Coumarin Content, Morphological Variation, and Molecular Phylogenetics of Melilotus
Abstract
1. Introduction
2. Results
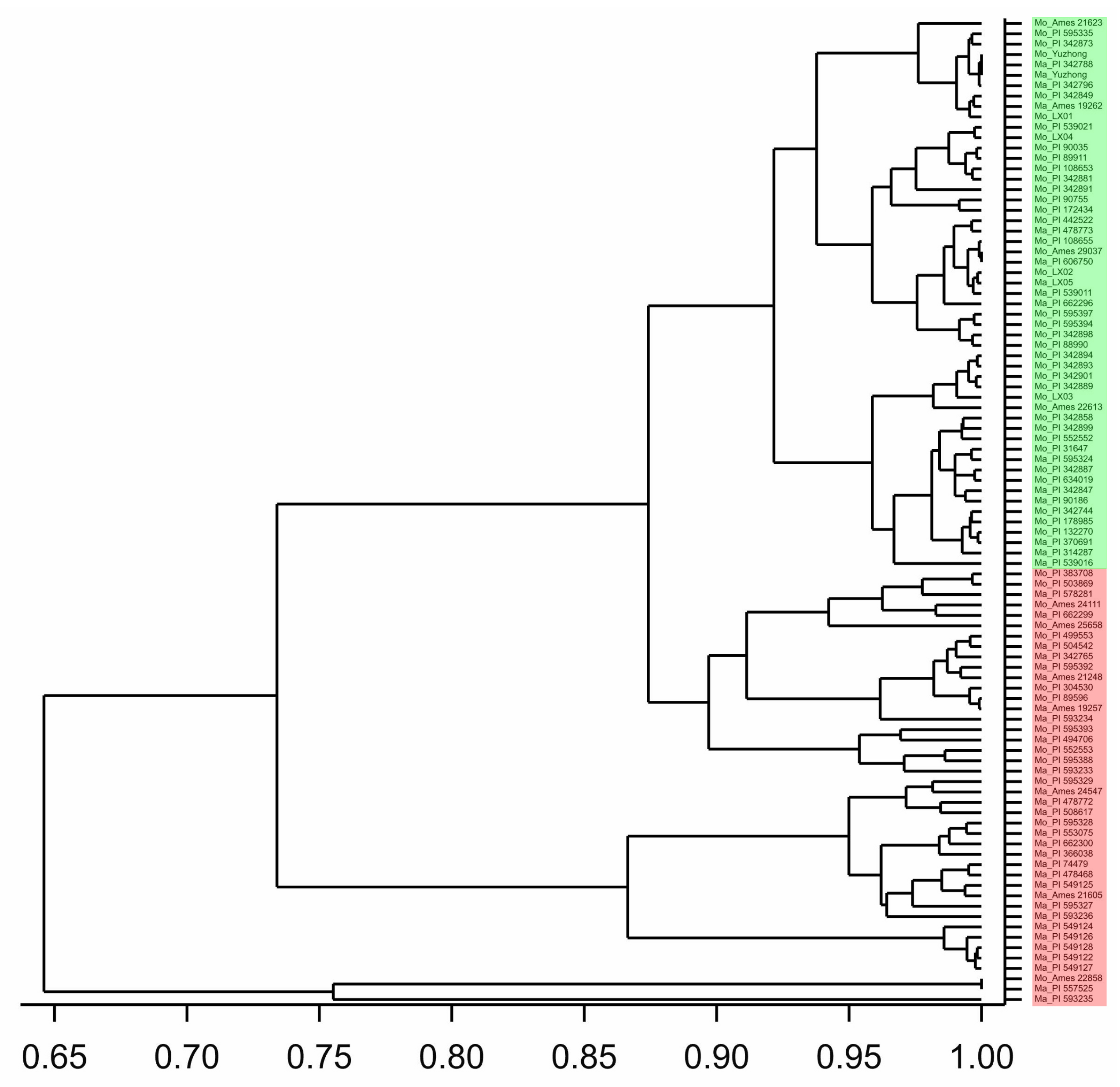
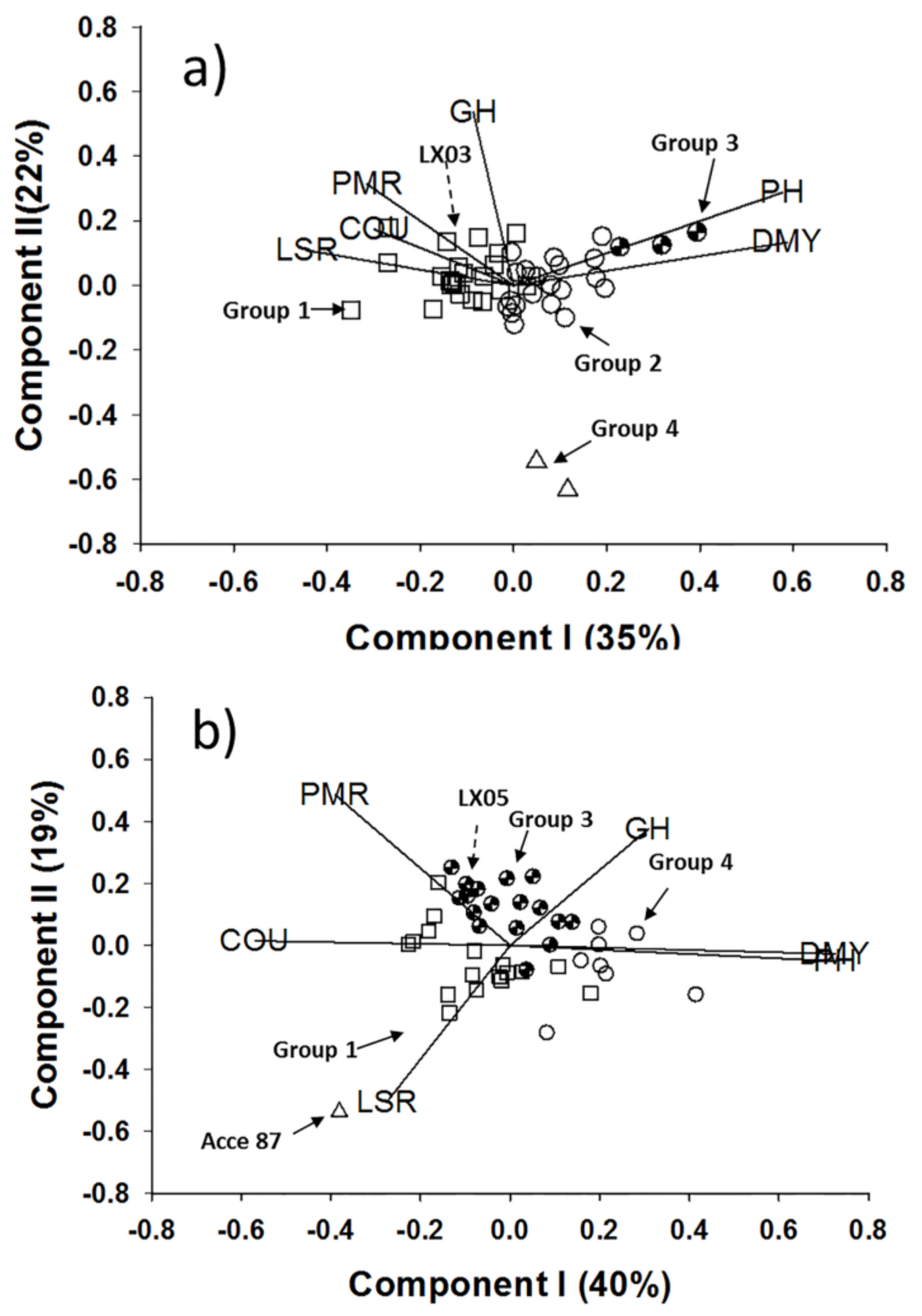
2.1. Coumarin Content and Morphological Variation
2.2. Alignments and Sequence Variation
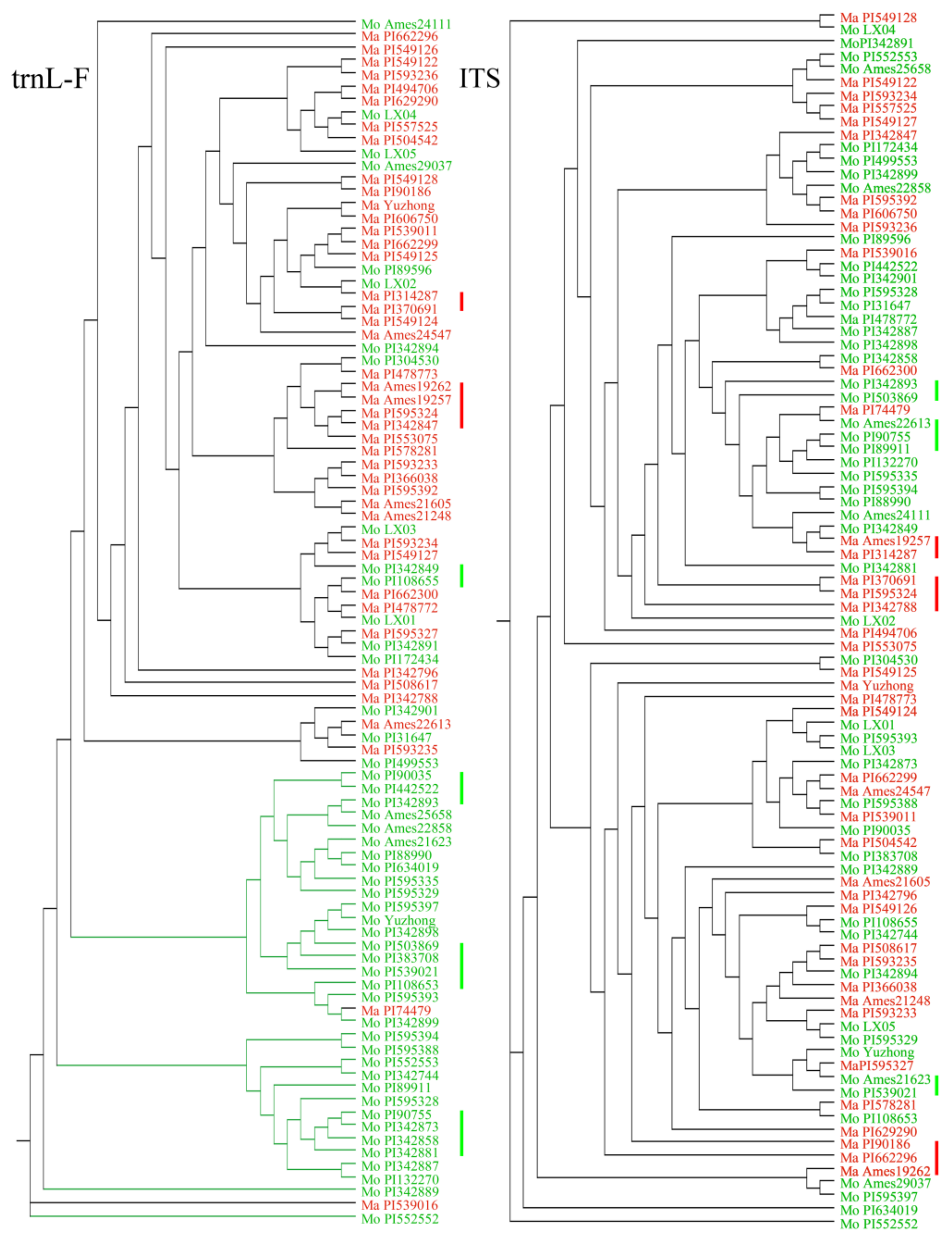
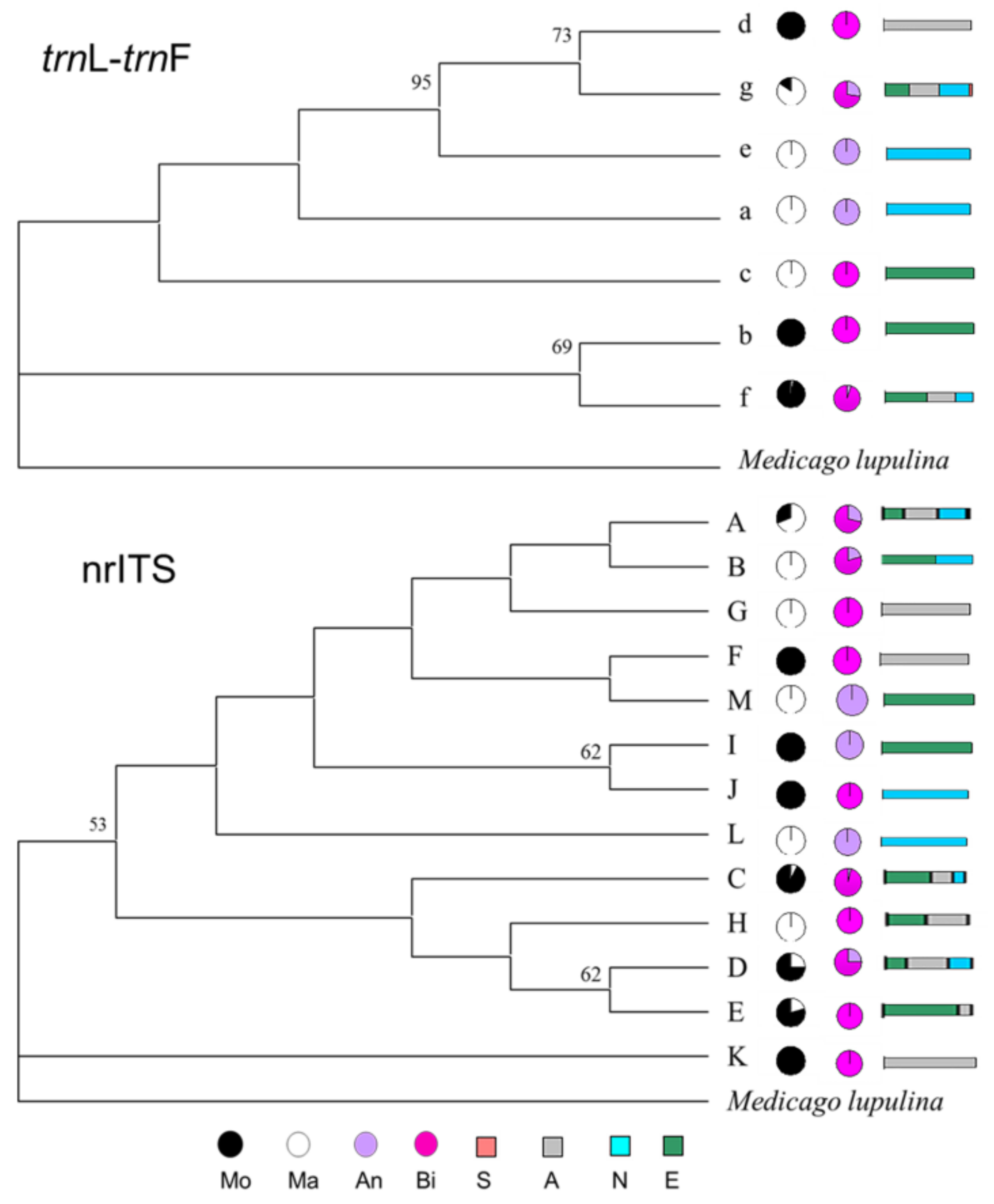
2.3. Phylogenetic Analyses
2.4. Phylogenetic Relationships of Haplotypes
2.5. Genetic Variation within and between Species
2.6. Molecular Variation within and between Areas
3. Discussion
4. Materials and Methods
4.1. Germplasm
4.2. Field Experiment
4.2.1. Preparation of Plant Material
4.2.2. Field Trial
4.2.3. Agronomic Traits
4.2.4. Analysis of Variance and Pattern Analysis
4.3. Molecular Phylogeny
4.3.1. DNA Extraction, Amplification, and Sequencing
4.3.2. Phylogenetic Analyses
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Hawryl, M.A.; Soczewinski, E.; Dzido, T.H. Separation of coumarins from Archangelica officinalis in high-performance liquid chromatography and thin-layer chromatography systems. J. Chromatogr. A 2000, 886, 75–81. [Google Scholar] [CrossRef]
- De Vincenzi, M.; Mancini, E.; Dessi, M. Monographs on botanical flavouring substances used in foods. Part VI. Fitoterapia 1997, 68, 49–61. [Google Scholar]
- Celeghini, R.; Vilegas, J.H.; Lanças, F.M. Extraction and quantitative HPLC analysis of coumarin in hydroalcoholic extracts of Mikania glomerata Spreng: (“guaco”) leaves. J. Brazil Chem. Soc. 2001, 12, 706–709. [Google Scholar] [CrossRef]
- Yarnell, E.; Abascal, K. Plant coumarins: Myths and realities. Altern. Complement. Ther. 2009, 15, 24–30. [Google Scholar] [CrossRef]
- Christensen, T.D.; Maegaard, M.; Sørensen, H.T.; Hjortdal, V.E.; Hasenkam, J.M. Self-versus conventional management of oral anticoagulant therapy: Effects on INR variability and coumarin dose in a randomized controlled trial. Am. J. Cardiovasc. Drug 2007, 7, 191–197. [Google Scholar] [CrossRef]
- Lee, C.M.; Jiang, L.M.; Shang, H.S.; Hon, P.M.; He, Y.; Wong, H.N. Prehispanolone, a novel platelet activating factor receptor antagonist from Leonurus heterophyllus. Br. J. Pharmacol. 1991, 103, 1719–1724. [Google Scholar] [CrossRef] [PubMed]
- Evans, P.; Kearney, G. Melilotus albus (Medik) is productive and regenerates well on saline soils of neutral to alkaline reaction in the high rainfall zone of south-western Victoria. Aust. J. Exp. Agric. 2003, 43, 349–355. [Google Scholar] [CrossRef]
- Nair, R.M.; Whittall, A.; Hughes, S.J.; Craig, A.D.; Revell, D.K.; Miller, S.M.; Powell, T.; Auricht, G.C. Variation in coumarin content of Melilotus species grown in South Australia. N. Z. J. Agric. Res. 2010, 53, 201–213. [Google Scholar] [CrossRef]
- Bowman, G.; Shirley, C.; Cramer, C. Managing cover crops profitably. In Sustainable Agriculture Network Handbook Series Book; Sustainable Agriculture Network: Beltsville, MD, USA, 1998; Volume 3, pp. 22–23. [Google Scholar]
- Aboel-Atta, A.M.I. Isozymes, RAPD and ISSR variation in Melilotus indica (L.) All. and M. siculus (Turra) BG Jacks.(Leguminosae). Acad. J. Plant Sci. 2009, 2, 113–118. [Google Scholar]
- Yang, Y.F. The pioneer pasture Melilotus that have drought and barren resistance. Beijing Agric. 2003, 8, 25–26. [Google Scholar]
- Hirsch, A.M.; Lum, M.R.; Krupp, R.S.N.; Yang, W.; Karlowski, W.M. Melilotus Alba Desr., White Sweetclover, a Mellifluous Model Legume; Horizon Scientific Press: Poole, UK, 2000; pp. 627–642. [Google Scholar]
- Pleşca-Manea, L.; Pârvu, A.E.; Pârvu, M.; Taămaş, M.; Buia, R.; Puia, M. Effects of Melilotus officinalis on acute inflammation. Phytother. Res. 2002, 16, 316–319. [Google Scholar] [CrossRef] [PubMed]
- Stevenson, G.A. An agronomic and taxonomic review of the genus Melilotus Mill. Can. J. Plant Sci. 1969, 49, 1–20. [Google Scholar] [CrossRef]
- Cong, J.M.; Chen, F.Q.; Sun, C.L. Study on Comprehensive Development of Metlilotus suaverolens L. J. Anhui Agric. Sci. 2012, 5, 2962–2963. [Google Scholar]
- Griffiths, B.S.; Donn, S.; Neilson, R.; Daniell, T.J. Molecular sequencing and morphological analysis of a nematode community. Appl. Soil Ecol. 2006, 32, 325–337. [Google Scholar] [CrossRef]
- Subbotin, S.A.; Halford, P.D.; Warry, A.; Perry, R.N. Variations in ribosomal DNA sequences and phylogeny of Globodera parasitising solanaceous plants. Nematology 2000, 2, 591–604. [Google Scholar] [CrossRef]
- Gozel, U.; Lamberti, F.; Duncan, L.; Agostinelli, A.; Rosso, L.; Nguyen, K.; Adams, B.J. Molecular and morphological consilience in the characterisation and delimitation of five nematode species from Florida belonging to the Xiphinema americanum-group. Nematology 2006, 8, 521–532. [Google Scholar] [CrossRef]
- Tian, B.; Liu, R.; Wang, L.; Qiu, Q.; Chen, K.; Liu, J. Phylogeographic analyses suggest that a deciduous species (Ostryopsis davidiana Decne., Betulaceae) survived in northern China during the Last Glacial Maximum. J. Biogeogr. 2009, 36, 2148–2155. [Google Scholar] [CrossRef]
- Hughes, C.E.; Bailey, C.D.; Harris, S.A. Divergent and reticulate species relationships in Leucaena (Fabaceae) inferred from multiple data sources: Insights into polyploid origins and nrDNA polymorphism. Am. J. Bot. 2002, 89, 1057–1073. [Google Scholar] [CrossRef] [PubMed]
- Arnheim, N.; Krystal, M.; Schmickel, R.; Wilson, G.; Ryder, O.; Zimmer, E. Molecular evidence for genetic exchanges among ribosomal genes on nonhomologous chromosomes in man and apes. Proc. Natl. Acad. Sci. USA 1980, 77, 7323–7327. [Google Scholar] [CrossRef] [PubMed]
- Zimmer, E.; Martin, S.; Beverley, S.; Kan, Y.; Wilson, A.C. Rapid duplication and loss of genes coding for the alpha chains of hemoglobin. Proc. Natl. Acad. Sci. USA 1980, 77, 2158–2162. [Google Scholar] [CrossRef] [PubMed]
- Hillis, D.M.; Dixon, M.T. Ribosomal DNA: Molecular evolution and phylogenetic inference. Q. Rev. Biol. 1991, 66, 411–453. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Abbott, R.J.; Zheng, W.; Chen, P.; Wang, Y.; Liu, J. History and evolution of alpine plants endemic to the Qinghai-Tibetan Plateau: Aconitum gymnandrum (Ranunculaceae). Mol. Ecol. 2009, 18, 709–721. [Google Scholar] [CrossRef] [PubMed]
- Oskoueiyan, R.; Kazempour Osaloo, S.; Maassoumi, A.A.; Nejadsattari, T.; Mozaffarian, V. Phylogenetic status of Vavilovia formosa (Fabaceae-Fabeae) based on nrDNA ITS and cpDNA sequences. Biochem. Syst. Ecol. 2010, 38, 313–319. [Google Scholar] [CrossRef]
- Zhang, W.; Kan, S.L.; Zhao, H.; Li, Z.Y.; Wang, X.Q. Molecular Phylogeny of Tribe Theeae (Theaceae ss.) and Its Implications for Generic Delimitation. PLoS ONE 2014, 9, e98133. [Google Scholar]
- Cardoso, D.; Paganucci de Queiroz, L.; Cavalcante de Lima, H.; Suganuma, E.; van den Berg, C.; Lavin, M. A molecular phylogeny of the vataireoid legumes underscores floral evolvability that is general to many early-branching papilionoid lineages. Am. J. Bot. 2013, 100, 403–421. [Google Scholar] [CrossRef] [PubMed]
- Kores, P.J.; Molvray, M.; Weston, P.H.; Hopper, S.D.; Brown, A.P.; Cameron, K.M.; Chase, M.W. A phylogenetic analysis of Diurideae (Orchidaceae) based on plastid DNA sequence data. Am. J. Bot. 2001, 88, 1903–1914. [Google Scholar] [CrossRef] [PubMed]
- Berg, C.V.D.; Higgins, W.E.; Dressler, R.L.; Whitten, W.M.; Soto Arenas, M.A.; Culham, A.; Chase, M.W.A. Phylogenetic analysis of Laeliinae (Orchidaceae) based on sequence data from internal transcribed spacers (ITS) of nuclear ribosomal DNA. Lindleyana 2000, 15, 96–114. [Google Scholar]
- Whitten, W.M.; Williams, N.H.; Chase, M.W. Subtribal and generic relationships of Maxillarieae (Orchidaceae) with emphasis on Stanhopeinae: Combined molecular evidence. Am. J. Bot. 2000, 87, 1842–1856. [Google Scholar] [CrossRef] [PubMed]
- Scheen, A.C.; Bendiksby, M.; Ryding, O.; Mathiesen, C.; Albert, V.A.; Lindqvist, C. Molecular phylogenetics, character evolution, and suprageneric classification of Lamioideae (Lamiaceae). Ann. Mo. Bot. Gard. 2016, 97, 191–217. [Google Scholar] [CrossRef]
- Faye, A.; Pintaud, J.C.; Baker, W.; Sonké, B.; Couvreur, T. A plastid phylogeny of the African rattans (Ancistrophyllinae, Arecaceae). Syst. Bot. 2014, 39, 1099–1107. [Google Scholar] [CrossRef]
- Sunderland, T.C.H. A taxonomic revision of the rattans of Africa (Arecaceae: Calamoideae). Phytotaxa 2012, 51, 1–76. [Google Scholar] [CrossRef][Green Version]
- Baker, W.J.; Dransfield, J.; Hedderson, T.A. Phylogeny, character evolution, and a new classification of the calamoid palms. Syst. Bot. 2000, 25, 297–322. [Google Scholar] [CrossRef]
- Di, H.Y.; Duan, Z.; Luo, K.; Zhang, D.Y.; Wu, F.; Zhang, J.Y.; Liu, W.X.; Wang, Y.R. Interspecific phylogenic relationships within genus Melilotus based on nuclear and chloroplast DNA. PLoS ONE 2015, 10, e0132596. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Zhang, D.Y.; Ma, J.X.; Luo, K.; Di, H.Y.; Liu, Z.P.; Zhang, J.Y.; Wang, Y.R. Analysis of genetic diversity and population structure in accessions of the genus Melilotus. Ind. Crop. Prod. 2016, 85, 84–92. [Google Scholar] [CrossRef]
- Yan, Z.Z.; Wu, F.; Luo, K.; Zhao, Y.F.; Yan, Q.; Zhang, Y.F.; Wang, Y.R.; Zhang, J.Y. Cross-species transferability of EST-SSR markers developed from the transcriptome ofMelilotusand their application to population genetics research. Sci. Rep. 2017, 7, 17959. [Google Scholar] [CrossRef] [PubMed]
- Bajerová, P.; Adam, M.; Bajer, T.; Ventura, K. Comparison of various techniques for the extraction and determination of antioxidants in plants. J. Sep. Sci. 2014, 37, 835–844. [Google Scholar] [CrossRef] [PubMed]
- Witaicenis, A.; Seito, L.N.; da Silveira Chagas, A.; de Almeida, L.D.; Luchini, A.C.; Rodrigues-Orsi, P.; Cestari, S.H.; Di Stasi, L.C. Antioxidant and intestinal anti-inflammatory effects of plant-derived coumarin derivatives. Phytomedicine 2014, 21, 240–246. [Google Scholar] [CrossRef] [PubMed]
- Céspedes, C.L.; Avila, J.G.; Martínez, A.; Serrato, B.; Calderón-Mugica, J.C.; Salgado-Garciglia, R. Antifungal and antibacterial activities of Mexican tarragon (Tagetes lucida). J. Agric. Food Chem. 2006, 54, 3521–3527. [Google Scholar] [CrossRef] [PubMed]
- Musa, M.A.; Cooperwood, J.S.; Khan, M.O.F. A review of coumarin derivatives in pharmacotherapy of breast cancer. Curr. Med. Chem. 2008, 15, 2664–2679. [Google Scholar] [CrossRef] [PubMed]
- Podolsky, R.H.; Holtsford, T.P. Population-Structure of Morphological Traits in Clarkia-Dudleyana. 1. Comparison of F-St between Allozymes and Morphological Traits. Genetics 1995, 140, 733–744. [Google Scholar] [PubMed]
- Luo, K.; Jahufer, M.Z.Z.; Wu, F.; Di, H.Y.; Zhang, D.Y.; Meng, X.C.; Zhang, J.Y.; Wang, Y.R. Genotypic variation in a breeding population of yellow sweet clover (Melilotus officinalis). Front. Plant Sci. 2016, 7, 972. [Google Scholar] [CrossRef] [PubMed]
- Wu, F.; Ma, J.X.; Meng, Y.Q.; Zhang, D.Y.; Muvunyi, B.P.; Luo, K.; Di, H.Y.; Guo, W.L.; Wang, Y.R.; Feng, B.C.; et al. Potential DNA barcodes for Melilotus species based on five single loci and their combinations. PLoS ONE 2017, 12, e0182693. [Google Scholar] [CrossRef] [PubMed]
- Fior, S.; Karis, P.O.; Casazza, G.; Minuto, L.; Sala, F. Molecular phylogeny of the Caryophyllaceae (Caryophyllales) inferred from chloroplast matK and nuclear rDNA ITS sequences. Am. J. Bot. 2006, 93, 399–411. [Google Scholar] [CrossRef] [PubMed]
- Mitsui, Y.; Chen, S.T.; Zhou, Z.K.; Peng, C.I.; Deng, Y.F.; Setoguchi, H. Phylogeny and biogeography of the genus Ainsliaea (Asteraceae) in the Sino-Japanese region based on nuclear rDNA and plastid DNA sequence data. Ann. Bot. 2008, 101, 111–124. [Google Scholar] [CrossRef] [PubMed]
- Winton, L.M.; Krohn, A.L.; Conn, J.S. Microsatellite markers for the invasive plant species white sweetclover (Melilotus alba) and yellow sweetclover (Melilotus officinalis). Mol. Ecol. Notes 2007, 7, 1296–1298. [Google Scholar] [CrossRef]
- Zhang, B.L.; Wen, C.R. Melilotus; Agriculture Press: Liaoning, China, 1978. [Google Scholar]
- Clarke, A.E. The Number and Morphology of Chromosomes in the Genus Melilotus; University of California Press: Berkeley, CA, USA, 1934. [Google Scholar]
- Maddison, W.P.; Knowles, L.L. Inferring phylogeny despite incomplete lineage sorting. Syst. Biol. 2006, 55, 21–30. [Google Scholar] [CrossRef] [PubMed]
- Käss, E.; Wink, M. Phylogenetic relationships in the Papilionoideae (family Leguminosae) based on nucleotide sequences of cpDNA (rbcL) and ncDNA (ITS 1 and 2). Mol. Phylogent. Evol. 1997, 8, 65–88. [Google Scholar] [CrossRef] [PubMed]
- Rieseberg, L.H. Hybrid origins of plant species. Annu. Rev. Ecol. Syst. 1997, 28, 359–389. [Google Scholar] [CrossRef]
- Rieseberg, L.H.; Carney, S.E. Plant hybridization. New Phytol. 1998, 140, 599–624. [Google Scholar] [CrossRef]
- Arnold, M.L.; Bulger, M.R.; Burke, J.M.; Hempel, A.L.; Williams, J.H. Natural hybridization: How low can you go and still be important? Ecology 1999, 80, 371–381. [Google Scholar] [CrossRef]
- Barton, N.H. The role of hybridization in evolution. Mol. Ecol. 2001, 10, 551–568. [Google Scholar] [CrossRef] [PubMed]
- Tang, C.N. Study on Extraction, Separation and Purification Technology of Coumarin from Melilotus. Master’s Thesis, Northwest University, Xian, China, 2010. [Google Scholar]
- Xie, C.; Sun, Q.; Ni, Z.; Yang, T.; Nevo, E.; Fahima, T. Chromosomal location of a Triticum dicoccoides-derived powdery mildew resistance gene in common wheat by using microsatellite markers. Theor. Appl. Genet. 2003, 106, 341–345. [Google Scholar] [CrossRef] [PubMed]
- Kroonenberg, P.M.K. The TUCKALS line: A suite of programs for three-way data analysis. Comput. Stat. Data Anal. 1994, 18, 73–96. [Google Scholar] [CrossRef]
- Gabriel, K.R. The biplot graphical display of matrices with applicationto principal component analysis. Biometrika 1971, 58, 453–467. [Google Scholar] [CrossRef]
- Zhang, J.Y.; Yuan, Q.H.; Zhang, W.S.; Lu, T. Genomic DNA extraction and optimizing RAPD procedure of Lespedeza floribunda. Acta Agrestia Sin. 2004, 12, 219–223. [Google Scholar]
- Williams, W.M.; Ansari, H.A.; Ellison, N.W.; Hussain, S.W. Evidence of Three Subspecies in Trifolium nigrescens Viv. Ann. Bot. 2001, 87, 683–691. [Google Scholar] [CrossRef]
- Taberlet, P.; Gielly, L.; Pautou, G.; Bouvet, J. Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol. Biol. 1991, 17, 1105–1109. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.; Abbott, R.J.; Milne, R.I.; Tian, X.M.; Liu, J. Phylogeography of Pinus tabulaeformis Carr. (Pinaceae), a dominant species of coniferous forest in northern China. Mol. Ecol. 2008, 17, 4276–4288. [Google Scholar] [CrossRef] [PubMed]
- Larkin, M.; Blackshields, G.; Brown, N.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Mark, P.V.D.; Huelsenbeck, J. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- Librado, P.; Rozas, J. DnaSP v5: A software for comprehensive analysis of DNA polymorphism data. Bioinformatics 2009, 25, 1451–1452. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.X. Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 1997, 147, 915–925. [Google Scholar] [PubMed]
- Excoffier, L.; Smouse, P.E.; Quattro, J.M. Analysis of molecular variance inferred from metric distances among DNA haplotypes: Application to human mitochondrial DNA restriction data. Genetics 1992, 131, 479–491. [Google Scholar] [PubMed]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. 2005, 1, 47–50. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |
| Trait | M. Officinalis | M. Albus | ||||||
|---|---|---|---|---|---|---|---|---|
| Range | Mean | Significance | Pst | Range | Mean | Significance | Pst | |
| PH | 20–100 | 62 | * | 0.4254 | 11–161 | 65 | * | 0.4103 |
| DMY | 4–41 | 17 | * | 0.4127 | 0.5–43 | 16 | * | 0.4642 |
| LSR | 0.7–1.8 | 1.17 | * | 0.4238 | 0.3–3.4 | 1.00 | * | 0.4138 |
| Cou | 0.3–1.5 | 0.83 | * | 0.4022 | 0.2–1.3 | 0.73 | * | 0.4094 |
| PMR | 1–4 | 2.75 | * | 0.5251 | 1–4 | 2.16 | * | 0.5434 |
| GH | 1–2 | 1.96 | * | 0.3405 | 1–2 | 1.56 | * | 0.3690 |
| Species | Groups | No. in Group | PH | DMY | LSR | COU | PMR | GH |
|---|---|---|---|---|---|---|---|---|
| M. officinalis | Group 1 | 23 | 55 | 12 | 1.255 | 0.908 | 3.39 | 2 |
| Group 2 | 20 | 43 | 20 | 1.104 | 0.79 | 2.15 | 2 | |
| Group 3 | 3 | 92 | 35 | 1.04 | 0.526 | 2.67 | 2 | |
| Group 4 | 2 | 48 | 17 | 0.98 | 0.673 | 1.5 | 1 | |
| M. albus | Group 1 | 17 | 58 | 13 | 0.974 | 0.769 | 2.29 | 1 |
| Acce 87 | 1 | 11 | 0.52 | 3.421 | 1.158 | 1 | 1 | |
| Group 3 | 17 | 63 | 14 | 0.828 | 0.771 | 2.53 | 2 | |
| Group 4 | 8 | 94 | 29 | 1.115 | 0.511 | 1.25 | 1.875 |
| Sequence | Dataset | Length | N | h | Hd ± SD | π ± SD |
|---|---|---|---|---|---|---|
| trnL-F | M. albus | 459 | 44 | 4 | 0.171 ± 0.075 | 0.0017 ± 0.0015 |
| M. officinalis | 439 | 49 | 3 | 0.357 ± 0.064 | 0.0050 ± 0.0016 | |
| All | 459 | 93 | 7 | 0.592 ± 0.021 | 0.0069 ± 0.0013 | |
| ITS | M. albus | 714 | 44 | 7 | 0.490 ± 0.091 | 0.0010 ± 0.0009 |
| M. officinalis | 714 | 49 | 9 | 0.703 ± 0.645 | 0.0017 ± 0.0014 | |
| All | 714 | 93 | 13 | 0.701 ± 0.036 | 0.0017 ± 0.0014 |
| Region | Sequence | Source of Variation | d.f. | Sum of Squares | Variance Components | Percentage of Variation | Fst |
|---|---|---|---|---|---|---|---|
| A | trnL-F | Among species | 1 | 3.199 | 0.206 | 55.820 | 0.558 |
| Among accessions | 31 | 4.737 | 0.163 | 44.180 | |||
| Total | 32 | 7.935 | 0.370 | ||||
| ITS | Among species | 1 | 0.549 | 0.016 | 4.710 | 0.047 | |
| Among accessions | 31 | 9.355 | 0.323 | 95.290 | |||
| Total | 32 | 9.903 | 0.339 | ||||
| E | trnL-F | Among species | 1 | 5.417 | 0.347 | 75.020 | 0.750 |
| Among accessions | 34 | 3.583 | 0.116 | 24.980 | |||
| Total | 35 | 9.000 | 0.463 | ||||
| ITS | Among species | 1 | 2.419 | 0.134 | 30.320 | 0.303 | |
| Among accessions | 34 | 10.152 | 0.308 | 69.680 | |||
| Total | 35 | 12.571 | 0.442 | ||||
| N | trnL-F | Among species | 1 | 3.841 | 0.400 | 78.650 | 0.786 |
| Among accessions | 26 | 2.714 | 0.109 | 21.350 | |||
| Total | 27 | 6.556 | 0.509 | ||||
| ITS | Among species | 1 | 0.882 | 0.065 | 18.850 | 0.188 | |
| Among accessions | 26 | 6.733 | 0.281 | 81.150 | |||
| Total | 27 | 7.615 | 0.346 | ||||
| A & E & N | trnL-F | Among regions | 3 | 1.463 | 0.012 | 3.850 | 0.038 |
| Among accessions | 92 | 23.295 | 0.268 | 96.150 | |||
| Total | 93 | 95.000 | 24.758 | ||||
| ITS | Among regions | 3 | 1.855 | 0.014 | 3.840 | 0.038 | |
| Among accessions | 92 | 29.808 | 0.339 | 96.160 | |||
| Total | 93 | 31.663 | 0.352 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, J.; Di, H.; Luo, K.; Jahufer, Z.; Wu, F.; Duan, Z.; Stewart, A.; Yan, Z.; Wang, Y. Coumarin Content, Morphological Variation, and Molecular Phylogenetics of Melilotus. Molecules 2018, 23, 810. https://doi.org/10.3390/molecules23040810
Zhang J, Di H, Luo K, Jahufer Z, Wu F, Duan Z, Stewart A, Yan Z, Wang Y. Coumarin Content, Morphological Variation, and Molecular Phylogenetics of Melilotus. Molecules. 2018; 23(4):810. https://doi.org/10.3390/molecules23040810
Chicago/Turabian StyleZhang, Jiyu, Hongyan Di, Kai Luo, Zulfi Jahufer, Fan Wu, Zhen Duan, Alan Stewart, Zhuanzhuan Yan, and Yanrong Wang. 2018. "Coumarin Content, Morphological Variation, and Molecular Phylogenetics of Melilotus" Molecules 23, no. 4: 810. https://doi.org/10.3390/molecules23040810
APA StyleZhang, J., Di, H., Luo, K., Jahufer, Z., Wu, F., Duan, Z., Stewart, A., Yan, Z., & Wang, Y. (2018). Coumarin Content, Morphological Variation, and Molecular Phylogenetics of Melilotus. Molecules, 23(4), 810. https://doi.org/10.3390/molecules23040810





