012, 24, 759–773. [Google Scholar] [CrossRef]
- Martin, C.; Oyen, E.; Van Wanseele, Y.; Haddou, T.B.; Schmidhammer, H.; Andrade, J.; Waddington, L.; Van Eeckhaut, A.; Van Mele, B.; Gardiner, J.; et al. Injectable peptide-based hydrogel formulations for the extended in vivo release of opioids. Mater. Today Chem. 2017, 3, 49–59. [Google Scholar] [CrossRef]
- Hiew, S.H.; Mohanram, H.; Ning, L.; Guo, J.; Sánchez-Ferrer, A.; Shi, X.; Pervushin, K.; Mu, Y.; Mezzenga, R.; Miserez, A. A short peptide hydrogel with high stiffness induced by 3 10 -helices to β-Sheet Transition in Water. Adv. Sci. 2019, 6, 1901173. [Google Scholar] [CrossRef]
- Montero, N.; Alhajj, M.J.; Sierra, M.; Oñate-Garzon, J.; Yarce, C.J.; Salamanca, C.H. Development of polyelectrolyte complex nanoparticles-PECNs loaded with ampicillin by means of polyelectrolyte complexation and ultra-high pressure homogenization (UHPH). Polymers 2020, 12, 1168. [Google Scholar] [CrossRef]
- Doll, T.A.P.F.; Dey, R.; Burkhard, P. Design and optimization of peptide nanoparticles. J. Nanobiotechnol. 2015, 13, 73. [Google Scholar] [CrossRef]
- Jeong, W.; Bu, J.; Kubiatowicz, L.J.; Chen, S.S.; Kim, Y.; Hong, S. Peptide–nanoparticle conjugates: A next generation of diagnostic and therapeutic platforms? Nano Converg. 2018, 5, 38. [Google Scholar] [CrossRef]
- Pudlarz, A.; Szemraj, J. Nanoparticles as carriers of proteins, peptides and other therapeutic molecules. Open Life Sci. 2018, 13, 285–298. [Google Scholar] [CrossRef]
- Recio, C.; Maione, F.; Iqbal, A.J.; Mascolo, N.; De Feo, V. The potential therapeutic application of peptides and peptidomimetics in cardiovascular disease. Front. Pharmacol. 2017, 7. [Google Scholar] [CrossRef]
- Richardson, A.; de Antueno, R.; Duncan, R.; Hoskin, D.W. Intracellular delivery of bovine lactoferricin’s antimicrobial core (RRWQWR) kills T-leukemia cells. Biochem. Biophys. Res. Commun. 2009, 388, 736–741. [Google Scholar] [CrossRef] [PubMed]
- Soman, N.R.; Baldwin, S.L.; Hu, G.; Marsh, J.N.; Lanza, G.M.; Heuser, J.E.; Arbeit, J.M.; Wickline, S.A.; Schlesinger, P.H. Molecularly targeted nanocarriers deliver the cytolytic peptide melittin specifically to tumor cells in mice, reducing tumor growth. J. Clin. Invest. 2009, 119, 2830–2842. [Google Scholar] [CrossRef] [PubMed]
- Hilchie, A.L.; Hoskin, D.W.; Power Coombs, M.R. Anticancer activities of natural and synthetic peptides. In Antimicrobial Peptides: Basics for Clinical Application; Matsuzaki, K., Ed.; Springer: Singapore, 2019; pp. 131–147. [Google Scholar]
- Hansel, W.; Enright, F.; Leuschner, C. Destruction of breast cancers and their metastases by lytic peptide conjugates in vitro and in vivo. Mol. Cell. Endocrinol. 2007, 260, 183–189. [Google Scholar] [CrossRef] [PubMed]
- Berge, G.; Eliassen, L.T.; Camilio, K.A.; Bartnes, K.; Sveinbjørnsson, B.; Rekdal, Ø. Therapeutic vaccination against a murine lymphoma by intratumoral injection of a cationic anticancer peptide. Cancer Immunol. Immunother. 2010, 59, 1285–1294. [Google Scholar] [CrossRef] [PubMed]
- Camilio, K.A.; Rekdal, Ø.; Sveinbjörnsson, B. LTX-315 (OncoporeTM): A short synthetic anticancer peptide and novel immunotherapeutic agent. OncoImmunology 2014, 3, e29181. [Google Scholar] [CrossRef] [PubMed]
- Chiangjong, W.; Chutipongtanate, S.; Hongeng, S. Anticancer peptide: Physicochemical property, functional aspect and trend in clinical application (Review). Int. J. Oncol. 2020, 57, 678–696. [Google Scholar] [CrossRef]
- Ganji, A.; Saeedi, A.M.; Ghazavi, A.; Mosayebi, G. An overview of antimicrobial peptides as anticancer agents. J. Arak Univ. Med. Sci. 2019, 22, 2–15. [Google Scholar] [CrossRef]
- Hanaoka, Y.; Yamaguchi, Y.; Yamamoto, H.; Ishii, M.; Nagase, T.; Kurihara, H.; Akishita, M.; Ouchi, Y. In vitro and in vivo anticancer activity of human β-defensin-3 and its mouse homolog. Anticancer Res. 2016, 36, 5999–6004. [Google Scholar] [CrossRef]
- Chu, H.-L.; Yip, B.-S.; Chen, K.-H.; Yu, H.-Y.; Chih, Y.-H.; Cheng, H.-T.; Chou, Y.-T.; Cheng, J.-W. Novel antimicrobial peptides with high anticancer activity and selectivity. PLoS ONE 2015, 10, e0126390. [Google Scholar] [CrossRef]
- Ghandehari, F.; Behbahani, M.; Pourazar, A.; Noormohammadi, Z. In silico and in vitro studies of cytotoxic activity of different peptides derived from vesicular stomatitis virus G protein. Iran J. Basic Med. Sci. 2015, 18, 47–52. [Google Scholar]
- Ma, R.; Wong, S.W.; Ge, L.; Shaw, C.; Siu, S.W.I.; Kwok, H.F. In vitro and MD simulation study to explore physicochemical parameters for antibacterial peptide to become potent anticancer peptide. Mol. Ther. Oncolyt. 2020, 16, 7–19. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Hao, X.; Liu, D.; Huang, Y.; Chen, Y. In vitro characterization of the rapid cytotoxicity of anticancer peptide HPRP-A2 through membrane destruction and intracellular mechanism against gastric cancer cell lines. PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Militão, G.C.G.; Dantas, I.N.F.; Ferreira, P.M.P.; Alves, A.P.N.N.; Chaves, D.C.; Monte, F.J.Q.; Pessoa, C.; Odorico de Moraes, M.; Costa-Lotufo, L.V. In vitro and in vivo anticancer properties of cucurbitacin isolated from Cayaponia racemose. Pharm. Biol. 2012, 50, 1479–1487. [Google Scholar] [CrossRef] [PubMed]
- Xu, Q.; Huang, K.-C.; Tendyke, K.; Marsh, J.; Liu, J.; Qiu, D.; Littlefield, B.A.; Nomoto, K.; Atasoylu, O.; Risatti, C.A.; et al. In vitro and in vivo anticancer activity of (+)-spongistatin. Anticancer Res. 2011, 31, 7. [Google Scholar]
- Yang, S.; Meng, J.; Yang, Y.; Liu, H.; Wang, C.; Liu, J.; Zhang, Y.; Wang, C.; Xu, H. A HSP60-targeting peptide for cell apoptosis imaging. Oncogenesis 2016, 5, e201. [Google Scholar] [CrossRef] [PubMed]
- Kim, I.-W.; Lee, J.H.; Kwon, Y.-N.; Yun, E.-Y.; Nam, S.-H.; Ahn, M.-Y.; Kang, D.-C.; Hwang, J.S. Anticancer activity of a synthetic peptide derived from harmoniasin, an antibacterial peptide from the ladybug Harmonia axyridis. Int. J. Oncol. 2013, 43, 622–628. [Google Scholar] [CrossRef] [PubMed]
- Prachayasittikul, V.; Worachartcheewan, A.; Shoombuatong, W.; Prachayasittikul, V.; Nantasenamat, C. Classification of P-glycoprotein-interacting compounds using machine learning methods. EXCLI J. 2015, 14, 1611–2156. [Google Scholar] [CrossRef]
- Bharath, E.N.; Manjula, S.N.; Vijaychand, A. In silico drug design tool for overcoming the innovation deficit in the drug discovery process. Chemestry 2011, 3, 5. [Google Scholar]
- Mustata, G.; Muftuoglu, Y. Computational strategies in cancer drug discovery. In Advances in Cancer Management; Mohan, R., Ed.; InTech: London, UK, 2012. [Google Scholar]
- Leelananda, S.P.; Lindert, S. Computational methods in drug discovery. Beilstein J. Org. Chem. 2016, 12, 2694–2718. [Google Scholar] [CrossRef]
- Kumar, S.; Li, H. In silico design of anticancer peptides. In Proteomics for Drug Discovery; Lazar, I.M., Kontoyianni, M., Lazar, A.C., Eds.; Springer: New York, NY, USA, 2017; Volume 1647, pp. 245–254. [Google Scholar]
- Brockmeier, E.K.; Hodges, G.; Hutchinson, T.H.; Butler, E.; Hecker, M.; Tollefsen, K.E.; Garcia-Reyero, N.; Kille, P.; Collette, T.W.; Cossins, A.; et al. The role of omics in the application of adverse outcome pathways for chemical risk assessment. Toxicol. Sci. 2017, 158, 252–262. [Google Scholar] [CrossRef]
- Gottlieb, A.; Weingart, U.; Horn, D. Data mining of protein families using common peptides. Nat. Prec. 2008. [Google Scholar] [CrossRef]
- Bernot, A. Genome Transcriptome and Proteome Analysis. Brief. Bioinform. 2004, 19, 286–302. [Google Scholar]
- Goldenberg, S.L.; Nir, G.; Salcudean, S.E. A new era: Artificial intelligence and machine learning in prostate cancer. Nat. Rev. Urol. 2019, 16, 391–403. [Google Scholar] [CrossRef]
- Wei, L.; Zhou, C.; Chen, H.; Song, J.; Su, R. ACPred-FL: A sequence-based predictor using effective feature representation to improve the prediction of anti-cancer peptides. Bioinformatics 2018. [Google Scholar] [CrossRef]
- Porto, W.F.; Silva, O.N.; Franco, O.L. Prediction and rational design of antimicrobial peptides. In Protein Structure; Faraggi, E., Ed.; InTech: London, UK, 2012. [Google Scholar]
- Giangaspero, A.; Sandri, L.; Tossi, A. Amphipathic α helical antimicrobial peptides: A systematic study of the effects of structural and physical properties on biological activity. Eur. J. Biochem. 2001, 268, 5589–5600. [Google Scholar] [CrossRef] [PubMed]
- Kumariya, R.; Sood, S.K.; Rajput, Y.S.; Saini, N.; Garsa, A.K. Increased membrane surface positive charge and altered membrane fluidity leads to cationic antimicrobial peptide resistance in Enterococcus faecalis. Biochim. Biophys. Acta (BBA) Biomembr. 2015, 1848, 1367–1375. [Google Scholar] [CrossRef]
- Nyström, L.; Malmsten, M. Membrane interactions and cell selectivity of amphiphilic anticancer peptides. Curr. Opin. Colloid Interface Sci. 2018, 38, 1–17. [Google Scholar] [CrossRef]
- Miyazaki, Y.; Aoki, M.; Yano, Y.; Matsuzaki, K. Interaction of antimicrobial peptide magainin 2 with gangliosides as a target for human cell binding. Biochemistry 2012, 51, 10229–10235. [Google Scholar] [CrossRef]
- Risso, A.; Zanetti, M.; Gennaro, R. Cytotoxicity and apoptosis mediated by two peptides of innate immunity. Cell. Immunol. 1998, 189, 107–115. [Google Scholar] [CrossRef]
- Ntwasa, M.; Goto, A.; Kurata, S. Coleopteran antimicrobial peptides: Prospects for clinical applications. Int. J. Microbiol. 2012, 2012, 1–8. [Google Scholar] [CrossRef]
- Zelezetsky, I.; Tossi, A. Alpha-helical antimicrobial peptides—Using a sequence template to guide structure–activity relationship studies. Biochim. Biophys. Acta (BBA) Biomembr. 2006, 1758, 1436–1449. [Google Scholar] [CrossRef] [PubMed]
- Feder, R.; Dagan, A.; Mor, A. Structure-activity relationship study of antimicrobial dermaseptin S4 showing the consequences of peptide oligomerization on selective cytotoxicity. J. Biol. Chem. 2000, 275, 4230–4238. [Google Scholar] [CrossRef] [PubMed]
- Yeaman, M.R.; Yount, N.Y. Mechanisms of antimicrobial peptide action and resistance. Pharmacol. Rev. 2003, 55, 27–55. [Google Scholar] [CrossRef]
- Ellerby, H.M.; Bredesen, D.E.; Fujimura, S.; John, V. Hunter−killer peptide (HKP) for targeted therapy. J. Med. Chem. 2008, 51, 5887–5892. [Google Scholar] [CrossRef]
- Rodríguez Plaza, J.G.; Villalón Rojas, A.; Herrera, S.; Garza-Ramos, G.; Torres Larios, A.; Amero, C.; Zarraga Granados, G.; Gutiérrez Aguilar, M.; Lara Ortiz, M.T.; Polanco Gonzalez, C.; et al. Moonlighting peptides with emerging function. PLoS ONE 2012, 7. [Google Scholar] [CrossRef]
- Almaaytah, A.; Alaraj, M. The design and anticancer activity of a citropin1.1 hybrid peptide with selective activity against highly invasive metastatic cell lines. Int. J. Res. Pharm. Sci. 2019, 10, 3544–3553. [Google Scholar] [CrossRef]
- Mor, A.; Nguyen, V.H.; Delfour, A.; Migliore-Samour, D.; Nicolas, P.; Van Huong, N. Isolation, amino acid sequence and synthesis of dermaseptin, a novel antimicrobial peptide of amphibian skin. Biochem. 1991, 30, 8824–8830. [Google Scholar] [CrossRef] [PubMed]
- Brogden, K.A. Antimicrobial peptides: Pore formers or metabolic inhibitors in bacteria? Nat. Rev. Microbiol. 2005, 3, 238–250. [Google Scholar] [CrossRef] [PubMed]
- Pasupuleti, M.; Schmidtchen, A.; Malmsten, M. Antimicrobial peptides: Key components of the innate immune system. Crit. Rev. Biotechnol. 2012, 32, 143–171. [Google Scholar] [CrossRef]
- Acosta, Y.A.S. Diseño de Péptidos Antimicrobianos Derivados de Dermaseptina S4. Master’s Thesis, Universidad Nacional de Colombia, Medellín, Colombia, 2016; p. 94. [Google Scholar]
- Yang, Q.-Z.; Wang, C.; Lang, L.; Zhou, Y.; Wang, H.; Shang, D.-J. Design of potent, non-toxic anticancer peptides based on the structure of the antimicrobial peptide, temporin-1CEa. Arch. Pharm. Res. 2013, 36, 1302–1310. [Google Scholar] [CrossRef]
- Dennison, S.R.; Harris, F.; Bhatt, T.; Singh, J.; Phoenix, D.A. A theoretical analysis of secondary structural characteristics of anticancer peptides. Mol. Cell. Biochem. 2010, 333, 129–135. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.-B.; Wang, X.-F.; Wang, H.-Y.; Liu, Y.; Chen, Y. Studies on mechanism of action of anticancer peptides by modulation of hydrophobicity within a defined structural framework. Mol. Cancer Ther. 2011, 10. [Google Scholar] [CrossRef]
- Huang, Y.-B.; He, L.-Y.; Jiang, H.-Y.; Chen, Y.-X. Role of helicity on the anticancer mechanism of action of cationic-helical peptides. Int. J. Mol. Sci. 2012, 13, 6849–6862. [Google Scholar] [CrossRef]
- Sinthuvanich, C.; Veiga, A.S.; Gupta, K.; Gaspar, D.; Blumenthal, R.; Schneider, J.P. Anticancer β-hairpin peptides: Membrane-induced folding triggers activity. J. Am. Chem. Soc. 2012, 134, 6210–6217. [Google Scholar] [CrossRef]
- Hu, C.; Chen, X.; Zhao, W. Design and modification of anticancer peptides. Drug Des. 2016, 5. [Google Scholar] [CrossRef]
- Cantor, S.; Vargas, L.; Rojas, O.; Yarce, A.; Salamanca, C.; Oñate-Garzón, J. Evaluation of the antimicrobial activity of cationic peptides loaded in surface-modified nanoliposomes against foodborne bacteria. Int. J. Mol. Sci. 2019, 20, 680. [Google Scholar] [CrossRef]
- Martins, S.; Sarmento, B.; Ferreira, D.C.; Souto, E.B. Lipid-based colloidal carriers for peptide and protein delivery—Liposomes versus lipid nanoparticles. Int. J. Nanomed. 2007, 2, 595–607. [Google Scholar]
- Arenas, I.; Villegas, E.; Walls, O.; Barrios, H.; Rodríguez, R.; Corzo, G. Antimicrobial activity and stability of short and long based arachnid synthetic peptides in the presence of commercial antibiotics. Molecules 2016, 21, 225. [Google Scholar] [CrossRef] [PubMed]
- Park, I.Y.; Cho, J.H.; Kim, K.S.; Kim, Y.-B.; Kim, M.S.; Kim, S.C. Helix stability confers salt resistance upon helical antimicrobial peptides. J. Boil. Chem. 2004, 279, 13896–13901. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Fang, Y.; Wu, J. Flexibility is a mechanical determinant of antimicrobial activity for amphipathic cationic α-helical antimicrobial peptides. Biochim. Biophys. Acta (BBA) Biomembr. 2013, 1828, 2479–2486. [Google Scholar] [CrossRef]
- Zou, R.; Zhu, X.; Tu, Y.; Wu, J.; Landry, M.P. Activity of antimicrobial peptide aggregates decreases with increased cell membrane embedding free energy cost. Biochemistry 2018, 57, 2606–2610. [Google Scholar] [CrossRef]
- Hao, X.; Yan, Q.; Zhao, J.; Wang, W.; Huang, Y.-B.; Chen, Y. TAT modification of alpha-helical anticancer peptides to improve specificity and efficacy. PLoS ONE 2015, 10, e0138911. [Google Scholar] [CrossRef] [PubMed]
- Lim, K.J.; Sung, B.H.; Shin, J.R.; Lee, Y.W.; Kim, D.J.; Yang, K.S.; Kim, S.C. A cancer specific cell-penetrating peptide, BR2, for the efficient delivery of an scFv into cancer cells. PLoS ONE 2013, 8, e66084. [Google Scholar] [CrossRef]
- Grissenberger, S.; Riedl, S.; Rinner, B.; Leber, R.; Zweytick, D. Design of human lactoferricin derived antitumor peptides-activity and specificity against malignant melanoma in 2D and 3D model studies. Biochim. Biophys. Acta (BBA) Biomembr. 2020, 1862, 183264. [Google Scholar] [CrossRef] [PubMed]
- Karbalaeemohammad, S.; Naderi-Manesh, H. Two novel anticancer peptides from aurein1.2. Int. J. Pept. Res. Ther. 2011, 17, 159–164. [Google Scholar] [CrossRef]
- Gupta, U.K.; Mahanta, S.; Paul, S. In silico design of small peptide-based Hsp90 inhibitor: A novel anticancer agent. Med. Hypoth. 2013, 81, 853–861. [Google Scholar] [CrossRef]
- Plescia, J.; Salz, W.; Xia, F.; Pennati, M.; Zaffaroni, N.; Daidone, M.G.; Meli, M.; Dohi, T.; Fortugno, P.; Nefedova, Y.; et al. Rational design of shepherdin, a novel anticancer agent. Cancer Cell 2005, 7, 457–468. [Google Scholar] [CrossRef]
- E-Kobon, T.; Thongararm, P.; Roytrakul, S.; Meesuk, L.; Chumnanpuen, P. Prediction of anticancer peptides against MCF-7 breast cancer cells from the peptidomes of Achatina fulica mucus fractions. Computat. Struct. Biotechnol. J. 2016, 14, 49–57. [Google Scholar] [CrossRef]
- Li, B.; Lyu, P.; Xi, X.; Ge, L.; Mahadevappa, R.; Shaw, C.; Kwok, H.F. Triggering of cancer cell cycle arrest by a novel scorpion venom-derived peptide—Gonearrestide. J. Cell. Mol. Med. 2018, 22, 4460–4473. [Google Scholar] [CrossRef]
- Midoux, P.; Kichler, A.; Boutin, V.; Maurizot, J.C.; Monsigny, M. Membrane permeabilization and efficient gene transfer by a peptide containing several histidines. Bioconjug. Chem. 1998, 9, 260–267. [Google Scholar] [CrossRef]
- Dai, Y.; Cai, X.; Shi, W.; Bi, X.; Su, X.; Pan, M.; Li, H.; Lin, H.; Huang, W.; Qian, H. Pro-apoptotic cationic host defense peptides rich in lysine or arginine to reverse drug resistance by disrupting tumor cell membrane. Amino Acids 2017, 49, 1601–1610. [Google Scholar] [CrossRef] [PubMed]
- Strömstedt, A.A.; Ringstad, L.; Schmidtchen, A.; Malmsten, M. Interaction between amphiphilic peptides and phospholipid membranes. Curr. Opin. Colloid Interface Sci. 2010, 15, 467–478. [Google Scholar] [CrossRef]
- Yamaguchi, Y.; Yamamoto, K.; Sato, Y.; Inoue, S.; Morinaga, T.; Hirano, E. Combination of aspartic acid and glutamic acid inhibits tumor cell proliferation. Biomed. Res. 2016, 37, 153–159. [Google Scholar] [CrossRef]
- Shamova, O.; Orlov, D.; Stegemann, C.; Czihal, P.; Hoffmann, R.; Brogden, K.; Kolodkin, N.; Sakuta, G.; Tossi, A.; Sahl, H.-G.; et al. ChBac3.4: A novel proline-rich antimicrobial peptide from goat leukocytes. Int. J. Pept. Res. Ther. 2009, 15, 31–42. [Google Scholar] [CrossRef]
- Ahmaditaba, M.A.; Tehrani, M.H.H.; Zarghi, A.; Daraei, B. Design, synthesis and biological evaluation of novel peptide-like analogues as selective COX-2 inhibitors. IJPR 2018, 17, 87–92. [Google Scholar]
- Radicioni, G.; Stringaro, A.; Molinari, A.; Nocca, G.; Longhi, R.; Pirolli, D.; Scarano, E.; Iavarone, F.; Manconi, B.; Cabras, T.; et al. Characterization of the cell penetrating properties of a human salivary proline-rich peptide. Biochim. Biophys. Acta (BBA) Biomembr. 2015, 1848, 2868–2877. [Google Scholar] [CrossRef]
- Bhunia, D.; Mondal, P.; Das, G.; Saha, A.; Sengupta, P.; Jana, J.; Mohapatra, S.; Chatterjee, S.; Ghosh, S. Spatial position regulates power of tryptophan: Discovery of a major-groove-specific nuclear-localizing, cell-penetrating tetrapeptide. J. Am. Chem. Soc. 2018, 140, 1697–1714. [Google Scholar] [CrossRef]
- Harris, F.; Dennison, S.R.; Singh, J.; Phoenix, D.A. On the selectivity and efficacy of defense peptides with respect to cancer cells: On the selectivity and efficacy of defense. Med. Res. Rev. 2013, 33, 190–234. [Google Scholar] [CrossRef]
- Conlon, J.M.; Mechkarska, M.; Prajeep, M.; Arafat, K.; Zaric, M.; Lukic, M.L.; Attoub, S. Transformation of the naturally occurring frog skin peptide, alyteserin-2a into a potent, non-toxic anti-cancer agent. Amino Acids 2013, 44, 715–723. [Google Scholar] [CrossRef]
- Akbar, S.; Rahman, A.U.; Hayat, M.; Sohail, M. cACP: Classifying anticancer peptides using discriminative intelligent model via Chou’s 5-step rules and general pseudo components. Chemometr. Intell. Lab. Syst. 2020, 196, 103912. [Google Scholar] [CrossRef]
- Agrawal, P.; Bhagat, D.; Mahalwal, M.; Sharma, N.; Raghava, G.P.S. AntiCP 2.0: An updated model for predicting anticancer peptides. Bioinformatics 2020. [Google Scholar] [CrossRef] [PubMed]
- Langlois, R.E.; Lu, H. Machine learning for protein structure and function prediction. In Annual Reports in Computational Chemistry; Elsevier: Amsterdam, The Netherlands, 2008; Volume 4, pp. 41–66. [Google Scholar]
- Yang, Z.R. Peptide bioinformatics—Peptide classification using peptide machines. In Artificial Neural Networks; Livingstone, D.J., Ed.; Humana Press: Totowa, NJ, USA, 2008; Volume 458, pp. 155–179. [Google Scholar]
- Tyagi, A.; Tuknait, A.; Anand, P.; Gupta, S.; Sharma, M.; Mathur, D.; Joshi, A.; Singh, S.; Gautam, A.; Raghava, G.P.S. CancerPPD: A database of anticancer peptides and proteins. Nucl. Acids Res. 2015, 43, D837–D843. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Li, X.; Wang, Z. APD3: The antimicrobial peptide database as a tool for research and education. Nucl. Acids Res. 2016, 44, D1087–D1093. [Google Scholar] [CrossRef] [PubMed]
- Kang, X.; Dong, F.; Shi, C.; Liu, S.; Sun, J.; Chen, J.; Li, H.; Xu, H.; Lao, X.; Zheng, H. DRAMP 2.0, an updated data repository of antimicrobial peptides. Sci. Data 2019, 6, 148. [Google Scholar] [CrossRef]
- Kapoor, P.; Singh, H.; Gautam, A.; Chaudhary, K.; Kumar, R.; Raghava, G.P.S. TumorHoPe: A database of tumor homing peptides. PLoS ONE 2012, 7, e35187. [Google Scholar] [CrossRef]
- Manavalan, B.; Subramaniyam, S.; Shin, T.H.; Kim, M.O.; Lee, G. Machine-learning-based prediction of cell-penetrating peptides and their uptake efficiency with improved accuracy. J. Proteome Res. 2018, 17, 2715–2726. [Google Scholar] [CrossRef]
- Berrar, D. Cross-validation. In Encyclopedia of Bioinformatics and Computational Biology; Elsevier: Amsterdam, The Netherlands, 2019; pp. 542–545. [Google Scholar]
- Ferdinandy, B.; Gerencsér, L.; Corrieri, L.; Perez, P.; Újváry, D.; Csizmadia, G.; Miklósi, Á. Challenges of machine learning model validation using correlated behaviour data: Evaluation of cross-validation strategies and accuracy measures. PLoS ONE 2020, 15, e0236092. [Google Scholar] [CrossRef]
- Li, F.-M.; Wang, X.-Q. Identifying anticancer peptides by using improved hybrid compositions. Sci. Rep. 2016, 6, 33910. [Google Scholar] [CrossRef]
- Manavalan, B.; Basith, S.; Shin, T.H.; Choi, S.; Kim, M.O.; Lee, G. MLACP: Machine-learning-based prediction of anticancer peptides. Oncotarget 2017, 8, 77121–77136. [Google Scholar] [CrossRef]
- Gautam, A.; Singh, H.; Tyagi, A.; Chaudhary, K.; Kumar, R.; Kapoor, P.; Raghava, G.P.S. CPPsite: A curated database of cell penetrating peptides. Database 2012, 2012, bas015. [Google Scholar] [CrossRef]
- Gautam, A.; Chaudhary, K.; Kumar, R.; Raghava, G.P.S. Computer-aided virtual screening and designing of cell-penetrating peptides. In Cell-Penetrating Peptides; Langel, Ü., Ed.; Springer: New York, NY, USA, 2015; Volume 1324, pp. 59–69. [Google Scholar]
- Thomas, S.; Karnik, S.; Barai, R.S.; Jayaraman, V.K.; Idicula-Thomas, S. CAMP: A useful resource for research on antimicrobial peptides. Nucl. Acids Res. 2010, 38 (Suppl. S1), D774–D780. [Google Scholar] [CrossRef] [PubMed]
- Waghu, F.H.; Gopi, L.; Barai, R.S.; Ramteke, P.; Nizami, B.; Idicula-Thomas, S. CAMP: Collection of sequences and structures of antimicrobial peptides. Nucl. Acids Res. 2014, 42, D1154–D1158. [Google Scholar] [CrossRef] [PubMed]
- Asgari, E.; Poerner, N.; McHardy, A.C.; Mofrad, M.R.K. DeepPrime2Sec: Deep learning for protein secondary structure prediction from the primary sequences. Bioinformatics 2019. [Google Scholar] [CrossRef]
- Wu, C.; Gao, R.; Zhang, Y.; De Marinis, Y. PTPD: Predicting therapeutic peptides by deep learning and word2vec. BMC Bioinform. 2019, 20, 456. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, A.; Kapoor, P.; Kumar, R.; Chaudhary, K.; Gautam, A.; Raghava, G.P.S. In silico models for designing and discovering Novel anticancer peptides. Sci. Rep. 2013, 3, 2984. [Google Scholar] [CrossRef]
- Vijayakumar, S.; Ptv, L. ACPP: A web server for prediction and design of anti-cancer peptides. Int. J. Pept. Res. Ther. 2015, 21, 99–106. [Google Scholar] [CrossRef]
- Hajisharifi, Z.; Piryaiee, M.; Mohammad Beigi, M.; Behbahani, M.; Mohabatkar, H. Predicting anticancer peptides with Chou′s pseudo amino acid composition and investigating their mutagenicity via Ames test. J. Theor. Biol. 2014, 341, 34–40. [Google Scholar] [CrossRef]
- Chen, W.; Ding, H.; Feng, P.; Lin, H.; Chou, K.-C. iACP: A sequence-based tool for identifying anticancer peptides. Oncotarget 2016, 7, 16895–16909. [Google Scholar] [CrossRef]
- Chou, K.-C. Some remarks on protein attribute prediction and pseudo amino acid composition. J. Theoret. Biol. 2011, 273, 236–247. [Google Scholar] [CrossRef]
- Schaduangrat, N.; Nantasenamat, C.; Prachayasittikul, V.; Shoombuatong, W. ACPred: A computational tool for the prediction and analysis of anticancer peptides. Molecules 2019, 24, 1973. [Google Scholar] [CrossRef]
- Yi, H.-C.; You, Z.-H.; Zhou, X.; Cheng, L.; Li, X.; Jiang, T.-H.; Chen, Z.-H. ACP-DL: A deep learning long short-term memory model to predict anticancer peptides using high-efficiency feature representation. Mol. Ther. Nucl. Acids 2019, 17, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Ge, R.; Feng, G.; Jing, X.; Zhang, R.; Wang, P.; Wu, Q. EnACP: An ensemble learning model for identification of anticancer peptides. Front. Genet. 2020, 11, 760. [Google Scholar] [CrossRef] [PubMed]
| Anticancer Peptides | Examined Cell Lines |
|---|---|
| Magainin II | Bladder cancer cells |
| Buforin IIb | Cervical carcinoma cells |
| BR2 | Cervical carcinoma cells |
| PNC-2 and PNC-7 | Pancreatic cancer cells |
| RGD-PEG-Suc-PD0325901 | Melanoma A375 cells |
| p16 | Pancreatic cancer cells |
| Defensin | Lung Carcinoma cells |
| LL-37 | Ovarian Carcinoma, Breast Cancer cells |
| Cecropin A y B | Bladder cancer cells |
| Bac-7-ELP-p21 | Ovarian Carcinoma cells |
| NRC-3 and NRC-7 | Breast Cancer cells |
| Test | Information |
|---|---|
| Antimicrobial activity | Used to find the Minimum Inhibitory Concentration and the Bactericidal Concentration that kills 99.9% of the bacterial population. At present, microdilution is frequently used in 96-well plates and the reading can be done visually or through the creation of a curve relating the percentage of inhibition by the peptide and the concentration. |
| Hemolytic activity | Used to find the hemolytic concentration 50, a useful parameter to determine the degree of cytotoxicity that the peptide can cause in eukaryotic cells. |
| Cytotoxicity test on tumor cells | This test is usually performed by screening with (3-(4,5-dimethylthiazolyl-2)-2,5-diphenyltetrazolium bromide) (MTT), this colorimetric test allows the evaluation of the cellular metabolic activity by reducing the MTT to its insoluble form formed by oxidoreductase enzymes, changing from yellow to purple with the appearance of the formazan in living cells. |
| Live imaging | For this test, the cell nucleus is marked with 4′,6-diamidine-2-phenylindol (DAPI) and the peptide with another marker such as Fluorescein isothiocyanate (FITC) and observed by fluorescence microscopy. By means of this test it is possible to have a vision of the mechanism of damage of the anticarcinogenic peptide. |
| Analysis of morphological changes by H/E staining | The cells are fixed with methanol for 1 min and stained with H/E to visualize the morphology of the cells. |
| Pgp sensitivity assay | Pgp is a drug transporter that plays important roles in multidrug resistance and drug pharmacokinetics. The inhibition of Pgp has become a notable strategy for combating multidrug-resistant cancers. |
| Western blotting | It is used to determine if there is caspase activation or not and also to determine whether the peptide damage was caused by necrosis or apoptosis. To determine apoptosis, antibody against caspase 3 is incubated and its expression is displayed every few minutes, 1 h and 24 h. |
| DNA fragmentation test | DNA fragmentation is characteristic of apoptosis. After the cells are exposed to the peptide, the DNA is extracted and placed in agarose gel in order to visualize the DNA fragmentation. |
| TUNEL assay | It is an assay used for the detection of DNA fragments due to the process of apoptosis. This assay consists of the ability of terminal deoxynucleotidyl transferase to mark blunt ends of double-stranded DNA breaks independently of a template. |
| Anti-angiogenesis assay | Anticancer peptides are recognized for stopping angiogenesis caused by tumor cells. In this assay, venous endothelial cells from the human umbilical cord are used and confronted with the anticancer peptide. Then it is observed if there is a formation of blood connections or not compared with the control, expecting an inhibition of these by the anticarcinogenic peptide. |
| Flow cytometry | This test can determine whether or not there is cell or mitochondrial membrane damage, DNA fragmentation and cell cycle alteration. It also allows differentiation between necrotic, apoptotic or healthy cells. |
| Release of lactate dehydrogenase (LDH) | LDH is a cytoplasmic enzyme present in all cells and released into the cell space when the membrane ruptures. The assay uses the supernatant of the cells that were treated with the peptide by measuring the absorbance at 450 nm in microplates and relating the peptide concentration to the percentage of LDH release |
| Reactive oxygen intermediates (ROS) assay | This assay is used to detect the generation of ROS, whose generation induces damages in DNA, proteins, and membrane lipids. Kits such as the ROS assay kit (BestBio, Shanghai Co., China) are used which have a fluorescent probe that allows the intensity of the fluorescence to be detected by flow cytometry and directly correlated to an increase in ROS concentration |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liscano, Y.; Oñate-Garzón, J.; Delgado, J.P. Peptides with Dual Antimicrobial–Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides. Molecules 2020, 25, 4245. https://doi.org/10.3390/molecules25184245
Liscano Y, Oñate-Garzón J, Delgado JP. Peptides with Dual Antimicrobial–Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides. Molecules. 2020; 25(18):4245. https://doi.org/10.3390/molecules25184245
Chicago/Turabian StyleLiscano, Yamil, Jose Oñate-Garzón, and Jean Paul Delgado. 2020. "Peptides with Dual Antimicrobial–Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides" Molecules 25, no. 18: 4245. https://doi.org/10.3390/molecules25184245
APA StyleLiscano, Y., Oñate-Garzón, J., & Delgado, J. P. (2020). Peptides with Dual Antimicrobial–Anticancer Activity: Strategies to Overcome Peptide Limitations and Rational Design of Anticancer Peptides. Molecules, 25(18), 4245. https://doi.org/10.3390/molecules25184245







