Deep Learning for Validating and Estimating Resolution of Cryo-Electron Microscopy Density Maps †
Abstract
1. Introduction
1.1. The Unresolved Problem of Resolution and Map Validation in Cryo-EM
1.2. Map Local Resolution Determination
1.3. Approach
2. Results
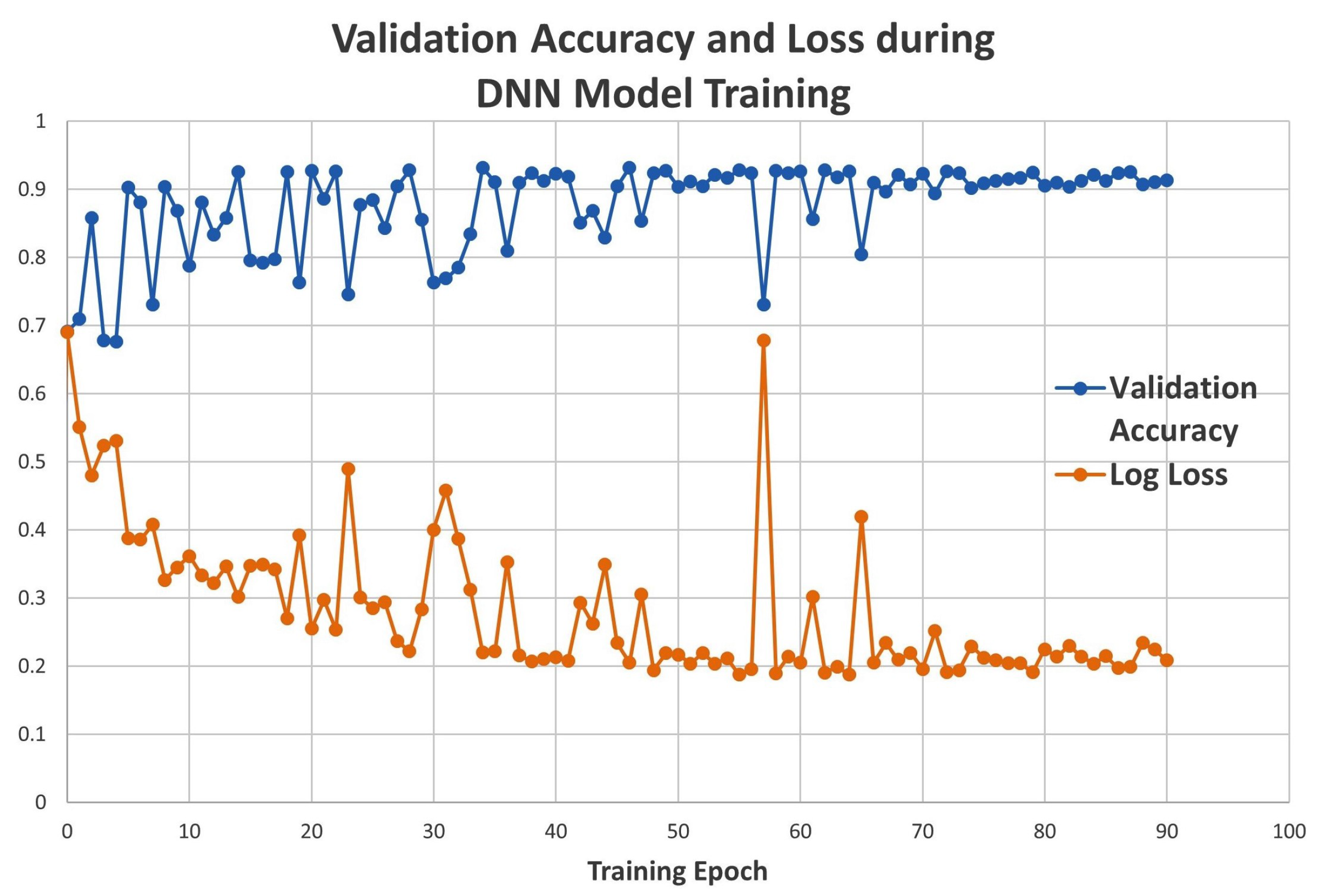
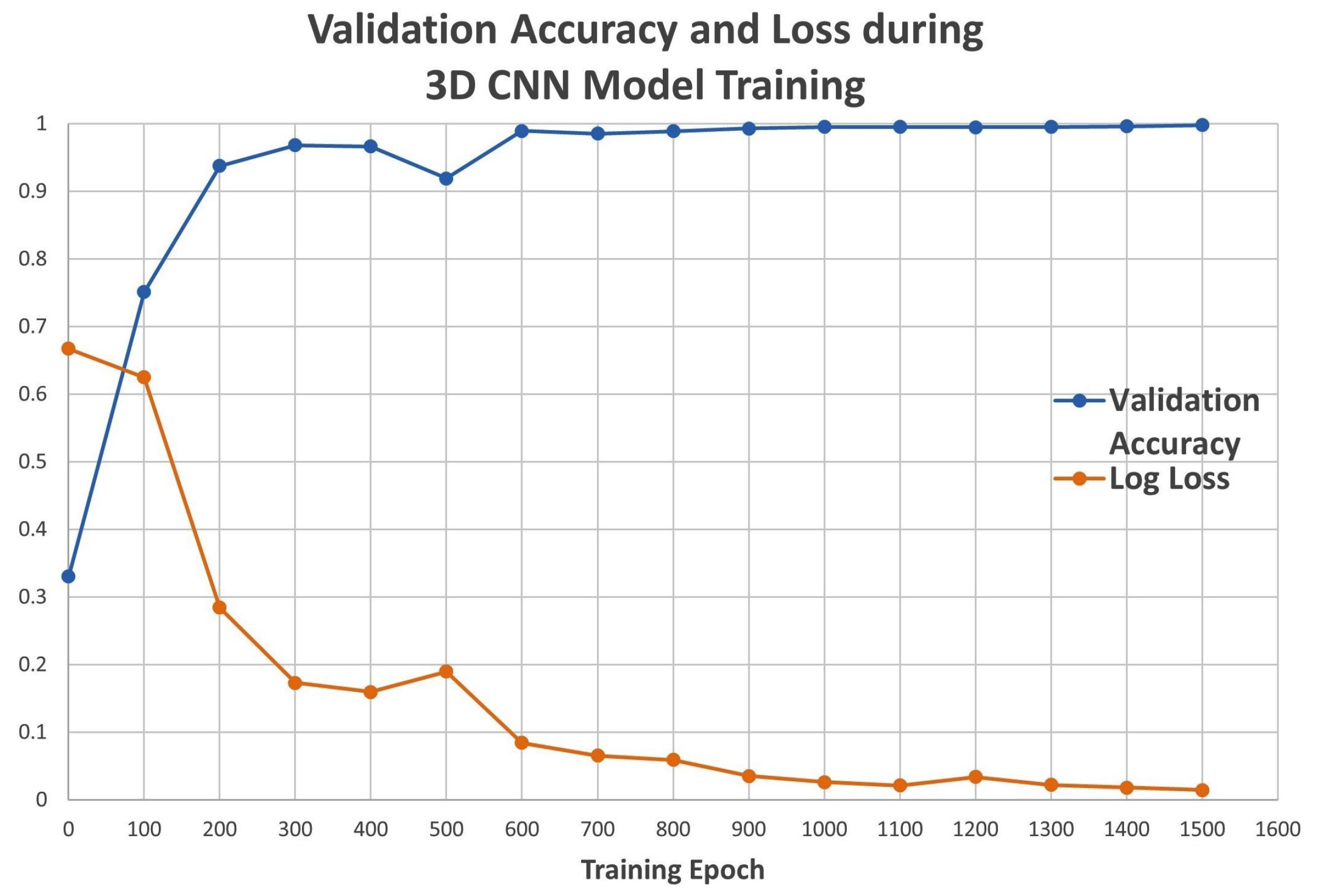
2.1. Model Metrics of DNN and 3D CNN Architectures
2.2. Performance of DNN and 3D CNN Models on Simulated Cryo-EM Density Maps at Different Resolutions
2.3. Performance of DNN and 3D CNN Models on Experimental Cryo-EM Density Maps
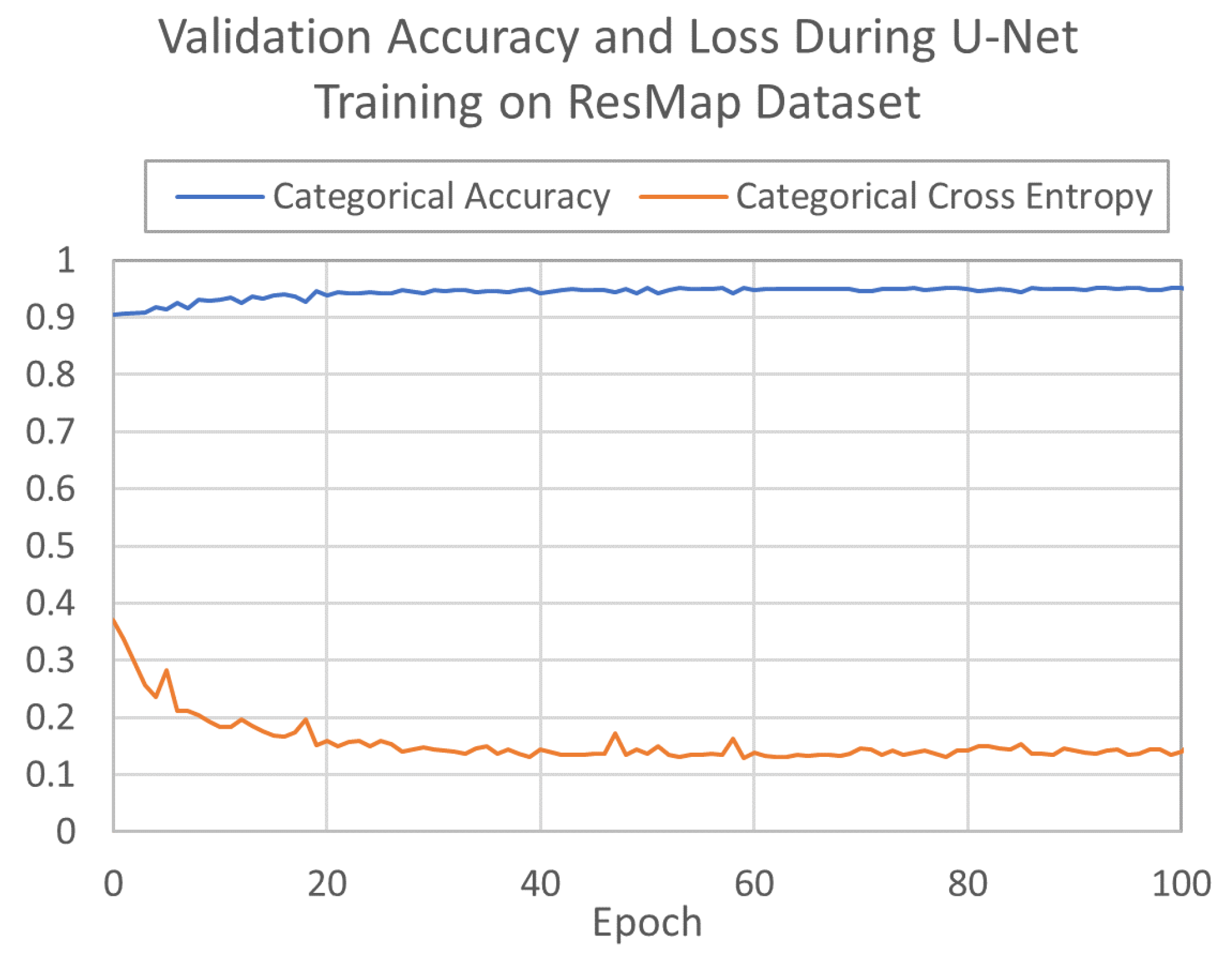
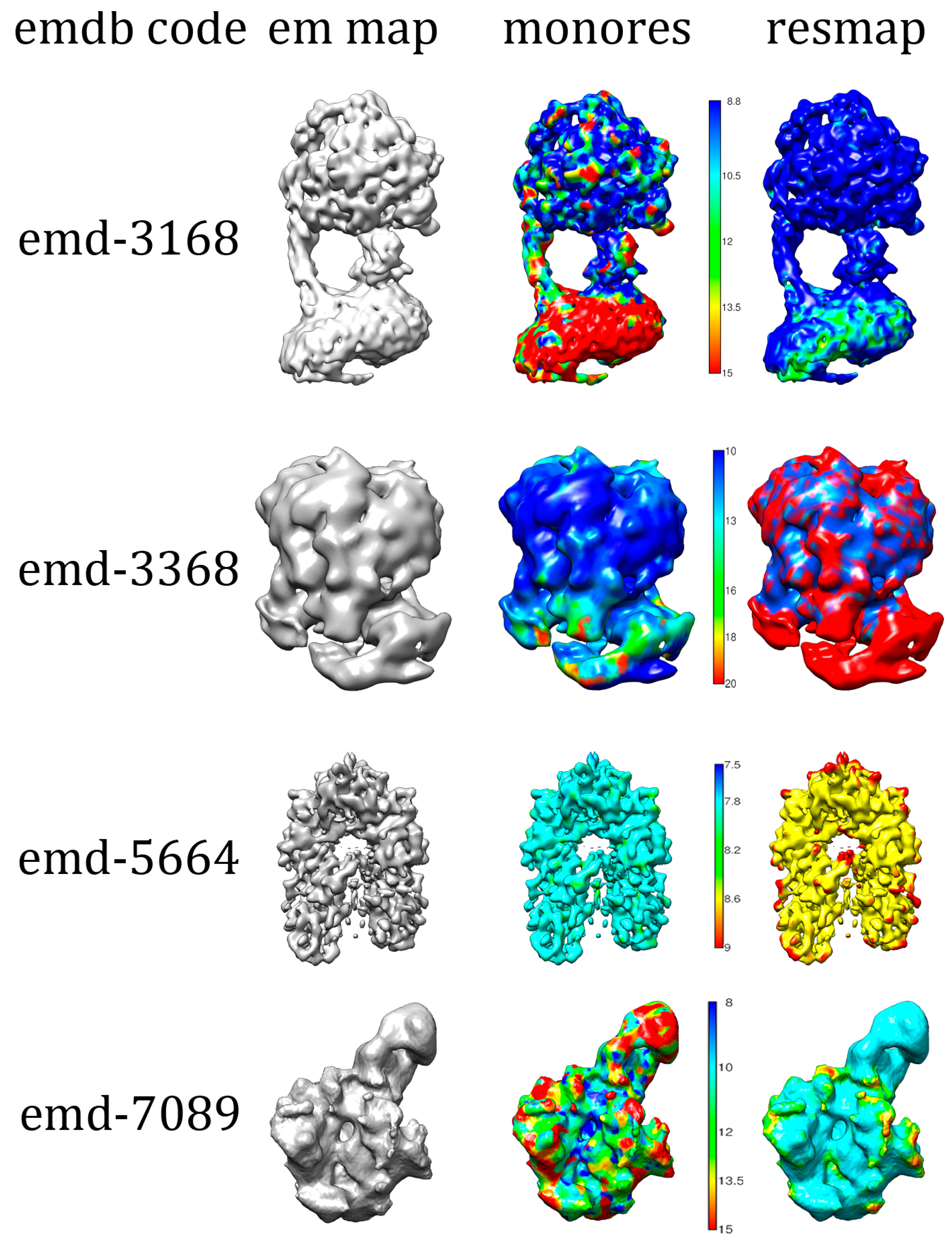
2.4. U-Net Model Performance and Metrics on Experimental Cryo-EM Density Maps
3. Discussion
4. Materials and Methods
4.1. Data Collection and Data Pre-Processing
4.2. Model Training
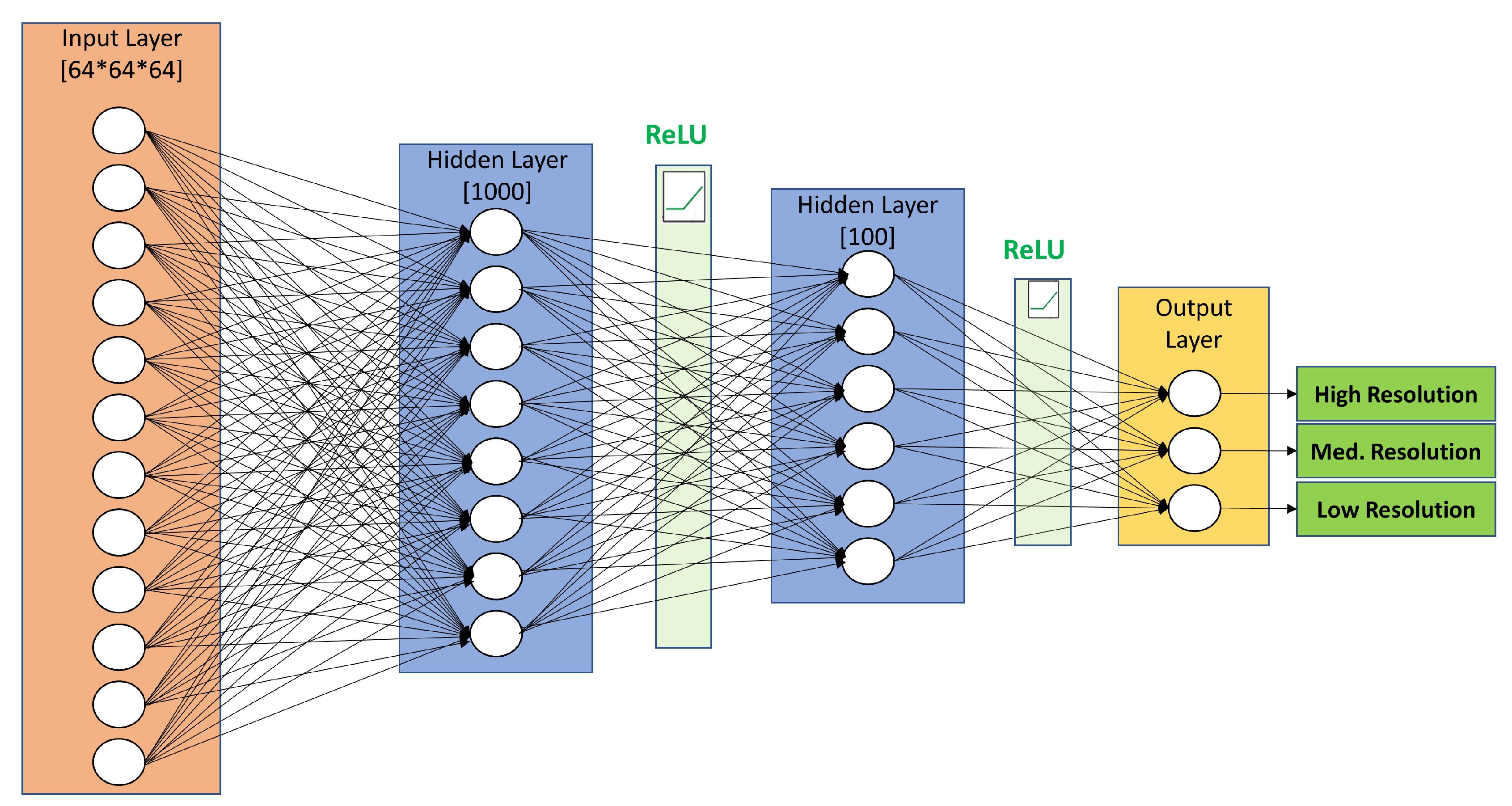
4.3. Dense Artificial Neural Network
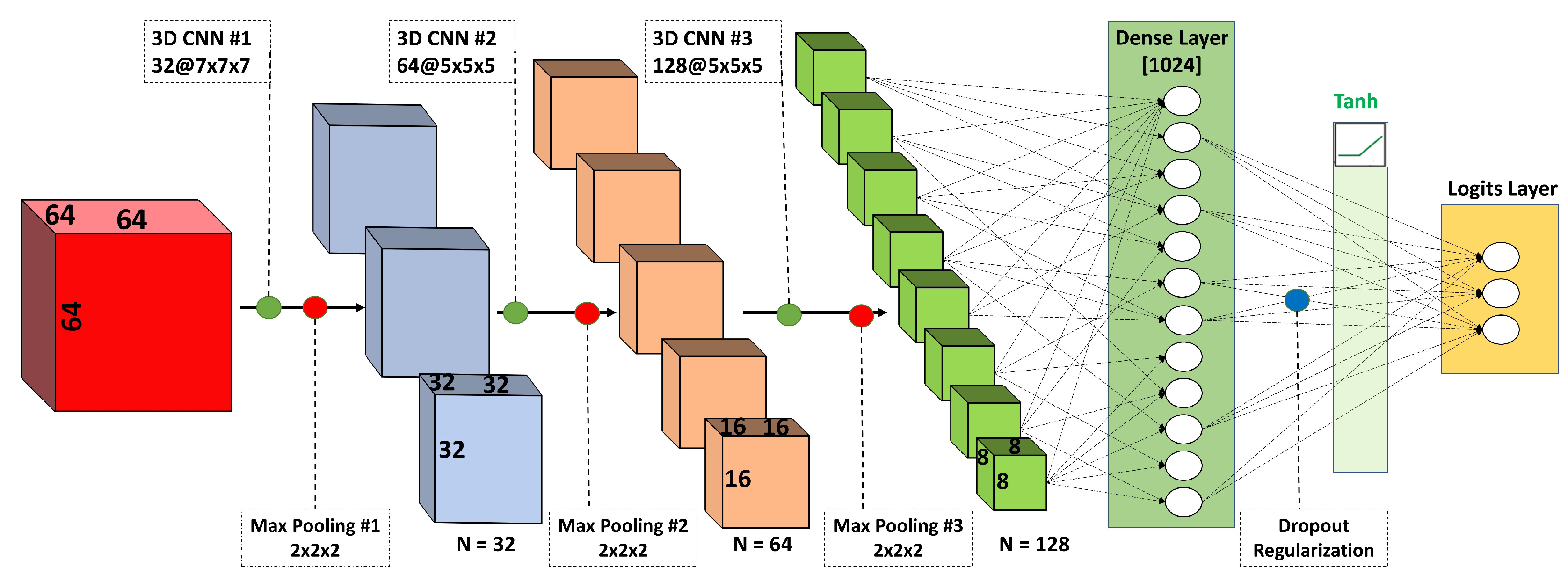
4.4. 3D Convolutional Neural Network
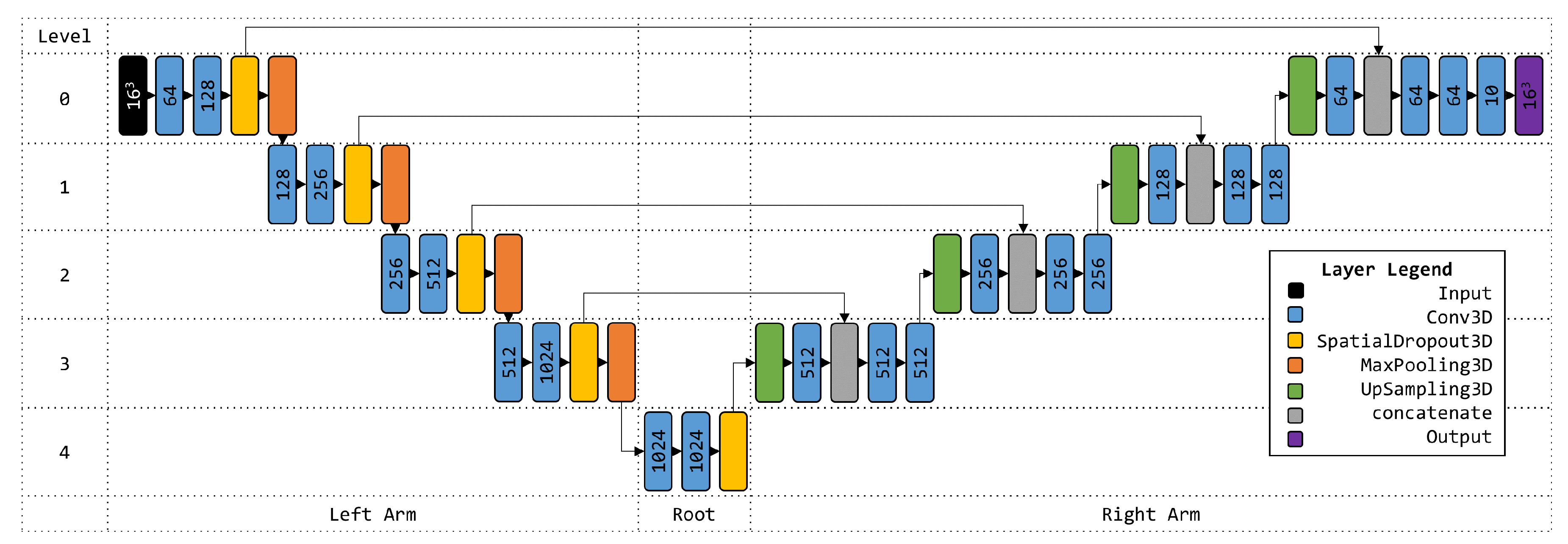
4.5. 3D U-Net Architecture
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| NMR | Nuclear Magnetic Resonance |
| cryo-EM | Cryo-Electron Microscopy |
| SSNR | Spectral Signal-to-Noise Ratio |
| FSC | Fourier Shell Correlation |
| SPA | Single Particle Analysis |
| DNN | Dense Neural Network |
| CNN | Convolutional Neural Network |
| PDB | Protein Data Bank |
| EMDB | Electron Microscopy Data Bank |
| kDa | KiloDalton |
| ReLU | Rectified Linear Units |
References
- Method of the Year 2015. Nat. Methods 2016, 13, 1. [CrossRef]
- Yusupova, G.; Yusupov, M. Ribosome biochemistry in crystal structure determination. RNA 2015, 21, 771–773. [Google Scholar] [CrossRef] [PubMed]
- Carpenter, E.P.; Beis, K.; Cameron, A.D.; Iwata, S. Overcoming the challenges of membrane protein crystallography. Curr. Opin. Struct. Biol. 2008, 18, 581–586. [Google Scholar] [CrossRef]
- Wang, L.; Sigworth, F.J. Cryo-EM and Single Particles. Physiology 2006, 21, 13–18. [Google Scholar] [CrossRef]
- Frank, J. Single-particle reconstruction of biological macromolecules in electron microscopy—30 years. Q. Rev. Biophys. 2009, 42, 139. [Google Scholar] [CrossRef]
- Kucukelbir, A.; Sigworth, F.J.; Tagare, H.D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 2014, 11, 63–65. [Google Scholar] [CrossRef] [PubMed]
- Vilas, J.L.; Gómez-Blanco, J.; Conesa, P.; Melero, R.; Miguel de la Rosa-Trevín, J.; Otón, J.; Cuenca, J.; Marabini, R.; Carazo, J.M.; Vargas, J.; et al. MonoRes: Automatic and Accurate Estimation of Local Resolution for Electron Microscopy Maps. Structure 2018, 26, 337–344.e4. [Google Scholar] [CrossRef]
- Cardone, G.; Heymann, J.B.; Steven, A.C. One number does not fit all: Mapping local variations in resolution in cryo-EM reconstructions. J. Struct. Biol. 2013, 184, 226–236. [Google Scholar] [CrossRef]
- Amunts, A.; Brown, A.; Toots, J.; Scheres, S.H.W.; Ramakrishnan, V. The structure of the human mitochondrial ribosome. Science 2015, 348, 95–98. [Google Scholar] [CrossRef]
- Khatter, H.; Myasnikov, A.G.; Natchiar, S.K.; Klaholz, B.P. Structure of the human 80S ribosome. Nature 2015, 520, 640–645. [Google Scholar] [CrossRef]
- Liu, Z.; Guo, F.; Wang, F.; Li, T.C.; Jiang, W. 2.9 Å Resolution Cryo-EM 3D Reconstruction of Close-Packed Virus Particles. Structure 2016, 24, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Kuhlbrandt, W. The Resolution Revolution. Science 2014, 343, 1443–1444. [Google Scholar] [CrossRef] [PubMed]
- Callaway, E. The revolution will not be crystallized: A new method sweeps through structural biology. Nature 2015, 525, 172–174. [Google Scholar] [CrossRef] [PubMed]
- Adrian, M.; Dubochet, J.; Lepault, J.; McDowall, A.W. Cryo-electron microscopy of viruses. Nature 1984, 308, 32–36. [Google Scholar] [CrossRef] [PubMed]
- Kühlbrandt, W. Cryo-EM enters a new era. eLife 2014, 3, e03678. [Google Scholar] [CrossRef] [PubMed]
- Scheres, S.H. Beam-induced motion correction for sub-megadalton cryo-EM particles. eLife 2014, 3, e03665. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Mooney, P.; Zheng, S.; Booth, C.R.; Braunfeld, M.B.; Gubbens, S.; Agard, D.A.; Cheng, Y. Electron counting and beam-induced motion correction enable near-atomic-resolution single-particle cryo-EM. Nat. Methods 2013, 10, 584–590. [Google Scholar] [CrossRef]
- Cressey, D.; Callaway, E. Cryo-electron microscopy wins chemistry Nobel. Nature 2017, 550, 167. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Hattne, J.; Reyes, F.E.; Sanchez-Martinez, S.; Jason de la Cruz, M.; Shi, D.; Gonen, T. Atomic resolution structure determination by the cryo-EM method MicroED: Atomic Resolution Structure Determination. Protein Sci. 2017, 26, 8–15. [Google Scholar] [CrossRef]
- Lawson, C.L.; Baker, M.L.; Best, C.; Bi, C.; Dougherty, M.; Feng, P.; van Ginkel, G.; Devkota, B.; Lagerstedt, I.; Ludtke, S.J.; et al. EMDataBank.org: Unified data resource for CryoEM. Nucleic Acids Res. 2011, 39, D456–D464. [Google Scholar] [CrossRef]
- Van Heel, M.; Schatz, M. REASSESSING THE REVOLUTIONS RESOLUTIONS. bioRxiv 2017. [Google Scholar] [CrossRef]
- Penczek, P.A. Resolution measures in molecular electron microscopy. Methods Enzymol. 2010, 482, 73–100. [Google Scholar] [CrossRef]
- Zhou, Z.H. Towards atomic resolution structural determination by single-particle cryo-electron microscopy. Curr. Opin. Struct. Biol. 2008, 18, 218–228. [Google Scholar] [CrossRef]
- Liao, H.Y.; Frank, J. Definition and Estimation of Resolution in Single-Particle Reconstructions. Structure 2010, 18, 768–775. [Google Scholar] [CrossRef]
- Banterle, N.; Bui, K.H.; Lemke, E.A.; Beck, M. Fourier ring correlation as a resolution criterion for super-resolution microscopy. J. Struct. Biol. 2013, 183, 363–367. [Google Scholar] [CrossRef]
- Van Heel, M.; Schatz, M. Fourier shell correlation threshold criteria. J. Struct. Biol. 2005, 151, 250–262. [Google Scholar] [CrossRef]
- Scheres, S.H.W.; Chen, S. Prevention of overfitting in cryo-EM structure determination. Nat. Methods 2012, 9, 853–854. [Google Scholar] [CrossRef] [PubMed]
- De la Rosa-Trevín, J.; Quintana, A.; del Cano, L.; Zaldívar, A.; Foche, I.; Gutiérrez, J.; Gómez-Blanco, J.; Burguet-Castell, J.; Cuenca-Alba, J.; Abrishami, V.; et al. Scipion: A software framework toward integration, reproducibility and validation in 3D electron microscopy. J. Struct. Biol. 2016, 195, 93–99. [Google Scholar] [CrossRef] [PubMed]
- Tang, G.; Peng, L.; Baldwin, P.R.; Mann, D.S.; Jiang, W.; Rees, I.; Ludtke, S.J. EMAN2: An extensible image processing suite for electron microscopy. J. Struct. Biol. 2007, 157, 38–46. [Google Scholar] [CrossRef] [PubMed]
- Beleites, C.; Salzer, R.; Sergo, V. Validation of soft classification models using partial class memberships: An extended concept of sensitivity & co. applied to grading of astrocytoma tissues. Chemom. Intell. Lab. Syst. 2013, 122, 12–22. [Google Scholar] [CrossRef]
- Shi, D.; Nannenga, B.L.; Iadanza, M.G.; Gonen, T. Three-dimensional electron crystallography of protein microcrystals. eLife 2013, 2, e01345. [Google Scholar] [CrossRef]
- Berman, H.M. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Shaikh, T.R.; Gao, H.; Baxter, W.T.; Asturias, F.J.; Boisset, N.; Leith, A.; Frank, J. SPIDER image processing for single-particle reconstruction of biological macromolecules from electron micrographs. Nat. Protoc. 2008, 3, 1941–1974. [Google Scholar] [CrossRef] [PubMed]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Couch, G.S.; Greenblatt, D.M.; Meng, E.C.; Ferrin, T.E. UCSF Chimera?A visualization system for exploratory research and analysis. J. Comput. Chem. 2004, 25, 1605–1612. [Google Scholar] [CrossRef]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Advances in Neural Information Processing Systems 25; Pereira, F., Burges, C.J.C., Bottou, L., Weinberger, K.Q., Eds.; Curran Associates, Inc.: Red Hook, NY, USA, 2012; pp. 1097–1105. [Google Scholar]
- Zeiler, M.D.; Fergus, R. Visualizing and Understanding Convolutional Networks. In Computer Vision—ECCV 2014; Fleet, D., Pajdla, T., Schiele, B., Tuytelaars, T., Eds.; Springer International Publishing: Cham, Switzerland, 2014; Volume 8689, pp. 818–833. [Google Scholar] [CrossRef]
- Clevert, D.A.; Unterthiner, T.; Hochreiter, S. Fast and Accurate Deep Network Learning by Exponential Linear Units (ELUs). arXiv, 2015; arXiv:1511.07289. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. arXiv, 2015; arXiv:1505.04597. [Google Scholar]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S.S.; Brox, T.; Ronneberger, O. 3D U-Net: Learning Dense Volumetric Segmentation from Sparse Annotation. arXiv, 2016; arXiv:1606.06650. [Google Scholar]
- Qi, C.R.; Su, H.; Mo, K.; Guibas, L.J. PointNet: Deep Learning on Point Sets for 3D Classification and Segmentation. arXiv, 2016; arXiv:1612.00593. [Google Scholar]
Sample Availability: A repository of accompanying computer code and data is available online at https://github.com/DrDongSi/3DEM-Resolution-Validation. |

| Simulated Resolution | DNN | 3D CNN | Simulated Resolution | DNN | 3D CNN |
|---|---|---|---|---|---|
| 1.5 Å | High | High | 8.5 Å | Medium | Medium |
| 2.0 Å | High | High | 9.0 Å | Low | Low |
| 2.5 Å | High | High | 9.5 Å | Low | Low |
| 3.0 Å | High | High | 10.0 Å | Low | Low |
| 3.5 Å | High | High | 10.5 Å | Medium | Low |
| 4.0 Å | High | High | 11.0 Å | Low | Low |
| 4.5 Å | Medium | High | 11.5 Å | Low | Low |
| 5.0 Å | Medium | Medium | 12.0 Å | Low | Low |
| 5.5 Å | Medium | Medium | 12.5 Å | Low | Low |
| 6.0 Å | Medium | Medium | 13.0 Å | Low | Low |
| 6.5 Å | Medium | Medium | 13.5 Å | Low | Low |
| 7.0 Å | Medium | Medium | 14.0 Å | Low | Low |
| 7.5 Å | Low | Medium | 14.5 Å | Low | Low |
| 8.0 Å | Medium | Medium | 15.0 Å | Low | Low |
| EMDB ID | Resolution | DNN | 3D CNN | EMDB ID | Resolution | DNN | 3D CNN |
|---|---|---|---|---|---|---|---|
| 8221 | 1.6 Å | High | High | 5751 | 8.0 Å | High | Medium |
| 2984 | 2.2 Å | High | High | 3340 | 7.2 Å | Medium | Medium |
| 2945 | 2.9 Å | High | High | 5101 | 8.0 Å | Low | Low |
| 6342 | 2.5 Å | High | High | 3168 | 7.4 Å | Medium | Medium |
| 8218 | 2.3 Å | High | High | 5653 | 7.3 Å | High | High |
| 8222 | 2.5 Å | High | High | 7089 | 13.2 Å | Low | Low |
| 8077 | 1.75 Å | High | High | 1601 | 14.1 Å | High | High |
| 8217 | 1.8 Å | High | High | 6013 | 14.8 Å | Medium | Medium |
| 8219 | 2.5 Å | High | High | 3368 | 13.0 Å | Medium | Medium |
| 6313 | 2.5 Å | High | High | 2862 | 12.5 Å | Low | Low |
| 3311 | 6.7 Å | Medium | High | 3098 | 11.0 Å | Medium | Medium |
| 5664 | 7.8 Å | High | High | 7471 | 12.5 Å | Medium | High |
| 6410 | 7.8 Å | High | High | 1711 | 13.0 Å | Low | Low |
| 4047 | 6.0 Å | Medium | High | 5169 | 11.0 Å | Medium | Medium |
| 1674 | 6.0 Å | Medium | Medium | 1981 | 14.2 Å | High | High |
| DNN Confusion Matrix | Predicted Resolution | |||
|---|---|---|---|---|
| High | Medium | Low | ||
| Published Resolution | High | 10 | 0 | 0 |
| Medium | 4 | 5 | 1 | |
| Low | 2 | 5 | 3 | |
| 3D CNN Confusion Matrix | Predicted Resolution | |||
|---|---|---|---|---|
| High | Medium | Low | ||
| Published Resolution | High | 10 | 0 | 0 |
| Medium | 5 | 4 | 1 | |
| Low | 3 | 4 | 3 | |
| Sensitivity | Specificity | PPV | NPV | ||
|---|---|---|---|---|---|
| DNN Model | High | 100.00% | 70.00% | 62.50% | 100.00% |
| Medium | 50.00% | 75.00% | 50.00% | 75.00% | |
| Low | 30.00% | 95.00% | 75.00% | 73.10% | |
| CNN Model | High | 100.00% | 60.00% | 55.60% | 100.00% |
| Medium | 40.00% | 80.00% | 50.00% | 72.70% | |
| Low | 30.00% | 95.00% | 75.00% | 73.10% | |
| U-Net Evaluation Accuracies | Model | ||
|---|---|---|---|
| MonoRes | ResMap | ||
| Set | MonoRes | 88.3% | 59.4% |
| ResMap | 88.9% | 94.7% | |
| Set | MonoRes | ResMap |
|---|---|---|
| Train | 9670 | 1400 |
| Validation | 1080 | 159 |
| Evaluation | 2694 | 393 |
| Total | 13,444 | 1952 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Avramov, T.K.; Vyenielo, D.; Gomez-Blanco, J.; Adinarayanan, S.; Vargas, J.; Si, D. Deep Learning for Validating and Estimating Resolution of Cryo-Electron Microscopy Density Maps †. Molecules 2019, 24, 1181. https://doi.org/10.3390/molecules24061181
Avramov TK, Vyenielo D, Gomez-Blanco J, Adinarayanan S, Vargas J, Si D. Deep Learning for Validating and Estimating Resolution of Cryo-Electron Microscopy Density Maps †. Molecules. 2019; 24(6):1181. https://doi.org/10.3390/molecules24061181
Chicago/Turabian StyleAvramov, Todor Kirilov, Dan Vyenielo, Josue Gomez-Blanco, Swathi Adinarayanan, Javier Vargas, and Dong Si. 2019. "Deep Learning for Validating and Estimating Resolution of Cryo-Electron Microscopy Density Maps †" Molecules 24, no. 6: 1181. https://doi.org/10.3390/molecules24061181
APA StyleAvramov, T. K., Vyenielo, D., Gomez-Blanco, J., Adinarayanan, S., Vargas, J., & Si, D. (2019). Deep Learning for Validating and Estimating Resolution of Cryo-Electron Microscopy Density Maps †. Molecules, 24(6), 1181. https://doi.org/10.3390/molecules24061181






