Development of Cell-Specific Aptamers: Recent Advances and Insight into the Selection Procedures
Abstract
1. Introduction
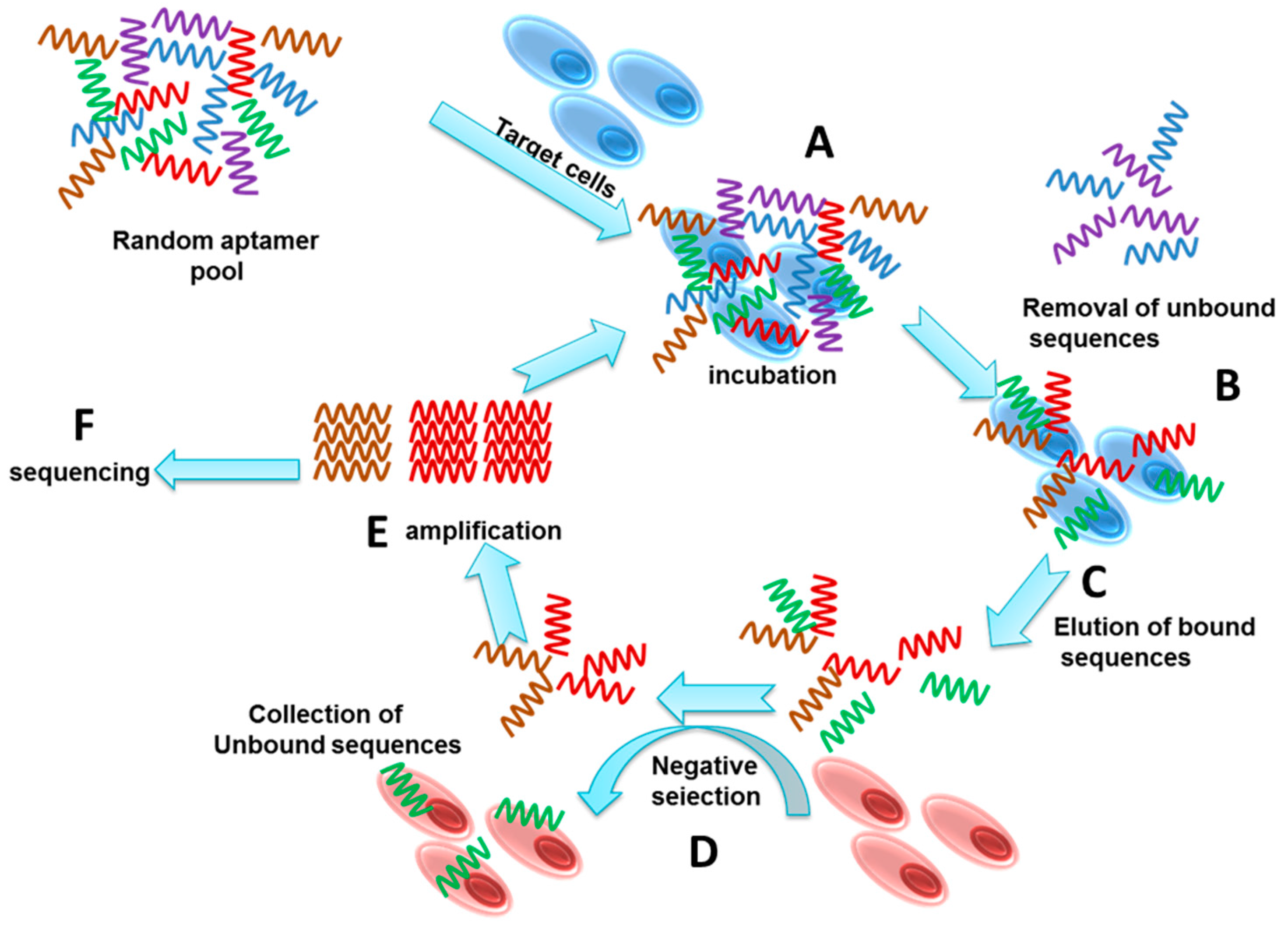
2. Cell Specific Aptamer Development by Cell-SELEX
3. Selection
4. Enrichment and Amplification
5. Regeneration of Selected Aptamers
5.1. Asymmetric PCR
5.2. Electrophoresis-Based Separation of DNA Strands
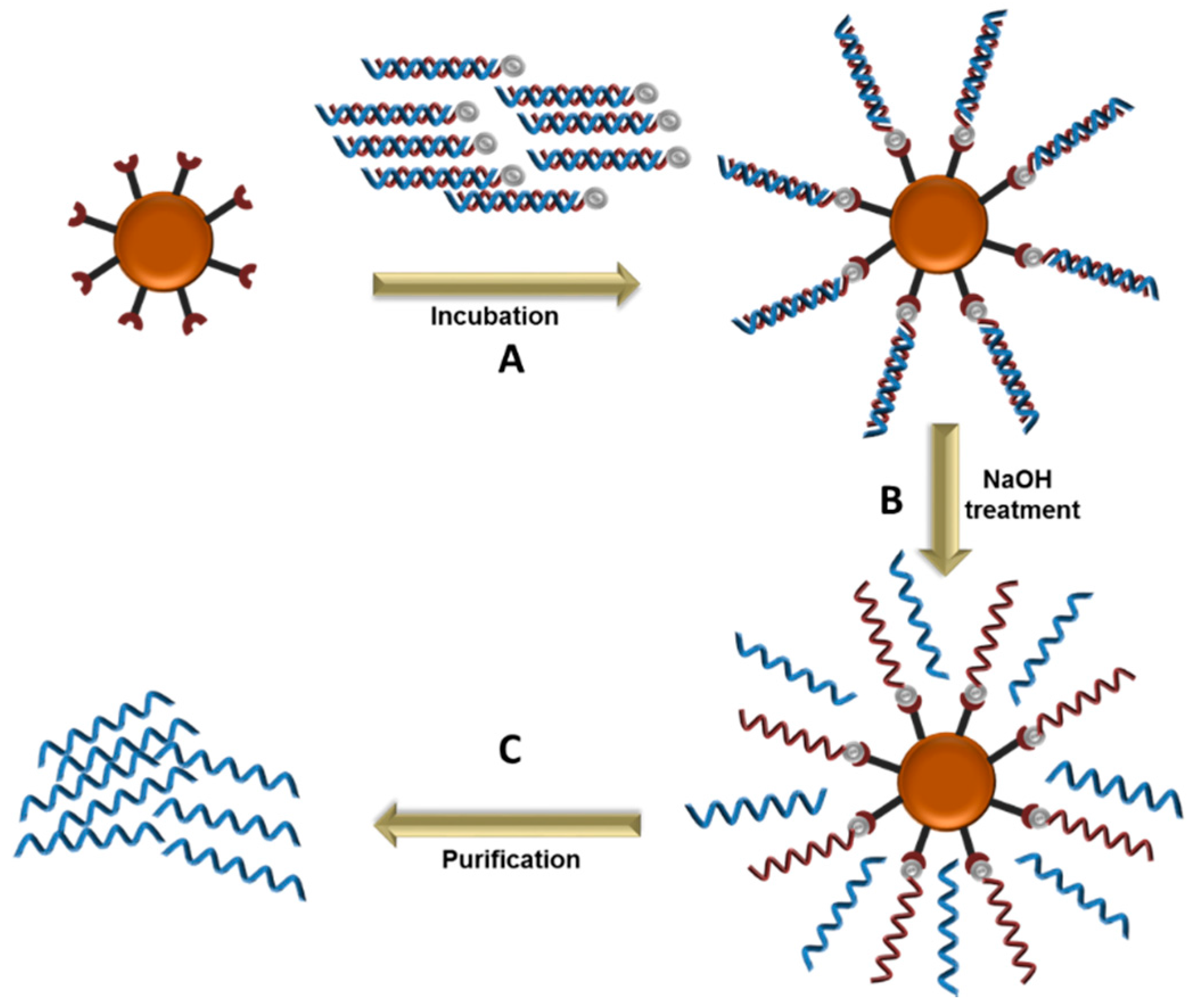
5.3. Magnetic Beads Separation
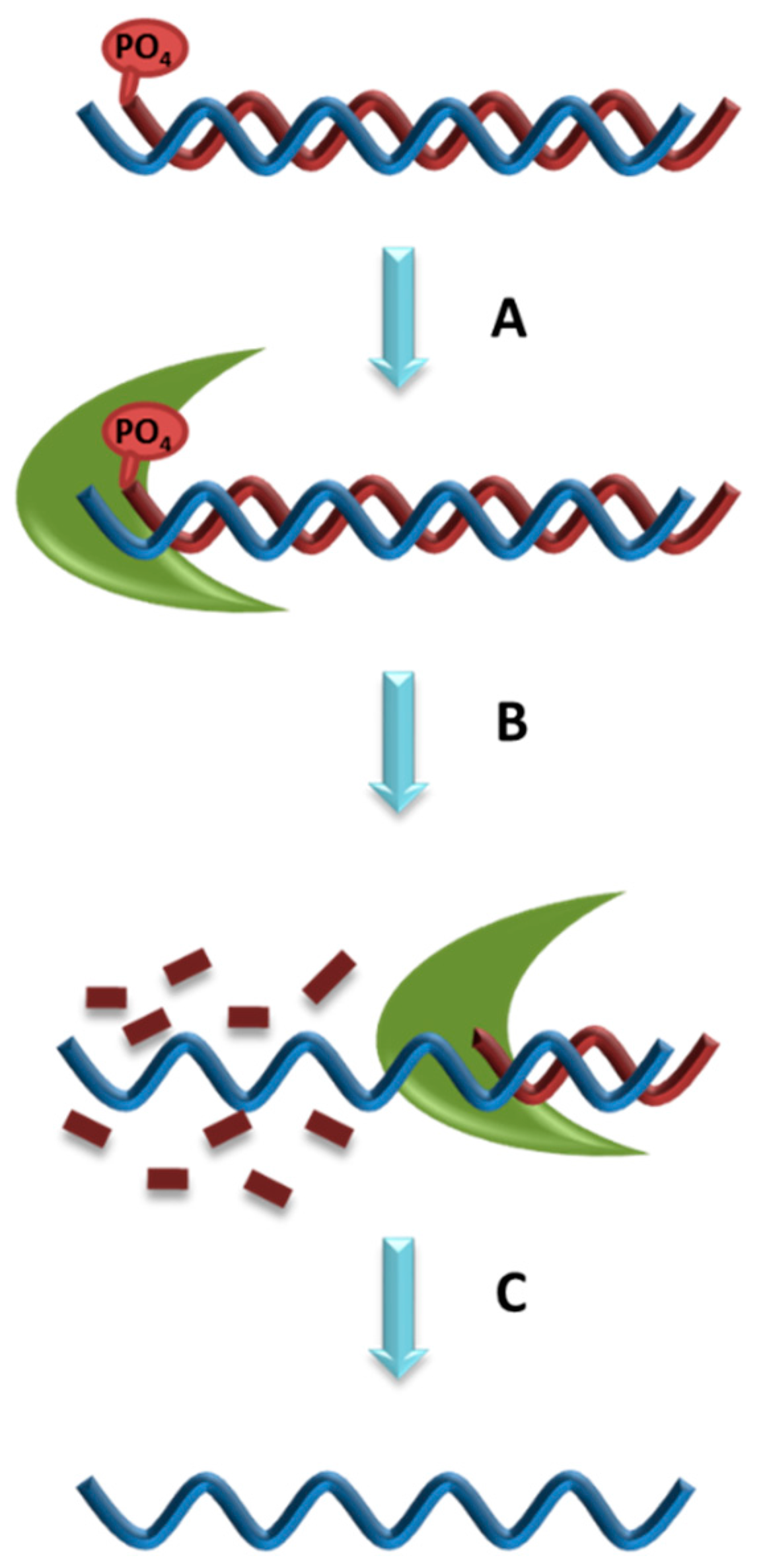
5.4. Lambda Exonuclease Digestion
6. Aptamer Binding Evaluation
6.1. Flow Cytometry
6.2. Enzyme-Linked Assays
6.3. Fluorescence and Confocal Microscopy
6.4. Radioactive Scintillation Counting
7. Aptamers Developed by Cell-SELEX
8. Aptamers Developed Against Cell-Specific Markers
9. Conclusions
Acknowledgments
Conflicts of Interest
References
- Fang, X.; Tan, W. Aptamers generated from cell-selex for molecular medicine: A chemical biology approach. Acc. Chem. Res. 2009, 43, 48–57. [Google Scholar] [CrossRef] [PubMed]
- Orava, E.W.; Cicmil, N.; Gariépy, J. Delivering cargoes into cancer cells using DNA aptamers targeting internalized surface portals. Biochim. Biophys. Acta 2010, 1798, 2190–2200. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, S.C.B. Methods developed for selex. Anal. Bioanal. Chem. 2007, 387, 171–182. [Google Scholar] [CrossRef] [PubMed]
- Stoltenburg, R.; Reinemann, C.; Strehlitz, B. Selex—A (r) evolutionary method to generate high-affinity nucleic acid ligands. Biomol. Eng. 2007, 24, 381–403. [Google Scholar] [CrossRef] [PubMed]
- Lipi, F.; Chen, S.; Chakravarthy, M.; Rakesh, S.; Veedu, R.N. In vitro evolution of chemically-modified nucleic acid aptamers: Pros and cons, and comprehensive selection strategies. RNA Biol. 2016, 13, 1232–1245. [Google Scholar] [CrossRef] [PubMed]
- Lauridsen, L.H.; Shamaileh, H.A.; Edwards, S.L.; Taran, E.; Veedu, R.N. Rapid one-step selection method for generating nucleic acid aptamers: Development of a DNA aptamer against α-bungarotoxin. PLoS ONE 2012, 7, e41702. [Google Scholar] [CrossRef] [PubMed]
- Gopinath, S.C.B.; Lakshmipriya, T.; Chen, Y.; Arshad, M.M.; Kerishnan, J.P.; Ruslinda, A.; Al-Douri, Y.; Voon, C.; Hashim, U. Cell-targeting aptamers act as intracellular delivery vehicles. Appl. Microbiol. Biotechnol. 2016, 100, 6955–6969. [Google Scholar] [CrossRef] [PubMed]
- Thiel, W.H.; Thiel, K.W.; Flenker, K.S.; Bair, T.; Dupuy, A.J.; McNamara, J.O.; Miller, F.J.; Giangrande, P.H. Cell-internalization selex: Method for identifying cell-internalizing rna aptamers for delivering sirnas to target cells. Methods Mol. Biol. 2015, 1218, 187–199. [Google Scholar] [CrossRef]
- Meyer, C.; Hahn, U.; Rentmeister, A. Cell-specific aptamers as emerging therapeutics. J. Nucleic Acids 2011, 2011, 904750. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Ali, Z.; Li, S.; Liu, B.; He, N. Aptamers generated from cell-systematic evolution of ligands through exponential enrichment and their applications. J. Nanosci. Nanotechnol. 2016, 16, 9346–9358. [Google Scholar] [CrossRef]
- Kunert, R.; Reinhart, D. Advances in recombinant antibody manufacturing. Appl. Microbiol. Biotechnol. 2016, 100, 3451–3461. [Google Scholar] [CrossRef] [PubMed]
- Jayasena, S.D. Aptamers: An emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar] [PubMed]
- Veedu, R.N. Aptamers: Tools for Nanotherapy and Molecular Imaging; CRC Press: Boca Raton, FL, USA, 2017. [Google Scholar]
- Teng, J.; Yuan, F.; Ye, Y.; Zheng, L.; Yao, L.; Xue, F.; Chen, W.; Li, B. Aptamer-based technologies in foodborne pathogen detection. Front. Microbiol. 2016, 7, 1426. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Yu, Y.; Jiang, F.; Zhou, J.; Li, Y.; Liang, C.; Dang, L.; Lu, A.; Zhang, G. Development of cell-selex technology and its application in cancer diagnosis and therapy. Int. J. Mol. Sci. 2016, 17, 2079. [Google Scholar] [CrossRef] [PubMed]
- Raddatz, M.S.L.; Dolf, A.; Endl, E.; Knolle, P.; Famulok, M.; Mayer, G. Enrichment of cell-targeting and population-specific aptamers by fluorescence-activated cell sorting. Angew. Chem. Int. Ed. Engl. 2008, 47, 5190–5193. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; Shangguan, D.; Xiong, X.; O’donoghue, M.B.; Tan, W. Development of DNA aptamers using cell-selex. Nat. Protoc. 2010, 5, 1169–1185. [Google Scholar] [CrossRef] [PubMed]
- Tolle, F.; Wilke, J.; Wengel, J.; Mayer, G. By-product formation in repetitive PCR amplification of DNA libraries during selex. PLoS ONE 2014, 9, e114693. [Google Scholar] [CrossRef] [PubMed]
- Citartan, M.; Tang, T.-H.; Tan, S.-C.; Hoe, C.-H.; Saini, R.; Tominaga, J.; Gopinath, S.C. Asymmetric PCR for good quality ssDNA generation towards DNA aptamer production. Songklanakarin J. Sci. Technol. 2012, 34, 125. [Google Scholar]
- Poddar, S. Symmetric vs asymmetric PCR and molecular beacon probe in the detection of a target gene of adenovirus. Mol. Cell. Probes 2000, 14, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Gyllensten, U.B.; Erlich, H.A. Generation of single-stranded DNA by the polymerase chain reaction and its application to direct sequencing of the HLA-DQA locus. Proc. Natl. Acad. Sci. USA 1988, 85, 7652–7656. [Google Scholar] [CrossRef] [PubMed]
- Tabarzad, M.; Kazemi, B.; Vahidi, H.; Aboofazeli, R.; Shahhosseini, S.; Nafissi-Varcheh, N. Challenges to design and develop of DNA aptamers for protein targets. I. Optimization of asymmetric PCR for generation of a single stranded DNA library. Iran. J. Pharm. Res. 2014, 13, 133. [Google Scholar] [PubMed]
- Sanchez, J.A.; Pierce, K.E.; Rice, J.E.; Wangh, L.J. Linear-after-the-exponential (late)–PCR: An advanced method of asymmetric PCR and its uses in quantitative real-time analysis. Proc. Natl. Acad. Sci. USA 2004, 101, 1933–1938. [Google Scholar] [CrossRef] [PubMed]
- Summer, H.; Grämer, R.; Dröge, P. Denaturing urea polyacrylamide gel electrophoresis (urea page). JoVE J. Vis. Exp. 2009, 32, e1485. [Google Scholar] [CrossRef] [PubMed]
- Keefe, A.D.; Pai, S.; Ellington, A. Aptamers as therapeutics. Nat. Rev. Drug Discov. 2010, 9, 537–550. [Google Scholar] [CrossRef] [PubMed]
- Hempelmann, E.; Wilson, R.J.M. Detection of glucose-6-phosphate dehydrogenase in malarial parasites. Mol. Biochem. Parasitol. 1981, 2, 197–204. [Google Scholar] [CrossRef]
- Walder, R.Y.; Hayes, J.R.; Walder, J.A. Use of PCR primers containing a 3′terminal ribose residuce to prevent cross-contamination of amplified sequences. Nucleic Acids Res. 1993, 21, 4339–4343. [Google Scholar] [CrossRef] [PubMed]
- Paul, A.; Avci-Adali, M.; Ziemer, G.; Wendel, H.P. Streptavidin-coated magnetic beads for DNA strand separation implicate a multitude of problems during cell-selex. Oligonucleotides 2009, 19, 243–254. [Google Scholar] [CrossRef] [PubMed]
- DeGrasse, J.A. A single-stranded DNA aptamer that selectively binds to staphylococcus aureus enterotoxin b. PLoS ONE 2012, 7, e33410. [Google Scholar] [CrossRef] [PubMed]
- Citartan, M.; Tang, T.-H.; Tan, S.-C.; Gopinath, S.C. Conditions optimized for the preparation of single-stranded DNA (ssDNA) employing lambda exonuclease digestion in generating DNA aptamer. World J. Microbiol. Biotechnol. 2011, 27, 1167–1173. [Google Scholar] [CrossRef]
- Avci-Adali, M.; Paul, A.; Wilhelm, N.; Ziemer, G.; Wendel, H.P. Upgrading selex technology by using lambda exonuclease digestion for single-stranded DNA generation. Molecules 2009, 15, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.S.; Ahmad, K.M.; Cho, M.; Kim, S.; Xiao, Y.; Soh, H.T. Improving aptamer selection efficiency through volume dilution, magnetic concentration, and continuous washing in microfluidic channels. Anal. Chem. 2011, 83, 6883–6889. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.Y.; Acquah, C.; Sidhu, A.; Ongkudon, C.M.; Yon, L.; Danquah, M.K. Selex modifications and bioanalytical techniques for aptamer–target binding characterization. Crit. Rev. Anal. Chem. 2016, 46, 521–537. [Google Scholar] [CrossRef] [PubMed]
- Shigdar, S.; Qiao, L.; Zhou, S.-F.; Xiang, D.; Wang, T.; Li, Y.; Lim, L.Y.; Kong, L.; Li, L.; Duan, W. Rna aptamers targeting cancer stem cell marker cd133. Cancer Lett. 2013, 330, 84–95. [Google Scholar] [CrossRef] [PubMed]
- Sefah, K.; Tang, Z.; Shangguan, D.; Chen, H.; Lopez-Colon, D.; Li, Y.; Parekh, P.; Martin, J.; Meng, L.; Phillips, J. Molecular recognition of acute myeloid leukemia using aptamers. Leukemia 2009, 23, 235–244. [Google Scholar] [CrossRef] [PubMed]
- Tang, Z.; Shangguan, D.; Wang, K.; Shi, H.; Sefah, K.; Mallikratchy, P.; Chen, H.W.; Li, Y.; Tan, W. Selection of aptamers for molecular recognition and characterization of cancer cells. Anal. Chem. 2007, 79, 4900–4907. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, H.P.; Smiley, R.D.; Jaykus, L.-A. Selection and characterization of DNA aptamers with binding selectivity to campylobacter jejuni using whole-cell selex. Appl. Microbiol. Biotechnol. 2010, 87, 2323–2334. [Google Scholar] [CrossRef] [PubMed]
- Kunii, T.; Ogura, S.-I.; Mie, M.; Kobatake, E. Selection of DNA aptamers recognizing small cell lung cancer using living cell-selex. Analyst 2011, 136, 1310–1312. [Google Scholar] [CrossRef] [PubMed]
- Drolet, D.W.; Moon-McDermott, L.; Romig, T.S. An enzyme-linked oligonucleotide assay. Nat. Biotechnol 1996, 14, 1021–1025. [Google Scholar] [CrossRef] [PubMed]
- Nagarkatti, R.; de Araujo, F.F.; Gupta, C.; Debrabant, A. Aptamer based, non-PCR, non-serological detection of chagas disease biomarkers in Trypanosoma cruzi infected mice. PLoS Negl. Trop. Dis. 2014, 8, e2650. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Liang, H.; Wu, X.; Gao, Y.; Zhang, X. Cell-ELA-based determination of binding affinity of DNA aptamer against U87-EGFRvIII cell. Sheng Wu Gong Cheng Xue Bao 2013, 29, 664–671. [Google Scholar] [PubMed]
- Daniels, D.A.; Chen, H.; Hicke, B.J.; Swiderek, K.M.; Gold, L. A tenascin-C aptamer identified by tumor cell SELEX: Systematic evolution of ligands by exponential enrichment. Proc. Natl. Acad. Sci. USA 2003, 100, 15416–15421. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.S.; Huh, Y.M.; Kim, S.; Lee, D.-K. Isolation of RNA aptamers targeting HER-2-overexpressing breast cancer cells using cell-SELEX. Bull. Korean Chem. Soc. 2009, 30, 1827–1831. [Google Scholar] [CrossRef]
- Huang, Y.Z.; Hernandez, F.J.; Gu, B.; Stockdale, K.R.; Nanapaneni, K.; Scheetz, T.E.; Behlke, M.A.; Peek, A.S.; Bair, T.; Giangrande, P.H. RNA aptamer-based functional ligands of the neurotrophin receptor, trkb. Mol. Pharmacol. 2012, 82, 623–635. [Google Scholar] [CrossRef] [PubMed]
- Shangguan, D.; Li, Y.; Tang, Z.; Cao, Z.C.; Chen, H.W.; Mallikaratchy, P.; Sefah, K.; Yang, C.J.; Tan, W. Aptamers evolved from live cells as effective molecular probes for cancer study. Proc. Natl. Acad. Sci. USA 2006, 103, 11838–11843. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Chen, Z.; Liu, M.; Deng, Y.; Li, S.; He, N. The aptamers generated from HepG2 cells. Sci. China Chem. 2017, 60. [Google Scholar] [CrossRef]
- Kim, Y.J.; Lee, H.S.; Jung, D.E.; Kim, J.M.; Song, S.Y. The DNA aptamer binds stemness-enriched cancer cells in pancreatic cancer. J. Mol. Recognit. 2016, 30, e2591. [Google Scholar] [CrossRef] [PubMed]
- Graham, J.C.; Zarbl, H. Use of cell-SELEX to generate DNA aptamers as molecular probes of HPV-associated cervical cancer cells. PLoS ONE 2012, 7, e36103. [Google Scholar] [CrossRef] [PubMed]
- Goringer, H.; Adler, A.; Forster, N.; Homann, M. Post-SELEX chemical optimization of a trypanosome-specific RNA aptamer. Comb. Chem. High Throughput Screen. 2008, 11, 16–23. [Google Scholar] [CrossRef]
- Pathania, P.; Sharma, A.; Kumar, B.; Rishi, P.; Suri, C.R. Selective identification of specific aptamers for the detection of non-typhoidal salmonellosis in an apta-impedimetric sensing format. Microchim. Acta 2017, 184, 1499–1508. [Google Scholar] [CrossRef]
- Pan, Q.; Zhang, X.-L.; Wu, H.-Y.; He, P.-W.; Wang, F.; Zhang, M.-S.; Hu, J.-M.; Xia, B.; Wu, J. Aptamers that preferentially bind type IVB pili and inhibit human monocytic-cell invasion by Salmonella enterica serovar typhi. Antimicrob. Agents Chemother. 2005, 49, 4052–4060. [Google Scholar] [CrossRef] [PubMed]
- Quang, N.N.; Miodek, A.; Cibiel, A.; Ducongé, F. Selection of aptamers against whole living cells: From cell-selex to identification of biomarkers. Synth. Antib. Methods Protoc. 2017, 253–272. [Google Scholar] [CrossRef]
- Wang, C.; Zhang, M.; Yang, G.; Zhang, D.; Ding, H.; Wang, H.; Fan, M.; Shen, B.; Shao, N. Single-stranded DNA aptamers that bind differentiated but not parental cells: Subtractive systematic evolution of ligands by exponential enrichment. J. Biotechnol. 2003, 102, 15–22. [Google Scholar] [CrossRef]
- Gao, H.; Qian, J.; Cao, S.; Yang, Z.; Pang, Z.; Pan, S.; Fan, L.; Xi, Z.; Jiang, X.; Zhang, Q. Precise glioma targeting of and penetration by aptamer and peptide dual-functioned nanoparticles. Biomaterials 2012, 33, 5115–5123. [Google Scholar] [CrossRef] [PubMed]
- Ninomiya, K.; Kaneda, K.; Kawashima, S.; Miyachi, Y.; Ogino, C.; Shimizu, N. Cell-SELEX based selection and characterization of DNA aptamer recognizing human hepatocarcinoma. Bioorg. Med. Chem. Lett. 2013, 23, 1797–1802. [Google Scholar] [CrossRef] [PubMed]
- Tan, Y.; Shi, Y.-S.; Wu, X.-D.; Liang, H.-Y.; Gao, Y.-B.; Li, S.-J.; Zhang, X.-M.; Wang, F.; Gao, T.-M. DNA aptamers that target human glioblastoma multiforme cells overexpressing epidermal growth factor receptor variant III in vitro. Acta Pharmacol. Sin. 2013, 34, 1491. [Google Scholar] [CrossRef] [PubMed]
- Hung, L.-Y.; Wang, C.-H.; Hsu, K.-F.; Chou, C.-Y.; Lee, G.-B. An on-chip cell-SELEX process for automatic selection of high-affinity aptamers specific to different histologically classified ovarian cancer cells. Lab Chip 2014, 14, 4017–4028. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Liang, H.; Tan, Y.; Yuan, C.; Li, S.; Li, X.; Li, G.; Shi, Y.; Zhang, X. Cell-SELEX aptamer for highly specific radionuclide molecular imaging of glioblastoma in vivo. PLoS ONE 2014, 9, e90752. [Google Scholar] [CrossRef] [PubMed]
- Li, W.-M.; Bing, T.; Wei, J.-Y.; Chen, Z.-Z.; Shangguan, D.-H.; Fang, J. Cell-SELEX-based selection of aptamers that recognize distinct targets on metastatic colorectal cancer cells. Biomaterials 2014, 35, 6998–7007. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Luo, Y.; Bing, T.; Chen, Z.; Lu, M.; Zhang, N.; Shangguan, D.; Gao, X. DNA aptamer evolved by cell-selex for recognition of prostate cancer. PLoS ONE 2014, 9, e100243. [Google Scholar] [CrossRef] [PubMed]
- Cibiel, A.; Quang, N.N.; Gombert, K.; Thézé, B.; Garofalakis, A.; Ducongé, F. From ugly duckling to swan: Unexpected identification from cell-SELEX of an anti-Annexin A2 aptamer targeting tumors. PLoS ONE 2014, 9, e87002. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.Y.; Kim, J.W.; Kim, W.K.; Han, B.S.; Park, S.G.; Chung, B.H.; Lee, S.C.; Bae, K.-H. Selection of aptamers for mature white adipocytes by cell SELEX using flow cytometry. PLoS ONE 2014, 9, e97747. [Google Scholar] [CrossRef] [PubMed]
- Souza, A.G.; Marangoni, K.; Fujimura, P.T.; Alves, P.T.; Silva, M.J.; Bastos, V.A.F.; Goulart, L.R.; Goulart, V.A. 3D cell-SELEX: Development of RNA aptamers as molecular probes for PC-3 tumor cell line. Exp. Cell Res. 2016, 341, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Teng, I.-T.; Zhang, L.; Delgado, S.; Champanhac, C.; Cansiz, S.; Wu, C.; Shan, H.; Tan, W. Molecular recognition of human liver cancer cells using DNA aptamers generated via cell-SELEX. PLoS ONE 2015, 10, e0125863. [Google Scholar] [CrossRef] [PubMed]
- Hung, L.-Y.; Wang, C.-H.; Che, Y.-J.; Fu, C.-Y.; Chang, H.-Y.; Wang, K.; Lee, G.-B. Screening of aptamers specific to colorectal cancer cells and stem cells by utilizing on-chip cell-SELEX. Sci. Rep. 2015, 5, 10326. [Google Scholar] [CrossRef] [PubMed]
- Rong, Y.; Chen, H.; Zhou, X.-F.; Yin, C.-Q.; Wang, B.-C.; Peng, C.-W.; Liu, S.-P.; Wang, F.-B. Identification of an aptamer through whole cell-SELEX for targeting high metastatic liver cancers. Oncotarget 2016, 7, 8282. [Google Scholar] [CrossRef] [PubMed]
- Chandrasekaran, R.; Lee, A.S.W.; Yap, L.W.; Jans, D.A.; Wagstaff, K.M.; Cheng, W. Tumor cell-specific photothermal killing by SELEX-derived DNA aptamer-targeted gold nanorods. Nanoscale 2016, 8, 187–196. [Google Scholar] [CrossRef] [PubMed]
- Jia, W.; Ren, C.; Wang, L.; Zhu, B.; Jia, W.; Gao, M.; Zeng, F.; Zeng, L.; Xia, X.; Zhang, X. Cd109 is identified as a potential nasopharyngeal carcinoma biomarker using aptamer selected by cell-SELEX. Oncotarget 2016, 7, 55328. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Zhang, Y.; Chen, Y.; Hong, S.; Sun, Y.; Sun, N.; Pei, R. In vitro selection of DNA aptamers against renal cell carcinoma using living cell-SELEX. Talanta 2017, 175, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Yuan, B.; Jiang, X.; Chen, Y.; Guo, Q.; Wang, K.; Meng, X.; Huang, Z.; Wen, X. Metastatic cancer cell and tissue-specific fluorescence imaging using a new DNA aptamer developed by cell-SELEX. Talanta 2017, 170, 56–62. [Google Scholar] [CrossRef] [PubMed]
- Mercier, M.-C.; Dontenwill, M.; Choulier, L. Selection of nucleic acid aptamers targeting tumor cell-surface protein biomarkers. Cancers 2017, 9, 69. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Li, G. Aptamers against cell surface receptors: Selection, modification and application. Curr. Med. Chem. 2011, 18, 4107–4116. [Google Scholar] [CrossRef] [PubMed]
- Shigdar, S.; Lin, J.; Yu, Y.; Pastuovic, M.; Wei, M.; Duan, W. Rna aptamer against a cancer stem cell marker epithelial cell adhesion molecule. Cancer Sci. 2011, 102, 991–998. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Erviti, L.; Seow, Y.; Yin, H.; Betts, C.; Lakhal, S.; Wood, M.J. Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 2011, 29, 341–345. [Google Scholar] [CrossRef] [PubMed]
- McNamara, J.O.; Kolonias, D.; Pastor, F.; Mittler, R.S.; Chen, L.; Giangrande, P.H.; Sullenger, B.; Gilboa, E. Multivalent 4–1BB binding aptamers costimulate CD8+ T cells and inhibit tumor growth in mice. J. Clin. Investig. 2008, 118, 376–386. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not Available. |
| Aptamer | Method | Target Cell | Kd (nM) | Reference |
|---|---|---|---|---|
| RNA | Cell-SELEX | HER-2-overexpressing breast cancer | 94.6 | [43] |
| DNA | Cell-SELEX | Acute myeloid leukemia cells | 5.4 ± 1.6 | [35] |
| RNA | Cell-SELEX | Mouse embryonic stem cells | - | [54] |
| DNA | Cell-SELEX | Human hepatocarcinoma | 19–450 | [55] |
| RNA | Cell-SELEX | AC133-epitope of CD133 expressed on HEK293T cells | 33.85–145 | [34] |
| DNA | Cell-SELEX | Over-expressing epidermal growth factor receptor variant III on human glioblastoma | ≤100 | [56] |
| DNA | on-chip Cell-SELEX | Different histologically classified ovarian cancer cells | 1.3 | [57] |
| DNA | Cell-SELEX | Epidermal growth factor receptor variant III on Glioblastoma | 3.37 ± 0.98 | [58] |
| DNA | Cell-SELEX | Metastatic colorectal cell | 8.1 ± 0.9 | [59] |
| DNA | Cell-SELEX | Prostate cancer cells | 73.59 ± 11.01 | [60] |
| RNA | Cell-SELEX | Annexin A2 | 10.5 ± 4.6 | [61] |
| DNA | Cell-SELEX By flow cytometry | 3T3-L1 adipocytes cells | 33.1 ± 2.9 | [62] |
| RNA | Cell-SELEX | PC3-prostate cancer cell | 71.57 ± 12.96 | [63] |
| DNA | Cell-SELEX | Human hepatoma cells HepG2 | 64–349 | [64] |
| DNA | On-chip Cell-SELEX | Colorectal cancer cells | 12.3 | [65] |
| DNA | Cell-SELEX | Hepatocellular carcinoma | 9.4 ± 2.0 | [66] |
| DNA | Cell internalization SELEX | MCF10CA1h human breast ductal carcinoma | [67] | |
| DNA | Cell-SELEX | Nasopharyngeal carcinoma cell lines | 11.93 ± 1.40 | [68] |
| DNA | Cell-SELEX | Renal cell carcinoma (768-O) | 9.4 ± 2.0 | [69] |
| DNA | Cell-SELEX | Metastatic colorectal carcinoma LOVO cells | 167.3 ± 30.2 | [70] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rahimizadeh, K.; AlShamaileh, H.; Fratini, M.; Chakravarthy, M.; Stephen, M.; Shigdar, S.; Veedu, R.N. Development of Cell-Specific Aptamers: Recent Advances and Insight into the Selection Procedures. Molecules 2017, 22, 2070. https://doi.org/10.3390/molecules22122070
Rahimizadeh K, AlShamaileh H, Fratini M, Chakravarthy M, Stephen M, Shigdar S, Veedu RN. Development of Cell-Specific Aptamers: Recent Advances and Insight into the Selection Procedures. Molecules. 2017; 22(12):2070. https://doi.org/10.3390/molecules22122070
Chicago/Turabian StyleRahimizadeh, Kamal, Hadi AlShamaileh, Milena Fratini, Madhuri Chakravarthy, Michelle Stephen, Sarah Shigdar, and Rakesh N. Veedu. 2017. "Development of Cell-Specific Aptamers: Recent Advances and Insight into the Selection Procedures" Molecules 22, no. 12: 2070. https://doi.org/10.3390/molecules22122070
APA StyleRahimizadeh, K., AlShamaileh, H., Fratini, M., Chakravarthy, M., Stephen, M., Shigdar, S., & Veedu, R. N. (2017). Development of Cell-Specific Aptamers: Recent Advances and Insight into the Selection Procedures. Molecules, 22(12), 2070. https://doi.org/10.3390/molecules22122070




