Anion Inhibition Studies of the Beta-Carbonic Anhydrase from Escherichia coli
Abstract
1. Introduction
2. Results and Discussion
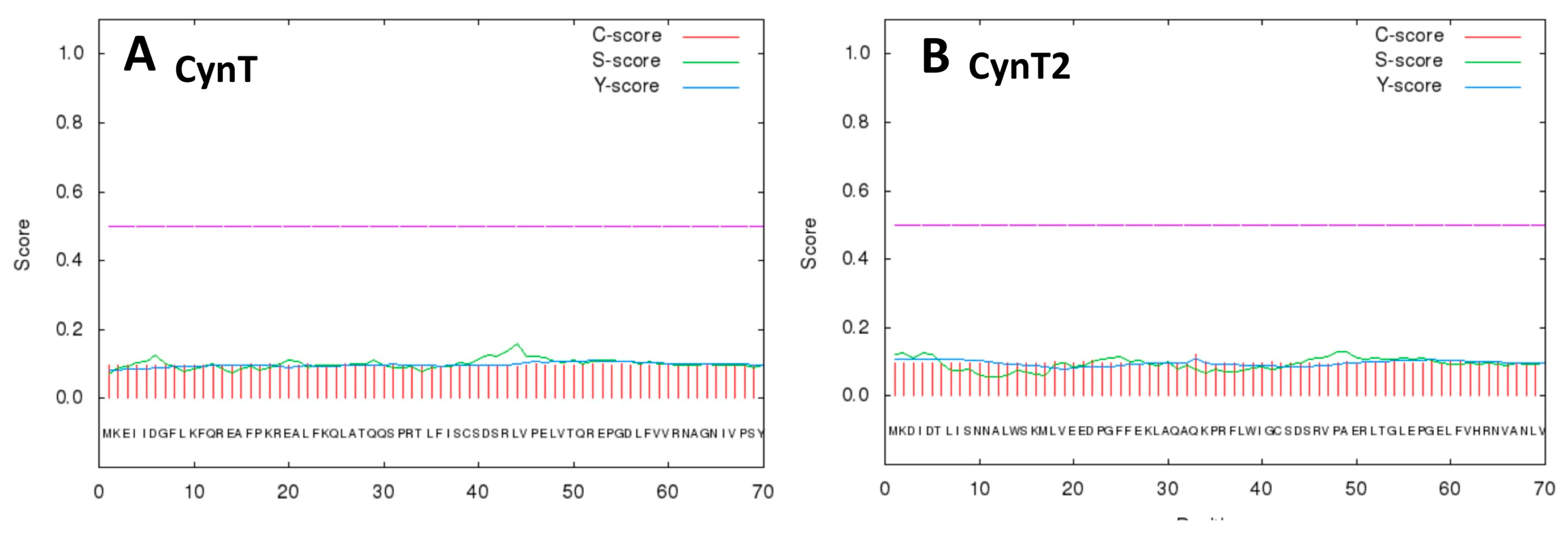
2.1. Primary Structure Analysis
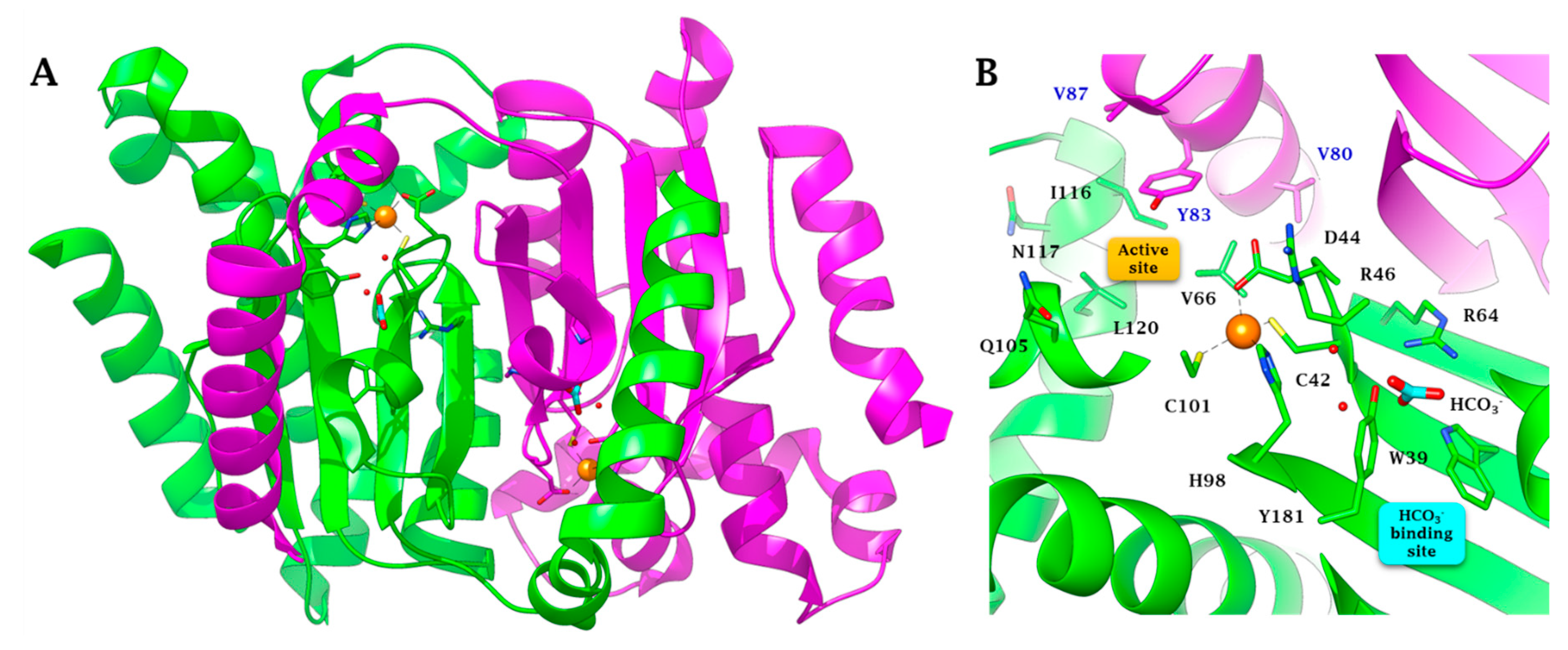
2.2. Three-Dimensional Structure Analysis
2.3. Production of Recombinat β-CA (CynT2)
2.4. Determination of the Kinetic Parameters Using the Stopped-Flow Technique
2.5. Inhibition Profile of Inorganic Metal-Complexing Anions
- A group of anions, such as iodide (I−), cyanide (CN−), azide (N3−), perchlorate (ClO4−) perosmate (OsO52−), pyrophosphate (P2O72−), divanadate (V2O72−) perrhenate (ReO4−), hexafluorophosphate (PF6−), and trifluoromethanesulfonate (CF3SO3−) were very weak inhibitors of the E.coli β-CA, with a KI > 10 mM (see Table 3). Within this group, VchCA_beta was inhibited with a KI ranging from 5.7 to 9.0 mM (Table 3), except for OsO52−, PF6−, and CF3SO3−, whose KIs were even not measurable (Table 3). Moreover, N3− and P2O72− showed a KI > 10 mM for the CynT2, while ClO4− with a KI > 200 mM was an ineffective inhibitor of CynT2 as well as the V. cholerae enzyme. Interesting to note that some of these anions resulted to be quite effective inhibitors, when tested on the two human CAs. For example, cyanide and azide inhibited the human isoform h CA I with KI of 0.5 and 12 µM, respectively (Table 3). Thus, low concentrations of N3− and CN− poison the two human enzymes as well as humans, whereas the bacterial CAs may better tolerate such toxic anions.
- Another group of anion inhibitors, which weakly inhibited CynT2, showed a KI in the range between 1.5 and 9.4 mM. This is the case of fluoride (F−), chloride (Cl−), bromide (Br−), thiocyanate (SCN−), nitrite (NO2−), nitrate (NO3−), bisulfite (HSO3−), sulfate (SO42−), hydrogensulfide (HS−), selenate (SeO42−), perruthenate (RuO4−), peroxydisulfate (S2O82−), selenocyanate (SeCN−), iminodisulfonate (NH(SO3)22−), and trithiocarbonate (CS32−). The inhibitory behavior of these anions was very similar to that exerted on the enzyme from V. cholerae (VchCA_beta); exceptions were HSO3−, SO42−, and NH(SO3)22−, which were ineffective inhibitors for VchCA_beta with a KI > 200 mM. At the same time, HS− inhibited the Vibrio enzyme with a KI > 20 mM. This last anion inhibited the human isoenzymes, hCA I and hCA II, with a KI = 0.6 µM and KI = 40 µM, respectively.
- The submillimolar inhibitors of CynT2 showing a KI in the range 0.25–0.89 mM were cyanate (CNO−), bicarbonate (HCO3−), carbonate (CO32−), stannate (SnO32−), tetraborate (B4O72−), and fluorosulfonate (FSO3−). These anions were millimolar inhibitors for the VchCA_beta (Table 1). The differences in the values of the inhibition constants can be due to the amino acid residues surrounding the catalytic pocket, which influence the interaction of the anion with the enzyme, even if the three-dimensional structure of the β-CAs from E. coli and V. cholerae are very similar. Intriguing is the result obtained using bicarbonate and carbonate as inhibitors. These two anions are not effective inhibitors (KI = 12–85 mM) for the two human isoforms (hCA I and hCA II), as well as for the β-CA from V. cholerae (KI = 5.9–6.7 mM). These results may reflect an evolutionary adaptation of the human and Vibrio CAs due to their continuous exposition to the high concentration of these two anions. The human enzymes are adapted to the high concentration of carbonate and bicarbonate present in the plasma, while V. cholerae colonizes the upper part of the small intestine characterized by high concentrations of bicarbonate, which is also a potent inducer of the expression of the genes involved in the virulence of the pathogen [45].
- The best inhibitors of CynT2 resulted to be the small molecules, such as sulfamide (NH2SO2NH2), sulfamate (NH2SO3H), phenylboronic acid (PhB(OH)2), phenylarsonic acid (PhAsO3H2) as well as diethyldithiocarbamate (EtNCS2−). As shown in Table 3, they showed KI values from 2.5 to 84 µM. These inhibitors were also effective against VchCA_beta (KI from 54 to 86 µM), except for EtNCS2−, which showed a KI = 730 µM. Interesting to note, that dithiocarbamates were recently reported as a potent new class of CAIs targeting both the α- and β-classes of such enzymes [60,61,62].
3. Materials and Methods
3.1. Chemicals and Instruments
3.2. Cloning, Expression, and Purification
3.3. Carbonic Anhydrase Assay for Monitoring the Recombinant Enzyme during the Purification Steps
3.4. SDS-PAGE
3.5. Western Blot
3.6. Protonography
3.7. Kinetic Parameters and Inhibition Constants Determined by the Stopped-Flow Technique
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mainil, J. Escherichia coli virulence factors. Vet. Immunol. Immunopathol. 2013, 152, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Allocati, N.; Masulli, M.; Alexeyev, M.F.; Di Ilio, C. Escherichia coli in Europe: An overview. Int. J. Environ. Res. Public Health 2013, 10, 6235–6254. [Google Scholar] [CrossRef] [PubMed]
- Conway, T.; Cohen, P.S. Commensal and Pathogenic Escherichia coli Metabolism in the Gut. In Metabolism and Bacterial Pathogenesis; Conway, T., Cohen, P., Eds.; ASM Press: Washington, WA, USA, 2015. [Google Scholar]
- Robins-Browne, R.M.; Holt, K.E.; Ingle, D.J.; Hocking, D.M.; Yang, J.; Tauschek, M. Are Escherichia coli Pathotypes Still Relevant in the Era of Whole-Genome Sequencing? Front. Cell Infect. Microbiol. 2016, 6, 141. [Google Scholar] [CrossRef] [PubMed]
- Stromberg, Z.R.; Van Goor, A.; Redweik, G.A.J.; Wymore Brand, M.J.; Wannemuehler, M.J.; Mellata, M. Pathogenic and non-pathogenic Escherichia coli colonization and host inflammatory response in a defined microbiota mouse model. Dis. Model Mech. 2018, 11, dmm035063. [Google Scholar] [CrossRef]
- Pitout, J.D. Extraintestinal Pathogenic Escherichia coli: A Combination of Virulence with Antibiotic Resistance. Front. Microbiol. 2012, 3, 9. [Google Scholar] [CrossRef] [PubMed]
- Nataro, J.P.; Kaper, J.B. Diarrheagenic Escherichia coli. Clin. Microbiol. Rev. 1998, 11, 142–201. [Google Scholar] [CrossRef] [PubMed]
- Moriel, D.G.; Bertoldi, I.; Spagnuolo, A.; Marchi, S.; Rosini, R.; Nesta, B.; Pastorello, I.; Corea, V.A.; Torricelli, G.; Cartocci, E.; et al. Identification of protective and broadly conserved vaccine antigens from the genome of extraintestinal pathogenic Escherichia coli. Proc. Natl. Acad. Sci. USA 2010, 107, 9072–9077. [Google Scholar] [CrossRef]
- Agus, A.; Massier, S.; Darfeuille-Michaud, A.; Billard, E.; Barnich, N. Understanding host-adherent-invasive Escherichia coli interaction in Crohn’s disease: Opening up new therapeutic strategies. Biomed. Res. Int. 2014, 2014, 567929. [Google Scholar] [CrossRef]
- Clermont, O.; Olier, M.; Hoede, C.; Diancourt, L.; Brisse, S.; Keroudean, M.; Glodt, J.; Picard, B.; Oswald, E.; Denamur, E. Animal and human pathogenic Escherichia coli strains share common genetic backgrounds. Infect. Genet. Evol. 2011, 11, 654–662. [Google Scholar] [CrossRef]
- Reid, S.D.; Herbelin, C.J.; Bumbaugh, A.C.; Selander, R.K.; Whittam, T.S. Parallel evolution of virulence in pathogenic Escherichia coli. Nature 2000, 406, 64–67. [Google Scholar] [CrossRef]
- Szych, J.; Wolkowicz, T.; La Ragione, R.; Madajczak, G. Impact of antibiotics on the intestinal microbiota and on the treatment of Shiga-toxin-producing Escherichia coli and Salmonella infections. Curr. Pharm. Des. 2014, 20, 4535–4548. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.; Capasso, C. An Overview of the Bacterial Carbonic Anhydrases. Metabolites 2017, 7, 56. [Google Scholar] [CrossRef] [PubMed]
- Vullo, D.; Del Prete, S.; Di Fonzo, P.; Carginale, V.; Donald, W.A.; Supuran, C.T.; Capasso, C. Comparison of the Sulfonamide Inhibition Profiles of the beta- and gamma-Carbonic Anhydrases from the Pathogenic Bacterium Burkholderia pseudomallei. Molecules 2017, 22, 421. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.S.; Ferry, J.G. Prokaryotic carbonic anhydrases. FEMS Microbiol. Rev. 2000, 24, 335–366. [Google Scholar] [CrossRef]
- Maeda, S.; Price, G.D.; Badger, M.R.; Enomoto, C.; Omata, T. Bicarbonate binding activity of the CmpA protein of the cyanobacterium Synechococcus sp. strain PCC 7942 involved in active transport of bicarbonate. J. Biol. Chem. 2000, 275, 20551–20555. [Google Scholar] [CrossRef] [PubMed]
- Joseph, P.; Ouahrani-Bettache, S.; Montero, J.L.; Nishimori, I.; Minakuchi, T.; Vullo, D.; Scozzafava, A.; Winum, J.Y.; Kohler, S.; Supuran, C.T. A new beta-carbonic anhydrase from Brucella suis, its cloning, characterization, and inhibition with sulfonamides and sulfamates, leading to impaired pathogen growth. Bioorg. Med. Chem. 2011, 19, 1172–1178. [Google Scholar] [CrossRef]
- Joseph, P.; Turtaut, F.; Ouahrani-Bettache, S.; Montero, J.L.; Nishimori, I.; Minakuchi, T.; Vullo, D.; Scozzafava, A.; Kohler, S.; Winum, J.Y.; et al. Cloning, characterization, and inhibition studies of a beta-carbonic anhydrase from Brucella suis. J. Med. Chem. 2010, 53, 2277–2285. [Google Scholar] [CrossRef]
- Supuran, C.T.; Capasso, C. New light on bacterial carbonic anhydrases phylogeny based on the analysis of signal peptide sequences. J. Enzym. Inhib. Med. Chem. 2016, 31, 1254–1260. [Google Scholar] [CrossRef]
- Capasso, C.; Supuran, C.T. An overview of the alpha-, beta- and gamma-carbonic anhydrases from Bacteria: Can bacterial carbonic anhydrases shed new light on evolution of bacteria? J. Enzym. Inhib. Med. Chem. 2015, 30, 325–332. [Google Scholar] [CrossRef]
- Merlin, C.; Masters, M.; McAteer, S.; Coulson, A. Why is carbonic anhydrase essential to Escherichia Coli. J. Bacteriol. 2003, 185, 6415–6424. [Google Scholar] [CrossRef]
- Supuran, C.T. Advances in structure-based drug discovery of carbonic anhydrase inhibitors. Expert Opin. Drug Discov. 2017, 12, 61–88. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T. Structure and function of carbonic anhydrases. Biochem. J. 2016, 473, 2023–2032. [Google Scholar] [CrossRef] [PubMed]
- Annunziato, G.; Angeli, A.; D’Alba, F.; Bruno, A.; Pieroni, M.; Vullo, D.; De Luca, V.; Capasso, C.; Supuran, C.T.; Costantino, G. Discovery of New Potential Anti-Infective Compounds Based on Carbonic Anhydrase Inhibitors by Rational Target-Focused Repurposing Approaches. ChemMedChem 2016, 11, 1904–1914. [Google Scholar] [CrossRef] [PubMed]
- Ozensoy Guler, O.; Capasso, C.; Supuran, C.T. A magnificent enzyme superfamily: Carbonic anhydrases, their purification and characterization. J. Enzym. Inhib. Med. Chem. 2016, 31, 689–694. [Google Scholar] [CrossRef]
- Del Prete, S.; Vullo, D.; De Luca, V.; Carginale, V.; Ferraroni, M.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Sulfonamide inhibition studies of the beta-carbonic anhydrase from the pathogenic bacterium Vibrio cholerae. Bioorg. Med. Chem. 2016, 24, 1115–1120. [Google Scholar] [CrossRef]
- Del Prete, S.; De Luca, V.; De Simone, G.; Supuran, C.T.; Capasso, C. Cloning, expression and purification of the complete domain of the eta-carbonic anhydrase from Plasmodium falciparum. J. Enzym. Inhib. Med. Chem. 2016, 31, 54–59. [Google Scholar] [CrossRef]
- Capasso, C.; Supuran, C.T. An Overview of the Carbonic Anhydrases from Two Pathogens of the Oral Cavity: Streptococcus mutans and Porphyromonas gingivalis. Curr. Top. Med. Chem. 2016, 16, 2359–2368. [Google Scholar] [CrossRef]
- Del Prete, S.; Nocentini, A.; Supuran, C.T.; Capasso, C. Bacterial iota-carbonic anhydrase: A new active class of carbonic anhydrase identified in the genome of the Gram-negative bacterium Burkholderia territorii. J. Enzym. Inhib. Med. Chem. 2020, 35, 1060–1068. [Google Scholar] [CrossRef]
- Jensen, E.L.; Clement, R.; Kosta, A.; Maberly, S.C.; Gontero, B. A new widespread subclass of carbonic anhydrase in marine phytoplankton. Isme. J. 2019, 13, 2094–2106. [Google Scholar] [CrossRef]
- Kikutani, S.; Nakajima, K.; Nagasato, C.; Tsuji, Y.; Miyatake, A.; Matsuda, Y. Thylakoid luminal theta-carbonic anhydrase critical for growth and photosynthesis in the marine diatom Phaeodactylum tricornutum. Proc. Natl. Acad. Sci. USA 2016, 113, 9828–9833. [Google Scholar] [CrossRef]
- Capasso, C.; Supuran, C.T. Bacterial, fungal and protozoan carbonic anhydrases as drug targets. Expert Opin. Ther. Targets 2015, 19, 1689–1704. [Google Scholar] [CrossRef] [PubMed]
- Supuran, C.T.; Capasso, C. The eta-class carbonic anhydrases as drug targets for antimalarial agents. Expert Opin. Ther. Targets 2015, 19, 551–563. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; Supuran, C.T. An Overview of the Selectivity and Efficiency of the Bacterial Carbonic Anhydrase Inhibitors. Curr. Med. Chem. 2015, 22, 2130–2139. [Google Scholar] [CrossRef]
- Capasso, C.; Supuran, C.T. Sulfa and trimethoprim-like drugs-antimetabolites acting as carbonic anhydrase, dihydropteroate synthase and dihydrofolate reductase inhibitors. J. Enzym. Inhib. Med. Chem. 2014, 29, 379–387. [Google Scholar] [CrossRef]
- Capasso, C.; Supuran, C.T. Anti-infective carbonic anhydrase inhibitors: A patent and literature review. Expert Opin. Ther. Pat. 2013, 23, 693–704. [Google Scholar] [CrossRef] [PubMed]
- De Smet, B.; Mayo, M.; Peeters, C.; Zlosnik, J.E.; Spilker, T.; Hird, T.J.; LiPuma, J.J.; Kidd, T.J.; Kaestli, M.; Ginther, J.L.; et al. Burkholderia stagnalis sp. nov. and Burkholderia territorii sp. nov., two novel Burkholderia cepacia complex species from environmental and human sources. Int. J. Syst. Evol. Microbiol. 2015, 65, 2265–2271. [Google Scholar] [CrossRef]
- Kozliak, E.I.; Guilloton, M.B.; Gerami-Nejad, M.; Fuchs, J.A.; Anderson, P.M. Expression of proteins encoded by the Escherichia coli cyn operon: Carbon dioxide-enhanced degradation of carbonic anhydrase. J. Bacteriol. 1994, 176, 5711–5717. [Google Scholar] [CrossRef]
- Guilloton, M.B.; Lamblin, A.F.; Kozliak, E.I.; Gerami-Nejad, M.; Tu, C.; Silverman, D.; Anderson, P.M.; Fuchs, J.A. A physiological role for cyanate-induced carbonic anhydrase in Escherichia coli. J. Bacteriol. 1993, 175, 1443–1451. [Google Scholar] [CrossRef]
- Guilloton, M.B.; Korte, J.J.; Lamblin, A.F.; Fuchs, J.A.; Anderson, P.M. Carbonic anhydrase in Escherichia coli. A product of the cyn operon. J. Biol. Chem. 1992, 267, 3731–3734. [Google Scholar]
- Cronk, J.D.; Endrizzi, J.A.; Cronk, M.R.; O’Neill, J.W.; Zhang, K.Y. Crystal structure of E. coli beta-carbonic anhydrase, an enzyme with an unusual pH-dependent activity. Protein Sci. 2001, 10, 911–922. [Google Scholar] [CrossRef]
- Modak, J.K.; Tikhomirova, A.; Gorrell, R.J.; Rahman, M.M.; Kotsanas, D.; Korman, T.M.; Garcia-Bustos, J.; Kwok, T.; Ferrero, R.L.; Supuran, C.T.; et al. Anti-Helicobacter pylori activity of ethoxzolamide. J. Enzym. Inhib. Med. Chem. 2019, 34, 1660–1667. [Google Scholar] [CrossRef] [PubMed]
- Ronci, M.; Del Prete, S.; Puca, V.; Carradori, S.; Carginale, V.; Muraro, R.; Mincione, G.; Aceto, A.; Sisto, F.; Supuran, C.T.; et al. Identification and characterization of the alpha-CA in the outer membrane vesicles produced by Helicobacter pylori. J. Enzym. Inhib. Med. Chem. 2019, 34, 189–195. [Google Scholar] [CrossRef] [PubMed]
- Buzas, G.M. Helicobacter pylori—2010. Orv. Hetil. 2010, 151, 2003–2010. [Google Scholar] [CrossRef] [PubMed]
- Abuaita, B.H.; Withey, J.H. Bicarbonate Induces Vibrio cholerae virulence gene expression by enhancing ToxT activity. Infect. Immun. 2009, 77, 4111–4120. [Google Scholar] [CrossRef] [PubMed]
- Kohler, S.; Ouahrani-Bettache, S.; Winum, J.Y. Brucella suis carbonic anhydrases and their inhibitors: Towards alternative antibiotics. J. Enzym. Inhib. Med. Chem. 2017, 32, 683–687. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Supuran, C.T. 3D-QSAR CoMFA studies on sulfonamide inhibitors of the Rv3588c beta-carbonic anhydrase from Mycobacterium tuberculosis and design of not yet synthesized new molecules. J. Enzym. Inhb. Med. Chem. 2014, 29, 449–455. [Google Scholar] [CrossRef]
- Ceruso, M.; Vullo, D.; Scozzafava, A.; Supuran, C.T. Sulfonamides incorporating fluorine and 1,3,5-triazine moieties are effective inhibitors of three beta-class carbonic anhydrases from Mycobacterium tuberculosis. J. Enzym. Inhib. Med. Chem. 2014, 29, 686–689. [Google Scholar] [CrossRef]
- Carta, F.; Maresca, A.; Covarrubias, A.S.; Mowbray, S.L.; Jones, T.A.; Supuran, C.T. Carbonic anhydrase inhibitors. Characterization and inhibition studies of the most active beta-carbonic anhydrase from Mycobacterium tuberculosis, Rv3588c. Bioorg. Med. Chem. Lett. 2009, 19, 6649–6654. [Google Scholar] [CrossRef]
- Rollenhagen, C.; Bumann, D. Salmonella enterica highly expressed genes are disease specific. Infect. Immun. 2006, 74, 1649–1660. [Google Scholar] [CrossRef]
- Lotlikar, S.R.; Kayastha, B.B.; Vullo, D.; Khanam, S.S.; Braga Reygan, E.; Murray, A.B.; McKenna, R.; Supuran, C.T.; Patrauchan, M.A. Pseudomonas aeruginosa β-carbonic anhydrase, psCA1, is required for calcium deposition and contributes to virulence. Cell Calcium 2019, 84, 102080. [Google Scholar] [CrossRef]
- Alonso-de Castro, S.; Terenzi, A.; Gurruchaga-Pereda, J.; Salassa, L. Catalysis Concepts in Medicinal Inorganic Chemistry. Chemistry 2019, 25, 6651–6660. [Google Scholar] [CrossRef] [PubMed]
- Mjos, K.D.; Orvig, C. Metallodrugs in medicinal inorganic chemistry. Chem. Rev. 2014, 114, 4540–4563. [Google Scholar] [CrossRef] [PubMed]
- Ferraroni, M.; Del Prete, S.; Vullo, D.; Capasso, C.; Supuran, C.T. Crystal structure and kinetic studies of a tetrameric type II beta-carbonic anhydrase from the pathogenic bacterium Vibrio cholerae. Acta Cryst. D Biol. Cryst. 2015, 71, 2449–2456. [Google Scholar] [CrossRef] [PubMed]
- Cronk, J.D.; Rowlett, R.S.; Zhang, K.Y.; Tu, C.; Endrizzi, J.A.; Lee, J.; Gareiss, P.C.; Preiss, J.R. Identification of a novel noncatalytic bicarbonate binding site in eubacterial beta-carbonic anhydrase. Biochemistry 2006, 45, 4351–4361. [Google Scholar] [CrossRef] [PubMed]
- Capasso, C.; De Luca, V.; Carginale, V.; Cannio, R.; Rossi, M. Biochemical properties of a novel and highly thermostable bacterial alpha-carbonic anhydrase from Sulfurihydrogenibium yellowstonense YO3AOP1. J. Enzym. Inhib. Med. Chem. 2012, 27, 892–897. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Tikhomirova, A.; Modak, J.K.; Hutton, M.L.; Supuran, C.T.; Roujeinikova, A. Antibacterial activity of ethoxzolamide against Helicobacter pylori strains SS1 and 26695. Gut Pathog. 2020, 12, 20. [Google Scholar] [CrossRef]
- Del Prete, S.; Vullo, D.; De Luca, V.; Carginale, V.; di Fonzo, P.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Anion inhibition profiles of alpha-, beta- and gamma-carbonic anhydrases from the pathogenic bacterium Vibrio cholerae. Bioorg. Med. Chem. 2016, 24, 3413–3417. [Google Scholar] [CrossRef]
- Supuran, C.T. How many carbonic anhydrase inhibition mechanisms exist. J. Enzym. Inhib. Med. Chem. 2016, 31, 345–360. [Google Scholar] [CrossRef]
- Aspatwar, A.; Hammaren, M.; Parikka, M.; Parkkila, S.; Carta, F.; Bozdag, M.; Vullo, D.; Supuran, C.T. In vitro inhibition of Mycobacterium tuberculosis beta-carbonic anhydrase 3 with Mono- and dithiocarbamates and evaluation of their toxicity using zebrafish developing embryos. J. Enzym. Inhib. Med. Chem. 2020, 35, 65–71. [Google Scholar] [CrossRef]
- Carta, F.; Aggarwal, M.; Maresca, A.; Scozzafava, A.; McKenna, R.; Supuran, C.T. Dithiocarbamates: A new class of carbonic anhydrase inhibitors. Crystallographic and kinetic investigations. Chem. Commun. 2012, 48, 1868–1870. [Google Scholar] [CrossRef]
- Maresca, A.; Carta, F.; Vullo, D.; Supuran, C.T. Dithiocarbamates strongly inhibit the beta-class carbonic anhydrases from Mycobacterium tuberculosis. J. Enzym. Inhib. Med. Chem. 2013, 28, 407–411. [Google Scholar] [CrossRef] [PubMed]
- Khalifah, R.G. The carbon dioxide hydration activity of carbonic anhydrase. I. Stop-flow kinetic studies on the native human isoenzymes B and C. J. Biol. Chem. 1971, 246, 2561–2573. [Google Scholar]
- Del Prete, S.; Vullo, D.; De Luca, V.; Carginale, V.; di Fonzo, P.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Anion inhibition profiles of the complete domain of the eta-carbonic anhydrase from Plasmodium falciparum. Bioorg. Anicmed. Chem. 2016, 24, 4410–4414. [Google Scholar] [CrossRef] [PubMed]
- De Luca, V.; Vullo, D.; Del Prete, S.; Carginale, V.; Osman, S.M.; AlOthman, Z.; Supuran, C.T.; Capasso, C. Cloning, characterization and anion inhibition studies of a gamma-carbonic anhydrase from the Antarctic bacterium Colwellia psychrerythraea. Bioorg. Med. Chem. 2016, 24, 835–840. [Google Scholar] [CrossRef] [PubMed]
- De Simone, G.; Supuran, C.T. (In)organic anions as carbonic anhydrase inhibitors. J. Inorg. Biochem. 2012, 111, 117–129. [Google Scholar] [CrossRef] [PubMed]
- Fox, J.M.; Kang, K.; Sherman, W.; Héroux, A.; Sastry, G.M.; Baghbanzadeh, M.; Lockett, M.R.; Whitesides, G.M. Interactions between Hofmeister anions and the binding pocket of a protein. J. Am. Chem. Soc. 2015, 137, 3859–3866. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; Vullo, D.; Ghobril, C.; Hitce, J.; Clavaud, C.; Marat, X.; Capasso, C.; Supuran, C.T. Cloning, Purification, and Characterization of a beta-Carbonic Anhydrase from Malassezia restricta, an Opportunistic Pathogen Involved in Dandruff and Seborrheic Dermatitis. Int. J. Mol. Sci. 2019, 20, 2447. [Google Scholar] [CrossRef]
- Bradford, M.M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- De Luca, V.; Del Prete, S.; Supuran, C.T.; Capasso, C. Protonography, a new technique for the analysis of carbonic anhydrase activity. J. Enzym. Inhib. Med. Chem. 2015, 30, 277–282. [Google Scholar] [CrossRef]
- Del Prete, S.; De Luca, V.; Iandolo, E.; Supuran, C.T.; Capasso, C. Protonography, a powerful tool for analyzing the activity and the oligomeric state of the gamma-carbonic anhydrase identified in the genome of Porphyromonas gingivalis. Bioorg. Med. Chem. 2015, 23, 3747–3750. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; De Luca, V.; Supuran, C.T.; Capasso, C. Protonography, a technique applicable for the analysis of eta-carbonic anhydrase activity. J. Enzym. Inhib. Med. Chem. 2015, 30, 920–924. [Google Scholar] [CrossRef] [PubMed]
- Del Prete, S.; Vullo, D.; Caminiti-Segonds, N.; Zoccola, D.; Tambutte, S.; Supuran, C.T.; Capasso, C. Protonography and anion inhibition profile of the alpha-carbonic anhydrase (CruCA4) identified in the Mediterranean red coral Corallium rubrum. Bioorg. Chem. 2018, 76, 281–287. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: All the samples of the compounds are available from the authors. |


| CA-Class | Presence in Genome | Acronym | Number of Amino Acid Residues |
|---|---|---|---|
| α | Absent | - | - |
| β | Present | CynT | 219 |
| CynT2 | 220 | ||
| γ | Present | Ecoli_gamma | 256 |
| ι | Present | Ecoli_iota | 150 |
| Organisms | CA-Class | Acronym | kcat 1 (s−1) | kcat/KM 1 (M−1 × s−1) | KI (Acetazolamide) 1 (nM) |
|---|---|---|---|---|---|
| Homo sapiens | α | hCA I 2 | 2.0 × 105 | 5.0 × 107 | 250 |
| α | hCA II 2 | 1.4 × 106 | 1.5 × 108 | 12 | |
| Vibrio cholerae | α | VchCA_alpha 2 | 8.2 × 105 | 7.0 × 107 | 6.8 |
| β | VchCA_beta 2 | 3.3 × 105 | 4.1 × 107 | 451 | |
| γ | VchCA_gamma 2 | 7.3 × 105 | 6.4 × 107 | 473 | |
| Burkholderia territorii | ι | BteCAι 3 | 3.0 × 105 | 9.7 × 107 | 65 |
| Escherichia coli | β | CynT2 | 5.3 × 105 | 4.1 × 107 | 227 |
| Anion | KI (mM) 1 | |||
|---|---|---|---|---|
| hCA I 2 | hCA II 2 | CynT2 | VchCA_beta 2 | |
| F− | >300 | >300 | 9.4 | 8.7 |
| Cl− | 6 | 200 | 6.7 | 8.1 |
| Br− | 4 | 63 | 3.8 | 7.4 |
| I− | 0.3 | 26 | >10 | 9.0 |
| CNO− | 0.0007 | 0.03 | 0.58 | 7.1 |
| SCN− | 0.2 | 1.6 | 5.7 | 9.5 |
| CN− | 0.0005 | 0.02 | >10 | 5.7 |
| N3− | 0.0012 | 1.51 | >10 | 20.5 |
| NO2− | 8.4 | 63 | 4.9 | 9.1 |
| NO3− | 7 | 35 | 2.4 | 8.4 |
| HCO3− | 12 | 85 | 0.81 | 5.9 |
| CO32− | 15 | 73 | 0.89 | 6.7 |
| HSO3− | 18 | 89 | 3.7 | >200 |
| SO42− | 63 | >200 | 1.7 | >200 |
| HS− | 0.0006 | 0.04 | 2.7 | 21.3 |
| NH2SO2NH2 | 0.31 | 1.13 | 0.011 | 0.054 |
| NH2SO3H | 0.021 | 0.39 | 0.0025 | 0.086 |
| PhAsO3H2 | 31.7 | 49 | 0.0061 | 0.079 |
| PhB(OH)2 | 58.6 | 23 | 0.0028 | 0.085 |
| ClO4− | >200 | >200 | >100 | >200 |
| SnO32− | 0.57 | 0.83 | 0.52 | 3.1 |
| SeO42− | 118 | 112 | 3.1 | 3.4 |
| TeO42− | 0.66 | 0.92 | 0.51 | 2.3 |
| OsO52− | 0.92 | 0.95 | >10 | nt |
| P2O72− | 25.8 | 48 | >10 | 15.1 |
| V2O72− | 0.54 | 0.57 | >10 | 7.9 |
| B4O72− | 0.64 | 0.95 | 0.25 | 3.4 |
| ReO4− | 0.11 | 0.75 | >10 | 6.3 |
| RuO4− | 0.101 | 0.69 | 9.5 | 8.4 |
| S2O82− | 0.107 | 0.084 | 6.4 | 3.4 |
| SeCN− | 0.085 | 0.086 | 3.1 | 5.3 |
| NH(SO3)22− | 0.31 | 0.76 | 1.5 | >200 |
| FSO3− | 0.79 | 0.46 | 0.83 | 8.9 |
| CS32− | 0.0087 | 0.0088 | 3.1 | 7.0 |
| EtNCS2− | 0.00079 | 0.0031 | 0.084 | 0.73 |
| PF6− | - | - | >10 | - |
| CF3SO3− | - | - | >10 | - |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Del Prete, S.; De Luca, V.; Nocentini, A.; Scaloni, A.; Mastrolorenzo, M.D.; Supuran, C.T.; Capasso, C. Anion Inhibition Studies of the Beta-Carbonic Anhydrase from Escherichia coli. Molecules 2020, 25, 2564. https://doi.org/10.3390/molecules25112564
Del Prete S, De Luca V, Nocentini A, Scaloni A, Mastrolorenzo MD, Supuran CT, Capasso C. Anion Inhibition Studies of the Beta-Carbonic Anhydrase from Escherichia coli. Molecules. 2020; 25(11):2564. https://doi.org/10.3390/molecules25112564
Chicago/Turabian StyleDel Prete, Sonia, Viviana De Luca, Alessio Nocentini, Andrea Scaloni, Margaret D. Mastrolorenzo, Claudiu T. Supuran, and Clemente Capasso. 2020. "Anion Inhibition Studies of the Beta-Carbonic Anhydrase from Escherichia coli" Molecules 25, no. 11: 2564. https://doi.org/10.3390/molecules25112564
APA StyleDel Prete, S., De Luca, V., Nocentini, A., Scaloni, A., Mastrolorenzo, M. D., Supuran, C. T., & Capasso, C. (2020). Anion Inhibition Studies of the Beta-Carbonic Anhydrase from Escherichia coli. Molecules, 25(11), 2564. https://doi.org/10.3390/molecules25112564










