Analytical and Clinical Assessment of a Portable, Isothermal Recombinase Polymerase Amplification (RPA) Assay for the Molecular Diagnosis of Urogenital Schistosomiasis
Abstract
1. Introduction
2. Results
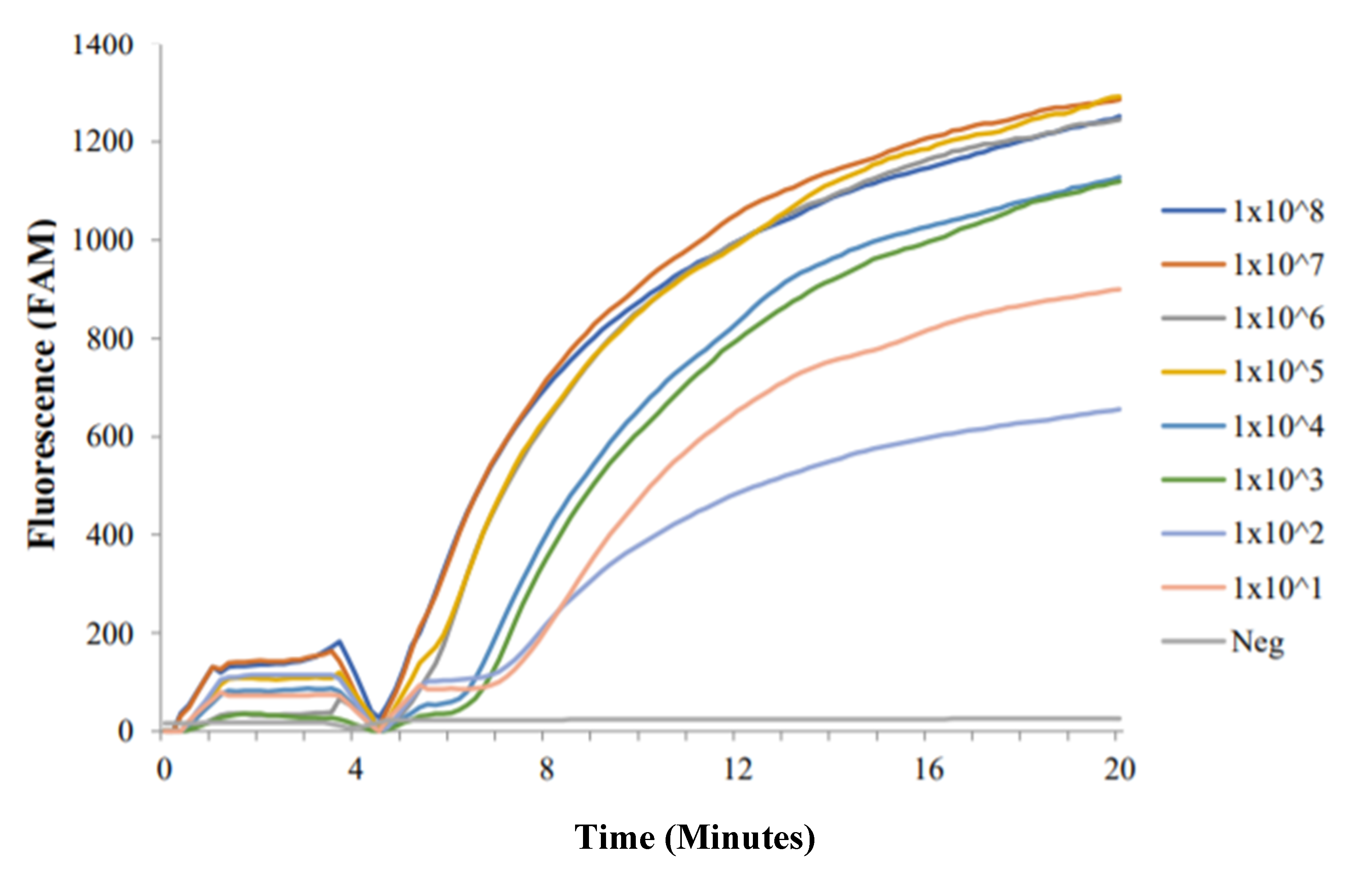
2.1. RT-ShDra1-RPA Analytical Sensitivity
2.1.1. Synthesised ShDra1 Copies
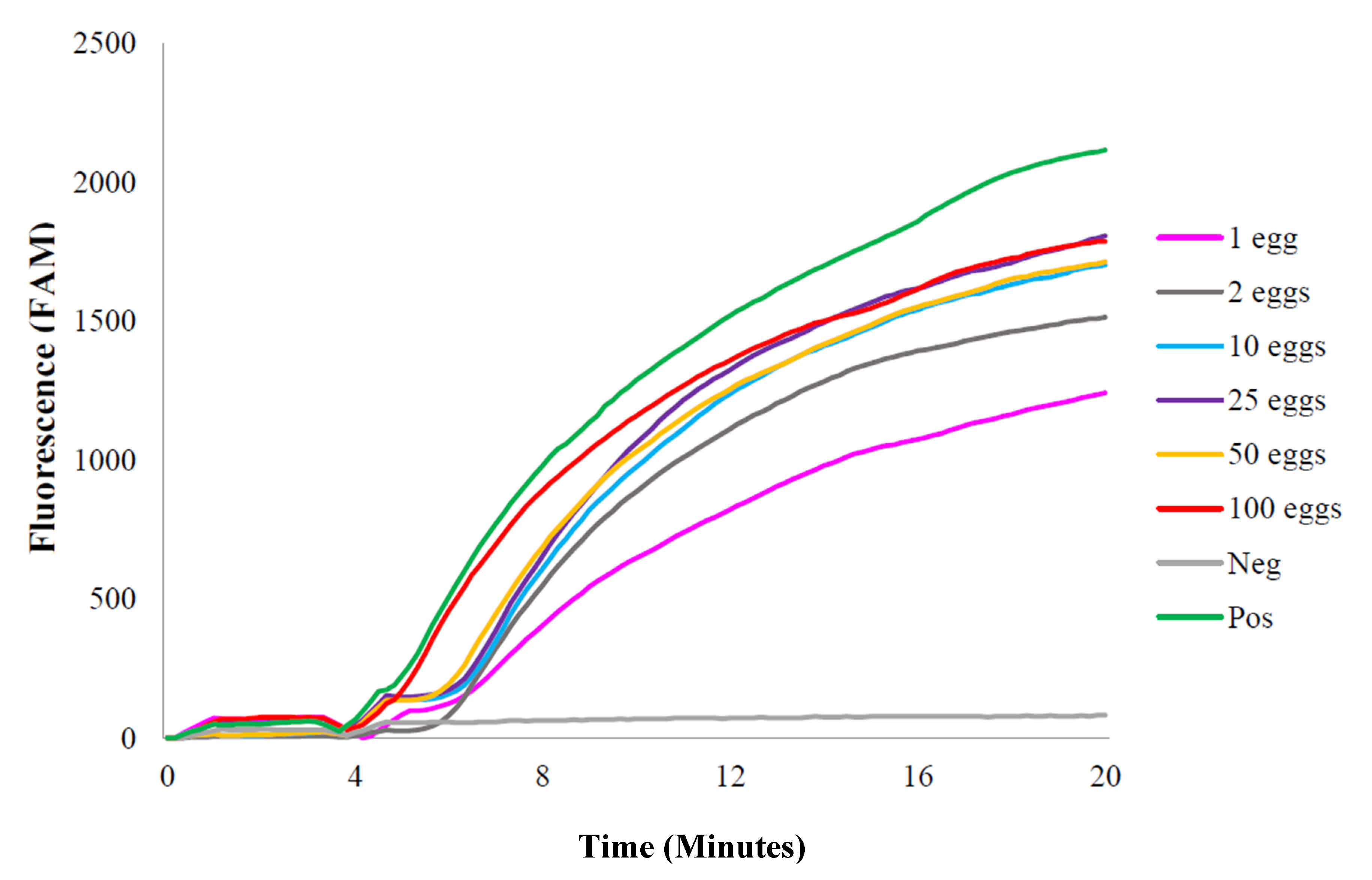
2.1.2. S. haematobium Eggs: Laboratory-Spiked Samples
2.2. RT-ShDra1-RPA Clinical Performance
3. Discussion
Study Limitations and Future Work
4. Materials and Methods
4.1. RPA Reactions
4.2. RT-ShDra1-RPA Analytical Sensitivity
4.2.1. Synthesised ShDra1 Copies
4.2.2. S. haematobium Eggs: Laboratory-Prepared Samples
4.3. RT-ShDra1-RPA Clinical Performance
4.3.1. Study Area and Urine Samples
4.3.2. DNA Isolation and RT-ShDra1-RPA Use with Clinical Urine Sample Extracts
4.4. Statistical Analysis
4.5. Ethical Approval and Consent to Participate
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| bp | Base pairs |
| CAA | Circulating anodic antigen |
| CCA | Circulating cathodic antigen |
| FAM | 6-carboxyflourescein |
| fg | Femtogram |
| gDNA | Genomic DNA |
| LoD | Limit of detection |
| LSHTM | London School of Hygiene and Tropical Medicine |
| MDA | Mass drug administration |
| NHM | Natural History Museum, London, UK |
| NPV | Negative predictive value |
| NTD | Neglected tropical disease |
| PC | Preventative chemotherapy |
| PCR | Polymerase chain reaction |
| PHL-IdC | Public Health Laboratory Ivo de Carneri |
| PPV | Positive predictive value |
| qPCR | Quantitative/real-time PCR |
| RPA | Recombinase polymerase amplification |
| Sh | Schistosoma haematobium |
| STI | Sexually transmitted infection |
| Th2 | T-helper type 2 |
References
- Colley, D.G.; Bustinduy, A.L.; Secor, W.E.; King, C.H. Human schistosomiasis. Lancet 2014, 383, 2253–2264. [Google Scholar] [CrossRef]
- Tchuem, T.L.A.; Rollinson, D.; Stothard, J.R.; Molyneux, D. Moving from control to elimination of schistosomiasis in sub-Saharan Africa: Time to change and adapt strategies. Infect. Dis. Poverty 2017, 6, 1–14. [Google Scholar] [CrossRef]
- Brindley, P.J.; Hotez, P.J. Break Out: Urogenital schistosomiasis and Schistosoma haematobium infection in the post-genomic era. PLoS Negl. Trop. Dis. 2013, 7, 2–4. [Google Scholar] [CrossRef] [PubMed]
- Hotez, P.J.; Kamath, A. Neglected tropical diseases in sub-Saharan Africa: Review of their prevalence, distribution, and disease burden. PLoS Negl. Trop. Dis. 2009, 3, 2–11. [Google Scholar] [CrossRef] [PubMed]
- Burke, M.L.; Jones, M.K.; Gobert, G.N.; Li, Y.S.; Ellis, M.K.; McManus, D.P. Immunopathogenesis of human schistosomiasis. Parasite Immunol. 2009, 31, 163–176. [Google Scholar] [CrossRef] [PubMed]
- Honeycutt, J.; Hammam, O.; Fu, C.L.; Hsieh, M.H. Controversies and challenges in research on urogenital schistosomiasis-associated bladder cancer. Trends Parasitol. 2014, 30, 324–332. [Google Scholar] [CrossRef]
- Kjetland, E.F.; Leutscher, P.D.C.; Ndhlovu, P.D. A review of female genital schistosomiasis. Trends Parasitol. 2012, 28, 58–65. [Google Scholar] [CrossRef]
- Christinet, V.; Lazdins-Helds, J.K.; Stothard, J.R.; Reinhard-Rupp, J. Female genital schistosomiasis (FGS): From case reports to a call for concerted action against this neglected gynaecological disease. Int. J. Parasitol. 2016, 46, 395–404. [Google Scholar] [CrossRef]
- Leutscher, P.D.C.; Ramarokoto, C.; Hoffmann, S.; Jensen, J.S.; Ramaniraka, V.; Randrianasolo, B.; Raharisolo, C.; Migliani, R.; Christensen, N. Coexistence of urogenital schistosomiasis and sexually transmitted infection in women and men living in an area where Schistosoma haematobium is endemic. Clin. Infect. Dis. 2008, 47, 775–782. [Google Scholar] [CrossRef]
- Bustinduy, A.; King, C.; Scott, J.; Appleton, S.; Sousa-Figueiredo, J.C.; Betson, M.; Stothard, J.R. HIV and schistosomiasis co-infection in African children. Lancet Infect. Dis. 2014, 14, 640–649. [Google Scholar] [CrossRef]
- Mbabazi, P.S.; Andan, O.; Fitzgerald, D.W.; Chitsulo, L.; Engels, D.; Downs, J.A. Examining the relationship between urogenital schistosomiasis and HIV infection. PLoS Negl. Trop. Dis. 2011, 5, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Fenwick, A.; Savioli, L.; Engels, D.; Bergquist, N.R.; Todd, M.H. Drugs for the control of parasitic diseases: Current status and development in schistosomiasis. Trends Parasitol. 2003, 19, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Doenhoff, M.J.; Hagan, P.; Cioli, D.; Southgate, V.; Pica-Mattoccia, L.; Botros, S.; Coles, G.; Tchuem, T.L.A.; Mbaye, A.; Engels, D. Praziquantel: Its use in control of schistosomiasis in sub-Saharan Africa and current research needs. Parasitology 2009, 136, 1825–1835. [Google Scholar] [CrossRef] [PubMed]
- Bergquist, R.; Utzinger, J.; Keiser, J. Controlling schistosomiasis with praziquantel: How much longer without a viable alternative? Infect. Dis. Poverty 2017, 6, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Cioli, D.; Pica-Mattoccia, L.; Basso, A.; Guidi, A. Molecular & Biochemical Parasitology. Schistosomiasis control: Praziquantel forever? Mol. Biochem. Parasitol. 2014, 195, 23–29. [Google Scholar] [CrossRef] [PubMed]
- Cribb, D.M.; Clarke, N.E.; Doi, S.A.; Nery, S.V. Differential impact of mass and targeted deworming campaigns for schistosomiasis control in children: A systematic review and meta-analysis. Am. J. Trop. Med. Hyg. 2018, 99, 415. [Google Scholar]
- Kabuyaya, M.; Chimbari, M.J.; Mukaratirwa, S. Efficacy of praziquantel treatment regimens in pre-school and school aged children infected with schistosomiasis in sub-Saharan Africa: A systematic review. Infect. Dis. Poverty 2018, 7, 1–7. [Google Scholar] [CrossRef]
- Le, L.; Hsieh, M.H. Diagnosing urogenital schistosomiasis: Dealing with diminishing returns. Trends Parasitol. 2017, 33, 378–387. [Google Scholar] [CrossRef]
- Stete, K.; Krauth, S.J.; Coulibaly, J.T.; Knopp, S.; Hattendorf, J.; Müller, I.; Lohourignon, L.K.; Kern, W.V.; N’Goran, E.K.; Utzinger, J. Dynamics of Schistosoma haematobium egg output and associated infection parameters following treatment with praziquantel in school-aged children. Parasites Vectors 2012, 5, 1–10. [Google Scholar] [CrossRef]
- Knopp, S.; Ame, S.M.; Hattendorf, J.; Ali, S.M.; Khamis, I.S.; Bakar, F.; Khamis, M.A.; Person, B.; Kabole, F.; Rollinson, D. Urogenital schistosomiasis elimination in Zanzibar: Accuracy of urine filtration and haematuria reagent strips for diagnosing light intensity Schistosoma haematobium infections. Parasites Vectors 2018, 11, 1–11. [Google Scholar] [CrossRef]
- King, C.H.; Bertsch, D. Meta-analysis of urine heme dipstick diagnosis of Schistosoma haematobium infection, including low-prevalence and previously-treated populations. PLoS Negl. Trop. Dis. 2013, 7. [Google Scholar] [CrossRef] [PubMed]
- Mbanefo, E.C.; Huy, N.T.; Wadagni, A.A.; Eneanya, C.I.; Nwaorgu, O.; Hirayama, K. Host determinants of reinfection with schistosomes in humans: A systematic review and meta-analysis. PLoS Negl. Trop. Dis. 2014, 8. [Google Scholar] [CrossRef] [PubMed]
- Tchuem, T.L.A.; Momo, S.C.; Stothard, J.R.; Rollinson, D. Efficacy of praziquantel and reinfection patterns in single and mixed infection foci for intestinal and urogenital schistosomiasis in Cameroon. Acta Trop. 2013, 128, 275–283. [Google Scholar] [CrossRef] [PubMed]
- Dejon-Agobé, J.C.; Edoa, J.R.; Honkpehedji, Y.J.; Zinsou, J.F.; Adégbitè, B.R.; Ngwese, M.M.; Mangaboula, A.; Lell, B.; Grobusch, M.P.; Mordmüller, B.; et al. Schistosoma haematobium infection morbidity, praziquantel effectiveness and reinfection rate among children and young adults in Gabon. Parasites Vectors 2019, 12, 1–11. [Google Scholar] [CrossRef]
- Bergquist, R.; Johansen, M.V.; Utzinger, J. Diagnostic dilemmas in helminthology: What tools to use and when? Trends Parasitol. 2009, 25, 151–156. [Google Scholar] [CrossRef]
- Rollinson, D.; Knopp, S.; Levitz, S.; Stothard, J.R.; Tchuem, T.L.A.; Garba, A.; Mohammed, K.A.; Schur, N.; Person, B.; Colley, D.G.; et al. Time to set the agenda for schistosomiasis elimination. Acta Trop. 2013, 128, 423–440. [Google Scholar] [CrossRef]
- McCarthy, J.S.; Lustigman, S.; Yang, G.J.; Barakat, R.M.; García, H.H.; Sripa, B.; Willingham, A.L.; Prichard, R.K.; Basáñez, M.G. A research agenda for helminth diseases of humans: Diagnostics for control and elimination programmes. PLoS Negl. Trop. Dis. 2012, 6. [Google Scholar] [CrossRef]
- Hawkins, K.R.; Cantera, J.L.; Storey, H.L.; Leader, B.T.; De Los Santos, T. Diagnostic tests to support late-stage control programs for schistosomiasis and soil-transmitted h=elminthiases. PLoS Negl. Trop. Dis. 2016, 10, 1–15. [Google Scholar] [CrossRef]
- Knopp, S.; Becker, S.L.; Ingram, K.J.; Keiser, J.; Utzinger, J. Diagnosis and treatment of schistosomiasis in children in the era of intensified control. Expert Rev. Anti. Infect. Ther. 2013, 11, 1237–1258. [Google Scholar] [CrossRef]
- Knopp, S.; Corstjens, P.L.A.M.; Koukounari, A.; Cercamondi, C.I.; Ame, S.M.; Ali, S.M.; De Dood, C.J.; Mohammed, K.A.; Utzinger, J.; Rollinson, D.; et al. Sensitivity and specificity of a urine circulating anodic antigen test for the diagnosis of Schistosoma haematobium in low endemic settings. PLoS Negl. Trop. Dis. 2015, 9, 1–19. [Google Scholar] [CrossRef]
- Peralta, J.M.; Cavalcanti, M.G. Is POC-CCA a truly reliable test for schistosomiasis diagnosis in low endemic areas? The trace results controversy. PLoS Negl. Trop. Dis. 2018, 12, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Hinz, R.; Schwarz, N.G.; Hahn, A.; Frickmann, H. Serological approaches for the diagnosis of schistosomiasis–A review. Mol. Cell. Probes 2017, 31, 2–21. [Google Scholar] [CrossRef] [PubMed]
- Gordon, C.A.; Gray, D.J.; Gobert, G.N.; McManus, D.P. DNA amplification approaches for the diagnosis of key parasitic helminth infections of humans. Mol. Cell. Probes 2011, 25, 143–152. [Google Scholar] [CrossRef]
- Archer, J.; Lacourse, E.J.; Webster, L.B.; Stothard, J.R. An update on non-invasive urine diagnostics for human-infecting parasitic helminths: What more could be done and how? Parasitology 2019. [Google Scholar] [CrossRef]
- Vinkeles, M.N.V.S.; Van Dam, G.J.; Shaproski, D.; Kahama, A.I.; Brienen, E.A.T.; Vennervald, B.J.; Van Lieshout, L. Diagnostic performance of Schistosoma real-time PCR in urine samples from Kenyan children infected with Schistosoma haematobium: Day-to-day variation and follow-up after praziquantel treatment. PLoS Negl. Trop. Dis. 2014, 8. [Google Scholar] [CrossRef]
- Lodh, N.; Naples, J.M.; Bosompem, K.M.; Quartey, J.; Shiff, C.J. Detection of parasite-specific DNA in urine sediment obtained by filtration differentiates between single and mixed infections of Schistosoma mansoni and S. haematobium from endemic areas in Ghana. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Ibironke, O.; Koukounari, A.; Asaolu, S.; Moustaki, I.; Shiff, C. Validation of a new test for Schistosoma haematobium based on detection of Dra1 DNA fragments in urine: Evaluation through latent class analysis. PLoS Negl. Trop. Dis. 2012, 6, 1–6. [Google Scholar] [CrossRef]
- Guegan, H.; Fillaux, J.; Charpentier, E.; Robert-Gangneux, F.; Chauvin, P.; Guemas, E.; Boissier, J.; Valentin, A.; Cassaing, S.; Gangneux, J.P.; et al. Real-time PCR for diagnosis of imported schistosomiasis. PLoS Negl. Trop. Dis. 2019, 13, 1–18. [Google Scholar] [CrossRef]
- Keller, D.; Rothen, J.; Dangy, J.; Saner, C.; Daubenberger, C.; Allan, F.; Ame, S.M.; Ali, S.M.; Kabole, F.; Hattendorf, J.; et al. Performance of a real-time PCR approach for diagnosing Schistosoma haematobium infections of different intensity in urine samples from Zanzibar. Infect. Dis. Poverty 2020, 9, 1–13. [Google Scholar] [CrossRef]
- Minetti, C.; LaCourse, E.J.; Reimer, L.; Stothard, J.R. Focusing nucleic acid-based molecular diagnostics and xenomonitoring approaches for human helminthiases amenable to preventive chemotherapy. Parasitol. Open 2016, 2. [Google Scholar] [CrossRef]
- Weerakoon, K.G.; Gordon, C.A.; McManus, D.P. DNA diagnostics for schistosomiasis control. Trop. Med. Infect. Dis. 2018, 3, 81. [Google Scholar] [CrossRef]
- Pai, N.P.; Vadnais, C.; Denkinger, C.; Engel, N.; Pai, M. Point-of-care testing for infectious diseases: Diversity, complexity, and barriers in low- and middle-income countries. PLoS Med. 2012, 9. [Google Scholar] [CrossRef] [PubMed]
- Abbasi, I.; King, C.H.; Muchiri, E.M.; Hamburger, J. Detection of Schistosoma mansoni and Schistosoma haematobium DNA by loop-mediated isothermal amplification: Identification of infected snails from early prepatency. Am. J. Trop. Med. Hyg. 2010, 83, 427–432. [Google Scholar] [CrossRef] [PubMed]
- Lodh, N.; Mikita, K.; Bosompem, K.M.; Anyan, W.K.; Quartey, J.K.; Otchere, J.; Shiff, C.J. Point-of-care diagnosis of multiple schistosome parasites: Species-specific DNA detection in urine by loop-mediated isothermal amplification (LAMP). Acta Trop. 2017, 173, 125–129. [Google Scholar] [CrossRef] [PubMed]
- Rosser, A.; Rollinson, D.; Forrest, M.; Webster, B.L. Isothermal recombinase polymerase amplification (RPA) of Schistosoma haematobium DNA and oligochromatographic lateral flow detection. Parasites Vectors 2015, 8, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Rostron, P.; Pennance, T.; Bakar, F.; Rollinson, D.; Knopp, S.; Allan, F.; Kabole, F.; Ali, S.M.; Ame, S.M.; Webster, B.L. Development of a recombinase polymerase amplification (RPA) fluorescence assay for the detection of Schistosoma haematobium. Parasites Vectors 2019, 1–7. [Google Scholar] [CrossRef]
- Gonzalez, S.J.M.; Bhattacharyya, T.; Alshehri, H.R.; Poulton, K.; Allen, S.; Miles, M.A.; Arianitwe, M.; Tukahebwa, E.M.; Webster, B.; Stothard, J.R. Application of a recombinase polymerase amplification (RPA) assay and pilot field testing for Giardia duodenalis at Lake Albert, Uganda. Parasites Vectors 2020, 1–9. [Google Scholar] [CrossRef]
- Emery, A.M.; Allan, F.E.; Rabone, M.E.; Rollinson, D. Schistosomiasis collection at NHM (SCAN). Parasites Vectors 2012, 5, 1. [Google Scholar] [CrossRef]
- Knopp, S.; Mohammed, K.A.; Ali, S.M.; Khamis, I.S.; Ame, S.M.; Albonico, M.; Gouvras, A.; Fenwick, A.; Savioli, L.; Colley, D.G.; et al. Study and implementation of urogenital schistosomiasis elimination in Zanzibar (Unguja and Pemba islands) using an integrated multidisciplinary approach. BMC Public Health 2012, 12. [Google Scholar] [CrossRef]
- Knopp, S.; Ame, S.M.; Person, B.; Hattendorf, J.; Id, M.R.; Juma, S.; Muhsin, J.; Khamis, I.S.; Hollenberg, E.; Mohammed, K.A.; et al. A 5 year intervention study on elimination of urogenital schistosomiasis in Zanzibar. PLoS Negl. Trop. Dis. 2019, 13, e0007268. [Google Scholar] [CrossRef]
- Knopp, S.; Person, B.; Ame, S.M.; Ali, S.M.; Hattendorf, J.; Juma, S.; Muhsin, J.; Khamis, I.S.; Mohammed, K.A.; Utzinger, J.; et al. Evaluation of integrated interventions layered on mass drug administration for urogenital schistosomiasis elimination: A cluster-randomised trial. Lancet Glob. Heal. 2019, 7, e1118–e1129. [Google Scholar] [CrossRef]
- Nunes, T.; Heuer, C.; Marshall, J.; Sanchez, J.; Thorn, R.; Reiczigel, J.; Robison-Cox, J.; Sebastiani, P.; Solymos, P.; Yoshida, K.; et al. Package ‘epiR’. Available online: https://cran.r-project.org/web/packages/epiR/epiR.pdf (accessed on 10 June 2020).
- R: A Language and Environment for Statistical Computing. Team, R. Development Core. Vienna: R Foundation for Statistical Computing. Available online: https://www.gbif.org/tool/81287/r-a-language-and-environment-for-statistical-computing (accessed on 10 June 2020).
Sample Availability: Samples (DNA extracts of parasites) are available from the authors upon appropriate request. |
| Urine-Egg Microscopy * (Reference) | |||||
|---|---|---|---|---|---|
| + | − | Total | |||
| RT-ShDra1-RPA (Index) | + | 148 | 0 | 148 | PPV % (±95% CI): 100 (97.5–100) (48/148) |
| − | 10 | 10 | 20 | NPV % (±95% CI): 50 (27.2–72.8) (10/20) | |
| Total | 158 | 10 | 168 | ||
| Sensitivity % (±95% CI): 93.7 (88.7–96.9) (148/158) | Specificity % (±95% CI): 100 (69.1–100) (10/10) | ||||
| Reference | Index | Analysis | Ultra-High (>400 Eggs/10 mL Urine) | High (50–399 Eggs/10 mL Urine) | Low (10–49 Eggs/10 mL Urine) | Ultra-Low (1–9 Eggs/10 mL Urine) | Low and Ultra-Low (1–49 Eggs/10 mL Urine) | Egg-Negative |
|---|---|---|---|---|---|---|---|---|
| Total number of samples within egg count category (% total) * | 9 (5%) | 27 (16%) | 52 (31%) | 70 (42%) | 122 (73%) | 10 (6%) | ||
| Urine-egg microscopy * | RT-ShDra1-RPA | Sensitivity % (±95% CI) ** | 100 [9/9] (66.4–1) | 96.3 [26/27] (81–99.9) | 94.2 [49/52] (84.1–98.8) | 91.4 [64/70] (82.2–96.8) | 92.6 [113/122] (86.5–96.6) | NA |
| Negative Predictive Value % (±95% CI) ** | 100 [10/10] (69.2–100) | 90.9 [10/11] (58.7–99.8) | 76.9 [10/13] (64.2–95) | 62.5 [10/16] (35.4–84.8) | 52.6 [10/19] (28.9–75.6) | NA |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Archer, J.; Barksby, R.; Pennance, T.; Rostron, P.; Bakar, F.; Knopp, S.; Allan, F.; Kabole, F.; Ali, S.M.; Ame, S.M.; et al. Analytical and Clinical Assessment of a Portable, Isothermal Recombinase Polymerase Amplification (RPA) Assay for the Molecular Diagnosis of Urogenital Schistosomiasis. Molecules 2020, 25, 4175. https://doi.org/10.3390/molecules25184175
Archer J, Barksby R, Pennance T, Rostron P, Bakar F, Knopp S, Allan F, Kabole F, Ali SM, Ame SM, et al. Analytical and Clinical Assessment of a Portable, Isothermal Recombinase Polymerase Amplification (RPA) Assay for the Molecular Diagnosis of Urogenital Schistosomiasis. Molecules. 2020; 25(18):4175. https://doi.org/10.3390/molecules25184175
Chicago/Turabian StyleArcher, John, Rebecca Barksby, Tom Pennance, Penelope Rostron, Faki Bakar, Stefanie Knopp, Fiona Allan, Fatma Kabole, Said M. Ali, Shaali M. Ame, and et al. 2020. "Analytical and Clinical Assessment of a Portable, Isothermal Recombinase Polymerase Amplification (RPA) Assay for the Molecular Diagnosis of Urogenital Schistosomiasis" Molecules 25, no. 18: 4175. https://doi.org/10.3390/molecules25184175
APA StyleArcher, J., Barksby, R., Pennance, T., Rostron, P., Bakar, F., Knopp, S., Allan, F., Kabole, F., Ali, S. M., Ame, S. M., Rollinson, D., & Webster, B. L. (2020). Analytical and Clinical Assessment of a Portable, Isothermal Recombinase Polymerase Amplification (RPA) Assay for the Molecular Diagnosis of Urogenital Schistosomiasis. Molecules, 25(18), 4175. https://doi.org/10.3390/molecules25184175






