Antibiotic Resistant Superbugs: Assessment of the Interrelationship of Occurrence in Clinical Settings and Environmental Niches
Abstract
:1. Introduction
2. The Soil Resistome as a Contributor to AMR
3. RAbs and AMR by Aquatic Microbiota
4. Emergence of AMR in Clinical and Sub-Clinical Settings
5. Reported Emerging Threat Level of AMR
- a
- LuxI/LuxR–type quorum sensing: The signal molecules utilized here are the acyl-homoserine lactones (AHL) and they are found in Gram-negative bacteria, for example the complex QS machinery in Acinetobacter is mediated by LuxI/LuxR system peculiar to Gram-negative bacteria. This cell signalling system is made up of AHL [145].
- b
- Oligopeptide-bicomponental quorum sensing: This utilizes small peptides as signal molecules and are found in Gram-positive bacteria.
6. Coordinated Approaches towards Addressing the Emergence and Spread of AMR
7. Conclusions
Acknowledgments
Conflicts of Interest
References
- Rizzo, L.; Manaia, C.; Merlin, C.; Schwartz, T.; Dagot, C.; Ploy, M.C.; Michael, I.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: A review. Sci. Total Environ. 2013, 447, 345–360. [Google Scholar] [CrossRef] [PubMed]
- Bérdy, J. Bioactive microbial metabolites. J. Antibiot. 2005, 58, 1–26. [Google Scholar] [CrossRef] [PubMed]
- Gros, M.; Petrović, M.; Barceló, D. Multi-residue analytical methods using LC-tandem MS for the determination of pharmaceuticals in environmental and wastewater samples: A review. Anal. Bioanal. Chem. 2006, 386, 941–952. [Google Scholar] [CrossRef] [PubMed]
- Segura, P.A.; François, M.; Gagnon, C.; Sauvé, S. Review of the occurrence of anti-infectives in contaminated wastewaters and natural drinking waters. Environ. Health Perspect. 2009, 117, 675–684. [Google Scholar] [CrossRef] [PubMed]
- Allen, H.K.; Donato, J.; Wang, H.H.; Cloud-Hansen, K.A.; Davies, J.; Handelsman, J. Call of the wild: Antibiotic resistance genes in natural environments. Nat. Rev. Microbiol. 2010, 8, 251–259. [Google Scholar] [CrossRef] [PubMed]
- D’Costa, V.M.; King, C.E.; Kalan, L.; Morar, M.; Sung, W.W.L.; Schwarz, C.; Froese, D.; Zazula, G.; Calmels, F.; Debruyne, R.; et al. Antibiotic resistance is ancient. Nature 2011, 477, 457–461. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, K.J.; Reyes, A.; Wang, B.; Selleck, E.M.; Sommer, M.O.A.; Dantas, G. The shared antibiotic resistome of soil bacteria and human pathogens. Science 2012, 337, 1107–1111. [Google Scholar] [CrossRef] [PubMed]
- Hughes, S.R.; Kay, P.; Brown, L.E. Global synthesis and critical evaluation of pharmaceutical data sets collected from river systems. Environ. Sci. Technol. 2013, 47, 661–677. [Google Scholar] [CrossRef] [PubMed]
- Gillings, M.R. Evolutionary consequences of antibiotic use for the resistome, mobilome and microbial pangenome. Front. Microbiol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Baquero, F.; Martínez, J.L.; Cantón, R. Antibiotics and antibiotic resistance in water environments. Curr. Opin. Biotechnol. 2008, 19, 260–265. [Google Scholar] [CrossRef] [PubMed]
- Pei, R.; Kim, S.C.; Carlson, K.H.; Pruden, A. Effect of river landscape on the sediment concentrations of antibiotics and corresponding antibiotic resistance genes (ARG). Water Res. 2006, 40, 2427–2435. [Google Scholar] [CrossRef] [PubMed]
- Pruden, A.; Arabi, M.; Storteboom, H.N. Correlation between upstream human activities and riverine antibiotic resistance genes. Environ. Sci. Technol. 2012, 46, 11541–11549. [Google Scholar] [CrossRef] [PubMed]
- Khan, G.A.; Berglund, B.; Khan, M.K.; Lindgren, P.E.; Fick, J. Occurrence and abundance of antibiotics and resistance genes in rivers, canal and near drug formulation facilities—A study in Pakistan. PLoS ONE 2013, 8, e62712. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Yu, T.; Zhang, Y.; Yang, M.; Li, Z.; Liu, M.; Qi, R. Antibiotic resistance characteristics of environmental bacteria from an oxytetracycline production wastewater treatment plant and the receiving river. Appl. Environ. Microbiol. 2010, 76, 3444–3451. [Google Scholar] [CrossRef] [PubMed]
- Kristiansson, E.; Fick, J.; Janzon, A.; Grabic, R.; Rutgersson, C.; Weijdegård, B.; Söderström, H.; Joakim Larsson, D.G. Pyrosequencing of antibiotic-contaminated river sediments reveals high levels of resistance and gene transfer elements. PLoS ONE 2011, 6, e17038. [Google Scholar] [CrossRef] [PubMed]
- Holmberg, S.D.; Solomon, S.L.; Blake, P.A. Health and economic impacts of antimicrobial resistance. Rev. Infect. Dis. 1987, 9, 1065–1078. [Google Scholar] [CrossRef] [PubMed]
- Cheng, G.; Hu, Y.; Yin, Y.; Yang, X.; Xiang, C.; Wang, B.; Chen, Y.; Yang, F.; Lei, F.; Wu, N.; et al. Functional screening of antibiotic resistance genes from human gut microbiota reveals a novel gene fusion. FEMS Microbiol. Lett. 2012, 336, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Gullberg, E.; Cao, S.; Berg, O.G.; Ilbäck, C.; Sandegren, L.; Hughes, D.; Andersson, D.I. Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 2011, 7, e1002158. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Global Action Plan on Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2015. [Google Scholar]
- Serrano, P.H. Responsible Use of Antibiotics in Aquaculture; In FAO Fisheries Technical Paper 469 (Rome; United Nations); Food and Agriculture Organization of the United Nations: Rome, Italy, 2005; p. 89. [Google Scholar]
- Smith, P.R.; LeBreton, A.; Horsberg, T.E.; Corsin, F. Guidelines for antimicrobial use in aquaculture. In Guide to Antimicrobial Use in Animals; Guardabassi, L., Jensen, L.B., Kruse, H., Eds.; Blackwell Publishing Ltd.: Oxford, UK, 2009; pp. 207–218. [Google Scholar]
- Waksman, S.A. Antagonistic relations of microorganisms. Bacteriol. Rev. 1941, 5, 231–291. [Google Scholar] [PubMed]
- Lewis, I.M. Bacterial antagonism with special reference to the effect of Pseudomonas fluorescens on spore forming bacteria of soils. J. Bacteriol. 1929, 17, 89–103. [Google Scholar] [PubMed]
- Waksman, S.A.; Woodruff, H.B. The soil as a source of microorganisms antagonistic to disease producing bacteria. J. Bacteriol. 1940, 40, 581–600. [Google Scholar] [PubMed]
- Florey, H.W. The use of micro-organisms for therapeutic purposes. Yale J. Biol. Med. 1946, 19, 101–117. [Google Scholar] [CrossRef] [PubMed]
- Smith, T. Antibiotics from soil bacteria. Nat. Struct. Biol. 2000, 7, 189–190. [Google Scholar] [CrossRef] [PubMed]
- Plough, H.H. Penicillin resistance of Staphylococcus aureus and its clinical implications. Am. J. Clin. Pathol. 1945, 15, 446–451. [Google Scholar] [CrossRef] [PubMed]
- Cox, G.; Wright, G.D. Intrinsic antibiotic resistance: mechanisms, origins, challenges and solutions. Int. J. Med. Microbiol. 2013, 303, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Benveniste, R.; Davies, J. Aminoglycoside antibiotic-inactivating enzymes in actinomycetes similar to those present in clinical isolates of antibiotic-resistant bacteria. Proc. Natl. Acad. Sci. USA 1973, 70, 2276–2280. [Google Scholar] [CrossRef] [PubMed]
- Nesme, J.; Cécillon, S.; Delmont, T.O.; Monier, J.-M.; Vogel, T.M.; Simonet, P. Large-scale metagenomics based study of antibiotic resistance in the environment. Curr. Biol. 2014, 10, 1096–1100. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, G.; Steinberg, N.; Oppenheimer-Shaanan, N.; Olender, T.; Doron, S.; Ben-Ari, J.; Sirota-Madi, A.; Bloom-Ackermann, Z.; Kolodkin-Gal, I. Not so simple, not so subtle: The interspecies competition between Bacillus simplex and Bacillus subtilis and its impact on the evolution of biofilms. NPJ Biofilms Microbiomes 2016, 2, 15027. [Google Scholar] [CrossRef]
- Vas-Moreira, I.; Nune, O.C.; Manaia, C.M. Bacterial diversity and antibiotic resistance in water habitats: searching the links with the human microbiome. FEM Microbiol. Rev. 2014, 38, 718–761. [Google Scholar] [CrossRef] [PubMed]
- Adegoke, A.A.; Awolusi, O.O.; Stenström, T.A. Organic Fertilizers: Public Health Intricacies. In Organic Fertilizers—From Basic Concepts to Applied Outcomes; Larramendy, M., Ed.; INTECH: Vienna, Austria, 2016. [Google Scholar]
- McEwen, S.A.; Fedorka-Cray, P.J. Antimicrobial use and resistance in animals. Clin. Infect. Dis. 2002, 34 (Suppl. S3), S93–S106. [Google Scholar] [CrossRef] [PubMed]
- Butaye, P.; Devriese, L.A.; Haesebrouck, F. Antimicrobial growth promoters used in animal feed: Effects of less well known antibiotics on gram-positive bacteria. Clin. Microbiol. Rev. 2003, 16, 175–188. [Google Scholar] [CrossRef] [PubMed]
- APUA’s International Surveillance of Reservoirs of Resistance (ISRAR) 2008–2010. Available online: http://emerald.tufts.edu/med/apua/research/israr_10_957998750.pdf (accessed on 11 October 2016).
- Wang, M.; Tang, J.C. Research of antibiotics pollution in soil environments and its ecological toxicity. J. Agro-Environ. Sci. 2010, 29, 261–266. [Google Scholar]
- McManus, P.S.; Stockwell, V.O.; Sundin, G.W.; Jones, A.L. Antibiotic use in plant agriculture. Ann. Rev. Phytopathol. 2002, 40, 443–465. [Google Scholar] [CrossRef] [PubMed]
- Pan, X.; Qiang, Z.M.; Ben, W.W.; Chen, M.X. Residual veterinary antibiotics in swine manure from concentrated animal feeding operations in Shandong Province, China. Chemosphere 2011, 84, 695–700. [Google Scholar] [CrossRef] [PubMed]
- Cheng, V.C.C.; Wong, S.C.Y.; Ho, P.L.; Yuen, K.Y. Strategic measures for the control of surging antimicrobial resistance in Hong Kong and mainland of China. Emerg. Microbes Infect. 2015, 4, e8. [Google Scholar] [CrossRef] [PubMed]
- Pace, N.R. Mapping the tree of life: Progress and prospects. Microbiol. Mol. Biol. Rev. 2009, 73, 565–576. [Google Scholar] [CrossRef] [PubMed]
- Hussain, M.; Zouhar, M.; Ryšánek, P.; Anwar, S.A. Relationship between meloidogyne incognita density and plant growth of okra. J. Anim. Plant Sci. 2016, 26, 739–744. [Google Scholar]
- Zhang, H.; Luo, Y.; Wu, L.; Huang, Y.; Christie, P. Residues and potential ecological risks of veterinary antibiotics in manures and composts associated with protected vegetable farming. Environ. Sci. Pollut. Res. Int. 2015, 22, 5908–5918. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Zhou, Y.; Huang, Y.; Wu, L.; Liu, X.; Luo, Y. Residues and risks of veterinary antibiotics in protected vegetable soils following application of different manures. Chemosphere 2016, 152, 229–237. [Google Scholar] [CrossRef] [PubMed]
- Furtula, V.; Farrell, E.G.; Diarrassouba, F.; Rempel, H.; Pritchard, J.; Diarra, M.S. Veterinary pharmaceuticals and antibiotic resistance of Escherichia coli isolates in poultry litter from commercial farms and controlled feeding trials. Poultry Sci. 2010, 89, 180–188. [Google Scholar] [CrossRef] [PubMed]
- Watkinson, A.J.; Murby, E.J.; Kolpin, D.W.; Costanzo, S.D. The occurrence of antibiotics in an urban watershed: from wastewater to drinking water. Sci. Total Environ. 2009, 407, 2711–2723. [Google Scholar] [CrossRef] [PubMed]
- Li, D.; Yang, M.; Hu, J.; Zhang, J.; Liu, R.; Gu, X.; Zhang, Y.; Wang, Z. Antibiotic-resistance profile in environmental bacteria isolated from penicillin production wastewater treatment plant and the receiving river. Environ. Microbiol. 2009, 11, 1506–1517. [Google Scholar] [CrossRef] [PubMed]
- Zuccato, E.; Castiglioni, S.; Fanelli, R. Identification of the pharmaceuticals for human use contaminating the Italian aquatic environment. J. Hazard. Mater. 2005, 122, 205–209. [Google Scholar] [CrossRef] [PubMed]
- Ternes, T.A.; Hirsch, R. Occurrence and behavior of X-ray contrast media in sewage facilities and the aquatic environment. Environ. Sci. Technol. 2000, 34, 2741–2748. [Google Scholar] [CrossRef]
- Castiglioni, S.; Fanelli, R.; Calamari, D.; Bagnati, R.; Zuccato, E. Methodological approaches for studying pharmaceuticals in the environment by comparing predicted and measured concentrations in River Po, Italy. Regul. Toxicol. Pharmacol. 2004, 39, 25–32. [Google Scholar] [CrossRef] [PubMed]
- Berto, J.; Rochenbach, G.C.; Barreiros, M.A.B.; Correa, A.X.R.; Peluso-Silva, S.; Radetski, C.M. Physicochemical, microbiological, and ecotoxicological evaluation of a septic tank/Fenton reaction combination for the treatment of hospital wastewaters. Ecotoxicol. Environ. Saf. 2009, 72, 1076–1081. [Google Scholar] [CrossRef] [PubMed]
- Heuer, H.; Smalla, K. Plasmids foster diversification and adaptation of bacterial populations in soil. FEMS Microbiol. Rev. 2012, 36, 1083–1104. [Google Scholar] [CrossRef] [PubMed]
- Gaze, W.H.; Zhang, L.; Abdouslam, N.A.; Hawkey, P.M.; Calvo-Bado, L.; Royle, J.; Brown, H.; Davis, S.; Kay, P.; Boxall, A.B.; et al. Impacts of anthropogenic activity on the ecology of class 1 integrons and integron-associated genes in the environment. ISME J. 2011, 5, 1253–1261. [Google Scholar] [CrossRef] [PubMed]
- Nardelli, M.; Scalzo, P.M.; Ramírez, M.S.; Quiroga, M.P.; Cassini, M.H.; Centrón, D. Class 1 integrons in environments with different degrees of urbanization. PLoS ONE 2012, 7, e39223. [Google Scholar] [CrossRef] [PubMed]
- British Society for Antimicrobial Chemotherapy. Standing Committee on Susceptibility Testing; Version 14.0; British Society for Antimicrobial Chemotherapy: Birmingham, UK, 2015. [Google Scholar]
- Lai, H.T.; Chien, Y.H.; Lin, J.S. Long-term transformation of oxolinic acid in water from an eel pond. Aquaculture 2008, 275, 96–101. [Google Scholar] [CrossRef]
- Zhang, Q.H.; Yang, W.N.; Ngo, H.H.; Guo, W.S.; Jin, P.K.; Dzakpasu, M.; Yang, S.J.; Wang, Q.; Wang, X.C.; Ao, D. Current status of urban wastewater treatment plants in China. Environ. Int. 2016, 92–93, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Beaber, J.W.; Hochhut, B.; Waldor, M.K. SOS response promotes horizontal dissemination of antibiotic resistance genes. Nature 2004, 427, 72–74. [Google Scholar] [CrossRef] [PubMed]
- Ubeda, C.; Maiques, E.; Knecht, E.; Lasa, I.; Novick, R.P.; Penadés, J.R. Antibiotic-induced SOS response promotes horizontal dissemination of pathogenicity island-encoded virulence factors in Staphylococci. Mol. Microbiol. 2005, 56, 836–844. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.; Davies, D. Origins and Evolution of Antibiotic Resistance. Microbiol. Mol. Biol. Rev. 2010, 74, 417–433. [Google Scholar] [CrossRef] [PubMed]
- Garcilla´n-Barcia, M.P.; Alvarado, A.; de la Cruz, F. Identification of bacterial plasmids based on mobility and plasmid population biology. FEMS Microbiol. Rev. 2011, 35, 936–956. [Google Scholar] [CrossRef] [PubMed]
- Partridge, S.R. Analysis of antibiotic resistance regions in Gram-negative bacteria. FEMS Microbiol. Rev. 2011, 35, 820–855. [Google Scholar] [CrossRef] [PubMed]
- Zhang, T.; Zhang, X.X.; Ye, L. Plasmid metagenome reveals high levels of antibiotic resistance genes and mobile genetic elements inactivated sludge. PLoS ONE 2011, 6, e26041. [Google Scholar] [CrossRef] [PubMed]
- Farias, T.; de Menorval, L.C.; Zajac, J.; Rivera, A. Benzalkonium chloride and sulfamethoxazole adsorption onto natural clinoptilolite: Effect of time, ionic strength, pH and temperature. J. Colloid Interface Sci. 2011, 363, 465–475. [Google Scholar] [CrossRef] [PubMed]
- Xi, C.; Zhang, Y.; Marrs, C.F.; Wen, Y.; Carl, S.; Betsy, F.; Jerome, N. Prevalence of antibiotic resistance in drinking water treatment and distribution systems. Appl. Environ. Microbiol. 2009, 75, 5714–5718. [Google Scholar] [CrossRef] [PubMed]
- Figueira, V.; Vaz-Moreira, I.; Silva, M.; Manaia, C.M. Diversity and antibiotic resistance of Aeromonas spp. In drinking and waste water treatment plants. Water Res. 2011, 45, 5599–5611. [Google Scholar] [CrossRef] [PubMed]
- Stenstrӧm, T.A.; Okoh, A.I.; Adegoke, A.A. Antibiogram of environmental isolates of Acinetobacter calcoaceticus from Nkonkobe Municipality, South Africa. Fresenius Environ. Bull. 2016, 25, 3059–3065. [Google Scholar]
- Brusselaers, N.; Vogelaers, D.; Blot, S. The rising problem of antimicrobial resistance in the intensive care unit. Ann. Intensive Care 2011, 1, 47. [Google Scholar] [CrossRef] [PubMed]
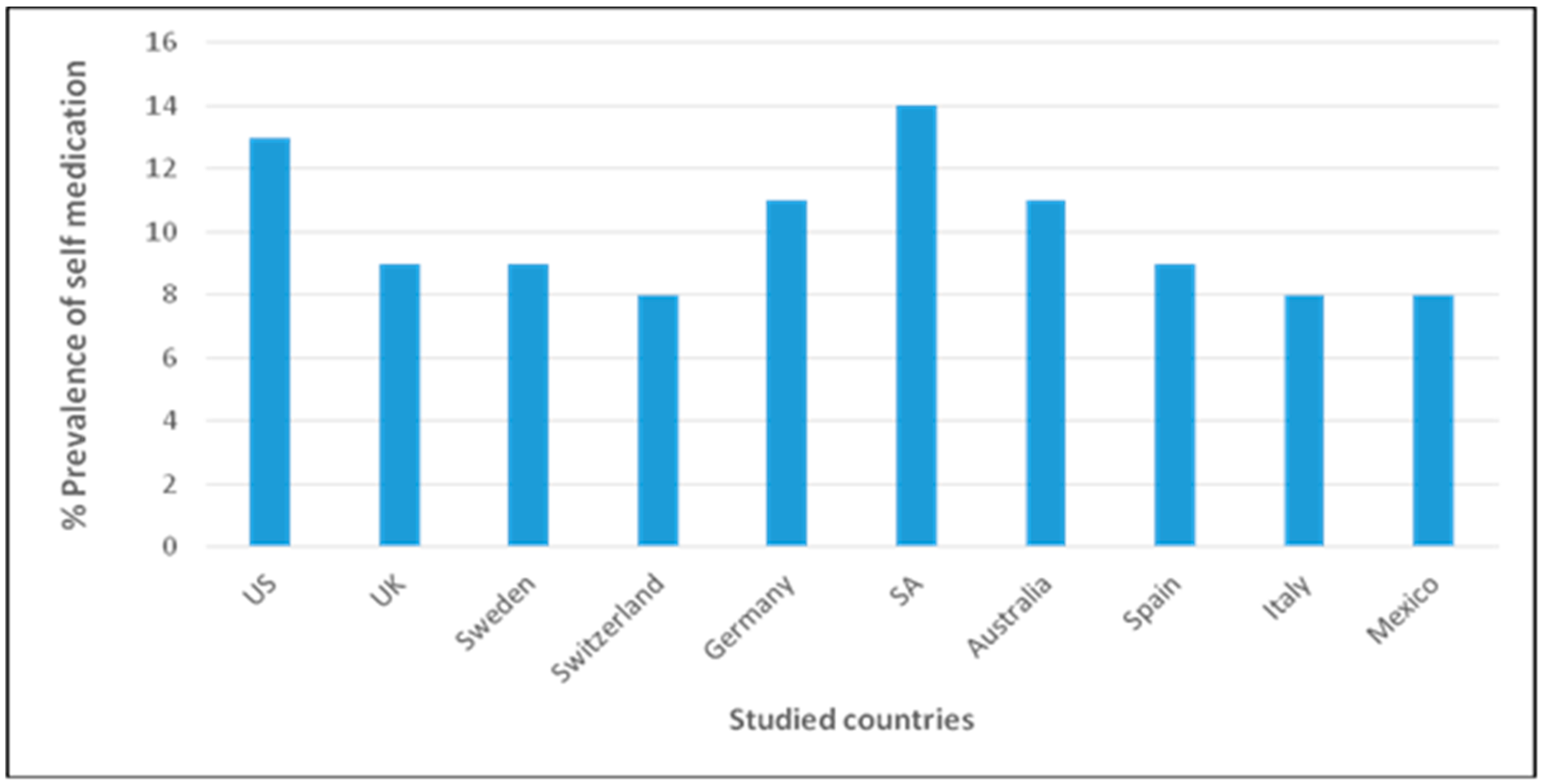
- Vizhi, S.K.; Senapathi, R. Evaluation of the perception, attitude and practice of self-medication among business students in 3 select Cities, South India. Int. J. Enterp. Innov. Manag. Stud. 2010, 1, 40–44. [Google Scholar]
- Bennadi, D. Self-medication: A current challenge. J. Basic Clin. Pharm. 2014, 5, 19–23. [Google Scholar] [CrossRef] [PubMed]
- Pagán, J.A.; Ross, S.; Yau, J.; Polsky, D. Self-medication and health insurance coverage in Mexico. Health Policy 2006, 75, 170–177. [Google Scholar] [CrossRef] [PubMed]
- Pan, H.; Cui, B.; Zhang, D.; Farrar, J.; Law, F.; Ba-Thein, W. Prior knowledge, older age, and higher allowance are risk factors for self-medication with antibiotics among university students in southern China. PLoS ONE 2012, 7, e41314. [Google Scholar] [CrossRef] [PubMed]
- Luyt, C.E.; Bréchot, N.; Trouillet, J.L.; Chastre, J. Antibiotic stewardship in the intensive care unit. Crit. Care 2014, 18, 480. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention, Office of Infectious Disease Antibiotic Resistance Threats in the United States, 2013. Available online: http://www.cdc.gov/drugresistance/threat-report-2013 (accessed on 28 June 2016).
- Ventola, L. The Antibiotic Resistance Crisis. Pharm. Ther. 2015, 40, 277–284. [Google Scholar]
- Senekal, M. Current resistance issues in antimicrobial therapy. CME 2010, 28, 54–57. [Google Scholar]
- Perez, F.; Hujer, A.M.; Hujer, K.M.; Decker, B.K.; Rather, P.N.; Bonomo, R.A. Global challenge of multidrug-resistant Acinetobacter baumannii. Antimicrob. Agents Chemother. 2007, 51, 3471–3484. [Google Scholar] [CrossRef] [PubMed]
- Falagas, M.E.; Karageorgopoulos, D.E. Pandrug Resistance (PDR), Extensive Drug Resistance (XDR), and Multidrug Resistance (MDR) among Gram-Negative Bacilli: Need for International Harmonization in Terminology. Clin. Infect. Dis. 2008, 46, 1121–1122. [Google Scholar] [CrossRef] [PubMed]
- Adegoke, A.A.; Mvuyo, T.; Okoh, A.I. Ubiquitous Acinetobacter species as beneficial commensals but gradually being emboldened with antibiotic resistance genes. J. Basic Microbiol. 2012, 52, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Shakibaie, M.R.; Adeli, S.; Salehi, M.H. Antibiotic resistance patterns and extended spectrum beta-lactamase production among Acinetobacter spp. isolated from anintensive care Unit of a hospital in Kerman, Iran. Antimicrob. Resist. Infect. Control 2012, 1, 1. [Google Scholar] [CrossRef] [PubMed]
- Sivaranjani, V.; Umadevi, S.; Srirangaraj, S.; Kali, A.; Seetha, K.S. Multi-drug resistant Acinetobacter species from various clinical samples in a tertiary care hospital from South India. AMJ 2013, 6, 697–700. [Google Scholar] [CrossRef] [PubMed]
- Souli, M.; Galani, I.; Giamarellou, H. Emergence of extensively drug—Resistant and pandrug—Resistant Gram—Negative bacilli in Europe. Eurosurveillance 2008, 13, 1–11. [Google Scholar]
- Luangtongkum, T.; Jeon, B.; Han, J.; Plummer, P.; Logue, C.M.; Zhang, Q. Antibiotic resistance in Campylobacter: Emergence, transmission and persistence. Future Microbiol. 2009, 4, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Lehtopolku, M.; Nakari, U.-M.; Kotilainen, P.; Huovinen, P.; Siitonen, A.; Hakanen, A.J. Antimicrobial Susceptibilities of Multidrug-Resistant Campylobacter jejuni and, C. coli Strains: In Vitro Activities of 20 Antimicrobial Agents. Antimicrob. Agents Chemother. 2010, 54, 1232–1236. [Google Scholar] [CrossRef] [PubMed]
- Wieczorek, K.; Osek, J. Antimicrobial Resistance Mechanisms among Campylobacter. BioMed Res. Int. 2013, 2013, 340605. [Google Scholar] [CrossRef] [PubMed]
- Sison, F.B.; Chaisowwong, W.; Alter, T.; Tiwananthagorn, S.; Pichpol, D.; Lampang, K.N.; Baumann, M.P.O.; Gölz, G. Loads and antimicrobial resistance. Poult. Sci. 2014, 93, 1270–1273. [Google Scholar] [CrossRef] [PubMed]
- Ngulukun, S.S.; Oboegbulem, S.S.; Fagbamila, I.; Barde, I. Antimicrobial resistance of Campylobacter jejuni and Campylobacter coli isolated from chickens in a diagnostic laboratory. Afr. J. Microbiol. Res. 2005, 9, 2197–2201. [Google Scholar]
- Garnacho-Montero, J.; Díaz-Martín, A.; García-Cabrera, P.; de Pipao, M.R.; Herna´ndez-Caballero, C. Risk Factors for Fluconazole-Resistant Candidemia. Antimicrob. Agent Chemother. 2010, 54, 3149–3154. [Google Scholar] [CrossRef] [PubMed]
- Spampinato, C.; Leonardi, D. Candida Infections, Causes, Targets, and Resistance Mechanisms: Traditional and Alternative Antifungal Agents. BioMed Res. Int. 2013, 2013, 204237. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Antifungal Resistance. August 2016. Available online: http://www.cdc.gov/fungal/antifungal-resistance.html (accessed on 9 September 2016). [Google Scholar]
- Liao, X.; Qiu, H.; Li, R.; Guo, F.; Liu, W.; Kang, M.; Kang, Y. Risk factors for fluconazole-resistant invasive candidiasis in intensive care unit patients: An analysis from the China Survey of Candidiasis study. J. Crit. Care 2015, 30, 862.e1–862.e5. [Google Scholar] [CrossRef] [PubMed]
- Owotade, F.J.; Gulube, Z.; Ramla, S.; Patel, M. Antifungal susceptibility of Candida albicans isolated from the oral cavities of patients with HIV infection and cancer. SADJ 2016, 71, 8–11. [Google Scholar]
- Kolar, M.; Bardon, J.; Chroma, M.; Hricova, K.; Stosova, T.; Sauer, P.; Koukalova, D. ESBL and AmpC beta-lactamase-producing Enterobacteriaceae in poultry in the Czech Republic. Vet. Med. 2010, 55, 119–124. [Google Scholar]
- Tärnberg, M. Extended-Spectrum Beta-Lactamase Producing Enterobacteriaceae; Linköping University Medical Dissertations; Linköping University: Linköping, Sweden, 2012. [Google Scholar]
- Zurfluh, K.; Hächler, H.; Nüesch-Inderbinen, M.; Stephan, R. Characteristics of Extended-Spectrum β-Lactamase-and Carbapenemase-Producing Enterobacteriaceae Isolates from Rivers and Lakes in Switzerland. Appl. Environ. Microbiol. 2013, 79, 3021–3026. [Google Scholar] [CrossRef] [PubMed]
- Kandeel, A. Prevalence and risk factors of extended-spectrum β-lactamases producing Enterobacteriaceae in a general hospital in Saudi Arabia. J. Microbiol. Infect. Dis. 2014, 4, 50–54. [Google Scholar] [CrossRef]
- Schmiedel, J.; Falgenhauer, L.; Domann, E.; Bauerfeind, R.; Prenger-Berninghoff, E.; Imirzalioglu, C.; Chakraborty, T. Multiresistant extended-spectrum β-lactamase-producing Enterobacteriaceae from humans, companion animals and horses in central Hesse, Germany. BMC Microbiol. 2014, 14, 187. [Google Scholar] [CrossRef] [PubMed]
- Pilmis, B.; Delory, T.; Groh, M.; Weiss, E.; Emirian, A.; Lecuyer, H.; Lesprit, P.; Zahar, J.-R. Extended-spectrum beta-lactamase-producing Enterobacteriaceae (ESBL-PE) infections: Are carbapenem alternatives achievable in daily practice? Int. J. Infect. Dis. 2015, 39, 62–67. [Google Scholar] [CrossRef] [PubMed]
- Lukac, P.J.; Bonomo, R.A.; Logan, L.K. Extended-Spectrum β-Lactamase–Producing Enterobacteriaceae in Children: Old Foe, Emerging Threat. Clin. Infect. Dis. 2015, 60, 1389–1397. [Google Scholar] [CrossRef] [PubMed]
- Willemsen, I.; Oome, S.; Verhulst, C.; Pettersson, A.; Verduin, K.; Kluytmans, J. Trends in Extended Spectrum Beta-Lactamase (ESBL) Producing Enterobacteriaceae and ESBL Genes in a Dutch Teaching Hospital, Measured in 5 Yearly Point Prevalence Surveys (2010–2014). PLoS ONE 2015, 10, e0141765. [Google Scholar] [CrossRef] [PubMed]
- Sahlström, L.; Rehbinder, V.; Albihn, A.; Aspan, A.; Bengtsson, B. Vancomycin resistant enterococci (VRE) in Swedish sewage sludge. Acta Vet. Scand. 2009, 51, 24. [Google Scholar] [CrossRef] [PubMed]
- Molton, J.S.; Tambyah, P.A.; Ang, B.S.P.; Ling, M.L.; Fisher, D.A. The Global Spread of Healthcare-Associated Multidrug-Resistant Bacteria: A Perspective from Asia. Clin. Infect. Dis. 2013, 56, 1310–1318. [Google Scholar] [PubMed]
- Varela, A.R.; Ferro, G.; Vredenburg, J.; Yanik, M.; Vieira, L.; Rizzo, L.; Lameiras, C.; Manaia, C.M. Vancomycin resistant enterococci: From the hospital effluent to the urban wastewater treatment plant. Sci. Total Environ. 2013, 450–451, 155–161. [Google Scholar] [CrossRef] [PubMed]
- Abebe, W.; Endris, M.; Tiruneh, M.; Moges, F. Prevalence of vancomycin resistant Enterococci and associated risk factors among clients with and without HIV in Northwest Ethiopia: A cross-sectional study. BMC Public Health 2014, 14, 185. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, R.E.R.; Micallef, S.A.; Gibbs, S.G.; George, A.; Claye, E.; Sapkota, E.; Josepha, S.W.; Sapkota, A.R. Detection of vancomycin-resistant enterococci (VRE) at four, U.S. wastewater treatment plants that provide effluent for reuse. Sci. Total Environ. 2014, 466–467, 404–411. [Google Scholar] [CrossRef] [PubMed]
- Daniel, D.S.; Lee, S.M.; Dykes, G.A.; Rahman, S. Public Health Risks of Multiple-Drug-Resistant Enterococcus spp. in Southeast Asia. Appl. Environ. Microbiol. 2015, 81, 6090–6097. [Google Scholar] [CrossRef] [PubMed]
- Lister, P.D.; Wolter, D.J.; Hanson, N.D. Antibacterial-Resistant Pseudomonas aeruginosa: Clinical Impact and Complex Regulation of Chromosomally Encoded Resistance Mechanisms. Clin. Microbiol. Rev. 2009, 22, 582–610. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, E.B.; Tam, V.H. Impact of multidrug-resistant Pseudomonas aeruginosa infection on patient outcomes. Expert Rev. Pharmacoecon. Outcomes Res. 2010, 10, 441–451. [Google Scholar] [CrossRef] [PubMed]
- Akingbade, O.A.; Balogun, S.A.; Ojo, D.A.; Afolabi, R.O.; Motayo, B.O.; Okerentugba, P.O.; Okonko, I.O. Plasmid Profile Analysis of Multidrug Resistant Pseudomonas aeruginosa isolated from Wound Infections in South West, Nigeria. World Appl. Sci. J. 2012, 20, 766–775. [Google Scholar]
- Meletis, G.; Bagkeri, M. Pseudomonas aeruginosa: Multi-Drug-Resistance Development and Treatment Options; INTECH: Vienna, Austria, 2013. [Google Scholar]
- Rizvi, M.; Ahmad, J.; Khan, F.; Shukla, I.; Malik, A.; Sami, H. Synergy of drug combinations in treating multidrug-resistant Pseudomonas aeruginosa. Australas. Med. J. 2015, 8, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Yayan, J.; Ghebremedhin, B.; Rasche, K. Antibiotic Resistance of Pseudomonas aeruginosa in Pneumonia at a Single University Hospital Center in Germany over a 10-Year Period. PLoS ONE 2015, 10, e0139836. [Google Scholar] [CrossRef] [PubMed]
- Mijovic, G.; Andric, B.; Terzic, D.; Lopicic, M.; Dupanovic, B. Antibiotic susceptibility of salmonella spp.: A comparison of two surveys with a 5 years interval. J. IMAB Annu. Proc. 2012, 18. [Google Scholar] [CrossRef]
- Fallah, S.H.; Asgharpour, F.; Naderian, Z.; Moulana, Z. Isolation and Determination of Antibiotic Resistance Patterns in Non-typhoid Salmonella spp. isolated from chicken. Int. J. Enteric Pathog. 2013, 1, 17–21. [Google Scholar] [CrossRef]
- Al Ayed, M.S.Z. Prevalence of nontyphoidal Salmonella serotypes and the antimicrobial resistance in pediatric patients in Najran Region, Saudi Arabia. Int. J. Curr. Microbiol. App. Sci. 2014, 3, 103–107. [Google Scholar]
- Khairy, R.M.M. Anti-Microbial Resistance of Non-Typhoid Salmonella in Egypt. Ferment. Technol. 2015, 4, 123. [Google Scholar]
- Eguale, T.; Gebreyes, W.A.; Asrat, D.; Alemayehu, H.; Gunn, J.S.; Engidawork, E. Non-typhoidal Salmonella serotypes, antimicrobial resistance and co-infection with parasites among patients with diarrhea and other gastrointestinal complaints in Addis Ababa, Ethiopia. BMC Infect. Dis 2015, 15, 497. [Google Scholar] [CrossRef] [PubMed]
- Hsu, C.Y.; Hsu, B.M.; Ji, B.M.; Chen, J.S.; Hsu, T.K.; Ji, D.D.; Tseng, S.F.; Chiu, Y.S.; Kao, P.M.; Huang, Y.L. Antibiotic resistance pattern and gene expression of non-typhoid Salmonella in riversheds. Environ. Sci. Pollut. Res. 2015, 22, 7843–7850. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Ke, B.; Deng, X.; Liang, J.; Ran, L.; Lu, L.; He, D.; Huang, Q.; Ke, C.; Li, Z.; et al. Serotypes, seasonal trends, and antibiotic resistance of non-typhoidal Salmonella from human patients in Guangdong Province, China, 2009–2012. BMC Infect. Dis. 2015, 15, 53. [Google Scholar] [CrossRef] [PubMed]
- Tu, L.T.; Hoang, N.V.; Cuong, N.V.; Campbell, J.; Bryant, J.E.; Hoa, N.T.; Kiet, B.T.; Thompson, C.; Duy, D.T.; Phat, V.V.; et al. High levels of contamination and antimicrobial-resistant non-typhoidal Salmonella serovars on pig and poultry farms in the Mekong Delta of Vietnam. Epidemiol. Infect. 2015, 143, 3074–3086. [Google Scholar] [CrossRef] [PubMed]
- Nagshetty, K.; Channappa, S.T.; Gaddad, S.M. Antimicrobial susceptibility of Salmonella typhi in India. J. Infect. Dev. Ctries 2010, 4, 70–73. [Google Scholar] [PubMed]
- Mandal, S.; DebMandal, M.; Pal, N.K. Antibiotic Resistance of Salmonella enterica Serovar Typhi in Kolkata, India, and In Vitro Experiments on Effect of Combined Chemotherapy. Sci. World J. 2012, 2012, 454059. [Google Scholar] [CrossRef] [PubMed]
- Adabara, N.U.; Ezugwu, B.U.; Momojimoh, A.; Madzu, A.; Hashiimu, Z.; Damisa, D. The Prevalence and Antibiotic Susceptibility Pattern of Salmonella typhi among Patients Attending a Military Hospital in Minna, Nigeria. Adv. Prev. Med. 2012, 2012, 875419. [Google Scholar] [CrossRef] [PubMed]
- Ugboko, H.; de, N. Mechanisms of Antibiotic resistance in Salmonella typhi. Int. J. Curr. Microbiol. Appl. Sci. 2014, 3, 461–476. [Google Scholar]
- Balcázar, J.L.; Subirats, J.; Borrego, C.M. The role of biofilms as environmental reservoirs of antibiotic resistance. Front. Microbiol. 2015, 6, 1216. [Google Scholar] [CrossRef] [PubMed]
- Holcomb, H.G.; Durbin, K.J.; Cho, M.; Choi, K.J.; Darling, N.D.; Angerio, A.D. Ethicillin-resistant Staphylococcus aureus as a threat to public health: a cellular approach. GU J. Health Sci. 2008, 5, 2. [Google Scholar]
- Adegoke, A.A.; Komolafe, A.O. Multidrug Resistant Staphylococcus aureus in Clinical Cases in Ile-Ife, Southwest Nigeria. Int. J. Med. Med. Sci. 2009, 1, 68–72. [Google Scholar]
- Frieden, T.R. Maximizing infection prevention in the next decade: Defining the unacceptable. Infect. Control Hosp. Epidemiol. 2010, 31, S1–S3. [Google Scholar] [CrossRef] [PubMed]
- Johnson, A.P. Methicillin-resistant Staphylococcus aureus: the European landscape. Antimicrob. Chemother. 2011, 66 (Suppl. S4), iv43–iv48. [Google Scholar] [CrossRef] [PubMed]
- Neel, R. Multidrug resistance of isolates of methicillin resistant Staphylococcus aureus (MRSA) in paper currency notes from restaurants and hotels in Lusaka in Zambia. Int. J. Pharm. Pharm. Sci. 2012, 5, 363–366. [Google Scholar]
- Adegoke, A.A.; Okoh, A.I. Species diversity and antibiotic resistance properties of Staphylococcus of farm animal origin in Nkonkobe Municipality. Folia Microbiol. 2014, 59, 133–140. [Google Scholar] [CrossRef] [PubMed]
- Reinert, R.R.; Reinert, S.; van der Linden, M.; Cil, M.; Al-Lahham, A.; Appelbaum, P. Antimicrobial Susceptibility of Streptococcus pneumoniae in Eight European Countries from 2001 to 2003. Antimicrob. Agents Chemother. 2005, 29, 2903–2913. [Google Scholar] [CrossRef] [PubMed]
- Dagan, R.; Klugman, K. Impact of Conjugate Pneumococcal Vaccine on Antibiotic Resistance. In Pneumococcal Vaccines; Siber, G., Klugman, K., Mäkelä, P., Eds.; ASM Press: Washington, DC, USA, 2008; pp. 369–385. [Google Scholar]
- Vila-Corcoles, A.; Bejarano-Romero, F.; Salsench, E. Drug-resistance in Streptococcus pneumoniae isolates among Spanish middle aged and older adults with community-acquired pneumonia. BMC Infect. Dis. 2009, 9, 36. [Google Scholar] [CrossRef] [PubMed]
- Chawla, K.; Gurung, B.; Mukhopadhyay, C.; Bairy, I. Reporting emerging resistance of Streptococcus pneumoniae from India. J. Glob. Infect. Dis. 2010, 2, 10–14. [Google Scholar] [CrossRef] [PubMed]
- Jenkins, T.C.; Sakai, J.; Knepper, B.C.; Swartwood, C.J. Risk Factors for Drug-resistant Streptococcus pneumoniae and Antibiotic Prescribing Practices in Outpatient Community-acquired Pneumonia. Acad. Emerg. Med. 2012, 19, 703–706. [Google Scholar] [CrossRef] [PubMed]
- Raju, A.; Santos, E.R.; Bakota, E.V.; Yang, B.; Arafat, R.R. Susceptibility Profile of Drug-Resistant Streptococcus pneumoniae Based on ELR. ISDS 2015 Conference Abstracts Online. J. Public Health Inform. 2015, 8, e154. [Google Scholar]
- Kim, L.; McGee, L.; Tomczyk, S.; Beall, B. Biological and epidemiological features of antibiotic-resistant Streptococcus pneumoniae in pre- and post-conjugate vaccine eras: a United States perspective. Clin. Microbiol. Rev. 2016, 29, 525–552. [Google Scholar] [CrossRef] [PubMed]
- Zignol, M.; Dean, A.S.; Falzon, D.; van Gemert, W.; Wright, A.; van Deun, A.; Portaels, F.; Laszlo, A.; Espinal, M.A.; Pablos-Méndez, A.; et al. Twenty Years of Global Surveillance of Antituberculosis-Drug Resistance. N. Engl. J. Med. 2016, 375, 1081–1089. [Google Scholar] [CrossRef] [PubMed]
- Udwadia, Z.F.; Amale, R.A.; Ajbani, K.K.; Rodrigues, C. Totally Drug-Resistant Tuberculosis in India. Clin. Infect. Dis. 2012, 54, 579–581. [Google Scholar] [CrossRef] [PubMed]
- Klopper, M.; Warren, R.M.; Hayes, C.; van Pittius, N.C.G.; Streicher, E.M.; Müller, B.; Sirgel, F.A.; Mamisa, C.-N.; Hoosain, E.; Coetzee, G.; et al. Emergence and Spread of Extensively and Totally Drug-Resistant Tuberculosis, South Africa. Emerg. Infect. Dis. 2013, 19, 449–455. [Google Scholar] [CrossRef] [PubMed]
- Velayati, A.A.; Farnia, P.; Masjedi, M.R. The totally drug resistant tuberculosis (TDR-TB). Int. J. Clin. Exp. Med. 2013, 6, 307–309. [Google Scholar] [PubMed]
- Parida, S.K.; Axelsson-Robertson, R.; Rao, M.V.; Singh, N.; Master, I.; Lutckii, A.; Keshavjee, S.; Andersson, J.; Zumla, A.; Maeurer, M. Totally drug-resistant tuberculosis and adjunct therapies (Review). J. Intern. Med. 2015, 277, 388–405. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, W.; Zhu, Y.; Gong, Q.; Yu, W.; Lu, X. Antibiotics at subinhibitory concentrations improve the quorum sensing behavior of Chromobacterium violaceum. FEMS Microbiol. Lett. 2013, 341, 37–44. [Google Scholar] [CrossRef] [PubMed]
- Subhadra, B.; Oh, M.H.; Choi, C.H. Quorum sensing in Acinetobacter: With special emphasis on antibiotic resistance, biofilm formation and quorum quenching. AIMS Microbiol. 2016, 2, 27–41. [Google Scholar]
- Anderson, G.G.; O’Toole, G.A. Innate and Induced Resistance Mechanisms of Bacterial Biofilms. Curr. Top. Microbiol. Immunol. 2008, 322, 85–105. [Google Scholar] [PubMed]
- Andersson, D.I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 2014, 12, 465–478. [Google Scholar] [CrossRef] [PubMed]
- Stewart, P.S.; Costerton, J.W. Antibiotic resistance of bacteria in biofilms. Lancet 2001, 358, 135–138. [Google Scholar] [CrossRef]
- Anderl, J.N.; Franklin, M.J.; Stewart, P.S. Role of antibiotic penetration limitation in Klebsiella pneumoniae biofilm resistance to ampicillin and ciprofloxacin. Antimicrob. Agents Chemother. 2000, 44, 1818–1824. [Google Scholar] [CrossRef] [PubMed]
- Røder, H.L.; Sørensen, S.J.; Burmølle, M. Studying bacterial multispecies biofilms: Where to start? Trends Microbiol. 2016, 24, 503–513. [Google Scholar] [CrossRef] [PubMed]
- Weitao, T. Multicellularity of a unicellular organism in response to DNA replication stress. Res. Microbiol. 2009, 160, 87–88. [Google Scholar] [CrossRef] [PubMed]
- Kostakioti, M.; Hadjifrangiskou, M.; Hultgren, S.J. Bacterial biofilms: development, dispersal, and therapeutic strategies in the dawn of the post antibiotic era. Cold Spring Harbor Perspect. Med. 2013, 3, a010306. [Google Scholar] [CrossRef] [PubMed]
- Mulcahy, L.R.; Isabella, V.M.; Lewis, K. Pseudomonas aeruginosa biofilms in disease. Microb. Ecol. 2014, 68, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Ray, P. Quorum sensing-mediated regulation of staphylococcal virulence and antibiotic resistance. Future Microbiol. 2014, 9, 669–681. [Google Scholar] [CrossRef] [PubMed]
- Canton, R.; Morosini, M. Emergence and spread of antibiotic resistance following exposure to antibiotics. FEMS Microbiol. Rev. 2011, 35, 977–991. [Google Scholar] [CrossRef] [PubMed]
- Davies, J.A.; Harrison, J.J.; Marques, L.L.; Foglia, G.R.; Stremick, C.A.; Storey, D.G.; Turner, R.J.; Olson, M.E.; Ceri, H. The GacS sensor kinase controls phenotypic reversion of small colony variants isolated from biofilms of Pseudomonas aeruginosa PA14. FEMS Microbiol. Ecol. 2007, 59, 32–46. [Google Scholar] [CrossRef] [PubMed]
- Paraje, M.G. Antimicrobial Resistance in Biofilms. In Science Against Microbial Pathogens: Communicating Current Research and Technological Advances; Méndez-Vilas, A., Ed.; Formatex Research Center: Badajoz, Spain, 2011; pp. 736–744. [Google Scholar]
- Anderson, S.W.; Stapp, J.R.; Burns, J.L.; Qin, X. Characterization of small-colony-variant Stenotrophomonas maltophilia isolated from the sputum specimens of five patients with cystic fibrosis. J. Clin. Microbiol. 2007, 45, 529–535. [Google Scholar] [CrossRef] [PubMed]
- Wright, G.D. The antibiotic resistome: the nexus of chemical and genetic diversity. Nat. Rev. Microbiol. 2007, 5, 175–186. [Google Scholar] [CrossRef] [PubMed]
- Stenström, T.A.; Seidu, R.; Ekane, N.; Zurbrügg, C. Microbial Exposure and Health Assessments in Sanitation Technologies and Systems; EcoSanRes Series; Stockholm Environment Institute (SEI): Stockholm, Sweden, 2011. [Google Scholar]
- Scallan, E.; Hoekstra, R.M.; Angulo, F.J.; Tauxe, R.V.; Widdowson, M.A.; Roy, S.L.; Jones, J.L.; Griffin, P.M. Foodborne illness acquired in the United States—Major pathogens. Emerg. Infect. Dis. 2011, 17, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Ashbolt, N.J.; Amezquita, A.; Backhaus, T.; Borrelio, P.; Brant, K.K.; Collignon, P.; Coors, A.; Finley, R.; Gaze, W.H.; Heberer, T.; et al. Human health risk assessment (HHRA) for environmental development and transfer of antibiotic resistance. Environ. Health Perspect. 2013, 121, 993–1001. [Google Scholar] [CrossRef] [PubMed]
- Michael, I.; Rizzo, L.; McArdell, C.S.; Manaia, C.M.; Merlin, C.; Schwartz, T.; Dagot, C.; Fatta-Kassinos, D. Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: A review. Water Res. 2013, 47, 957–995. [Google Scholar] [CrossRef] [PubMed]
- Shakya, R.; Thapa, P.; Saha, R.N. In vitro and in vivo evaluation of gastro retentive floating drug delivery system of ofloxacin. Asian J. Pharm. Sci. 2013, 8, 191–198. [Google Scholar] [CrossRef]
- Ling, L.L.; Schneider, T.; Peoples, A.J.; Spurring, A.L.; Engels, I. A new antibiotic kills pathogens without detectable resistance. Nature 2015, 517, 455–459. [Google Scholar] [CrossRef] [PubMed]
- Palombo, E.A. Traditional Medicinal Plant Extracts and Natural Products with Activity against Oral Bacteria: Potential Application in the Prevention and Treatment of Oral Diseases. Evid. Based Complement. Altern. Med. 2011, 2011, 680354. [Google Scholar] [CrossRef] [PubMed]
- Adak, S.; Upadrasta, L.; Kumar, S.P.J.; Soni, R.; Banerjee, R. Quorum Quenching—An Alternative Antimicrobial Therapeutics. Available online: http://www.formatex.info/microbiology3/book/586–593.pdf (accessed on 11 October 2016).
- Tang, K.; Zhang, X.H. Quorum Quenching Agents: Resources for Antivirulence Therapy. Mar. Drugs 2014, 12, 3245–3282. [Google Scholar] [CrossRef] [PubMed]
- Bordi, C.; de Bentzmann, S. Hacking into bacterial biofilms: a new therapeutic challenge. Ann. Intensive Care 2011, 1, 19. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. Step-by-Step Approach for Development and Implementation of Hospital Antibiotic Policy and Standard Treatment Guidelines 2011; World Health Organization: Geneva, Switzerland, 2011. [Google Scholar]
- Sample Availability: Samples of the materials (journals used) are available from the authors.
| Environment | Source | RAb/ARGs | Reported Concentration | Country | References |
|---|---|---|---|---|---|
| Soil | Soil | CIP | 2.77 μg/kg | Pakistan | [42] |
| OFL | 2.98 μg/kg | ||||
| LEV | 3.35 μg/kg | ||||
| OXT | 4.53 μg/kg | ||||
| DOX | 3.12 μg/kg | ||||
| Grape soil | Sul 1 | (39.19 ± 0.77) × 10−2 | China | [43] | |
| sulII | (0.42 ± 0.08) × 10−3 | ||||
| sulIII | (0.48 ± 0.10) × 10−3 | ||||
| tetA | (0.02 ± 0.00) × 10−3 | ||||
| tetB | (0.44 ± 0.07) × 10−3 | ||||
| tetO | (10.55 ± 1.23) × 10−2 | ||||
| Soil | SMT | 0.01 μg/g | China | [43] | |
| OTC | 0.02 μg/g | ||||
| Vegetable soil | TET | 8400 μg/kg | China | [44] | |
| Animal manure | BAC | 0.01–1.76 mg/kg | Canada | [45] | |
| Aquatic Environment | Wastewater | CIP | 3.0–5.25 mg/L | Pakistan | [42] |
| LEV | 0–6.20 mg/L | ||||
| OFL | 2.45–4.12 mg/L | ||||
| OTC | 0–9.40 mg/L | ||||
| DOX | 1.58–6.75 mg/L | ||||
| AMX | 6.94 μg/L | Australia | [46] | ||
| CIP | 0.72 μg/L | Hong Kong | [47] | ||
| OFL | 0.60 μg/L | Italy | [48] | ||
| ERY | 2.5–6.0 μg/L | Germany | [49] | ||
| Surface water | OFL | 0.31 μg/L | Italy | [48,50] | |
| Hospital effluents | AMX | 35.12 μg/L | Brazil | [51] | |
| AMP | 389.13 μg/L | ||||
| CFX | 300.1 μg/L | ||||
| PEN G | 434.46 μg/L |
| Bacteria Threat Level | Examples of Reported Antibiotics/Antibiotic Groups to Which Resistance Occurred | Countries Where This Has Been Reported | References |
|---|---|---|---|
| Pan drug resistant (PDR)/Extended spectrum drug resistant (XDR) Acinetobacter spp. | Resistant to at least 3 classes + Carbapenems, polymyxins, tigecycline or fluoroquinolones | Greece, US, India, South Africa, Iran, Greece | [77,78,79,80,81,82] |
| Drug resistant Campylobacter spp. | Range of 45% to 94.7% resistant to Erythromycin, azithromycin, clindamycin, telithromycin, ciprofloxacin, | US; Finland; Poland; Philippines; China; Nigeria | [83,84,85,86,87] |
| Fluconazole-resistant Candida spp. | 8.0%–98.8% resistant to Itraconazole, voriconazole, caspofungin, echinocandin, amphotericin B deoxycholate, fluconazole | US, UK, Argentina, Spain, China, South Africa | [88,89,90,91,92] |
| Extended spectrum β-lactamase producing Enterobacteriaceae (ESBLs) | 23% to 85.1% resistant to cephalosporins, gentamicin, kanamycin, streptomycin, nalidixic acid, ciprofloxacin, tetracycline, chloramphen-icol, sulfamethoxazole | US, Switzerland, Netherland, Saudi Arabia, France, Germany, Czech Republic, Sweden | [93,94,95,96,97,98,99,100,101,102] |
| Vancomycin-resistant Enterococcus (VRE) | ≤90% ampicillin, chloramphen-icol, clindamycin, ciproflo-xin, erythromycin, neomycin, penicillin, rifampicin, tetracycline and vancomycin | US, Spain, Portugal Sweden, UK, Australia, Iran, Ethiopia | [103,104,105,106] |
| Multidrug-resistant Pseudomonas aeruginosa | 20% to 85.7% Cefepime, piperacillin-tazobactam, piperacillin, amikacin, levofloxacin, ciprofloxacin, Ofloxacin, meropenem, etc. | US, India, Germany South African, Nigeria, Greece | [107,108,109,110,111,112] |
| Drug-resistant Non-typhoidal Salmonella spp. | ≤100% resistant to nalidixic acid, tetracycline, streptomycin, ciprofloxacin, azithromycin and cefotaxime | US, Iran, Egypt, Ethiopia, UK, China, Congo Republic, Saudi Arabia, Greece | [113,114,115,116,117,118,119,120] |
| Drug-resistant Salmonella | Resistant to ceftriaxone, cefuroxime, amoxicillin, ampicillin, ciprofloxacin and augmentin | US, Nigeria, India, Southern Asia and Kenya | [121,122,123,124,125] |
| Methicillin-resistant Staphylococcus aureus (MRSA) | Usually resistant to wide range of beta lactam antibiotics to ≤100% | US, Nigeria, South Africa, Tanzania, several countries in Europe | [126,127,128,129,130,131] |
| Drug-resistant Streptococcus pneumoniae | e.g., 37% were resistant to erythromycin, 29.6% to cefotaxime, 7.4% to levofloxacin, and 14.8% were identified as multidrug resistant | US, Spain, India, Austria Belgium, France, Germany, Italy, Portugal, Spain and Switzerland | [132,133,134,135,136,137,138] |
| Total Drug-resistant Mycobacterium tuberculosis | >30 cases of TDR-TB reported. 32% of patients with MDR-TB exhibited resistance to a fluoroquinolone | India, Iran, Italy and South Africa | [139,140,141,142,143] |
| Attributes/Mechanism | Application/Example (s) | Reference (s) |
|---|---|---|
| Quorum sensing | Mediated by accessory gene regulator (agr) | [154] |
| Biofilm formation | Increased interaction of high population densities and close distant cells in biofilms for genetic exchange among mixed microbial communities converting biofilms to hotspots for antibiotic resistance GacS-GacA system is associated with the production of small-colony variants that affect motility, biofilm formation, and antibiotic resistance | [150,155,156] |
| Enzyme production | Beta lactamases, extended spectrum beta lactamase, metallo beta lactamase, etc. induced by exposure to imipenem and piperacillin in P. aeruginosa biofilms | [157] |
| Mutation | The evolution of AMR under the sub-MIC arises progressively as low-cost mutations (e.g., duplications and amplifications) in high frequency (Canton and Morosini, 2011) | [155] |
| Small colony variant (SMV) | Down-regulation of the bacterial electron transport and/or dihydrofolate reductase (DHFR) pathway sulfamethoxazole resistance, bringing about small colonial form GacS-GacA system | [156,157,158] |
| Target change | C1 metabolism e.g., Trimethoprim, Sulfamethoxazole, Daptomycin, Colistin, Gentamicin, streptomycin, spectinomycin etc. | [159] |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adegoke, A.A.; Faleye, A.C.; Singh, G.; Stenström, T.A. Antibiotic Resistant Superbugs: Assessment of the Interrelationship of Occurrence in Clinical Settings and Environmental Niches. Molecules 2017, 22, 29. https://doi.org/10.3390/molecules22010029
Adegoke AA, Faleye AC, Singh G, Stenström TA. Antibiotic Resistant Superbugs: Assessment of the Interrelationship of Occurrence in Clinical Settings and Environmental Niches. Molecules. 2017; 22(1):29. https://doi.org/10.3390/molecules22010029
Chicago/Turabian StyleAdegoke, Anthony Ayodeji, Adekunle Christopher Faleye, Gulshan Singh, and Thor Axel Stenström. 2017. "Antibiotic Resistant Superbugs: Assessment of the Interrelationship of Occurrence in Clinical Settings and Environmental Niches" Molecules 22, no. 1: 29. https://doi.org/10.3390/molecules22010029
APA StyleAdegoke, A. A., Faleye, A. C., Singh, G., & Stenström, T. A. (2017). Antibiotic Resistant Superbugs: Assessment of the Interrelationship of Occurrence in Clinical Settings and Environmental Niches. Molecules, 22(1), 29. https://doi.org/10.3390/molecules22010029





