Characterization of a Cis-Prenyltransferase from Lilium longiflorum Anther
Abstract
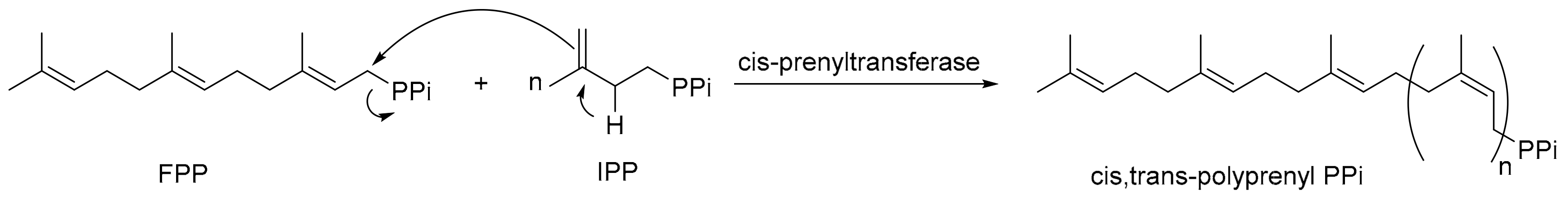
1. Introduction
2. Results and Discussion
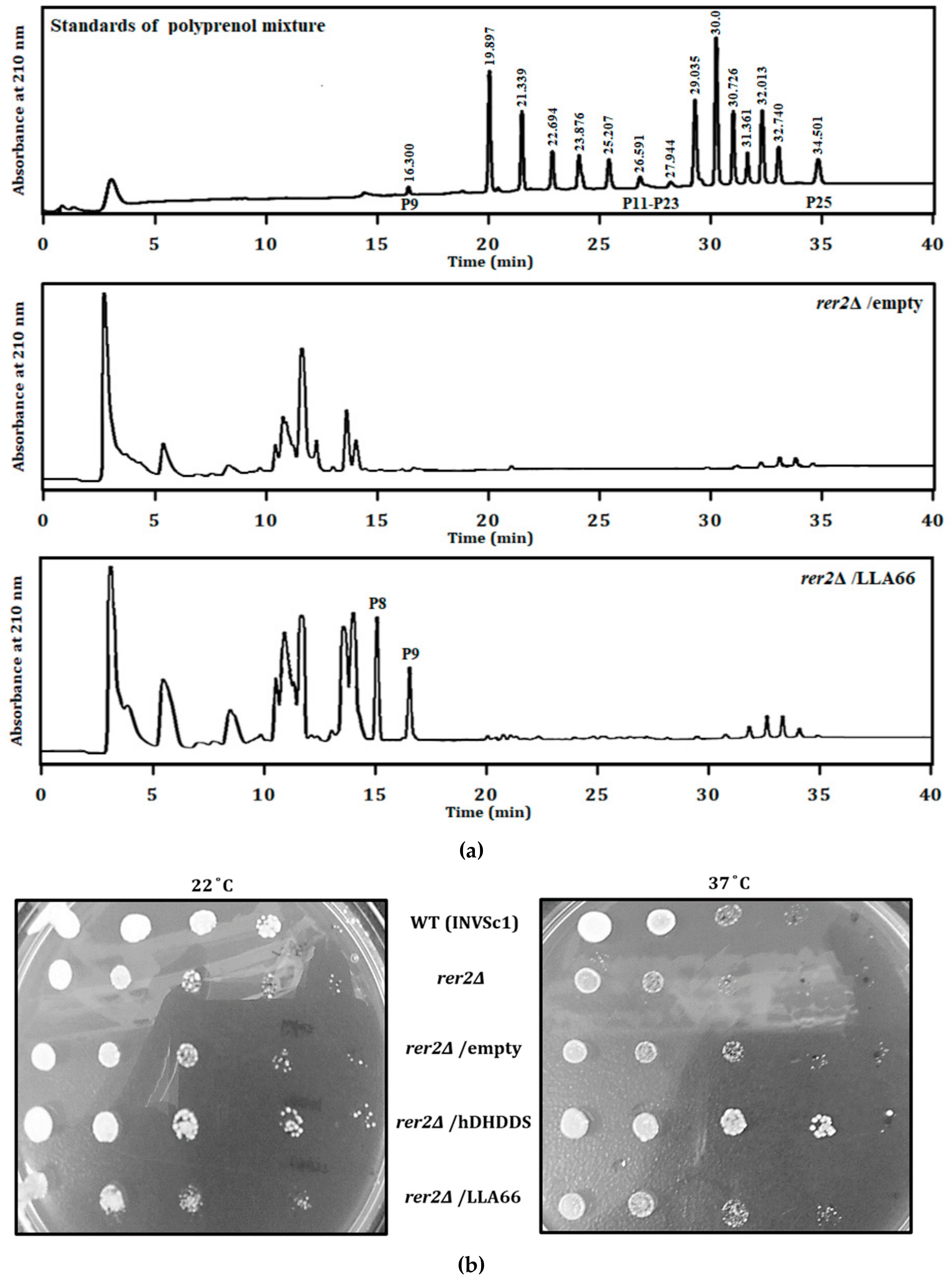
2.1. Products and Functional Analysis of LLA66 in S. cerevisiae
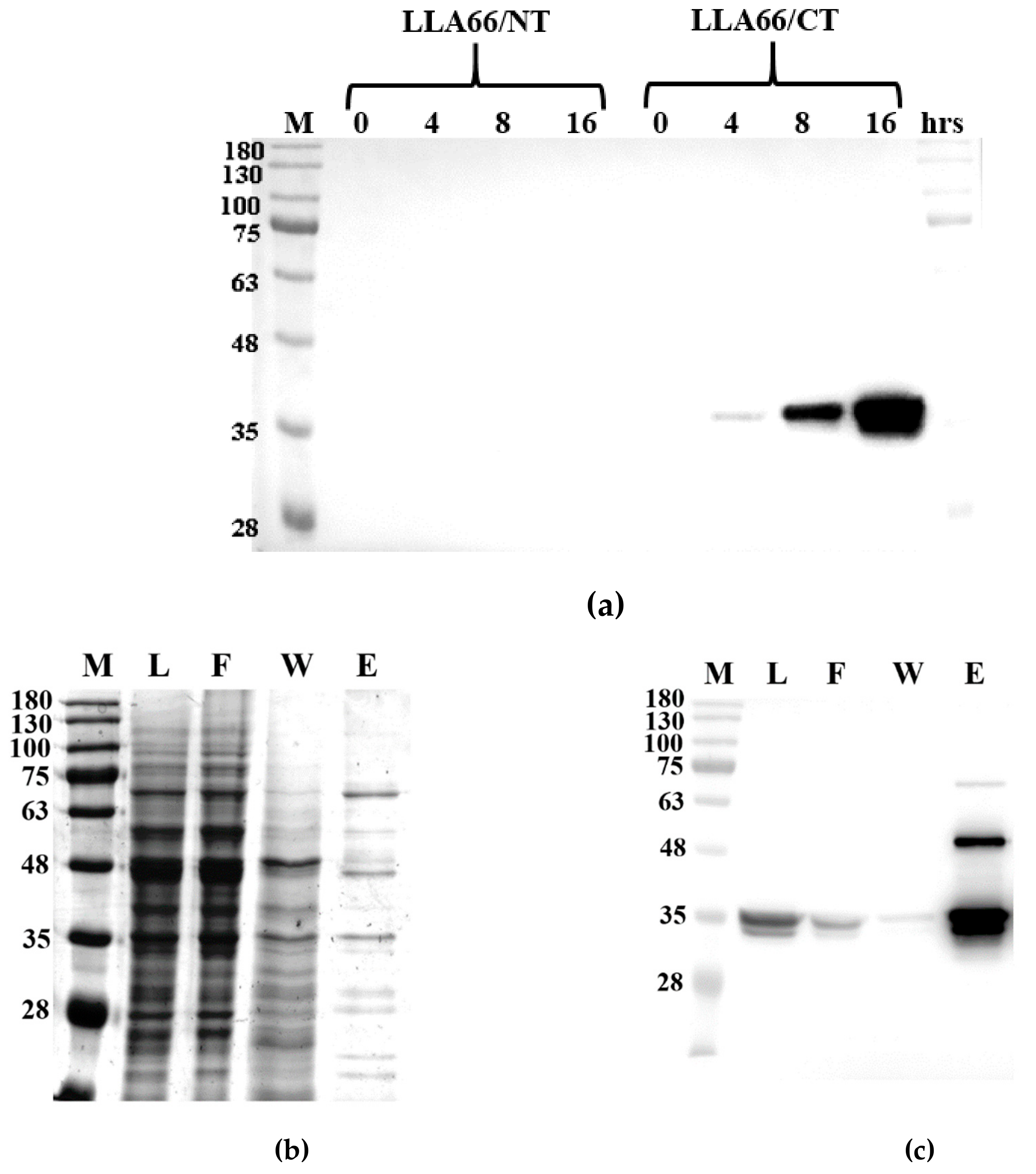
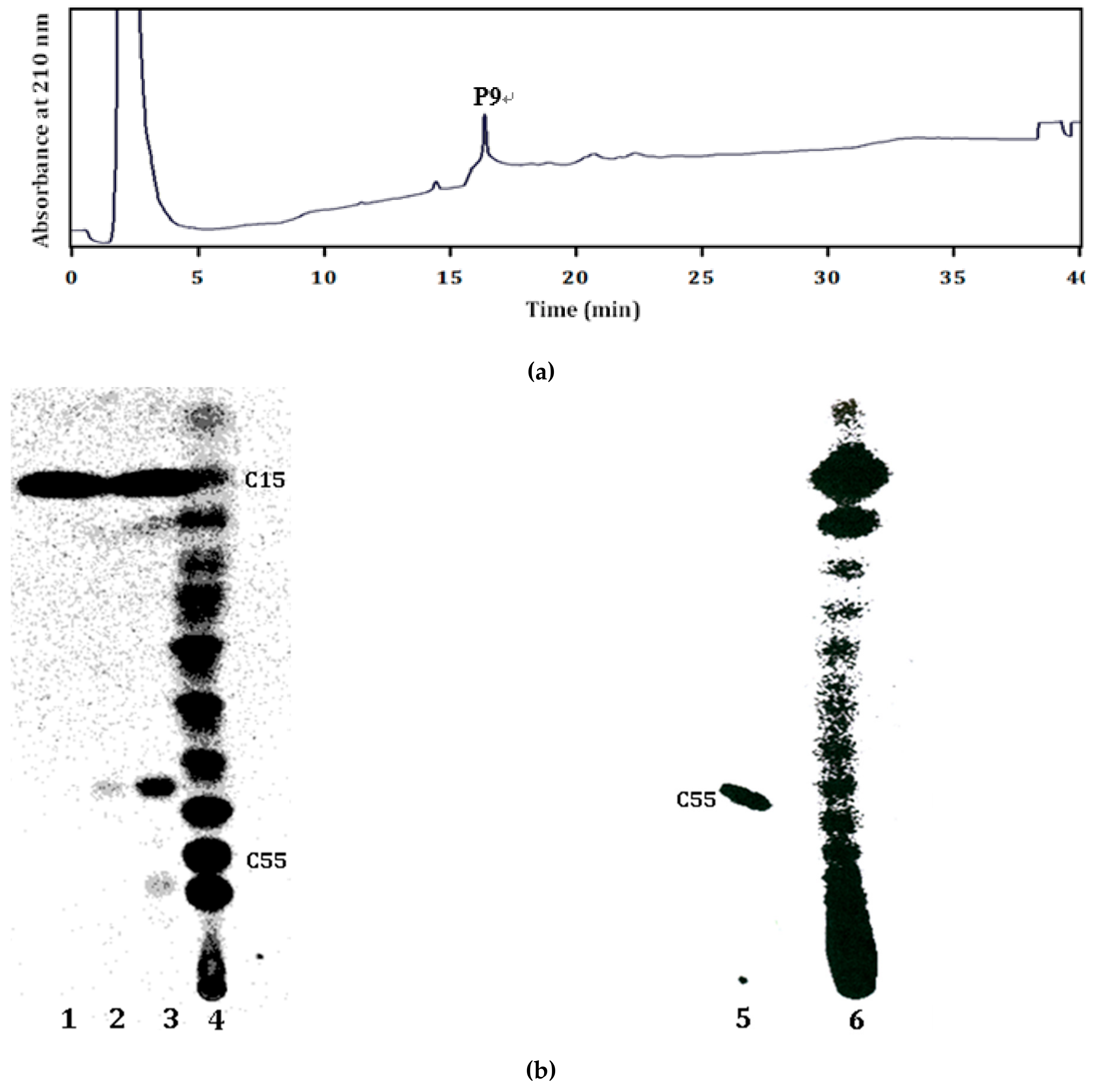
2.2. Products of the Recombinant LLA66
2.3. Identification of S. cerevisiae Enzymes for Unexpected IPP-Utilizing Activity
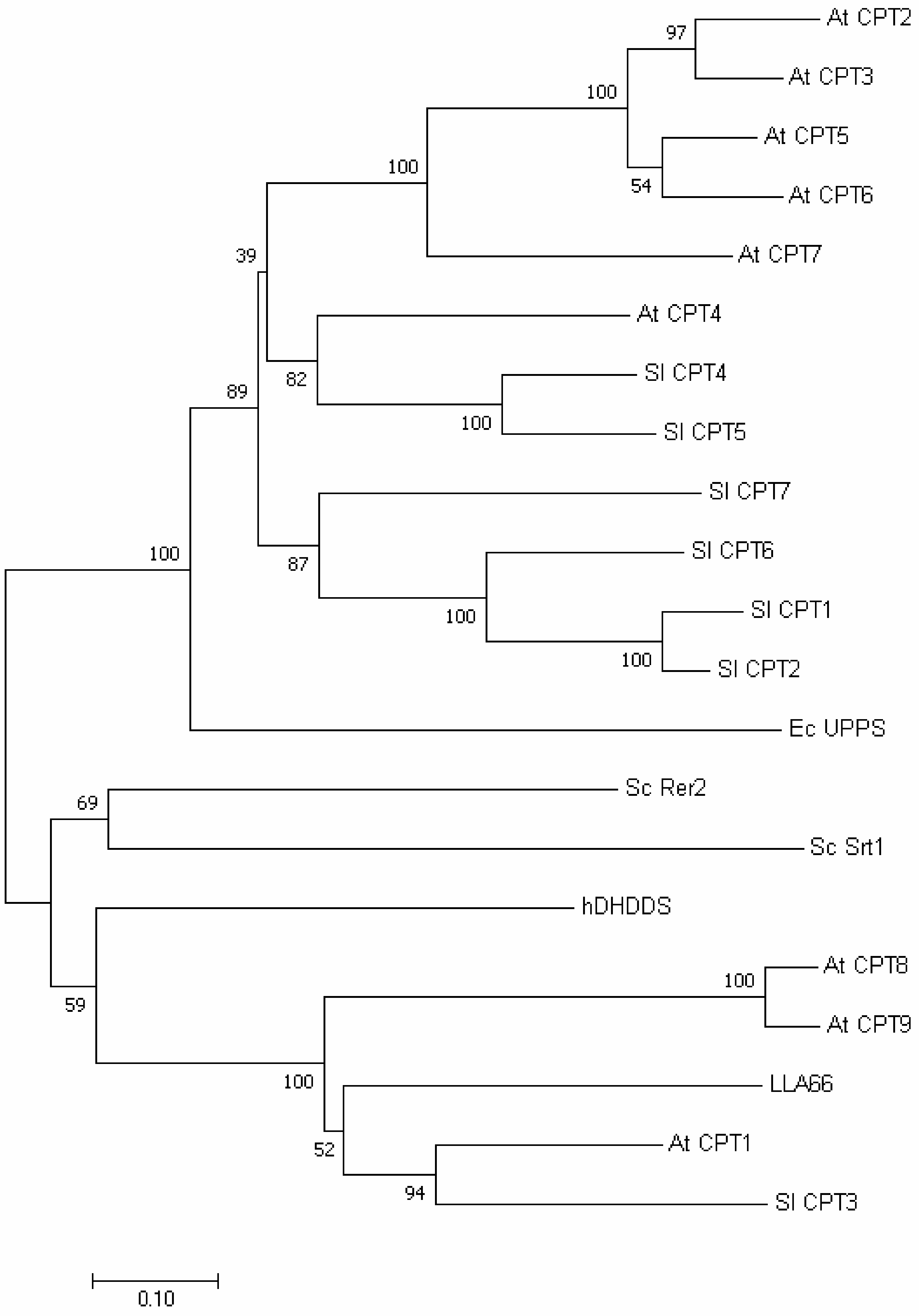
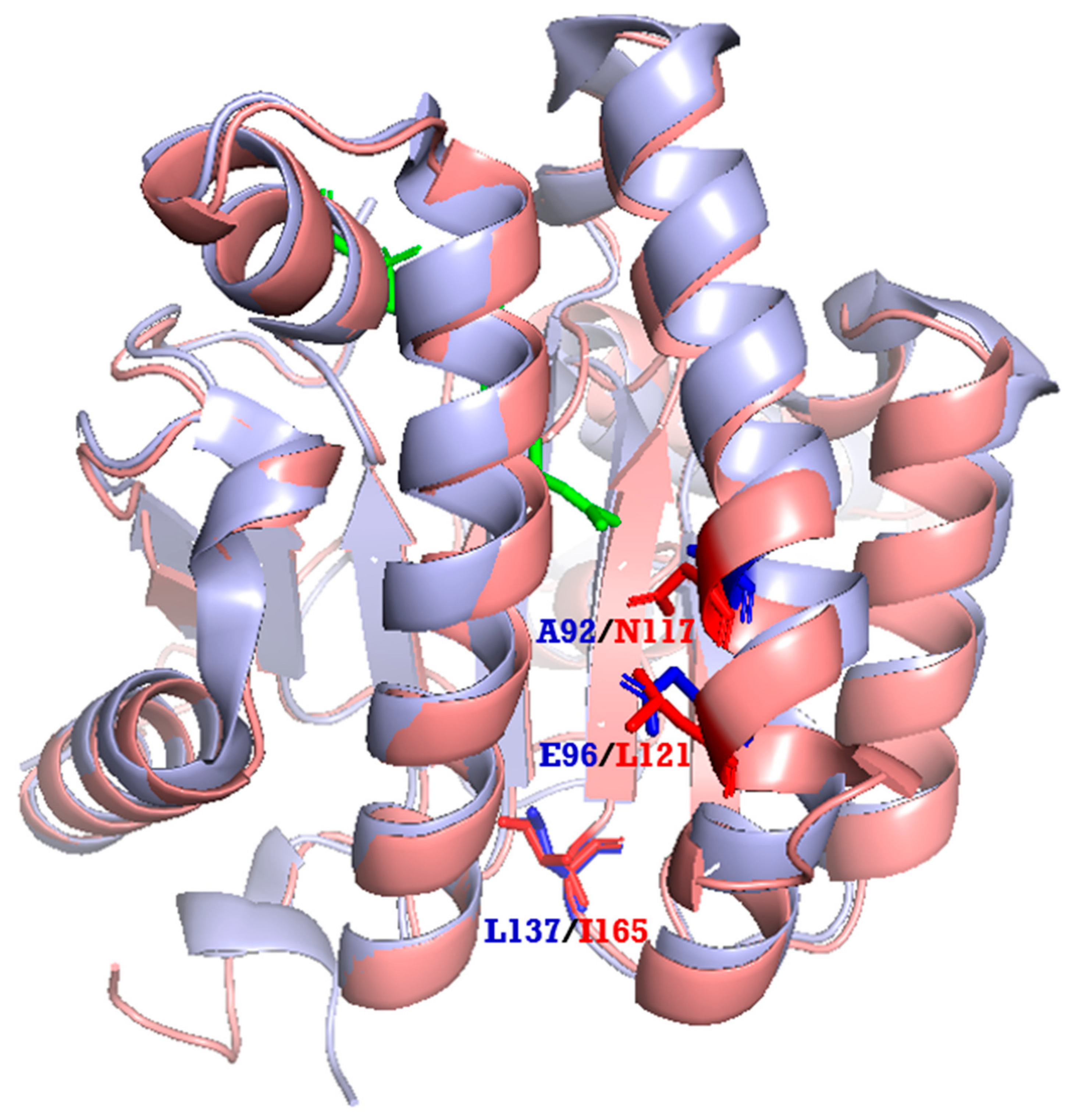
2.4. Phylogenetic Tree and Homology-Based Model of LLA66
3. Materials and Methods
3.1. Materials
3.2. Preparation of LLA66 Construct
3.3. Extraction of Polyisoprenoids from S. cerevisiae
3.4. HPCL Analysis of the LLA66 Products in S. cerevisiae
3.5. Expression and Purification of the Recombinant LLA66
3.6. TLC Analysis of the Recombinant LLA66 Products
3.7. Phylogenetic Tree Analysis of LLA66
3.8. Homology Modeling of LLA66
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| IPP | isopentenyl diphosphate |
| DMAPP | dimethylallyl diphosphate |
| FPP | farnesyl diphosphate |
| FPPS | farnesyl diphosphate synthase |
| UPPS | undecaprenyl diphosphate synthase |
| PCR | polymerase chain reaction |
| Ni-NAT | nickel nitrilo-tri-acetic acid |
| SDS-PAGE | sodium dodecyl sulfate-polyacrylamide |
| Da. | Dalton |
| IPTG | isopropyl-β-d-thiogalactopytanoside |
| Tris | tris(hydroxymethyl)aminomethane |
| Hepes | 4-(2-hydroxyethyl)-1-piperazineethanesufonic acid |
| HPLC | high performance liquid chromatography |
| TLC | thin layer chromatography |
References
- Poulter, C.D.; Rilling, H.C. Biosynthesis of Isoprenoid Compounds; Porter, J.W., Spurgeon, S.L., Eds.; John Wiley & Sons: New York, NY, USA, 1981; Volume 1, pp. 1–282. [Google Scholar]
- Ogura, K.; Koyama, T.; Sagami, H. Subcellular Biochemistry; Bittman, R., Ed.; Plenum: New York, NY, USA, 1997; Volume 28, pp. 57–88. [Google Scholar]
- Sacchettini, J.C.; Poulter, C.D. Creating isoprenoid diversity. Science 1997, 277, 1788–1789. [Google Scholar] [CrossRef] [PubMed]
- Rohmer, M.; Knani, M.; Simonin, P.; Sutter, B.; Sahm, H. Isoprenoid biosynthesis in bacteria: A novel pathway for the early steps leading to isopentenyl diphosphate. Biochem. J. 1993, 295, 517–524. [Google Scholar] [CrossRef] [PubMed]
- Arigoni, D.; Sagner, S.; Latzel, C.; Eisenreich, W.; Bacher, A.; Zenk, M.H. Terpenoid biosynthesis from 1-deoxy-d-xylulose in higher plants by intramolecular skeletal rearrangement. Proc. Natl. Acad. Sci. USA 1997, 94, 10600–10605. [Google Scholar] [CrossRef] [PubMed]
- Durbecq, V.; Sainz, G.; Oudjama, Y.; Clantin, B.; Bompard-Gilles, C.; Tricot, C.; Caillet, J.; Stalon, V.; Droogmans, L.; Villeret, V. Crystal structure of isopentenyl diphosphate:dimethylallyl diphosphate isomerase. EMBO J. 2001, 20, 1530–1537. [Google Scholar] [CrossRef] [PubMed]
- Liang, P.H. Reaction kinetics, catalytic mechanisms, conformational changes, and inhibitor design for prenyltransferases. Biochemistry 2009, 48, 6562–6570. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Ohnuma, S. Chain-length determination mechanism of isoprenyl diphosphate synthases and implications for molecular evolution. Trend Biochem. Sci. 1999, 24, 445–451. [Google Scholar] [CrossRef]
- Teng, K.H.; Liang, P.H. Undecaprenyl diphosphate synthase, a cis-prenyltransferase synthesizing lipid carrier for bacterial cell wall biosynthesis. Mol. Membr. Biol. 2012, 29, 267–273. [Google Scholar] [CrossRef]
- Sato, M.; Sato, K.; Nishikawa, S.; Hirata, A.; Kato, J.; Nakano, A. The yeast RER2 gene, identified by endoplasmic reticulum protein localization mutations, encodes cis-prenyltransferase, a key enzyme in dolichol synthesis. Mol. Cell. Biol. 1999, 19, 471–483. [Google Scholar] [CrossRef]
- Endo, S.; Zhang, Y.W.; Takahashi, S.; Koyama, T. Identification of human dehydrodolichyl diphosphate synthase gene. Biochim. Biophys. Acta 2003, 21625, 291–295. [Google Scholar] [CrossRef]
- Jozwiak, A.; Gutkowska, M.; Gawarecka, K.; Surmacz, L.; Buczkowska, A.; Lichocka, M.; Nowakowska, J.; Swiezewska, E. Polyprenol Reductase2 deficiency is lethal in Arabidopsis due to male sterility. Plant Cell 2015, 27, 3336–3353. [Google Scholar] [CrossRef]
- Surmacz, L.; Swiezewska, E. Polyisoprenoids: Secondary metabolites or physiologically important superlipids? Biochem. Biophys. Res. Commun. 2011, 407, 627–632. [Google Scholar] [CrossRef] [PubMed]
- Akhtar, T.A.; Matsuba, Y.; Schauvinhold, I.; Yu, G.; Lees, H.A.; Klein, S.E.; Pichersky, E. The tomato cis-prenyltransferase gene family. Plant J. 2013, 73, 640–652. [Google Scholar] [CrossRef] [PubMed]
- Swiezewska, E.; Danikiewicz, W. Polyisoprenoids: Structure, biosynthesis and function. Prog. Lipid Res. 2005, 44, 235–258. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.C.; Wang, B.J.; Huang, J.K.; Wang, C.S. Expression, localization and function of a cis-prenyltransferase in the Tapetum and Microspores of Lily Anthers. Plant Cell Physiol. 2011, 5, 1487–1500. [Google Scholar] [CrossRef] [PubMed]
- Ko, T.P.; Chen, Y.K.; Robinson, H.; Tsai, P.C.; Gao, Y.G.; Chen, A.P.; Wang, A.H.; Liang, P.H. Mechanism of Product Chain Length Determination and the Role of a Flexible Loop in Escherichia coli Undecaprenyl-pyrophosphate Synthase Catalysis. J. Biol. Chem. 2001, 276, 47474–47482. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.K.; Han, K.H.; Ryu, S.B.; Kang, H. Molecular cloning, expression, and functional analysis of a cis-prenyltransferase from Arabidopsis thaliana. Implications in rubber biosynthesis. J. Biol. Chem. 2000, 275, 18482–18488. [Google Scholar] [CrossRef]
- Zhang, H.; Ohyama, K.; Boudet, J.; Chen, Z.; Yang, J.; Zhang, M.; Muranaka, T.; Maurel, C.; Zhu, J.K.; Gong, Z. Dolichol biosynthesis and its effects on the unfolded protein response and abiotic stress resistance in Arabidopsis. Plant Cell 2008, 20, 1879–1898. [Google Scholar] [CrossRef] [PubMed]
- Kera, K.; Takahashi, S.; Sutoh, T.; Koyama, T.; Nakayama, T. Identification and characterization of a cis, trans-mixed heptaprenyl diphosphate synthase from Arabidopsis thaliana. FEBS J. 2012, 279, 3813–3827. [Google Scholar] [CrossRef]
- Surmacz, L.; Plochocka, D.; Kania, M.; Danikiewicz, W.; Swiezewska, E. cis-Prenyltransferase atCPT6 produces a family of very short-chain polyisoprenoids in planta. Biochim. Biophys. Acta 2014, 1841, 240–250. [Google Scholar] [CrossRef]
- Brasher, M.I.; Surmacz, L.; Leong, B.; Pitcher, J.; Swiezewska, E.; Pichersky, E.; Akhtar, T.A. A two-component enzyme complex is required for dolichol biosynthesis in tomato. Plant J. 2015, 82, 903–914. [Google Scholar] [CrossRef]
- Schilmiller, A.L.; Schauvinhold, I.; Larson, M.; Xu, R.; Charbonneau, A.L.; Schmidt, A.; Wilkerson, C.; Last, R.L.; Pichersky, E. Monoterpenes in the glandular trichomes of tomato are synthesized from a neryl diphosphate precursor rather than geranyl diphosphate. Proc. Natl. Acad. Sci. USA 2009, 106, 10865–10870. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.F.; Tzeng, J.D.; Liu, M.C.; Yei, F.L.; Chung, M.C.; Wang, C.S. Identification of anther-specific/predominant genes regulated by gibberellin during development of lily anthers. J. Plant Physiol. 2008, 165, 553–563. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not Available. |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yao, J.-Y.; Teng, K.-H.; Liu, M.-C.; Wang, C.-S.; Liang, P.-H. Characterization of a Cis-Prenyltransferase from Lilium longiflorum Anther. Molecules 2019, 24, 2728. https://doi.org/10.3390/molecules24152728
Yao J-Y, Teng K-H, Liu M-C, Wang C-S, Liang P-H. Characterization of a Cis-Prenyltransferase from Lilium longiflorum Anther. Molecules. 2019; 24(15):2728. https://doi.org/10.3390/molecules24152728
Chicago/Turabian StyleYao, Jyun-Yu, Kuo-Hsun Teng, Ming-Che Liu, Co-Shine Wang, and Po-Huang Liang. 2019. "Characterization of a Cis-Prenyltransferase from Lilium longiflorum Anther" Molecules 24, no. 15: 2728. https://doi.org/10.3390/molecules24152728
APA StyleYao, J.-Y., Teng, K.-H., Liu, M.-C., Wang, C.-S., & Liang, P.-H. (2019). Characterization of a Cis-Prenyltransferase from Lilium longiflorum Anther. Molecules, 24(15), 2728. https://doi.org/10.3390/molecules24152728






