Analysis of Metabolite Accumulation Related to Pod Color Variation of Caragana intermedia
Abstract
1. Introduction
2. Results
2.1. Multivariate Analysis of Metabolites Related to Pod Color and Growth Stage
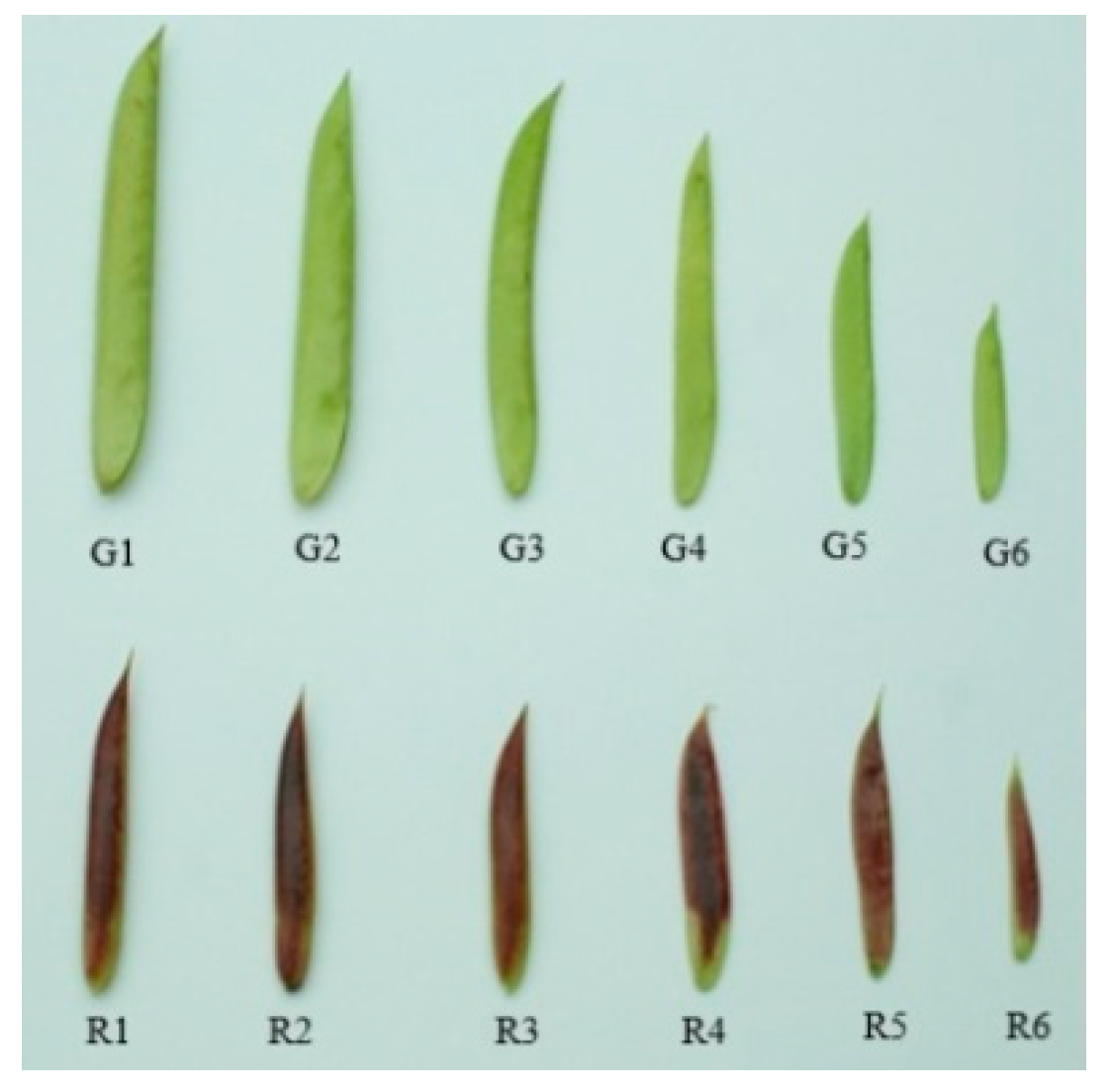
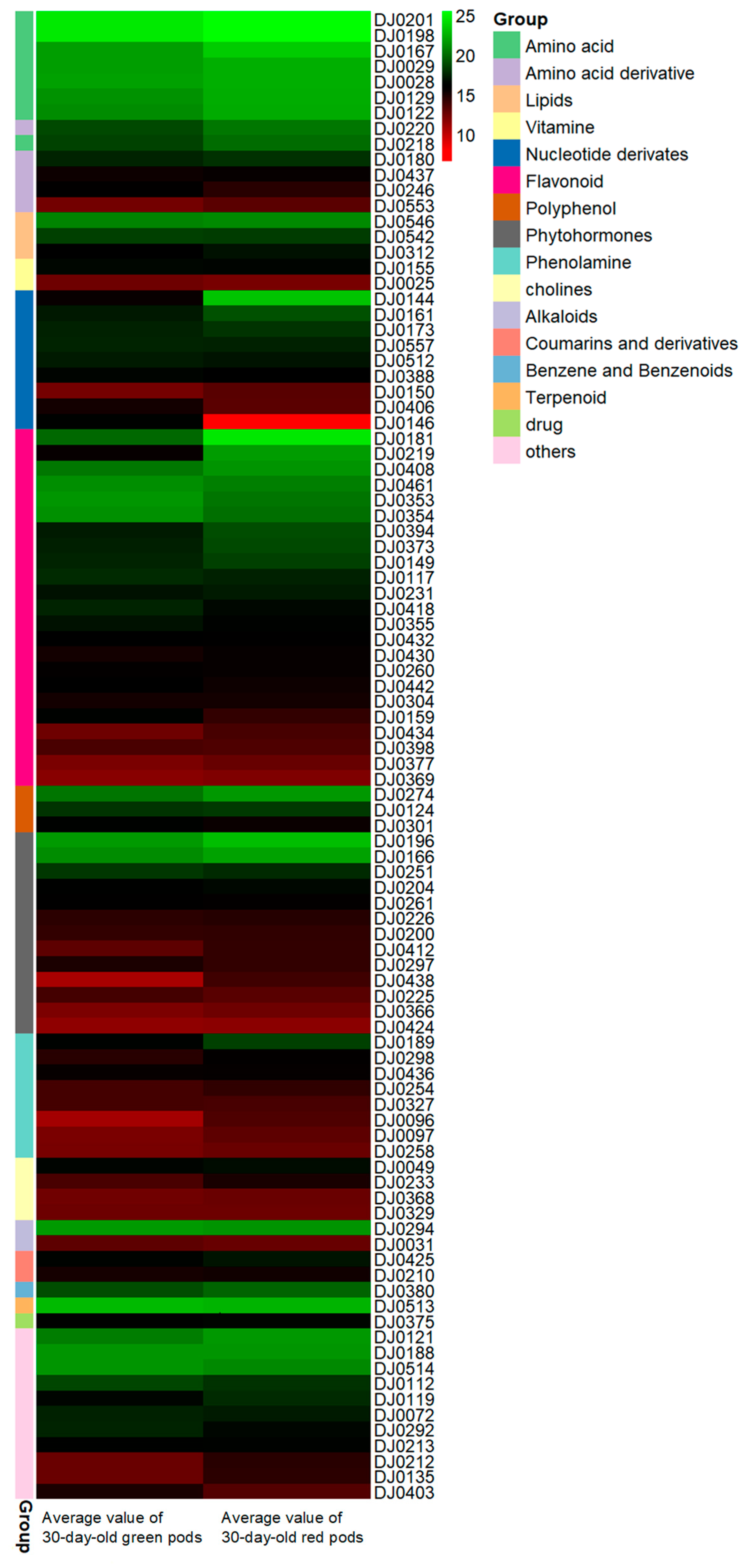
2.2. Differential Metabolite Profiling in Red and Green Pods of C. intermedia
2.3. Analysis of Metabolite Accumulation Patterns in Red and Green Pods of C. intermedia
3. Discussion
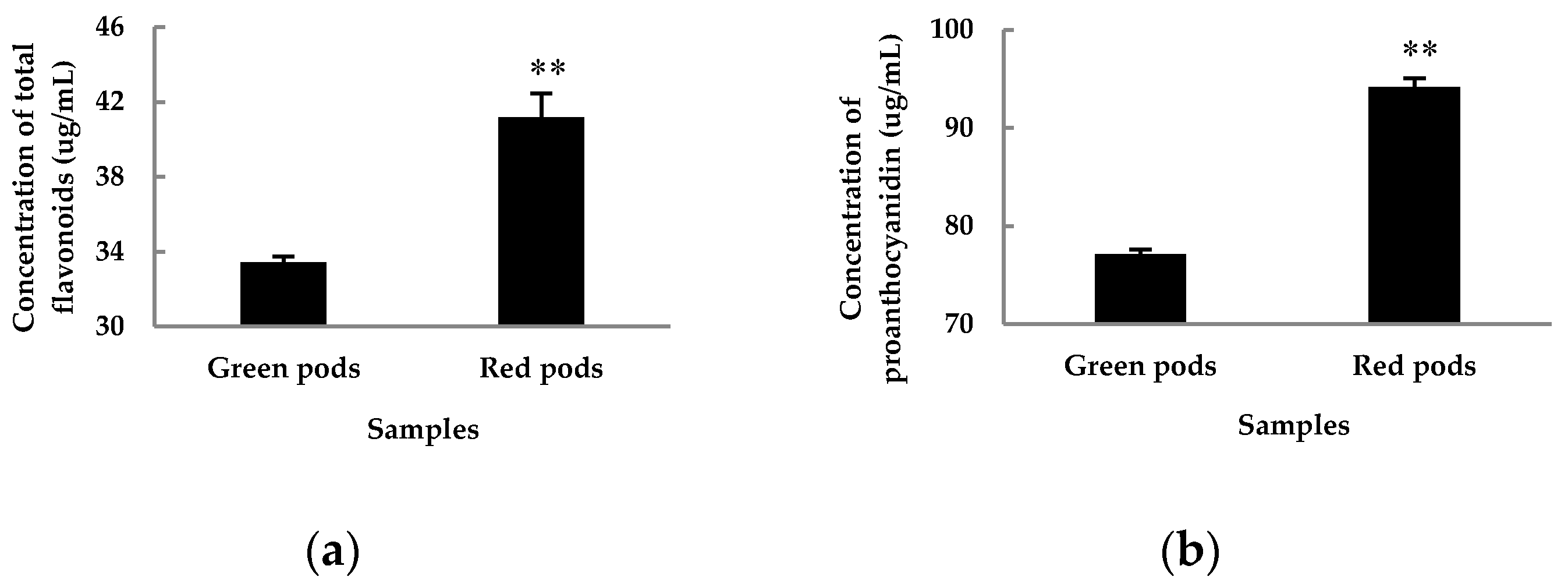
3.1. Flavonoids Are Highly Correlated with Pod Color
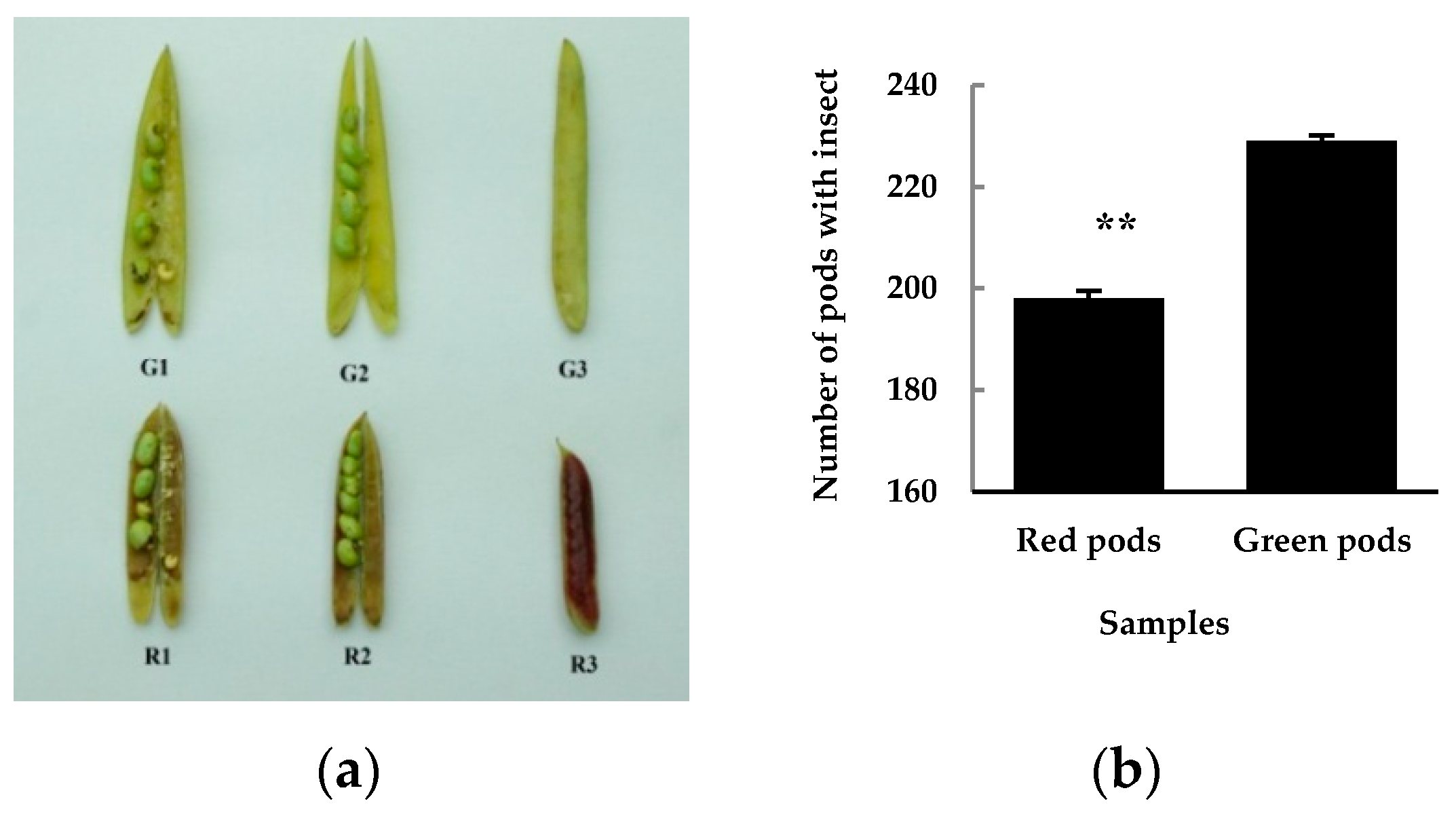
3.2. Differential Insect Feeding Rate in Red and Green Pods of C. intermedia
3.3. Possible Metabolic Regulation Mechanism of Pod Color Variation in C. intermedia
4. Materials and Methods
4.1. Plant Materials
4.2. Sample Preparation and Extraction
4.3. High Performance Liquid Chromatography (HPLC) Conditions
4.4. ESI-Q TRAP-MS/MS
4.5. Metabolite Identification and Quantification
4.6. Statistical Analysis
4.7. Determination of Total Flavonoids
4.8. Determination of Proanthocyanidin
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wan, Y.Q.; Mao, M.Z.; Wan, D.L.; Yang, Q.; Yang, F.Y.; Mandlaa; Li, G.J.; Wang, R.G. Identification of the WRKY gene family and functional analysis of two genes in Caragana intermedia. BMC Plant Biol. 2018, 18, 31. [Google Scholar] [CrossRef] [PubMed]
- Yang, Q.; Yin, J.J.; Li, G.; Qi, L.W.; Yang, F.Y.; Wang, R.G.; Li, G.J. Reference gene selection for qRT-PCR in Caragana korshinskii Kom. under different stress conditions. Mol. Biol. Rep. 2014, 41, 2325–2334. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.M.; Dong, J.; Liu, Y.; Gao, H.W. A novel dehydration-responsive element-binding protein from Caragana korshinskii is involved in the response to multiple abiotic stresses and enhances stress tolerance in transgenic tobacco. Plant Mol. Biol. Rep. 2010, 28, 664–675. [Google Scholar] [CrossRef]
- Zhu, J.F.; Li, W.F.; Yang, W.H.; Qi, L.W.; Han, S.Y. Identification of microRNAs in Caragana intermedia by high-throughput sequencing and expression analysis of 12 microRNAs and their targets under salt stress. Plant Cell Rep. 2013, 32, 1339–1349. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.W.; Chen, G.X.; Zhao, P.S.; Zhou, Q.; Zhao, X. The abundance of certain metabolites responds to drought stress in the highly drought tolerant plant Caragana korshinskii. Acta Physiol. Plant. 2017, 39, 116. [Google Scholar] [CrossRef]
- Cui, Y.Y.; Yang, C.X.; Yang, X.D.; Yan, X.P. Zeolitic imidazolate framework-8 for selective extraction of a highly active anti-oxidant flavonoid from Caragana Jubata. J. Chromatogr. A 2018, 1544, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Zheng, C.J.; Wang, L.; Han, T.; Xin, H.L.; Jiang, Y.P.; Pan, L.; Jia, X.G.; Qin, L.P. Pruinosanones A-C, antiinflammatory isoflavone derivatives from Caragana pruinosa. Sci. Rep. 2016, 6, 31743. [Google Scholar] [CrossRef] [PubMed]
- He, Q.S.; Zhang, L.; Fan, Z.Y.; Feng, G.; Wang, F.J.; Liu, Z.Q.; Tang, T.; Kuang, S.X. Protective effects of total flavonoids in Caragana against hypoxia/reoxygenation- induced injury in human brain microvascular endothelial cells. Biomed. Pharmacother. 2017, 89, 316–322. [Google Scholar] [CrossRef] [PubMed]
- Peng, Q.Q.; Wang, K.; Cheng, K.J.; Yang, H.G.; Qiu, J.G.; Zhang, W.J.; Jiang, Q.W.; Yang, Y.; Zheng, D.W.; Huang, J.R.; et al. Caragaphenol A induces reactive oxygen species related apoptosis in human gastric cancer cells. Am. J. Transl. Res. 2017, 9, 3804–3815. [Google Scholar]
- Luan, G.X.; Tie, F.F.; Yuan, Z.Z.; Li, G.; He, J.; Wang, Z.H.; Wang, H.L. Hypaphorine, an indole alkaloid isolated from Caragana korshinskii Kom., inhibites 3T3-L1 adipocyte differentiation and improves insulin sensitivity in vitro. Chem. Biodivers. 2017, 14. [Google Scholar] [CrossRef]
- Ma, K.Q.; Miao, Y.H.; Gao, Y.; Tian, J.S.; Gao, L.; Ye, D.Y.; Qin, X.M. Increasing the Level of IRS-1 and Insulin Pathway Sensitivity by Natural Product Carainterol A. Molecules 2016, 21, 1303. [Google Scholar] [CrossRef] [PubMed]
- Haberer, G.; Kieber, J.J. Cytokinins. New insights into a classic phytohormone. Plant Physiol. 2002, 128, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Nishiyama, R.; Watanabe, Y.; Fujita, Y.; Le, D.T.; Kojima, M.; Werner, T.; Vankova, R.; Yamaguchi-Shinozaki, K.; Shinozaki, K.; Kakimoto, T.; et al. Analysis of cytokinin mutants and regulation of cytokinin metabolic genes reveals important regulatory roles of cytokinins in drought, salt and abscisic acid responses, and abscisic acid biosynthesis. Plant Cell 2011, 23, 2169–2183. [Google Scholar] [CrossRef] [PubMed]
- Hichri, I.; Barrieu, F.; Bogs, J.; Kappel, C.; Delrot, S.; Lauvergeat, V. Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J. Exp. Bot. 2011, 62, 2465–2483. [Google Scholar] [CrossRef] [PubMed]
- Dixon, R.A.; Xie, D.Y.; Sharma, S.B. Proanthocyanidins—Afinal frontier in flavonoid research? New Phytol. 2005, 165, 9–28. [Google Scholar] [CrossRef] [PubMed]
- Hirasuna, T.J.; Shuler, M.L.; Lackney, V.K.; Spanswick, R.M. Enhanced anthocyanin production in grape cell cultures. Plant Sci. 1991, 78, 107–120. [Google Scholar] [CrossRef]
- Meyer, H.J.; Staden, J.V. The in vitro production of an anthocyanin from callus cultures of Oxalis linearis. Plant Cell Tiss. Org. 1995, 40, 55–58. [Google Scholar] [CrossRef]
- Maharik, N.; Elgengaihi, S.; Taha, H. Anthocyanin production in callus cultures of Crataegus Sinaica Boiss. Int. J. Acad. Res. 2009, 1, 30–34. [Google Scholar]
- Ozeki, Y.; Komamine, A. Induction of anthocyanin synthesis in relation toembryogenesis in a carrot suspension culture: Correlation ofmetabolic differentiation with morphological differentiation. Physiol. Plantarum 1981, 53, 570–577. [Google Scholar] [CrossRef]
- Ozeki, Y.; Komamine, A. Effects of growth regulators on the induction of anthocyanin synthesis in carrot suspension cultures. Plant Cell Physiol. 1986, 27, 1361–1368. [Google Scholar] [CrossRef]
- Wang, Y.C.; Wang, N.; Xu, H.F.; Jiang, S.H.; Fang, H.C.; Su, M.Y.; Zhang, Z.Y.; Zhang, T.L.; Chen, X.S. Auxin regulates anthocyanin biosynthesis through the Aux/IAA–ARF signaling pathway in apple. Hortic. Res. 2018, 5, 59. [Google Scholar] [CrossRef] [PubMed]
- KEGG: Kyoto Encyclopedia of Genes and Genomes. Available online: http://kegg.jp/ (accessed on 16 February 2019).
- Xu, W.J.; Dubos, C.; Lepiniec, L. Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci. 2015, 20, 176–185. [Google Scholar] [CrossRef] [PubMed]
- Nakabayashi, R.; Yonekura-Sakakibara, K.; Urano, K.; Suzuki, M.; Yamada, Y.; Nishizawa, T.; Matsuda, F.; Kojima, M.; Sakakibara, H.; Shinozaki, K.; et al. Enhancement of oxidative and drought tolerance in Arabidopsis by overaccumulation of antioxidant flavonoids. Plant J. 2014, 77, 367–379. [Google Scholar] [CrossRef] [PubMed]
- Gould, K.S. Nature’s Swiss Army Knife: The diverse protective roles of anthocyanins in leaves. J. Biomed. Biotechnol. 2004, 5, 314–320. [Google Scholar] [CrossRef] [PubMed]
- Lane, G.A.; Sutherland, O.R.W.; Skipp, R.A. Isoflavonoids as insect feeding deterents and antifungal components from root of Lupinus augustifolius. J. Chem. Ecol. 1987, 13, 771–783. [Google Scholar] [CrossRef] [PubMed]
- Jackowski, J.; Popłoński, J.; Twardowska, K.; Magiera-Dulewicz, J.; Hurej, M.; Huszcza, E. Deterrent activity of hops flavonoids and their derivatives against stored product pests. Bull. Entomol. Res. 2017, 107, 592–597. [Google Scholar] [CrossRef] [PubMed]
- Osugi, A.; Sakakibara, H. Q&A: How do plants respond to cytokinins and what is their importance? BMC Biol. 2015, 13, 102. [Google Scholar]
- European MassBank (NORMAN MassBank). Available online: http://www.massbank.jp/ (accessed on 16 February 2019).
- KNApSAcK: A Comprehensive Species-Metabolite Relationship Database. Available online: http:// kanaya.naist.jp/KNApSAcK/ (accessed on 16 February 2019).
- HMDB Version 4.0. Available online: http://www.hmdb.ca/ (accessed on 16 February 2019).
- Moto V 1.0. Available online: http://www.ab.wur.nl/moto/ (accessed on 16 February 2019).
- METLIN: Metabolite Link. Available online: http://metlin.scripps.edu/ (accessed on 16 February 2019).
- Chen, W.; Gong, L.; Guo, Z.L.; Wang, W.S.; Zhang, H.Y.; Liu, X.Q.; Yu, S.B.; Xiong, L.Z.; Luo, J. A novel integrated method for large-scale detection, identification, and quantification of widely targeted metabolites: Application in the study of rice metabolomics. Mol. Plant 2013, 6, 1769–1780. [Google Scholar] [CrossRef]
- Chukwumah, Y.; Walker, L.T.; Verghese, M. Peanut skin color: A biomarker for total polyphenolic content and antioxidative capacities of peanut cultivars. Int. J. Mol. Sci. 2009, 10, 4941–4952. [Google Scholar] [CrossRef]
- Mitsunaga, T.; Doi, T.; Kondo, Y.; Abe, I. Color development of proanthocyanidins in vanillin- hydrochloric acid reaction. J. Wood Sci. 1998, 44, 125–130. [Google Scholar] [CrossRef]
Sample Availability: Not available. |



| Index | Component Name | Peak Area of Green Pods | Peak Area of Red Pods | Fold (Red/Green) | VIP 1 |
|---|---|---|---|---|---|
| DJ0219 | Apigenin-7-O-glucoside | 49,167 | 3,540,000 | 72.000 | 2.42218 |
| DJ0181 | Quercetin-3-O-rhamnoside | 970,667 | 24,600,000 | 25.343 | 2.03162 |
| DJ0394 | Luteolin O-malonylhexoside | 154,000 | 507,000 | 3.292 | 1.31471 |
| DJ0373 | Apigenin 7-O-glucoside | 174,333 | 451,333 | 2.589 | 1.24127 |
| DJ0434 | SelginO-hexoside derivative | 5987 | 14,143 | 2.362 | 1.02893 |
| DJ0408 | Apigenin O-malonylhexoside | 1,373,333 | 2,820,333 | 2.054 | 1.03458 |
| DJ0353 | Peonidin O-hexoside | 2,923,333 | 1,323,333 | 0.453 | 1.00394 |
| DJ0354 | Chrysoeriol 5-O-hexoside | 2,586,667 | 1,136,667 | 0.439 | 1.04638 |
| DJ0159 | Chlorogenic acid | 75,300 | 20,800 | 0.276 | 1.1813 |
| Metabolic Pathway | Total Number of Metabolites | Detected Metabolites in Green Pods (10- and 30-Day-Old) (p Value) | Detected Metabolites in Red Pods (10- and 30-Day-Old) (p Value) | Detected Metabolites in 30-Day-OldGreen and Red Pods (p Value) |
|---|---|---|---|---|
| Aminoacyl-tRNA biosynthesis | 75 | 7 (0.00002) | - | - |
| ABC transporters | 90 | 6 (0.0005) | - | 4 (0.006) |
| Cyanoamino acid metabolism | 41 | 5 (0.00008) | 4 (0.001) | - |
| Cysteine and methionine metabolism | 56 | 5 (0.0004) | 4 (0.004) | - |
| Purine metabolism | 92 | 4 (0.02) | 7 (0.00009) | 3 (0.04) |
| Tyrosine metabolism | 76 | 4 (0.01) | - | 3 (0.02) |
| Sulfur metabolism | 18 | 2 (0.02) | 2 (0.02) | - |
| Sphingolipid metabolism | 25 | 2 (0.03) | 2 (0.04) | - |
| Phenylalanine, tyrosine and Tryptophan biosynthesis | 27 | 2 (0.04) | 2 (0.04) | - |
| Selenoamino acid metabolism | 30 | 2 (0.046) | - | - |
| Thiamine metabolism | 26 | - | 2 (0.04) | - |
| β-Alanine metabolism | 31 | - | - | 2 (0.03) |
| Component Name | Class | Peak Area of 10-Day-Old Red Pods | Peak Area of 30-Day-OldRed Pods | Peak Area of 10-Day-OldGreen Pods | Peak Area of 30-Day-OldGreen Pods | Fold Change 1 (Red/Green) | VIP (Red/Green) |
|---|---|---|---|---|---|---|---|
| Inosine | Nucleotide derivates | 2.53 × 105 | 1.00 × 102 | -- | -- | 2533.67/-- | 3.17/-- |
| iPR | Nucleotide derivates | 2.96 × 106 | 9.33 × 103 | 8.08 × 105 | 3.90 × 104 | 316.79/20.69 | 3.90/3.04 |
| tZR | Phytohormones | 8.13 × 106 | 2.18 × 105 | 5.69 × 106 | 2.97 × 105 | 37.31/19.15 | 3.05/2.78 |
| iP | Phytohormones | 5.86 × 104 | 6.10 × 103 | 2.78 × 104 | 4.55 × 103 | 9.60/6.12 | 2.42/2.17 |
| IAA | Phytohormones | 3.06 × 104 | 3.34 × 103 | 3.16 × 104 | 3.02 × 103 | 9.14/10.45 | 2.38/2.47 |
| trans-Zeatin riboside-O-glucoside | Phytohormones | 7.54 × 104 | 9.70 × 103 | 1.14 × 105 | 1.48 × 104 | 7.78/7.72 | 2.32/2.30 |
| DZR | Phytohormones | 3.38 × 105 | 5.69 × 104 | -- | -- | 5.94/-- | 2.14/-- |
| tZROG | Phytohormones | 1.46 × 105 | 2.64 × 104 | 1.83 × 105 | 2.39 × 104 | 5.53/7.68 | 2.09/2.29 |
| Caffeoyl shikimic acid | Polyphenol | 1.94 × 105 | 4.46 × 104 | -- | -- | 4.36/-- | 1.94/-- |
| LPC (1-acyl 18:1) | Lipids | 1.24 × 106 | 3.25 × 105 | -- | -- | 3.83/-- | 1.15/-- |
| Guanosine | Nucleotide derivates | -- | -- | 8.06 × 106 | 4.74 × 104 | --/169.95 | --/2.04 |
| Crotonoside | others | -- | -- | 3.62 × 104 | 5.67 × 102 | --/63.94 | --/2.15 |
| Spermidine derivative | Phenolamine | -- | -- | 1.38 × 105 | 2.62 × 104 | --/5.28 | --/1.75 |
| Dopamine | others | -- | -- | 1.42 × 104 | 3.24 × 103 | --/4.38 | --/2.09 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, F.; Yang, T.; Liu, K.; Yang, Q.; Wan, Y.; Wang, R.; Li, G. Analysis of Metabolite Accumulation Related to Pod Color Variation of Caragana intermedia. Molecules 2019, 24, 717. https://doi.org/10.3390/molecules24040717
Yang F, Yang T, Liu K, Yang Q, Wan Y, Wang R, Li G. Analysis of Metabolite Accumulation Related to Pod Color Variation of Caragana intermedia. Molecules. 2019; 24(4):717. https://doi.org/10.3390/molecules24040717
Chicago/Turabian StyleYang, Feiyun, Tianrui Yang, Kun Liu, Qi Yang, Yongqing Wan, Ruigang Wang, and Guojing Li. 2019. "Analysis of Metabolite Accumulation Related to Pod Color Variation of Caragana intermedia" Molecules 24, no. 4: 717. https://doi.org/10.3390/molecules24040717
APA StyleYang, F., Yang, T., Liu, K., Yang, Q., Wan, Y., Wang, R., & Li, G. (2019). Analysis of Metabolite Accumulation Related to Pod Color Variation of Caragana intermedia. Molecules, 24(4), 717. https://doi.org/10.3390/molecules24040717




