On the Spatial Distribution of Temporal Complexity in Resting State and Task Functional MRI
Abstract
:1. Introduction
2. Materials and Methods
2.1. Data and Preprocessing
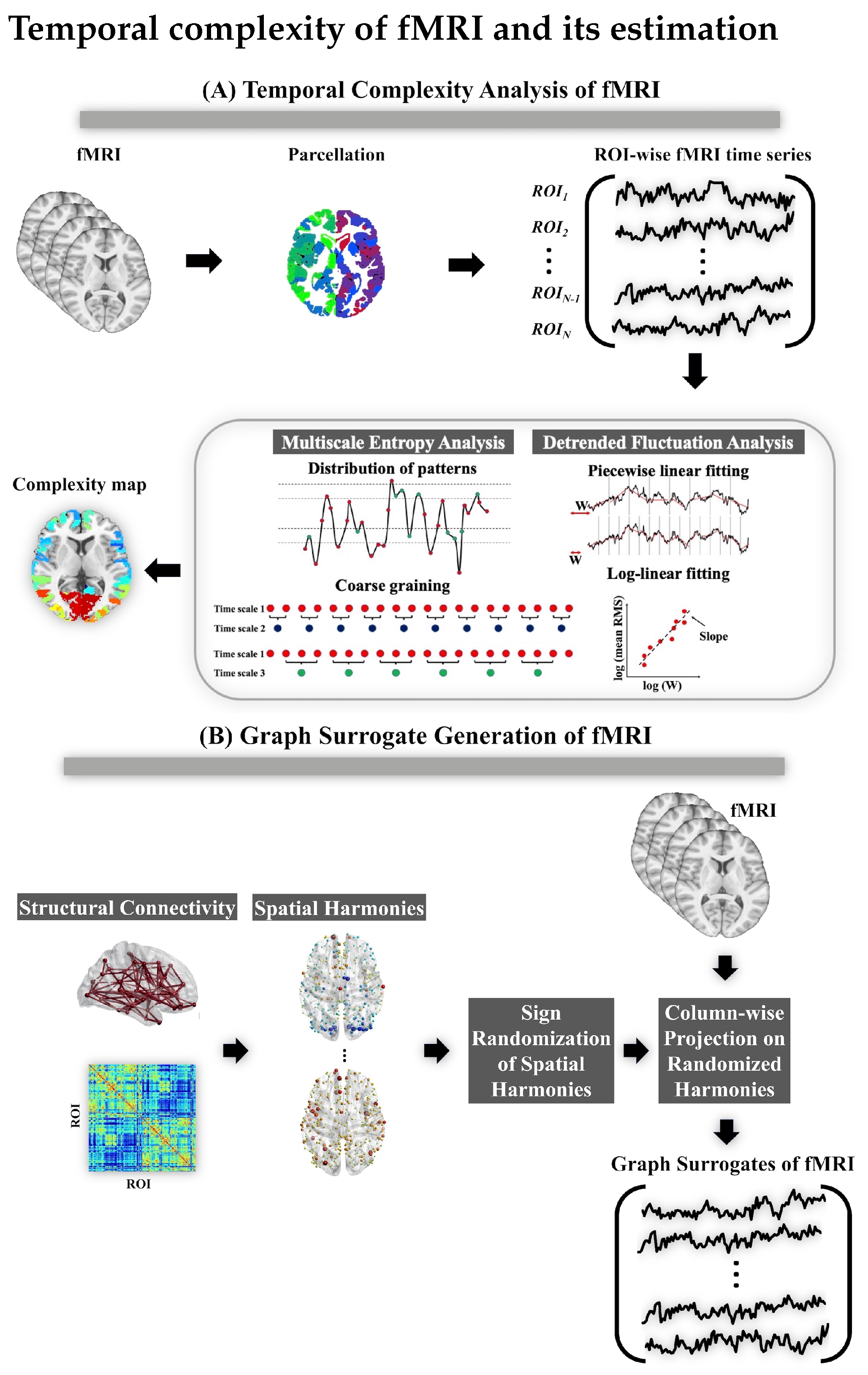
2.2. Temporal Complexity Analysis of fMRI
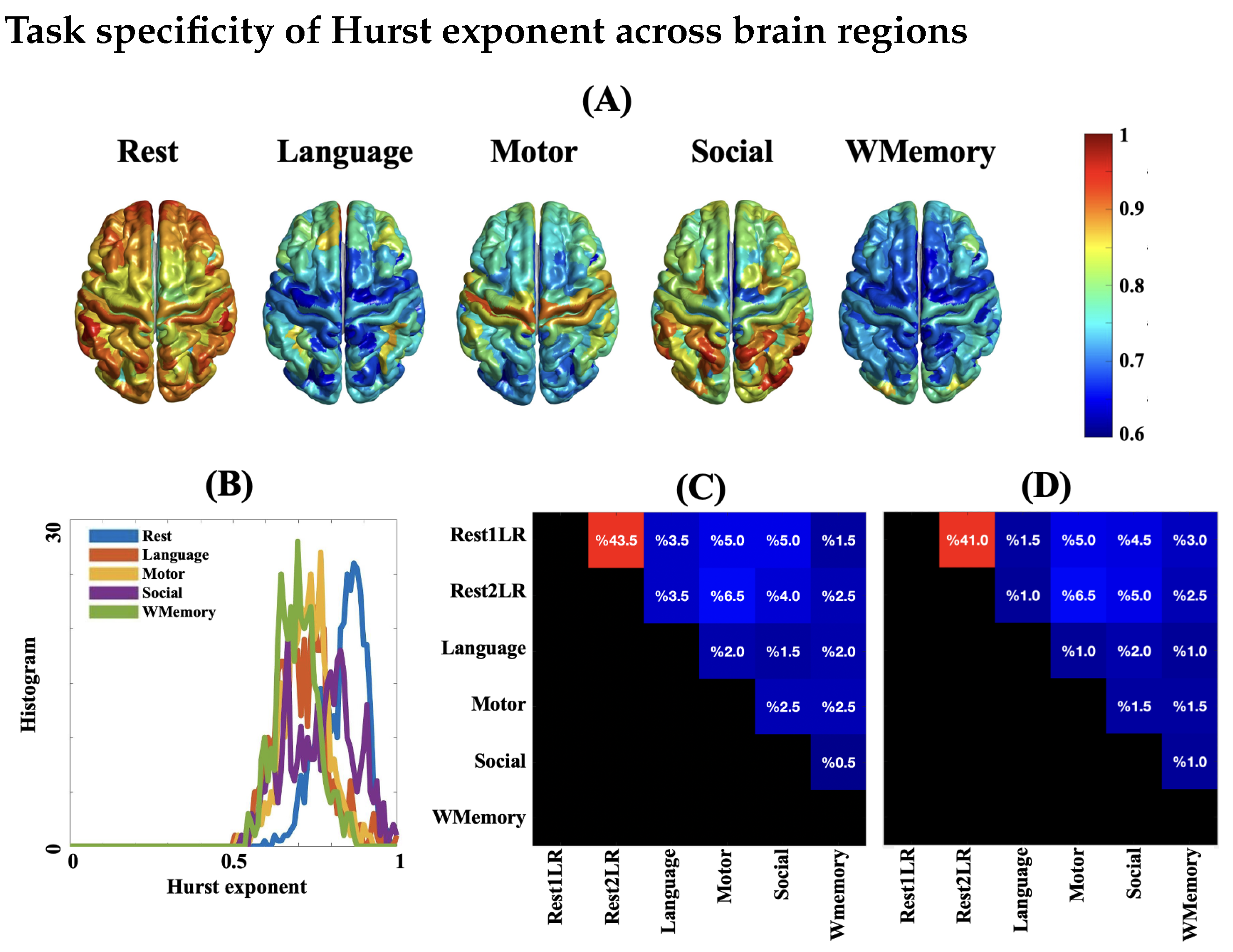
2.3. Task Specificity of fMRI Complexity
2.4. Spatial Distribution of fMRI Complexity across Grey Matter
3. Results
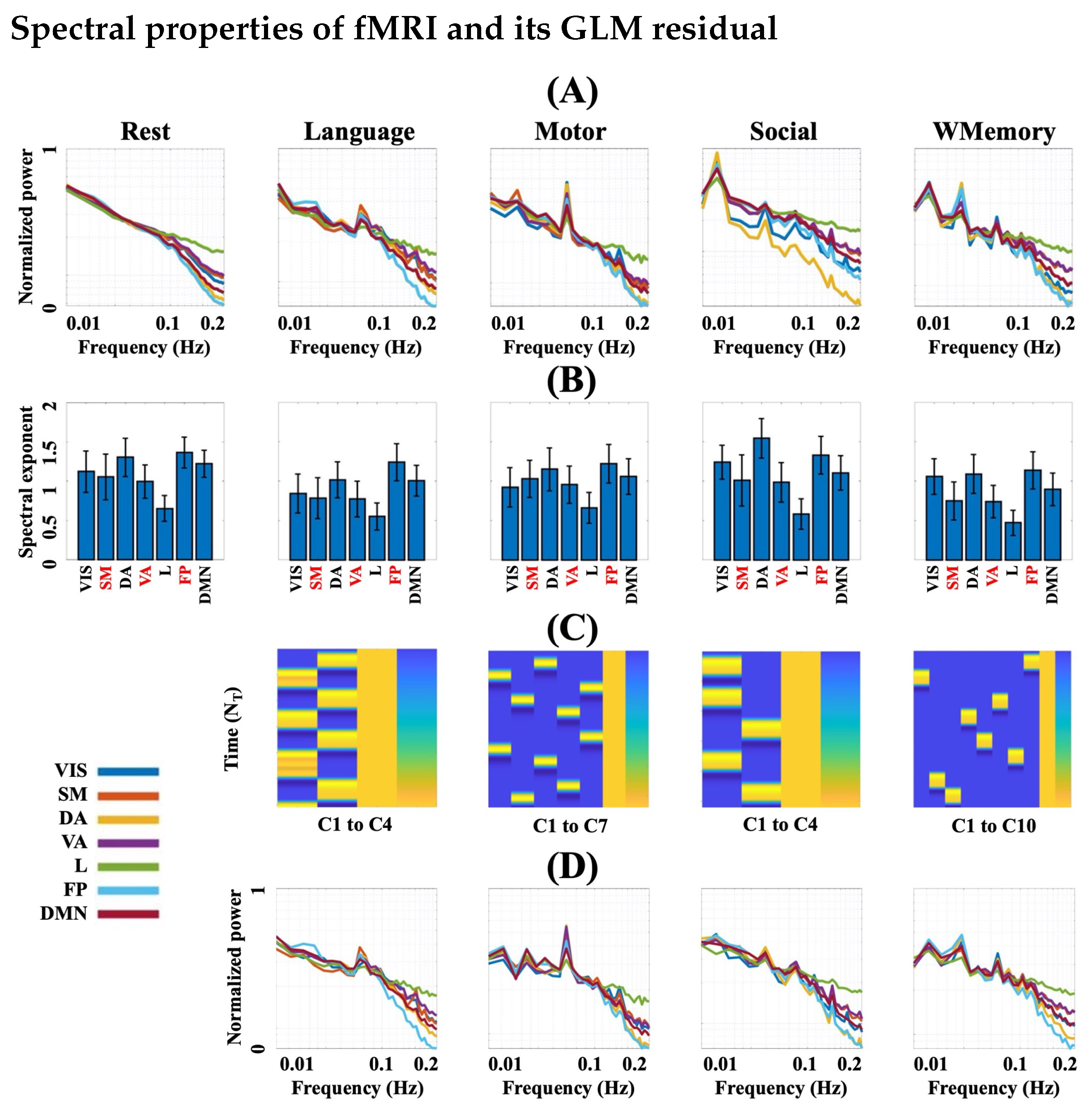
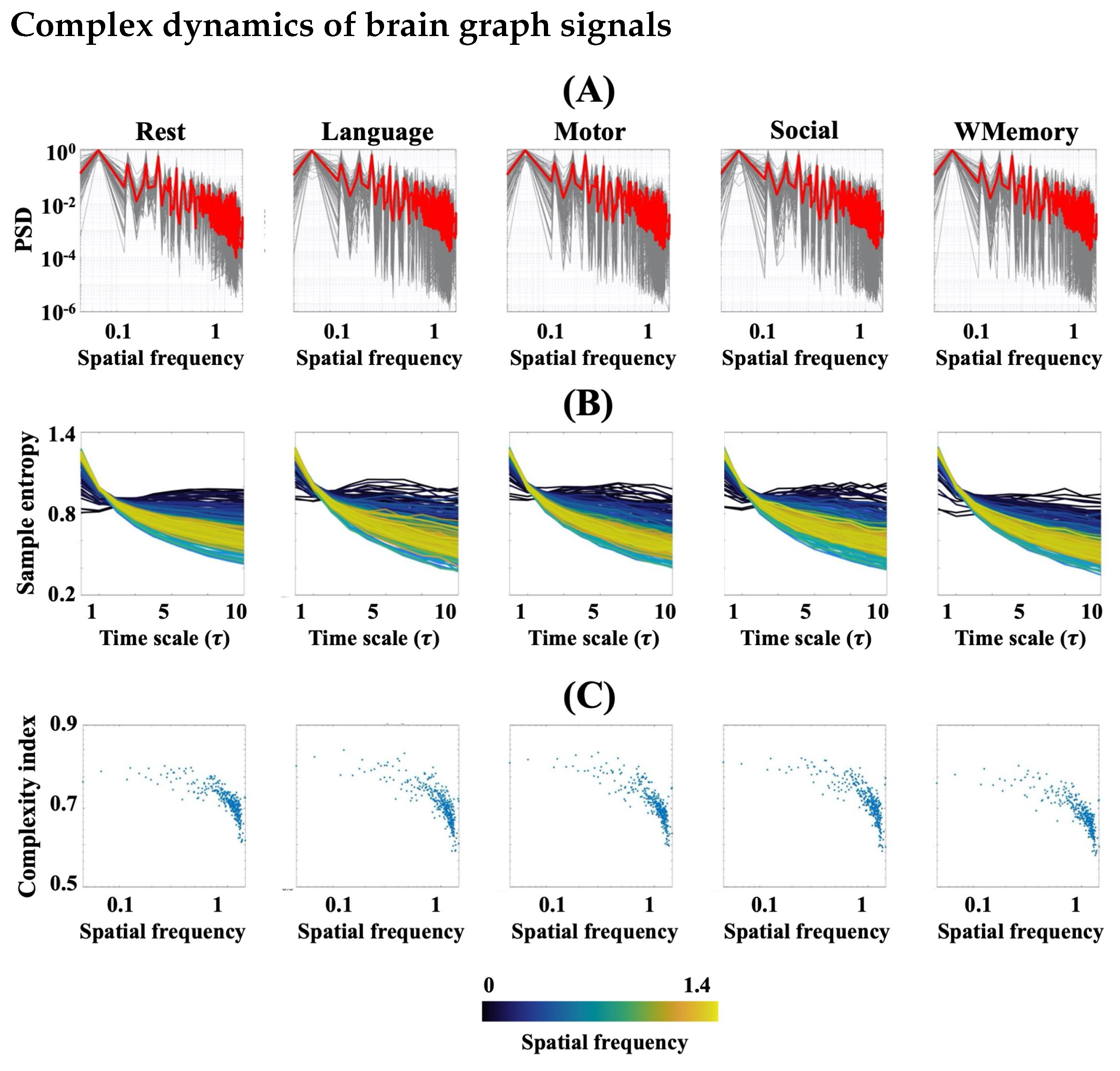
3.1. FMRI Represents Complex Behaviour during Rest and Task
3.2. Task Engagement Lowers Complexity of BOLD Activity
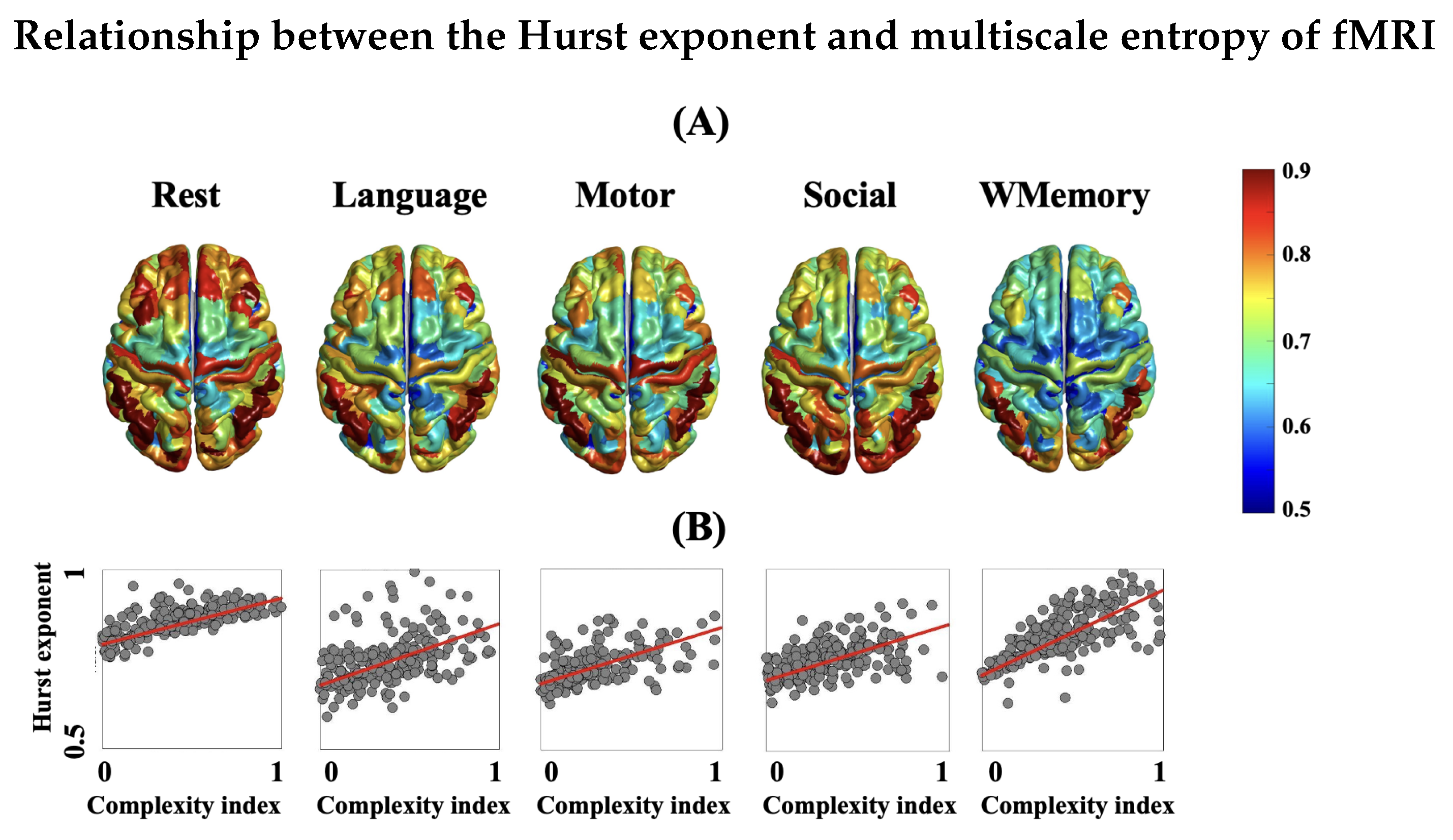
3.3. Complex Dynamics Exist in the Brain Structural-Functional Coupling
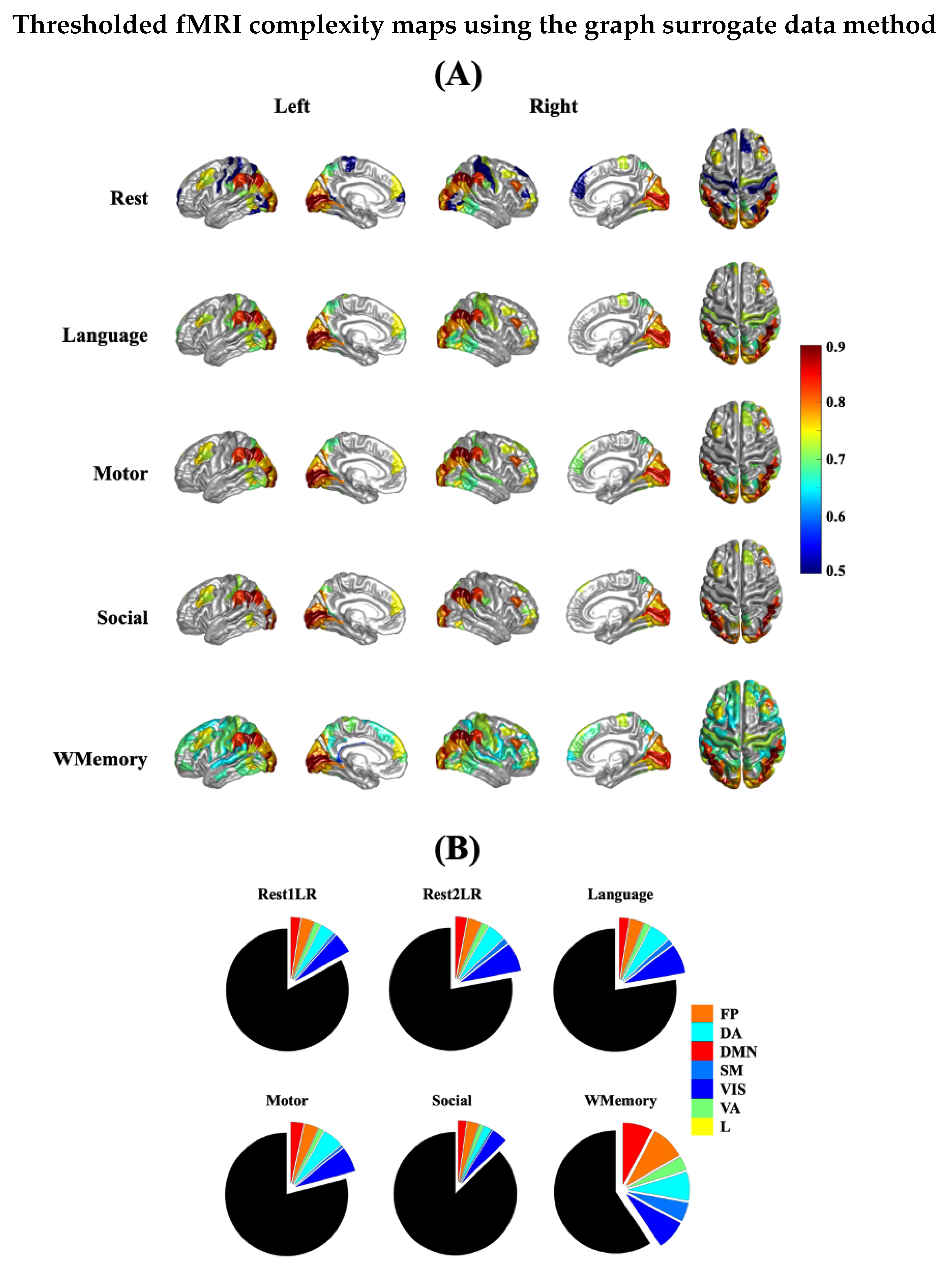
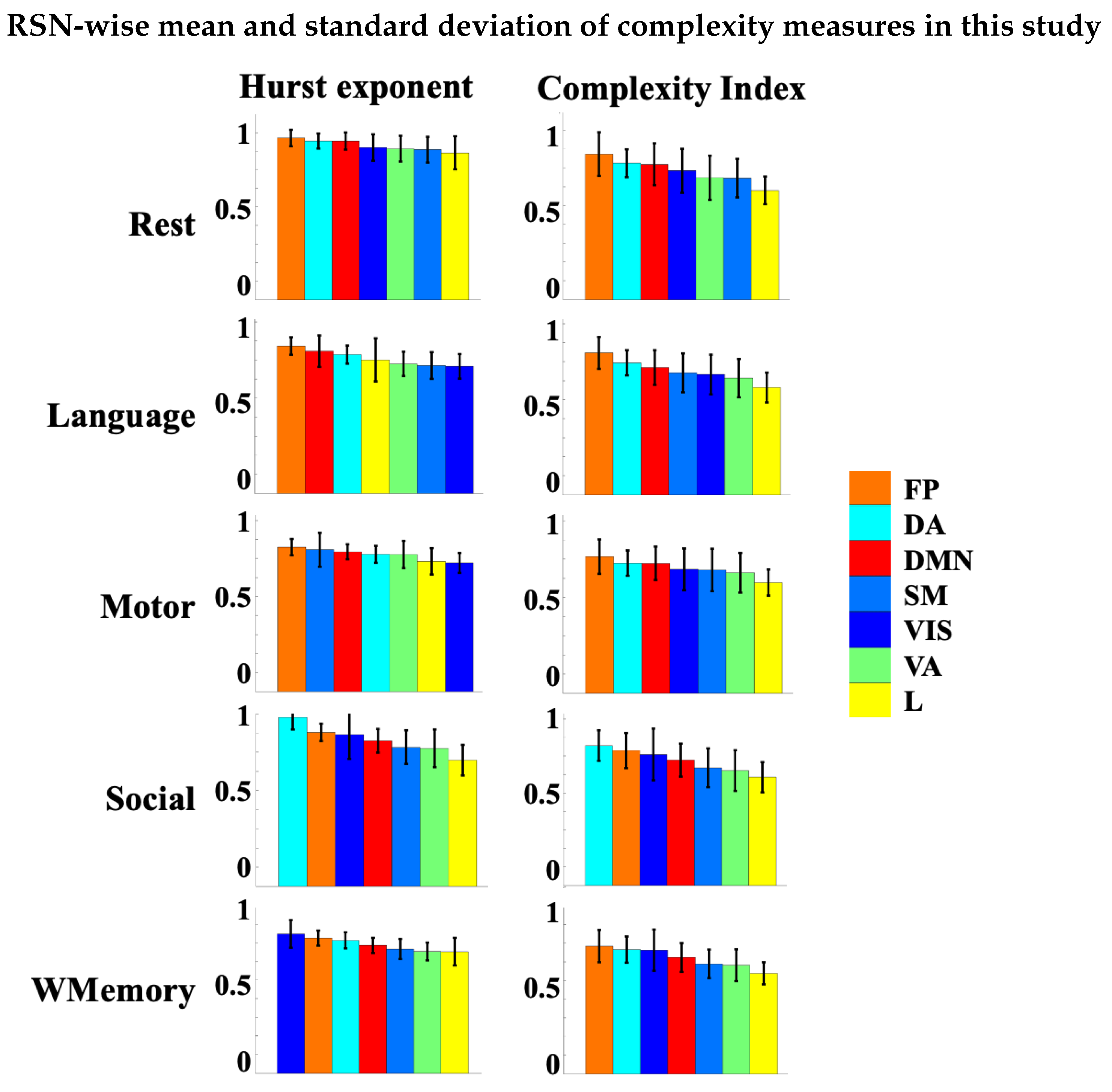
3.4. Spatial Patterns of Complex Dynamics in fMRI
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Appendix A. Multiscale Entropy Analysis
Appendix B. The Graph Surrogate Method for fMRI Complexity Analysis
Appendix B.1. Combining Brain Structure and Function
Appendix B.2. Spatial Harmonics of Brain Structure
Appendix B.3. Graph Surrogate Generation
Appendix B.4. Regarding Linearity
Appendix B.5. Regarding Functional Connectivity
Appendix B.6. Regarding the fMRI Temporal Correlation Matrix
Appendix B.7. Regarding Temporal Complexity
References
- Villecco, F.; Pellegrino, A. Evaluation of Uncertainties in the Design Process of Complex Mechanical Systems. Entropy 2017, 19, 475. [Google Scholar] [CrossRef]
- Villecco, F.; Pellegrino, A. Entropic Measure of Epistemic Uncertainties in Multibody System Models by Axiomatic Design. Entropy 2017, 19, 291. [Google Scholar] [CrossRef]
- Glynn, C.C.; Konstantinou, K.I. Reduction of randomness in seismic noise as a short-term precursor to a volcanic eruption. Sci. Rep. 2016, 6, 37733. [Google Scholar] [CrossRef]
- Shao, Z.G. Contrasting the complexity of the climate of the past 122,000 years and recent 2000 years. Sci. Rep. 2017, 7, 4143. [Google Scholar] [CrossRef]
- Min, L.; Guang, M.; Sarkar, N. Complexity Analysis of 2010 Baja California Earthquake Based on Entropy Measurements. In Proceedings of the Vulnerability, Uncertainty, and Risk, Liverpool, UK, 13–16 July 2014; American Society of Civil Engineers: Liverpool, UK, 2014; pp. 1815–1822. [Google Scholar] [CrossRef]
- Zhao, X.; Shang, P.; Wang, J. Measuring information interactions on the ordinal pattern of stock time series. Phys. Rev. E 2013, 87, 022805. [Google Scholar] [CrossRef] [PubMed]
- Lin, P.F.; Tsao, J.; Lo, M.T.; Lin, C.; Chang, Y.C. Symbolic Entropy of the Amplitude rather than the Instantaneous Frequency of EEG Varies in Dementia. Entropy 2015, 17, 560–579. [Google Scholar] [CrossRef]
- Rodríguez-Sotelo, J.L.; Osorio-Forero, A.; Jiménez-Rodríguez, A.; Cuesta-Frau, D.; Cirugeda-Roldán, E.; Peluffo, D. Automatic Sleep Stages Classification Using EEG Entropy Features and Unsupervised Pattern Analysis Techniques. Entropy 2014, 16, 6573–6589. [Google Scholar] [CrossRef]
- Pan, W.Y.; Su, M.C.; Wu, H.T.; Lin, M.C.; Tsai, I.T.; Sun, C.K. Multiscale Entropy Analysis of Heart Rate Variability for Assessing the Severity of Sleep Disordered Breathing. Entropy 2015, 17, 231–243. [Google Scholar] [CrossRef]
- Liu, Q.; Ma, L.; Fan, S.; Abbod, M.; Shieh, J. Sample entropy analysis for the estimating depth of anaesthesia through human EEG signal at different levels of unconsciousness during surgeries. PeerJ 2018, 6, e4817. [Google Scholar] [CrossRef]
- Chen, C.; Jin, Y.; Lo, I.; Zhao, H.; Sun, B.; Zhao, Q.; Zheng, J.; Zhang, X. Complexity Change in Cardiovascular Disease. Int. J. Biol. Sci. 2017, 13, 1320–1328. [Google Scholar] [CrossRef]
- Lake, D.; Richman, J.; Griffin, M.; Moorman, J. Sample entropy analysis of neonatal heart rate variability. Am. J. Physiol. Regul. Integr. Comp. Physiol. 2002, 283, R789–R797. [Google Scholar] [CrossRef] [PubMed]
- McDonough, I.M.; Nashiro, K. Network complexity as a measure of information processing across resting-state networks: Evidence from the Human Connectome Project. Front. Hum. Neurosci. 2014, 8, 409. [Google Scholar] [CrossRef] [PubMed]
- Omidvarnia, A.; Zalesky, A.; Mansour L, S.; Van De Ville, D.; Jackson, G.D.; Pedersen, M. Temporal complexity of fMRI is reproducible and correlates with higher order cognition. NeuroImage 2021, 230, 117760. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, M.; Omidvarnia, A.; Walz, J.; Zalesky, A.; Jackson, G. Spontaneous brain network activity: Analysis of its temporal complexity. Netw. Neurosci. 2017, 1, 100–115. [Google Scholar] [CrossRef]
- McIntosh, A.; Vakorin, V.; Kovacevic, N.; Wang, H.; Diaconescu, A.; Protzner, A. Spatiotemporal dependency of age-related changes in brain signal variability. Cereb. Cortex 2014, 24, 1806–1817. [Google Scholar] [CrossRef]
- Saxe, G.N.; Calderone, D.; Morales, L.J. Brain entropy and human intelligence: A resting-state fMRI study. PLoS ONE 2018, 13, e0191582. [Google Scholar] [CrossRef]
- Biswal, B.; Zerrin Yetkin, F.; Haughton, V.M.; Hyde, J.S. Functional connectivity in the motor cortex of resting human brain using echo-planar mri. Magn. Reson. Med. 1995, 34, 537–541. [Google Scholar] [CrossRef]
- van den Heuvel, M.P.; Sporns, O. An Anatomical Substrate for Integration among Functional Networks in Human Cortex. J. Neurosci. 2013, 33, 14489–14500. [Google Scholar] [CrossRef]
- Damoiseaux, J.S.; Rombouts, S.A.R.B.; Barkhof, F.; Scheltens, P.; Stam, C.J.; Smith, S.M.; Beckmann, C.F. Consistent resting-state networks across healthy subjects. Proc. Natl. Acad. Sci. USA 2006, 103, 13848–13853. [Google Scholar] [CrossRef]
- Ciuciu, P.; Varoquaux, G.; Abry, P.; Sadaghiani, S.; Kleinschmidt, A. Scale-free and multifractal properties of fMRI signals during rest and task. Front. Physiol. 2012, 3, 186. [Google Scholar] [CrossRef]
- Nezafati, M.; Temmar, H.; Keilholz, S.D. Functional MRI Signal Complexity Analysis Using Sample Entropy. Front. Neurosci. 2020, 14, 700. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.; Glover, G.H. Time–frequency dynamics of resting-state brain connectivity measured with fMRI. NeuroImage 2010, 50, 81–98. [Google Scholar] [CrossRef] [PubMed]
- Hutchison, R.M.; Womelsdorf, T.; Allen, E.A.; Bandettini, P.A.; Calhoun, V.D.; Corbetta, M.; Della Penna, S.; Duyn, J.H.; Glover, G.H.; Gonzalez-Castillo, J.; et al. Dynamic functional connectivity: Promise, issues, and interpretations. NeuroImage 2013, 80, 360–378. [Google Scholar] [CrossRef] [PubMed]
- Deco, G.; Kringelbach, M.L. Turbulent-like Dynamics in the Human Brain. Cell Rep. 2020, 33, 108471. [Google Scholar] [CrossRef] [PubMed]
- Vohryzek, J.; Cabral, J.; Vuust, P.; Deco, G.; Kringelbach, M.L. Understanding brain states across spacetime informed by whole-brain modelling. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2022, 380, 20210247. [Google Scholar] [CrossRef]
- Luppi, A.; Vohryzek, J.; Kringelbach, M.; Mediano, P.; Craig, M.; Adapa, R.; Carhart-Harris, R.; Roseman, L.; Pappas, I.; Finoia, P.; et al. Connectome Harmonic Decomposition of Human Brain Dynamics Reveals a Landscape of Consciousness. bioRxiv 2020. [Google Scholar] [CrossRef]
- Zamora-López, G.; Chen, Y.; Deco, G.; Kringelbach, M.L.; Zhou, C. Functional complexity emerging from anatomical constraints in the brain: The significance of network modularity and rich-clubs. Sci. Rep. 2016, 6, 38424. [Google Scholar] [CrossRef]
- Lynn, C.W.; Bassett, D. The physics of brain network structure, function and control. Nat. Rev. Phys. 2019, 1, 318–332. [Google Scholar] [CrossRef]
- Churchill, N.W.; Spring, R.; Grady, C.; Cimprich, B.; Askren, M.K.; Reuter-Lorenz, P.A.; Jung, M.S.; Peltier, S.; Strother, S.C.; Berman, M.G. The suppression of scale-free fMRI brain dynamics across three different sources of effort: Aging, task novelty and task difficulty. Sci. Rep. 2016, 6, 30895. [Google Scholar] [CrossRef]
- Jia, Y.; Gu, H.; Luo, Q. Sample entropy reveals an age-related reduction in the complexity of dynamic brain. Sci. Rep. 2017, 7, 7990. [Google Scholar] [CrossRef]
- Bassett, D.; Gazzaniga, M. Understanding complexity in the human brain. Trends Cogn. Sci. 2011, 15, 200–209. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, M.; Omidvarnia, A.; Curwood, E.K.; Walz, J.M.; Rayner, G.; Jackson, G.D. The dynamics of functional connectivity in neocortical focal epilepsy. NeuroImage Clin. 2017, 15, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Meskaldji, D.E.; Preti, M.G.; Bolton, T.A.; Montandon, M.L.; Rodriguez, C.; Morgenthaler, S.; Giannakopoulos, P.; Haller, S.; Van De Ville, D. Prediction of long-term memory scores in MCI based on resting-state fMRI. NeuroImage Clin. 2016, 12, 785–795. [Google Scholar] [CrossRef] [PubMed]
- Damaraju, E.; Allen, E.; Belger, A.; Ford, J.; McEwen, S.; Mathalon, D.; Mueller, B.; Pearlson, G.; Potkin, S.; Preda, A.; et al. Dynamic functional connectivity analysis reveals transient states of dysconnectivity in schizophrenia. NeuroImage Clin. 2014, 5, 298–308. [Google Scholar] [CrossRef]
- Zalesky, A.; Fornito, A.; Cocchi, L.; Gollo, L.L.; Breakspear, M. Time-resolved resting-state brain networks. Proc. Natl. Acad. Sci. USA 2014, 111, 10341–10346. [Google Scholar] [CrossRef]
- Dong, J.; Jing, B.; Ma, X.; Liu, H.; Mo, X.; Li, H. Hurst Exponent Analysis of Resting-State fMRI Signal Complexity across the Adult Lifespan. Front. Neurosci. 2018, 12, 34. [Google Scholar] [CrossRef]
- Dhamala, E.; Jamison, K.W.; Sabuncu, M.R.; Kuceyeski, A. Sex classification using long-range temporal dependence of resting-state functional MRI time series. Hum. Brain Mapp. 2020, 41, 3567–3579. [Google Scholar] [CrossRef]
- Liégeois, R.; Laumann, T.O.; Snyder, A.Z.; Zhou, J.; Yeo, B.T. Interpreting temporal fluctuations in resting-state functional connectivity MRI. NeuroImage 2017, 163, 437–455. [Google Scholar] [CrossRef]
- Liégeois, R.; Yeo, B.T.; Van De Ville, D. Interpreting null models of resting-state functional MRI dynamics: Not throwing the model out with the hypothesis. NeuroImage 2021, 243, 118518. [Google Scholar] [CrossRef]
- Prichard, D.; Theiler, J. Generating surrogate data for time series with several simultaneously measured variables. Phys. Rev. Lett. 1994, 73, 951–954. [Google Scholar] [CrossRef]
- Huang, W.; Bolton, T.A.W.; Medaglia, J.D.; Bassett, D.S.; Ribeiro, A.; de Ville, D.V. A Graph Signal Processing Perspective on Functional Brain Imaging. Proc. IEEE 2018, 106, 868–885. [Google Scholar] [CrossRef]
- Pirondini, E.; Vybornova, A.; Coscia, M.; Van De Ville, D. A Spectral Method for Generating Surrogate Graph Signals. IEEE Signal Process. Lett. 2016, 23, 1275–1278. [Google Scholar] [CrossRef]
- Van Essen, D.; Ugurbil, K.; Auerbach, E.; Barch, D.; Behrens, T.; Bucholz, R.; Chang, A.; Chen, L.; Corbetta, M.; Curtiss, S.; et al. The Human Connectome Project: A data acquisition perspective. NeuroImage 2012, 62, 2222–2231. [Google Scholar] [CrossRef] [PubMed]
- Barch, D.M.; Burgess, G.C.; Harms, M.P.; Petersen, S.E.; Schlaggar, B.L.; Corbetta, M.; Glasser, M.F.; Curtiss, S.; Dixit, S.; Feldt, C.; et al. Function in the human connectome: Task-fMRI and individual differences in behavior. NeuroImage 2013, 80, 169–189. [Google Scholar] [CrossRef]
- Smith, S.M.; Beckmann, C.F.; Andersson, J.; Auerbach, E.J.; Bijsterbosch, J.; Douaud, G.; Duff, E.; Feinberg, D.A.; Griffanti, L.; Harms, M.P.; et al. Resting-state fMRI in the Human Connectome Project. NeuroImage 2013, 80, 144–168. [Google Scholar] [CrossRef]
- Liégeois, R.; Santos, A.; Matta, V.; Van De Ville, D.; Sayed, A.H. Revisiting correlation-based functional connectivity and its relationship with structural connectivity. Netw. Neurosci. 2020, 4, 1235–1251. [Google Scholar] [CrossRef]
- Glasser, M.F.; Coalson, T.S.; Robinson, E.C.; Hacker, C.D.; Harwell, J.; Yacoub, E.; Ugurbil, K.; Andersson, J.; Beckmann, C.F.; Jenkinson, M.; et al. A multi-modal parcellation of human cerebral cortex. Nature 2016, 536, 171–178. [Google Scholar] [CrossRef]
- Hurst, H. Long-Term Storage Capacity of Reservoirs. Trans. Am. Soc. Civ. Eng. 1951, 116, 770–799. [Google Scholar] [CrossRef]
- Schaefer, A.; Brach, J.S.; Perera, S.; Sejdić, E. A comparative analysis of spectral exponent estimation techniques for 1/fβ processes with applications to the analysis of stride interval time series. J. Neurosci. Methods 2014, 222, 118–130. [Google Scholar] [CrossRef]
- Peng, C.; Havlin, S.; Stanley, H.E.; Goldberger, A.L. Quantification of scaling exponents and crossover phenomena in nonstationary heartbeat time series. Chaos Interdiscip. J. Nonlinear Sci. 1995, 5, 82–87. [Google Scholar] [CrossRef]
- He, B.J. Scale-Free Properties of the Functional Magnetic Resonance Imaging Signal during Rest and Task. J. Neurosci. 2011, 31, 13786–13795. [Google Scholar] [CrossRef] [PubMed]
- Omidvarnia, A.; Mesbah, M.; Pedersen, M.; Jackson, G. Range Entropy: A Bridge between Signal Complexity and Self-Similarity. Entropy 2018, 20, 962. [Google Scholar] [CrossRef] [PubMed]
- Costa, M.; Goldberger, A.L.; Peng, C.K. Multiscale Entropy Analysis of Complex Physiologic Time Series. Phys. Rev. Lett. 2002, 89, 068102. [Google Scholar] [CrossRef]
- Richman, J.; Moorman, J. Physiological time-series analysis using approximate entropy and sample entropy. Am. J. Physiol. Heart Circ. Physiol. 2000, 278, H2039–H2049. [Google Scholar] [CrossRef] [PubMed]
- Weng, W.C.; Jiang, G.J.A.; Chang, C.F.; Lu, W.Y.; Lin, C.Y.; Lee, W.T.; Shieh, J.S. Complexity of Multi-Channel Electroencephalogram Signal Analysis in Childhood Absence Epilepsy. PLoS ONE 2015, 10, e0134083. [Google Scholar] [CrossRef]
- Waschke, L.; Kloosterman, N.A.; Obleser, J.; Garrett, D.D. Behavior needs neural variability. Neuron 2021, 109, 751–766. [Google Scholar] [CrossRef]
- Faisal, A.A.; Selen, L.P.J.; Wolpert, D.M. Noise in the nervous system. Nat. Rev. Neurosci. 2008, 9, 292–303. [Google Scholar] [CrossRef]
- Garrett, D.D.; Samanez-Larkin, G.R.; MacDonald, S.W.; Lindenberger, U.; McIntosh, A.R.; Grady, C.L. Moment-to-moment brain signal variability: A next frontier in human brain mapping? Neurosci. Biobehav. Rev. 2013, 37, 610–624. [Google Scholar] [CrossRef]
- Friston, K.; Breakspear, M.; Deco, G. Perception and self-organized instability. Front. Comput. Neurosci. 2012, 6, 44. [Google Scholar] [CrossRef]
- McDonnell, M.D.; Ward, L.M. The benefits of noise in neural systems: Bridging theory and experiment. Nat. Rev. Neurosci. 2011, 12, 415–425. [Google Scholar] [CrossRef]
- Ghanbari, Y.; Bloy, L.; Edgar, J.C.; Blaskey, L.; Verma, R.; Roberts, T.P.L. Joint Analysis of Band-Specific Functional Connectivity and Signal Complexity in Autism. J. Autism Dev. Disord. 2015, 45, 444–460. [Google Scholar] [CrossRef] [PubMed]
- Delignieres, D.; Ramdani, S.; Lemoine, L.; Torre, K.; Fortes, M.; Ninot, G. Fractal analyses for ‘short’ time series: A re-assessment of classical methods. J. Math. Psychol. 2006, 50, 525–544. [Google Scholar] [CrossRef]
- Eke, A.; Herman, P.; Sanganahalli, B.G.; Hyder, F.; Mukli, P.; Nagy, Z. Pitfalls in Fractal Time Series Analysis: fMRI BOLD as an Exemplary Case. Front. Physiol. 2012, 3, 417. [Google Scholar] [CrossRef] [PubMed]
- He, B.J. Scale-free brain activity: Past, present, and future. Trends Cogn. Sci. 2014, 18, 480–487. [Google Scholar] [CrossRef]
- Ciuciu, P.; Abry, P.; He, B.J. Interplay between functional connectivity and scale-free dynamics in intrinsic fMRI networks. NeuroImage 2014, 95, 248–263. [Google Scholar] [CrossRef] [PubMed]
- Campbell, O.L.; Weber, A.M. Monofractal analysis of functional magnetic resonance imaging: An introductory review. Hum. Brain Mapp. 2022, 43, 2693–2706. [Google Scholar] [CrossRef]
- Fadili, M.; Bullmore, E. Wavelet-Generalized Least Squares: A New BLU Estimator of Linear Regression Models with 1/f Errors. NeuroImage 2002, 15, 217–232. [Google Scholar] [CrossRef]
- von Wegner, F.; Laufs, H.; Tagliazucchi, E. Mutual information identifies spurious Hurst phenomena in resting state EEG and fMRI data. Phys. Rev. E 2018, 97, 022415. [Google Scholar] [CrossRef]
- Buckner, R.L.; Sepulcre, J.; Talukdar, T.; Krienen, F.M.; Liu, H.; Hedden, T.; Andrews-Hanna, J.R.; Sperling, R.A.; Johnson, K.A. Cortical Hubs Revealed by Intrinsic Functional Connectivity: Mapping, Assessment of Stability, and Relation to Alzheimer’s Disease. J. Neurosci. 2009, 29, 1860–1873. [Google Scholar] [CrossRef]
- Lancaster, G.; Iatsenko, D.; Pidde, A.; Ticcinelli, V.; Stefanovska, A. Surrogate data for hypothesis testing of physical systems. Phys. Rep. 2018, 748, 1–60. [Google Scholar] [CrossRef]
- Weron, A.; Burnecki, K.; Mercik, S.; Weron, K. Complete description of all self-similar models driven by Lévy stable noise. Phys. Rev. E 2005, 71, 016113. [Google Scholar] [CrossRef] [PubMed]
- Hertrich, I.; Dietrich, S.; Blum, C.; Ackermann, H. The Role of the Dorsolateral Prefrontal Cortex for Speech and Language Processing. Front. Hum. Neurosci. 2021, 15, 645209. [Google Scholar] [CrossRef] [PubMed]
- Bockstaele, E.J.V.; Aston-Jones, G. Integration in the Ventral Medulla and Coordination of Sympathetic, Pain and Arousal Functions. Clin. Exp. Hypertens. 1995, 17, 153–165. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.U.; Mandic, D.P. Multivariate multiscale entropy: A tool for complexity analysis of multichannel data. Phys. Rev. E 2011, 84, 061918. [Google Scholar] [CrossRef]
- Pellé, H.; Ciuciu, P.; Rahim, M.; Dohmatob, E.; Abry, P.; van Wassenhove, V. Multivariate hurst exponent estimation in FMRI. Application to brain decoding of perceptual learning. In Proceedings of the 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI), Prague, Czech Republic, 13–16 April 2016; pp. 996–1000. [Google Scholar] [CrossRef]
- Schütze, H.; Martinetz, T.; Anders, S.; Madany Mamlouk, A. A Multivariate Approach to Estimate Complexity of FMRI Time Series. In Proceedings of the Artificial Neural Networks and Machine Learning—ICANN 2012, Lausanne, Switzerland, 11–14 September 2012; Villa, A.E.P., Duch, W., Érdi, P., Masulli, F., Palm, G., Eds.; Springer: Berlin/Heidelberg, Germany, 2012; pp. 540–547. [Google Scholar]
- Preti, M.G.; Van De Ville, D. Decoupling of brain function from structure reveals regional behavioral specialization in humans. Nat. Commun. 2019, 10, 4747. [Google Scholar] [CrossRef]
| Run | Session | Length in Minutes | No. of Conditions | No. of Trials | |
|---|---|---|---|---|---|
| 1 | Rest1LR | 399 | 4.8 | - | - |
| 2 | Rest2LR | 399 | 4.8 | - | - |
| 5 | Language | 305 | 3.67 | 2 | 11 |
| 6 | Motor | 273 | 3.29 | 5 | 10 |
| 7 | Social | 263 | 3.17 | 2 | 5 |
| 8 | Working Memory | 395 | 4.74 | 8 | 8 |
| Task Name | VIS | SM | DA | VA | L | FP | DMN |
|---|---|---|---|---|---|---|---|
| Rest1LR | 5.3% | 0.6% | 3.6% | 1.7% | 0% | 3.3% | 2.5% |
| Rest2LR | 7.5% | 1.4% | 4.7% | 1.7% | 0% | 3.6% | 3.1% |
| Language | 7.8% | 1.4% | 5.3% | 1.7% | 0% | 3.6% | 2.5% |
| Motor | 6.7% | 0.6% | 5.3% | 1.4% | 0% | 3.6% | 3.3% |
| Social | 4.2% | 0.3% | 1.9% | 1.1% | 0% | 3.1% | 2.2% |
| Working Memory | 7.8% | 5% | 7.5% | 3.6% | 0% | 8.9% | 7.8% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Omidvarnia, A.; Liégeois, R.; Amico, E.; Preti, M.G.; Zalesky, A.; Van De Ville, D. On the Spatial Distribution of Temporal Complexity in Resting State and Task Functional MRI. Entropy 2022, 24, 1148. https://doi.org/10.3390/e24081148
Omidvarnia A, Liégeois R, Amico E, Preti MG, Zalesky A, Van De Ville D. On the Spatial Distribution of Temporal Complexity in Resting State and Task Functional MRI. Entropy. 2022; 24(8):1148. https://doi.org/10.3390/e24081148
Chicago/Turabian StyleOmidvarnia, Amir, Raphaël Liégeois, Enrico Amico, Maria Giulia Preti, Andrew Zalesky, and Dimitri Van De Ville. 2022. "On the Spatial Distribution of Temporal Complexity in Resting State and Task Functional MRI" Entropy 24, no. 8: 1148. https://doi.org/10.3390/e24081148
APA StyleOmidvarnia, A., Liégeois, R., Amico, E., Preti, M. G., Zalesky, A., & Van De Ville, D. (2022). On the Spatial Distribution of Temporal Complexity in Resting State and Task Functional MRI. Entropy, 24(8), 1148. https://doi.org/10.3390/e24081148







