Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis
Abstract
1. Introduction
2. Results
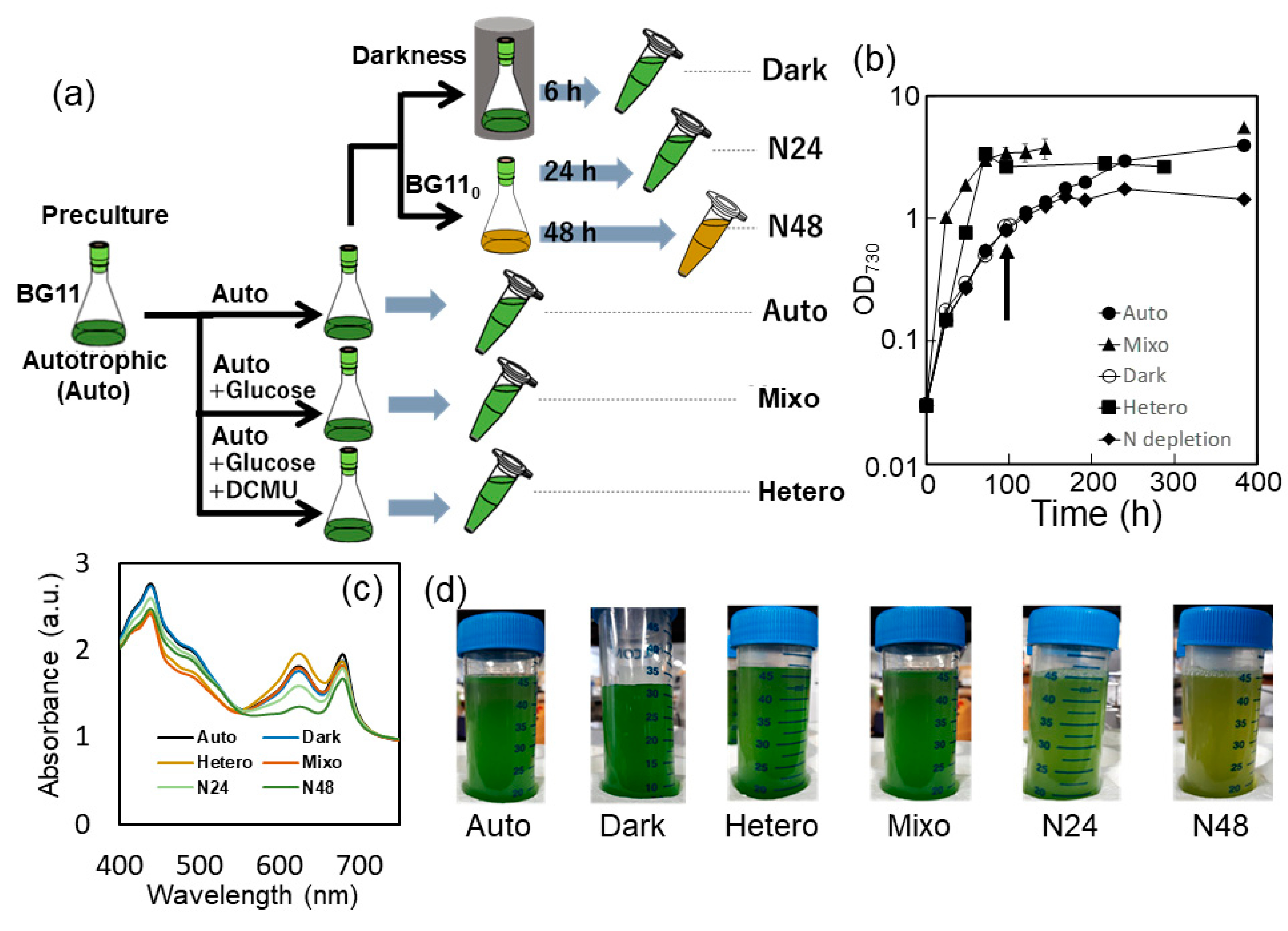
2.1. Synechocystis sp. PCC 6803 Culture under Various Growth Conditions
2.2. Acquisition of Targeted Proteomic and Phosphoproteomic Data
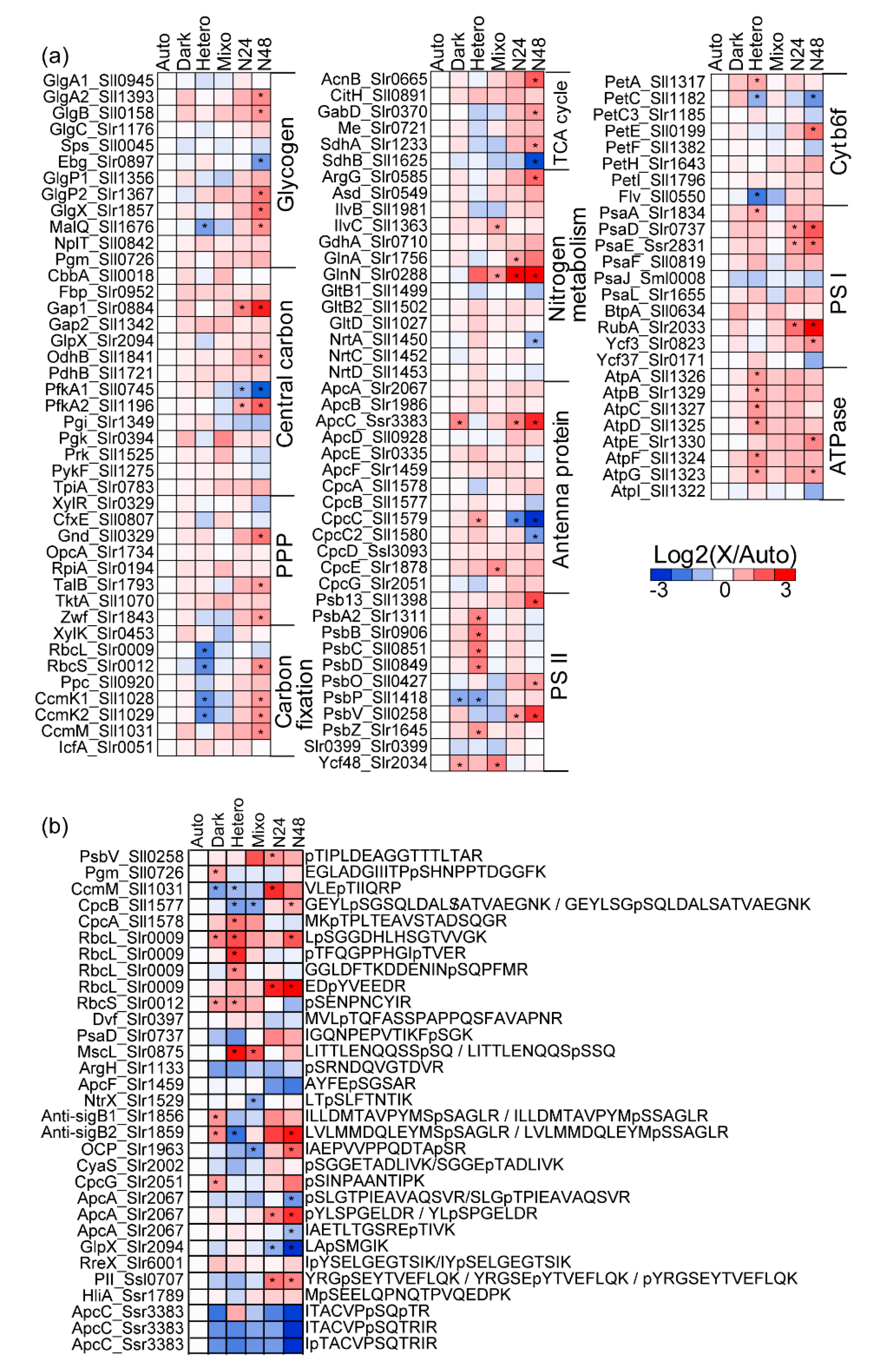
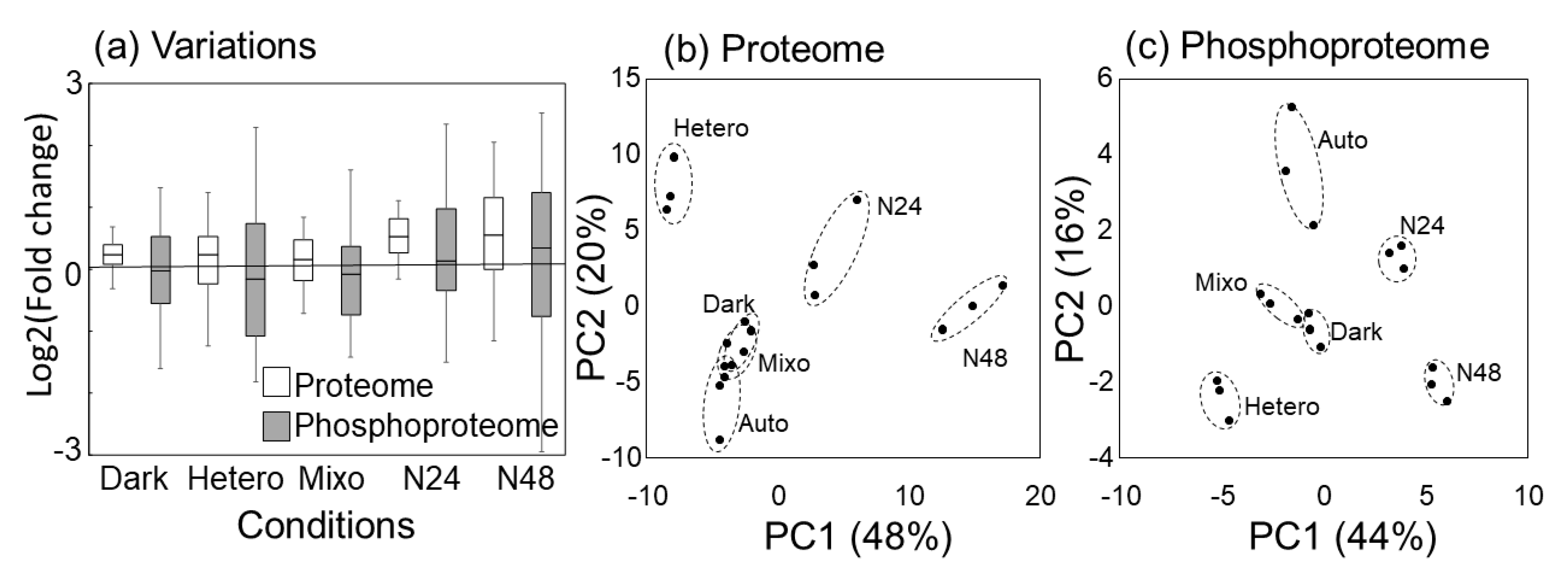
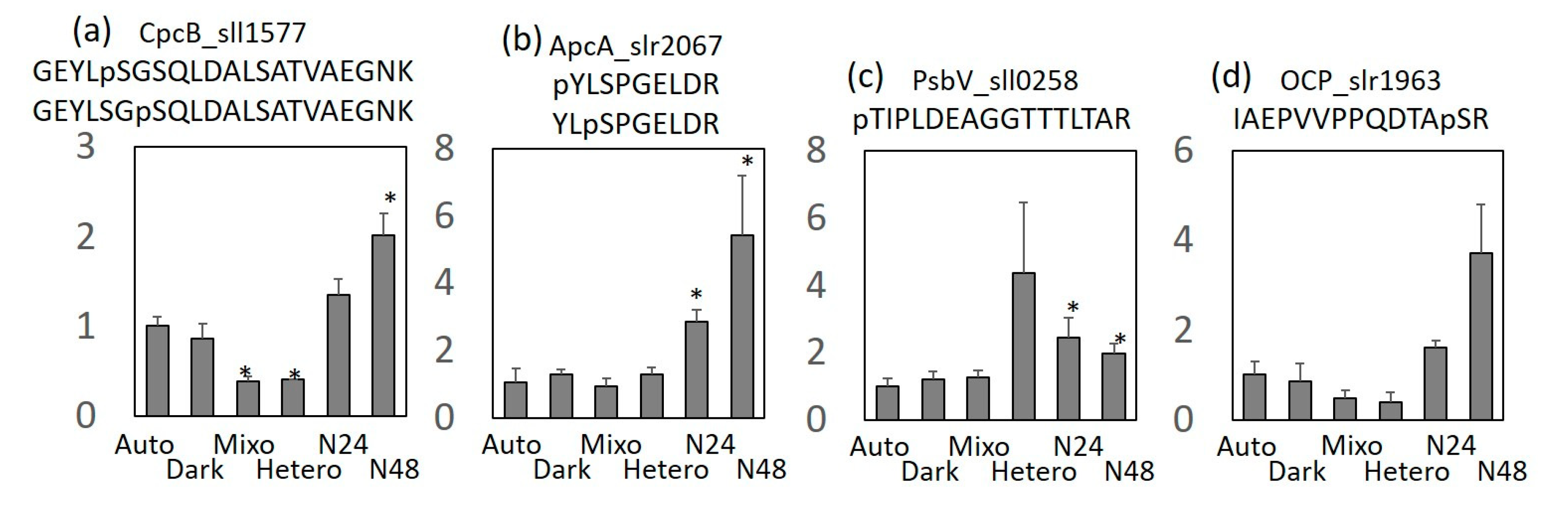
2.3. Comparison of Protein Expression and Phosphorylation Profiles
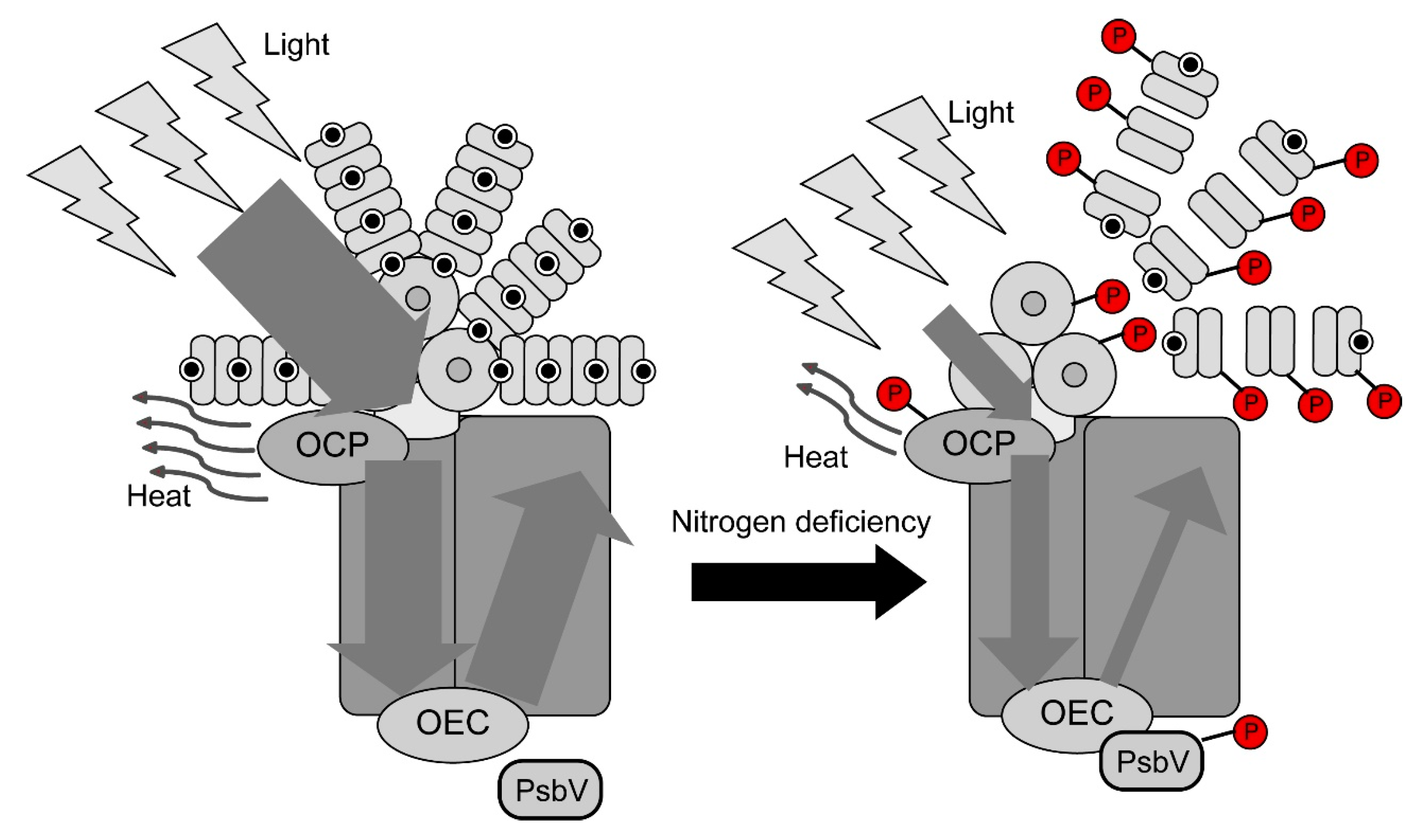
2.4. Changes in Photosynthetic Parameters under Nitrogen Deficiency
3. Discussion
4. Materials and Methods
4.1. Culture Conditions
4.2. Sample Preparation for Proteomic Analysis
4.3. Reduction and Alkylation/Methanol Chloroform Precipitation
4.4. Trypsin/LysC Digestion
4.5. Samples Desalination
4.6. Phosphopeptide Enrichment
4.7. Design of MRM Assays
4.8. NanoLC–MS/MS Using MRM Assays
4.9. Measurement of Ultraviolet–Visible (UV-VIS) Spectra
4.10. Proteomic Data Analysis
4.11. Chlorophyll Fluorescence Measurement
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Huang, S.; Chen, L.; Te, R.; Qiao, J.; Wang, J.; Zhang, W. Complementary iTRAQ proteomics and RNA-seq transcriptomics reveal multiple levels of regulation in response to nitrogen starvation in Synechocystis sp. PCC 6803. Mol. Biosyst. 2013, 9, 2565–2574. [Google Scholar] [CrossRef]
- Mann, M.; Jensen, O.N. Proteomic analysis of post-translational modifications. Nat. Biotechnol. 2003, 21, 255–261. [Google Scholar] [CrossRef]
- Rogers, L.D.; Foster, L.J. Phosphoproteomics-finally fulfilling the promise? Mol. Biosyst. 2009, 5, 1122–1129. [Google Scholar] [CrossRef]
- Cozzone, A.J. Protein phosphorylation in prokaryotes. Annu. Rev. Microbiol. 1988, 42, 97–125. [Google Scholar] [CrossRef]
- Thorsness, P.E.; Koshland, D.E. Inactivation of isocitrate dehydrogenase by phosphorylation is mediated by the negative charge of the phosphate. J. Biol. Chem. 1987, 262, 10422–10425. [Google Scholar] [PubMed]
- Forchhammer, K. Global carbon/nitrogen control by PII signal transduction in cyanobacteria: From signals to targets. FEMS Microbiol. Rev. 2004, 28, 319–333. [Google Scholar] [CrossRef] [PubMed]
- Osanai, T.; Kanesaki, Y.; Nakano, T.; Takahashi, H.; Asayama, M.; Shirai, M.; Kanehisa, M.; Suzuki, I.; Murata, N.; Tanaka, K. Positive regulation of sugar catabolic pathways in the cyanobacterium Synechocystis sp. PCC 6803 by the group 2 σ factor SigE. J. Biol. Chem. 2005, 280, 30653–30659. [Google Scholar] [CrossRef] [PubMed]
- Iwai, M.; Takahashi, Y.; Minagawa, J. Molecular remodeling of photosystem II during state transitions in Chlamydomonas reinhardtii. Plant Cell 2008, 20, 2177–2189. [Google Scholar] [CrossRef]
- Kato, Y.; Sakamoto, W. Phosphorylation of photosystem II core proteins prevents undesirable cleavage of D1 and contributes to the fine-tuned repair of photosystem II. Plant J. 2014, 79, 312–321. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.C.; Jang, J.; Sakr, S.; Wang, L. Protein phosphorylation on Ser, Thr and Tyr residues in cyanobacteria. J. Mol. Microbiol. Biotechnol. 2006, 9, 154–166. [Google Scholar] [CrossRef]
- Mikkat, S.; Fulda, S.; Hagemann, M. A 2D gel electrophoresis-based snapshot of the phosphoproteome in the cyanobacterium Synechocystis sp. strain PCC 6803. Microbiology 2014, 160, 296–306. [Google Scholar] [CrossRef]
- Spät, P.; Macek, B.; Forchhammer, K. Phosphoproteome of the cyanobacterium Synechocystis sp. PCC 6803 and its dynamics during nitrogen starvation. Front. Microbiol. 2015, 6, 248. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Zhan, J.; Chen, Y.; Yang, M.; He, C.; Ge, F.; Wang, Q. Effects of phosphorylation of β subunits of phycocyanins on state transition in the model cyanobacterium Synechocystis sp. PCC 6803. Plant Cell Physiol. 2015, 56, 1997–2013. [Google Scholar] [CrossRef] [PubMed]
- Angeleri, M.; Muth-Pawlak, D.; Aro, E.M.; Battchikova, N. Study of O-phosphorylation sites in proteins involved in photosynthesis-related processes in Synechocystis sp. strain PCC 6803: Application of the SRM approach. J. Proteome Res. 2016, 15, 4638–4652. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.-G.; Kwon, J.; Eom, C.-Y.; Kang, Y.-M.; Roh, S.W.; Lee, K.-B.; Choi, J.-S. Directed analysis of cyanobacterial membrane phosphoproteome using stained phosphoproteins and titanium-enriched phosphopeptides. J. Microbiol. 2015, 53, 279–287. [Google Scholar] [CrossRef]
- Xiong, Q.; Chen, Z.; Ge, F. Proteomic analysis of post translational modifications in cyanobacteria. J. Proteomics 2016, 134, 57–64. [Google Scholar] [CrossRef]
- Fang, L.; Ge, H.; Huang, X.; Liu, Y.; Lu, M.; Wang, J.; Chen, W.; Xu, W.; Wang, Y. Trophic mode-dependent proteomic analysis reveals functional significance of light-independent chlorophyll synthesis in Synechocystis sp. PCC 6803. Mol. Plant 2017, 10, 73–85. [Google Scholar] [CrossRef]
- Tokumaru, Y.; Uebayashi, K.; Toyoshima, M.; Osanai, T.; Matsuda, F.; Shimizu, H. Comparative targeted proteomics of the central metabolism and photosystems in SigE mutant strains of Synechocystis sp. PCC 6803. Molecules 2018, 23, 1051. [Google Scholar] [CrossRef]
- Yoshikawa, K.; Hirasawa, T.; Ogawa, K.; Hidaka, Y.; Nakajima, T.; Furusawa, C.; Shimizu, H. Integrated transcriptomic and metabolomic analysis of the central metabolism of Synechocystis sp. PCC 6803 under different trophic conditions. Biotechnol. J. 2013, 8, 571–580. [Google Scholar] [CrossRef]
- Nakajima, T.; Kajihata, S.; Yoshikawa, K.; Matsuda, F.; Furusawa, C.; Hirasawa, T.; Shimizu, H. Integrated metabolic flux and omics analysis of Synechocystis sp. PCC 6803 under mixotrophic and photoheterotrophic conditions. Plant Cell Physiol. 2014, 55, 1606–1612. [Google Scholar] [CrossRef]
- Osanai, T.; Azuma, M.; Tanaka, K. Sugar catabolism regulated by light- and nitrogen-status in the cyanobacterium Synechocystis sp. PCC 6803. Photochem. Photobiol. Sci. 2007, 6, 508. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, H.; Kusama, Y.; Li, X.; Takaichi, S.; Nishiyama, Y. Overexpression of orange carotenoid protein protects the repair of PSII under strong light in Synechocystis sp. PCC 6803. Plant Cell Physiol. 2019, 60, 367–375. [Google Scholar] [CrossRef] [PubMed]
- Ogawa, T.; Sonoike, K. Effects of bleaching by nitrogen deficiency on the quantum yield of Photosystem II in Synechocystis sp. PCC 6803 revealed by Chl fluorescence measurements. Plant Cell Physiol. 2016, 57, 558–567. [Google Scholar] [CrossRef] [PubMed]
- Richaud, C.; Zabulon, G.; Joder, A.; Thomas, J.C. Nitrogen or sulfur starvation differentially affects phycobilisome degradation and expression of the nblA gene in Synechocystis strain PCC 6803. J. Bacteriol. 2001, 183, 2989–2994. [Google Scholar] [CrossRef]
- Saha, R.; Liu, D.; Connor, A.H.; Liberton, M.; Yu, J.; Bhattacharyya-pakrasi, M.; Balassy, A.; Zhang, F.; Moon, T.S.; Maranas, C.D.; et al. Diurnal regulation of cellular processes in the cyanobacterium. MBio 2016, 7. [Google Scholar] [CrossRef]
- Sendersky, E.; Kozer, N.; Levi, M.; Moizik, M.; Garini, Y.; Shav-Tal, Y.; Schwarz, R. The proteolysis adaptor, NblA, is essential for degradation of the core pigment of the cyanobacterial light-harvesting complex. Plant J. 2015, 83, 845–852. [Google Scholar] [CrossRef]
- Nguyen, A.Y.; Bricker, W.P.; Zhang, H.; Weisz, D.A.; Gross, M.L.; Pakrasi, H.B. The proteolysis adaptor, NblA, binds to the N-terminus of β-phycocyanin: Implications for the mechanism of phycobilisome degradation. Photosynth. Res. 2017, 132, 95–106. [Google Scholar] [CrossRef]
- Knoop, H.; Steuer, R. A computational analysis of stoichiometric constraints and trade-offs in cyanobacterial biofuel production. Front. Bioeng. Biotechnol. 2015, 3, 47. [Google Scholar] [CrossRef]
- Baier, A.; Winkler, W.; Korte, T.; Lockau, W.; Karradt, A. Degradation of phycobilisomes in Synechocystis sp. PCC6803: Evidence for essential formation of an NblA1/NblA2 heterodimer and its codegradation by a Clp protease complex. J. Biol. Chem. 2014, 289, 11755–11766. [Google Scholar] [CrossRef]
- Bienert, R.; Baier, K.; Volkmer, R.; Lockau, W.; Heinemann, U. Crystal structure of NblA from Anabaena sp. PCC 7120, a small protein playing a key role in phycobilisome degradation. J. Biol. Chem. 2006, 281, 5216–5223. [Google Scholar] [CrossRef]
- McGregor, A.; Klartag, M.; David, L.; Adir, N. Allophycocyanin trimer stability and functionality are primarily due to polar enhanced hydrophobicity of the phycocyanobilin binding pocket. J. Mol. Biol. 2008, 384, 406–421. [Google Scholar] [CrossRef] [PubMed]
- Collier, J.L.; Herbert, S.K.; Fork, D.C.; Grossman, A.R. Changes in the cyanobacterial photosynthetic apparatus during acclimation to macronutrient deprivation. Photosynth. Res. 1994, 42, 173–183. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Sherman, L.A. Characterization of Synechocystis sp. strain PCC 6803 and Δnbl mutants under nitrogen-deficient conditions. Arch. Microbiol. 2002, 178, 256–266. [Google Scholar] [CrossRef] [PubMed]
- Krasikov, V.; Aguirre von Wobeser, E.; Dekker, H.L.; Huisman, J.; Matthijs, H.C.P. Time-series resolution of gradual nitrogen starvation and its impact on photosynthesis in the cyanobacterium Synechocystis PCC 6803. Physiol. Plant. 2012, 145, 426–439. [Google Scholar] [CrossRef]
- Williams, J.G.K. Construction of specific mutations in photosystem II photosynthetic reaction center by genetic engineering methods in Synechocystis 6803. Methods Enzymol. 1988, 167, 766–778. [Google Scholar] [CrossRef]
- Rippka, R.; Deruelles, J.; Waterbury, J.B.; Herdman, M.; Stanier, R.Y. Generic assignments, strain histories and properties of pure cultures of cyanobacteria. Microbiology 1979, 111, 1–61. [Google Scholar] [CrossRef]
- Picotti, P.; Bodenmiller, B.; Mueller, L.N.; Domon, B.; Aebersold, R. Full Dynamic range proteome analysis of S. cerevisiae by targeted proteomics. Cell 2009, 138, 795–806. [Google Scholar] [CrossRef]
- Wessel, D.; Flügge, U.I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 1984, 138, 141–143. [Google Scholar] [CrossRef]
- Rappsilber, J.; Ishihama, Y.; Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 2003, 75, 663–670. [Google Scholar] [CrossRef]
- Ishihama, Y.; Rappsilber, J.; Mann, M. Modular stop and go extraction tips with stacked disks for parallel and multidimensional peptide fractionation in proteomics. J. Proteome Res. 2006, 5, 988–994. [Google Scholar] [CrossRef]
- Rappsilber, J.; Mann, M.; Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2007, 2, 1896–1906. [Google Scholar] [CrossRef] [PubMed]
- Kyoto Encyclopedia of Genes and Genomes (KEGG). Available online: http://www.genome.jp/kegg/kegg2.html (accessed on 30 July 2020).
- CyanoBase. Available online: http://genome.microbedb.jp/CyanoBase (accessed on 30 July 2020).
- MacLean, B.; Tomazela, D.M.; Shulman, N.; Chambers, M.; Finney, G.L.; Frewen, B.; Kern, R.; Tabb, D.L.; Liebler, D.C.; MacCoss, M.J. Skyline: An open source document editor for creating and analyzing targeted proteomics experiments. Bioinformatics 2010, 26, 966–968. [Google Scholar] [CrossRef] [PubMed]
- Matsuda, F.; Tomita, A.; Shimizu, H. Prediction of hopeless peptides unlikely to be selected for targeted proteome analysis. Mass Spectrom. 2017, 6, A0056. [Google Scholar] [CrossRef] [PubMed]
- Sato, N.; Misumi, O.; Toyoshima, M.; Moriyama, T.; Mori, N. Analysis of triacylglycerol accumulation under nitrogen deprivation in the red alga Cyanidioschyzon Merolae. Microbiology 2016, 162, 803–812. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are available from the authors. |
| qP | qN | ΦII | NPQ | Fv/Fm | Fv’/Fm’ | |
|---|---|---|---|---|---|---|
| Auto | 0.194 ± 0.069 | 0.282 ± 0.032 | 0.077 ± 0.031 | 0.065 ± 0.021 | 0.515 ± 0.012 | 0.393 ± 0.018 |
| N24 | 0.085 ± 0.032 * | 0.185 ± 0.046 * | 0.038 ± 0.015 | 0.032 ± 0.011 * | 0.528 ± 0.019 | 0.444 ± 0.019 * |
| N48 | 0.146 ± 0.055 | 0.167 ± 0.090 * | 0.051 ± 0.020 | 0.026 ± 0.004 * | 0.416 ± 0.063 * | 0.356 ± 0.073 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Toyoshima, M.; Tokumaru, Y.; Matsuda, F.; Shimizu, H. Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis. Molecules 2020, 25, 3582. https://doi.org/10.3390/molecules25163582
Toyoshima M, Tokumaru Y, Matsuda F, Shimizu H. Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis. Molecules. 2020; 25(16):3582. https://doi.org/10.3390/molecules25163582
Chicago/Turabian StyleToyoshima, Masakazu, Yuma Tokumaru, Fumio Matsuda, and Hiroshi Shimizu. 2020. "Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis" Molecules 25, no. 16: 3582. https://doi.org/10.3390/molecules25163582
APA StyleToyoshima, M., Tokumaru, Y., Matsuda, F., & Shimizu, H. (2020). Assessment of Protein Content and Phosphorylation Level in Synechocystis sp. PCC 6803 under Various Growth Conditions Using Quantitative Phosphoproteomic Analysis. Molecules, 25(16), 3582. https://doi.org/10.3390/molecules25163582






