Information Processing Using Networks of Chemical Oscillators
Abstract
1. Introduction
2. Information Processing with Oscillator Networks
2.1. Classification Type Problems
2.2. The Node Model
2.3. The Model of a Network
2.4. Top-Down Design of Computing Networks
- -
- The observation time ;
- -
- All parameters for a model of chemical oscillations inside a node; for the Oregonator model, they are , q and f;
- -
- Parameters and that translate an input value into the illumination of an input oscillator (cf. Equation (5));
- -
- The rates for reactions responsible for interactions between oscillators (, );
- -
- Location of input and normal oscillators;
- -
- Finally, the illumination times for all normal oscillators .
- (1)
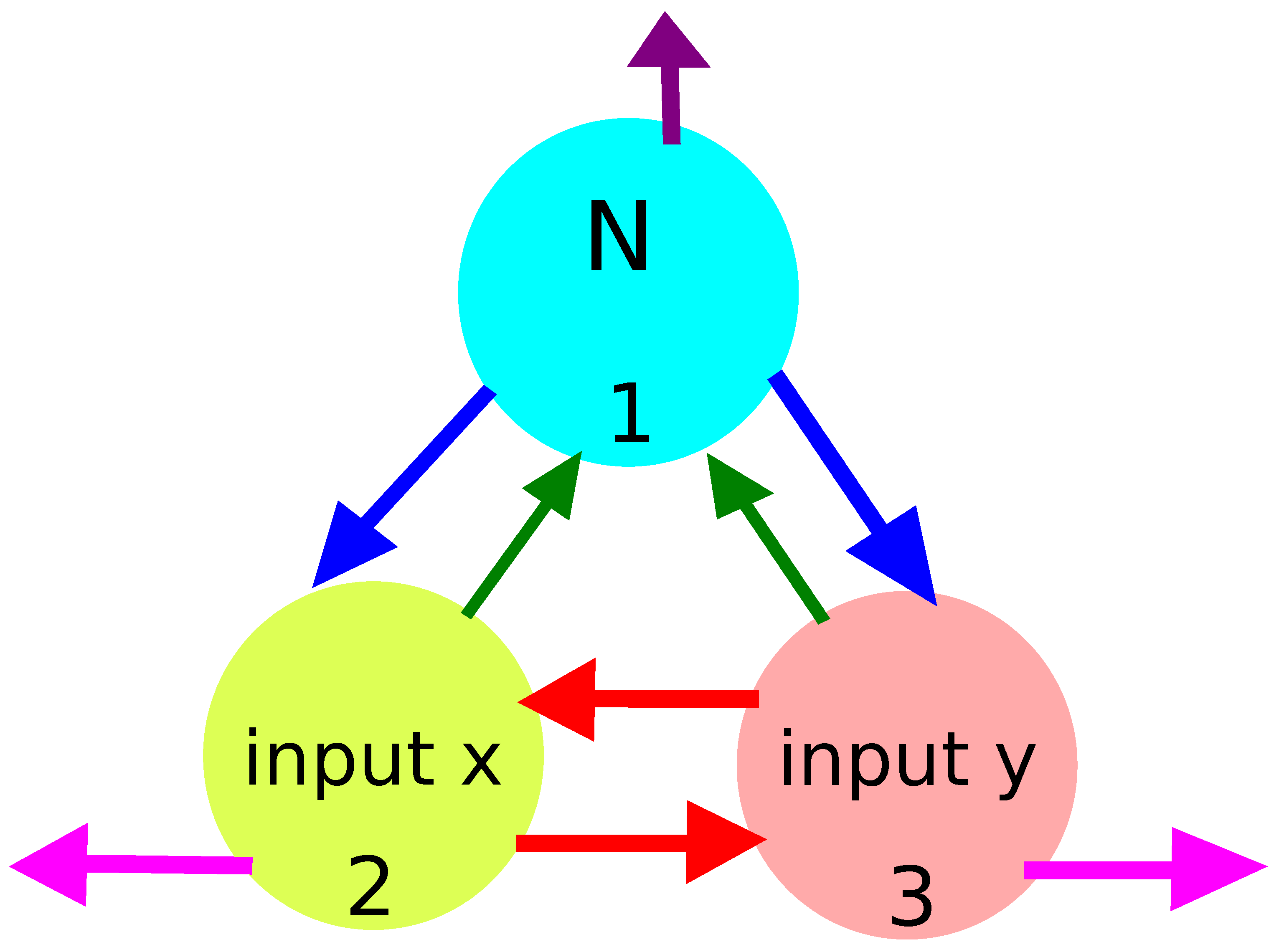
- My attention is restricted to classifiers formed by oscillators;
- (2)
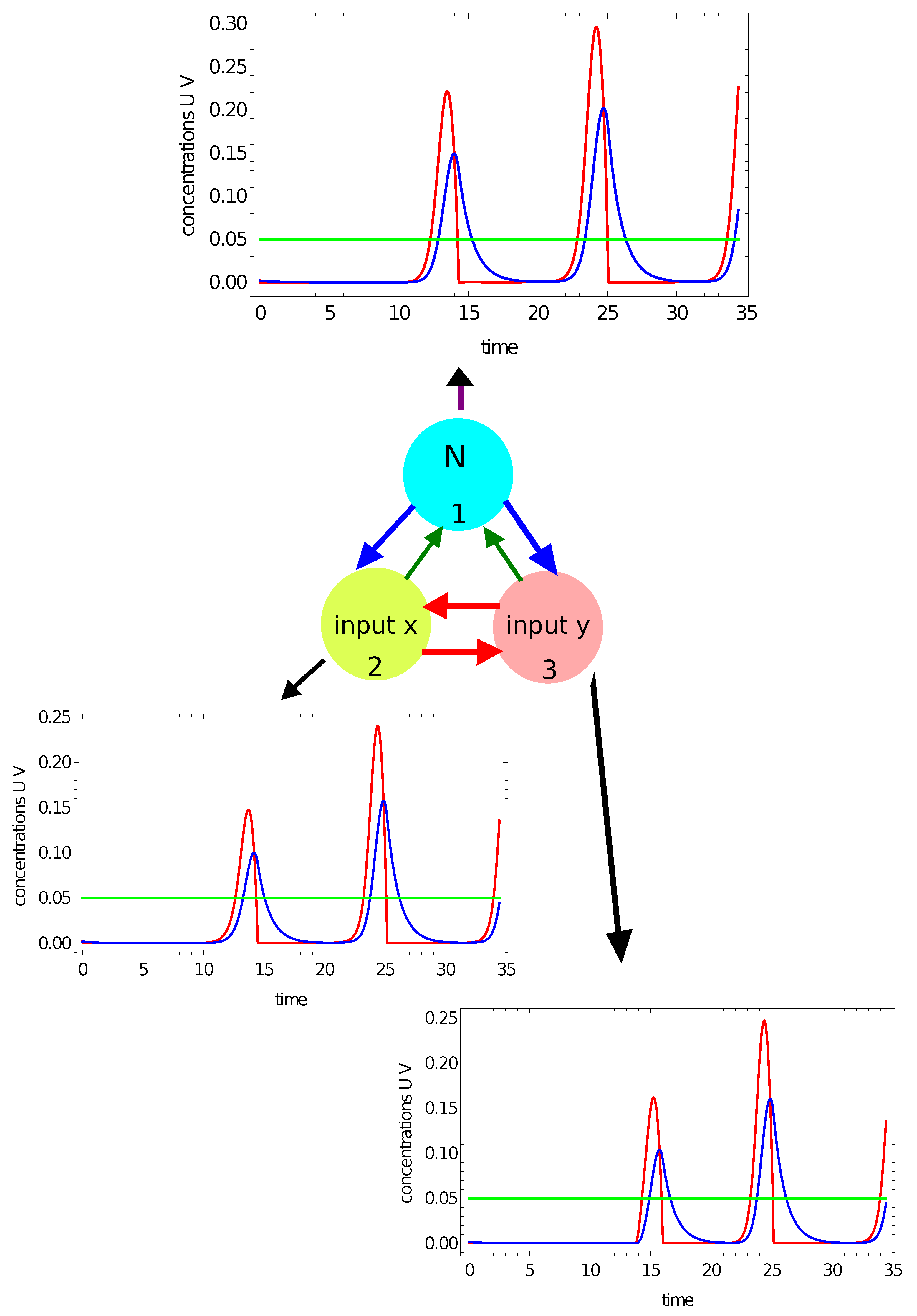
- There have to be input oscillators for each coordinate in the network and a normal oscillator. Keeping in mind the symmetry of the considered network, we can assume that node #1 is the normal oscillator and nodes #2 and #3 are the inputs of x- and y-coordinates, respectively;
- (3)
- The system symmetry reduces the number of parameters in the networks because: , , and .
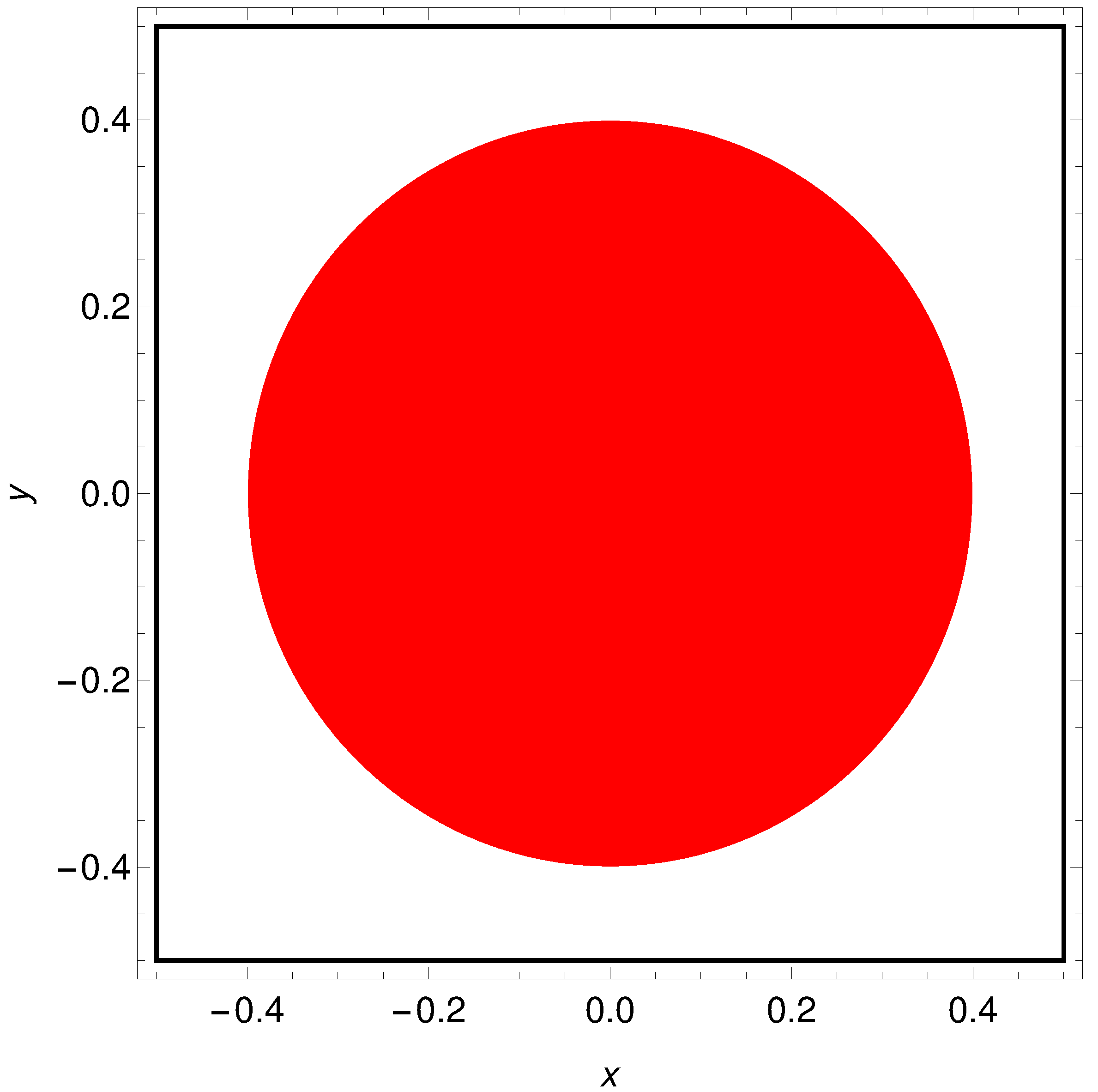
3. What Is the Color of a Point on the Japanese Flag? (As Seen by the Networks)
- -
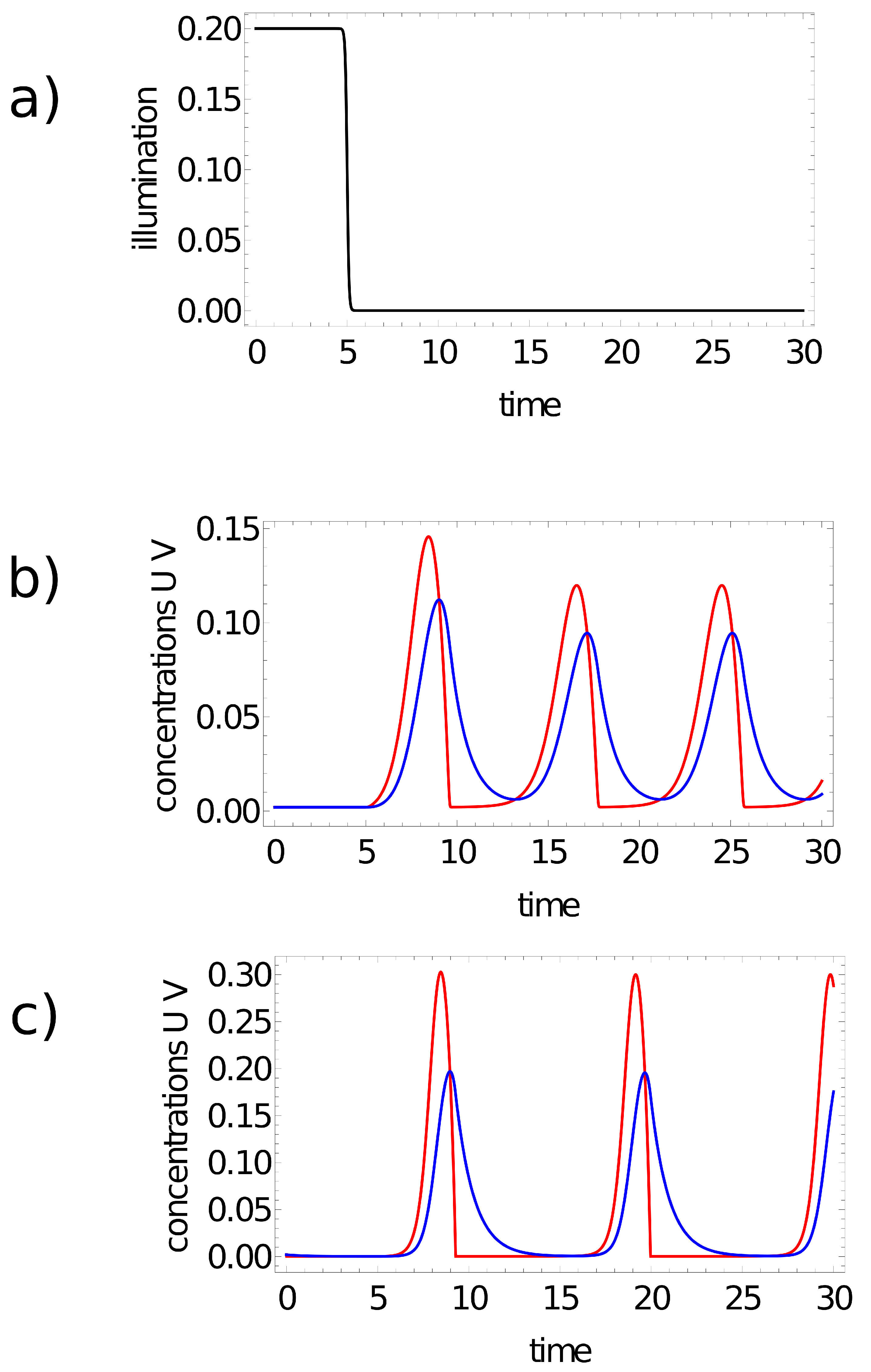
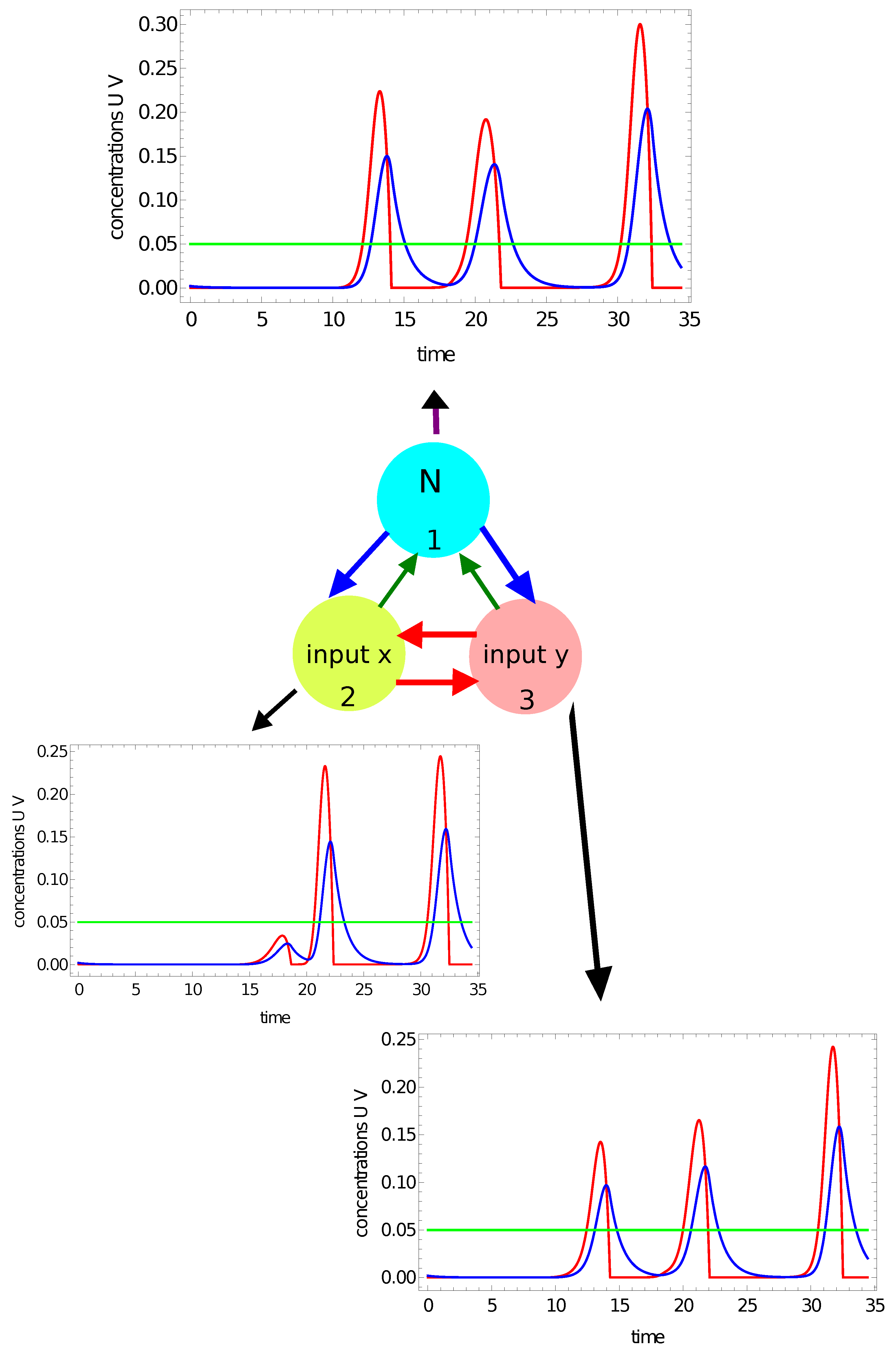
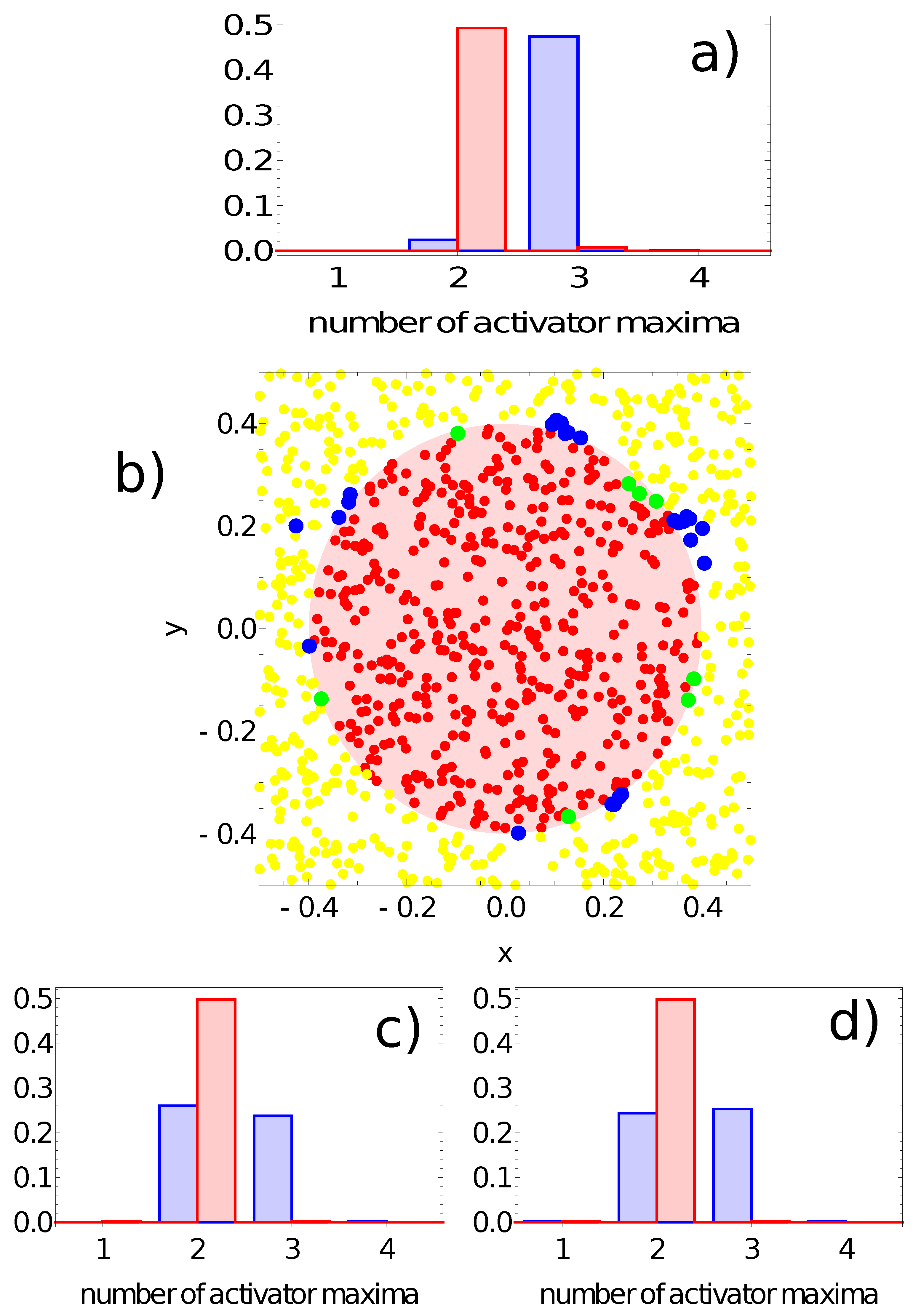
- If one or two maxima are observed, then the record represents a point in the sun area of the training dataset;
- -
- If three or more maxima are observed, then the record represents a point outside the sun area.
- -
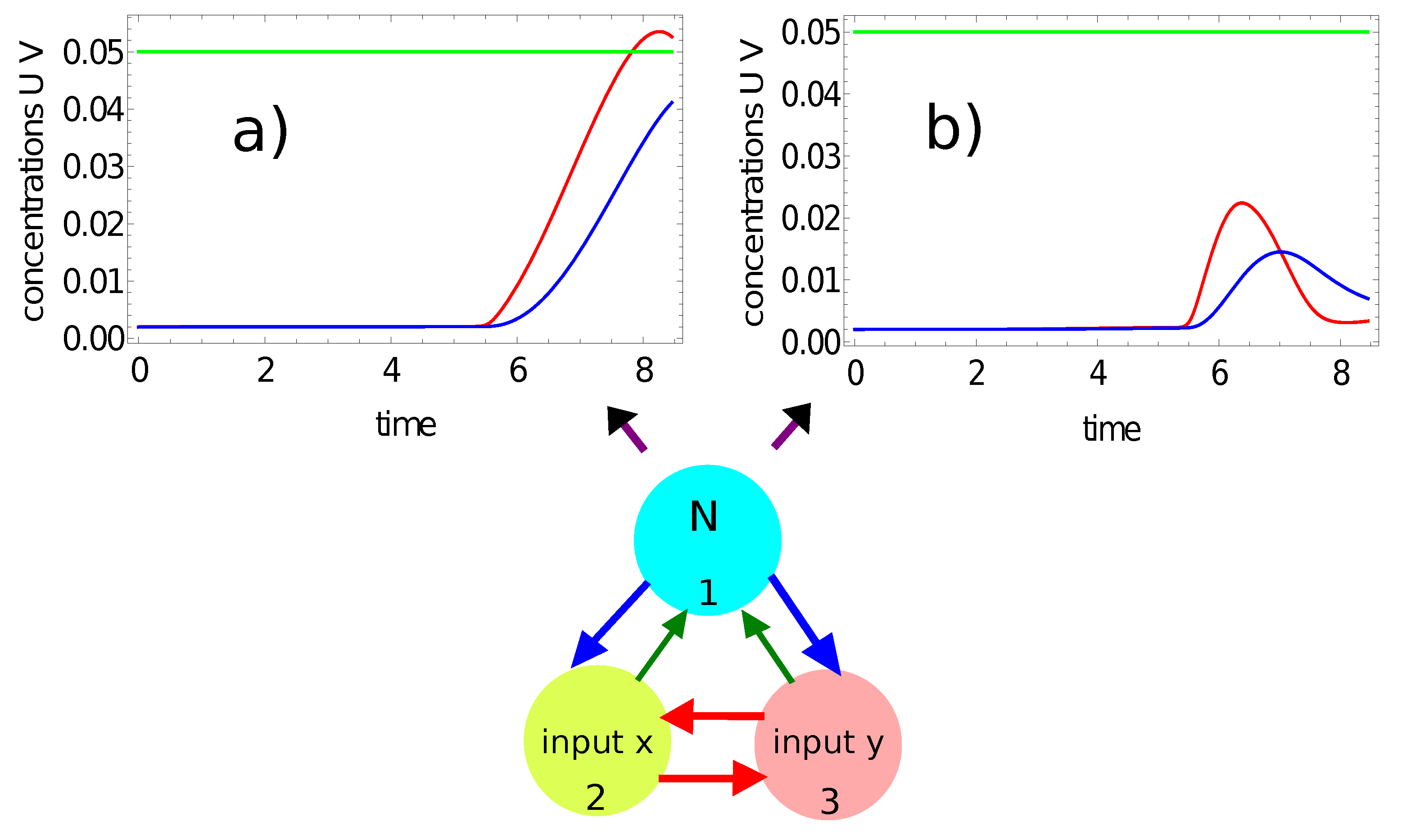
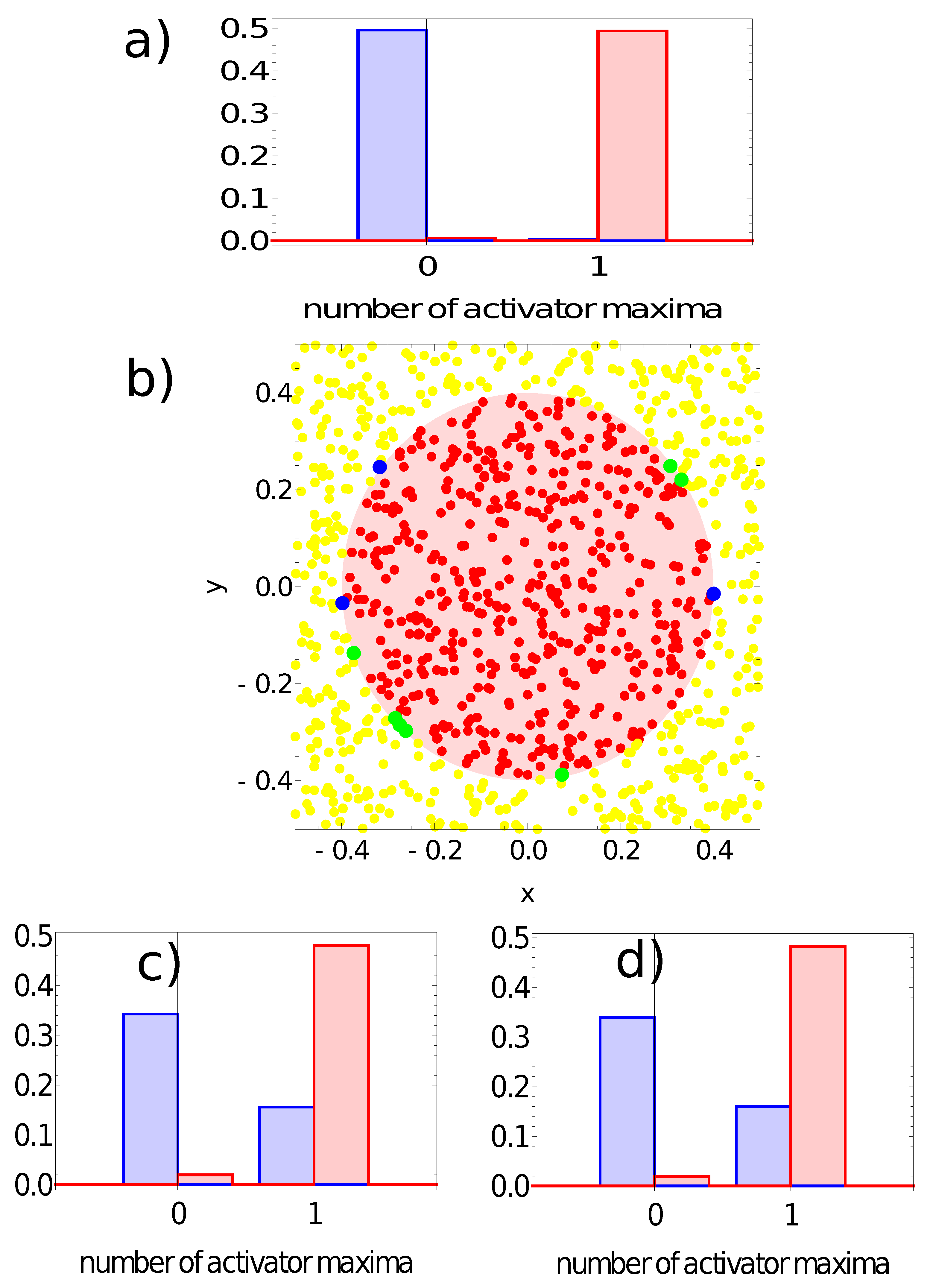
- If a single maximum of the activator is observed, then the record represents a point in the sun area of the training dataset;
- -
- If we record no maxima, then the processed data represent a point outside the sun area.
- -
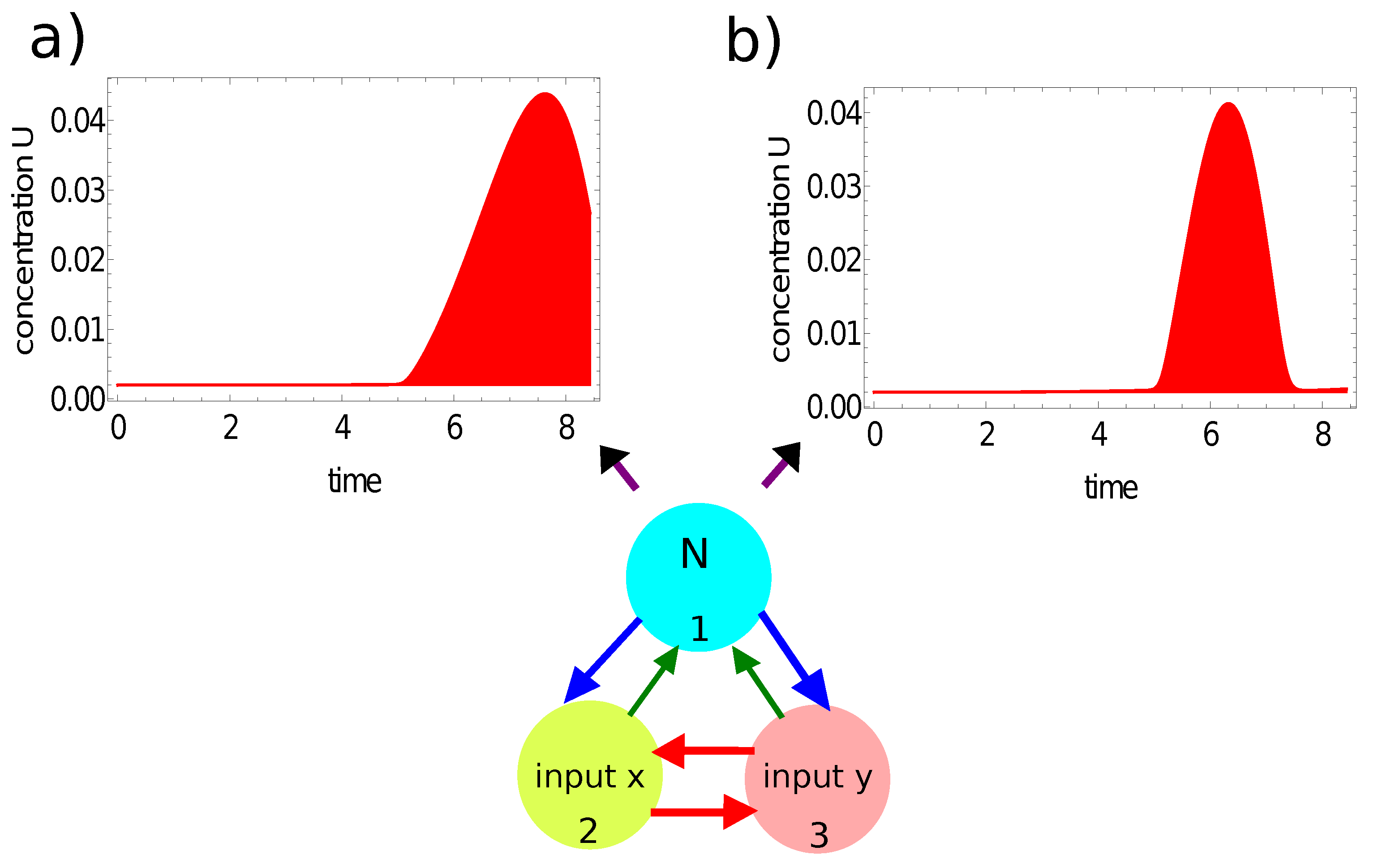
- If the value of , then the record represents a point in the sun area of the training dataset;
- -
- If the value of , then the record represents a point outside the sun area.
- -
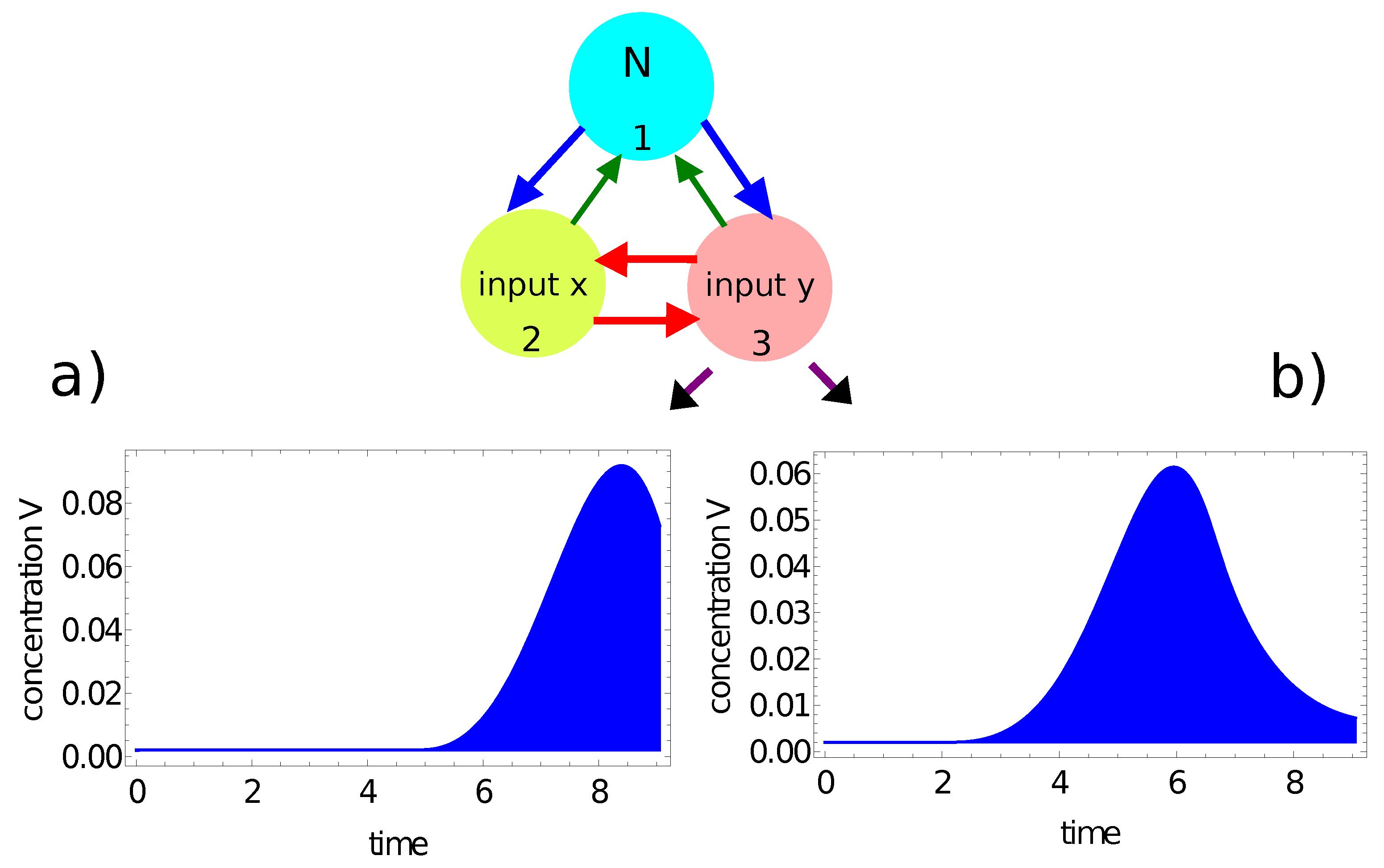
- If the value of , then the record represents a point in the sun area of the training dataset;
- -
- If the value of , then the record represents a point outside the sun area.
4. Discussion and Conclusions
- -
- If one or two maxima are observed, then the point is within the sun area;
- -
- If three or more maxima are observed, then the point is outside the sun area.
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviation
| BZ | Belousov–Zhabotinsky |
References
- Haken, H. Brain Dynamics; Springer Series in Synergetics; Springer: Berlin/Heidelberg, Germany, 2002. [Google Scholar]
- Moore, G.E. Cramming More Components onto Integrated Circuits. Electronics 1965, 38, 114–117. [Google Scholar] [CrossRef]
- Waldrop, M.M. The chips are down for Moore’s law. Nature 2016, 530, 144–147. [Google Scholar] [CrossRef] [PubMed]
- Feynman, R.P.; Hey, T.; Allen, R. Feynman Lectures on Computation; CRC Press: Boulder, CO, USA, 2000; ISBN 978-0738202969. [Google Scholar]
- Adamatzky, A. (Ed.) Advances in Uncnoventional Computing; Springer: Cham, Switzerland, 2018; Volume 2, ISBN 978-3-319-33920-7. [Google Scholar]
- Hjelmfelt, A.; Weinberger, E.D.; Ross, J. Chemical implementation of finite-state machines. Proc. Natl. Acad. Sci. USA 1992, 89, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Toth, A.; Showalter, K. Logic gates in excitable media. J. Chem. Phys. 1995, 103, 2058–2066. [Google Scholar] [CrossRef]
- Steinbock, O.; Kettunen, P.; Showalter, K. Chemical wave logic gates. J. Phys. Chem. 1996, 100, 18970–18975. [Google Scholar] [CrossRef]
- Adamatzky, A.; De Lacy Costello, B. Experimental logical gates in a reaction diffusion medium: The XOR gate and beyond. Phys. Rev. E 2002, 66, 046112. [Google Scholar] [CrossRef] [PubMed]
- Gorecki, J.; Gorecka, J.N.; Igarashi, Y. Information processing with structured excitable medium. Nat. Comput. 2009, 8, 473–492. [Google Scholar] [CrossRef]
- Magri, D.C.; Brown, G.J.; McClean, G.D.; de Silva, A.P. Communicating chemical congregation: A molecular AND logic gate with three chemical inputs as a “lab-on-a-molecule” prototype. J. Am. Chem. Soc. 2006, 128, 4950–4951. [Google Scholar] [CrossRef]
- de Silva, A.P.; Uchiyama, S. Molecular logic and computing. Nat. Nanotechnol. 2007, 2, 399–410. [Google Scholar] [CrossRef]
- Yoshikawa, K.; Motoike, I.N.; Ichino, T.; Yamaguchi, T.; Igarashi, Y.; Gorecki, J.; Gorecka, J.N. Basic information processing operations with pulses of excitation in a reaction-diffusion system. Int. J. Unconv. Comput. 2009, 5, 3–37. [Google Scholar]
- McKinney, B.O.F.; Daly, B.; Yao, C.; Schroeder, M.; de Silva, A.P. Consolidating Molecular Logic with New Solid-Bound YES and PASS 1 Gates and Their Combinations. ChemPhysChem 2017, 18, 1760. [Google Scholar] [CrossRef] [PubMed]
- Adamatzky, A.; De Lacy Costello, B.; Asai, T. Reaction-Diffusion Computers; Elsevier: New York, NY, USA, 2005. [Google Scholar]
- Lin, X.; Liu, Y.; Deng, J.; Lyu, Y.; Qian, P.; Lia, Y.; Wang, S. Multiple advanced logic gates made of DNA-Ag nanocluster and the application for intelligent detection of pathogenic bacterial genes. Chem. Sci. 2018, 9, 1774–1781. [Google Scholar] [CrossRef]
- Belousov, B.P. Collection of Short Papers on Radiation Medicine; Medgiz: Moscow, Russia, 1959; pp. 145–152. [Google Scholar]
- Zhabotinsky, A.M. Periodic liquid phase reactions. Proc. Acad. Sci. USSR 1964, 157, 392–395. [Google Scholar]
- Field, R.J.; Burger, M. Oscillations and Traveling Waves in Chemical Systems; Field, R.J., Burger, M., Eds.; Wiley: New York, NY, USA, 1985; ISBN 0-471-89384-6. [Google Scholar]
- Epstein, I.R.; Pojman, J.A. An Introduction to Nonlinear Chemical Dynamics: Oscillations, Waves, Patterns, and Chaos; Oxford University Press: New York, NY, USA, 1998; ISBN 9780195096705. [Google Scholar]
- Motoike, I.; Yoshikawa, K. Information Operations with an Excitable Field. Phys. Rev. E 1999, 59, 5354–5360. [Google Scholar] [CrossRef] [PubMed]
- Gorecka, J.N.; Gorecki, J. Multiargument logical operations performed with excitable chemical medium. J. Chem. Phys. 2006, 124, 084101. [Google Scholar] [CrossRef]
- Gentili, P.L.; Giubila, M.S.; Germani, R.; Romani, A.; Nicoziani, A.; Spalletti, A.; Heron, B.M. Optical Communication among Oscillatory Reactions and Photo-Excitable Systems: UV and Visible Radiation Can Synchronize Artificial Neuron Models. Angew. Chem. Int. Ed. 2017, 56, 7535. [Google Scholar] [CrossRef] [PubMed]
- Kuhnert, L. A new optical photochemical memory device in a light-sensitive chemical active medium. Nature 1986, 319, 393–394. [Google Scholar] [CrossRef]
- Kádár, S.; Amemiya, T.; Showalter, K. Reaction Mechanism for Light Sensitivity of the Ru(bpy)2+-Catalyzed Belousov–Zhabotinsky Reaction. J. Phys. Chem. A 1997, 101, 8200–8206. [Google Scholar] [CrossRef]
- Agladze, K.; Aliev, R.; Yamaguchi, T. Chemical Diode. J. Phys. Chem. 1996, 100, 13895–13897. [Google Scholar] [CrossRef]
- Motoike, I.N.; Yoshikawa, K.; Iguchi, Y.; Nakata, S. Real-Time Memory on an Excitable Field. Phys. Rev. E 2001, 63, 036220. [Google Scholar] [CrossRef] [PubMed]
- Gorecki, J.; Gorecka, J.N. On mathematical description of information processing in chemical systems. In Mathematical Approach to Nonlinear Phenomena: Modeling, Analysis and Simulations; GAKUTO International Series: Mathematical Sciences and Applications; Gakkotosho: Tokyo, Japan, 2005; Volume 23, pp. 73–90. ISBN 4762504327. [Google Scholar]
- Adleman, L.M. Molecular Computation of Solutions to Combinatorial Problems. Science 1994, 266, 1021–1024. [Google Scholar] [CrossRef] [PubMed]
- Calude, C.S.; Păun, G. Computing with Cells and Atoms; Taylor & Francis Publisher: London, UK, 2002. [Google Scholar]
- Steinbock, O.; Toth, A.; Showalter, K. Navigating complex labyrinths—Optimal paths from chemical waves. Science 1995, 267, 868–871. [Google Scholar] [CrossRef] [PubMed]
- Agladze, K.; Magome, N.; Aliev, R.; Yamaguchi, T.; Yoshikawa, K. Finding the optimal path with the aid of chemical wave. Physica D 1997, 106, 247–254. [Google Scholar] [CrossRef]
- Kuhnert, L.; Agladze, K.I.; Krinsky, V.I. Image processing using light-sensitive chemical waves. Nature 1989, 337, 244–247. [Google Scholar] [CrossRef]
- Rambidi, N.G.; Maximychev, A.V. Towards a biomolecular computer. Information processing capabilities of biomolecular nonlinear dynamic media. Biosystems 1997, 41, 195–211. [Google Scholar] [CrossRef]
- Szymanski, J.; Gorecka, J.N.; Igarashi, Y.; Gizynski, K.; Gorecki, J.; Zauner, K.P.; de Planque, M. Droplets with information processing ability. Int. J. Unconv. Comput. 2011, 7, 185–200. [Google Scholar]
- Gorecki, J.; Gizynski, K.; Guzowski, J.; Gorecka, J.N.; Garstecki, P.; Gruenert, G.; Dittrich, P. Chemical Computing with Reaction-Diffusion processes. Phil. Trans. R. Soc. A. 2015, 373, 20140219. [Google Scholar] [CrossRef]
- Guzowski, J.; Gizynski, K.; Gorecki, J.; Garstecki, P. Microfluidic platform for reproducible self-assembly of chemically communicating droplet networks with predesigned number and type of the communicating compartments. Lab Chip 2016, 16, 764–772. [Google Scholar] [CrossRef] [PubMed]
- Buisman, H.J.; ten Eikelder, H.M.M.; Hilbers, P.A.J.; Liekens, A.M.L. Computing Algebraic Functions with Biochemical Reaction Networks. Artif. Life 2009, 15, 5–19. [Google Scholar] [CrossRef] [PubMed]
- Hjelmfelt, A.; Weinberger, E.D.; Ross, J. Chemical implementation of neural networks and Turing machines. Proc. Nati. Acad. Sci. USA 1991, 88, 10983–10987. [Google Scholar] [CrossRef] [PubMed]
- Hjelmfelt, A.; Ross, J. Chemical implementation and thermodynamics of collective neural networks. Proc. Nati. Acad. Sci. USA 1992, 89, 388–391. [Google Scholar] [CrossRef] [PubMed]
- Hjelmfelt, A.; Ross, J. Pattern recognition, chaos, and multiplicity in neural networks of excitable systems. Proc. Natl. Acad. Sci. USA 1994, 91, 63–67. [Google Scholar] [CrossRef] [PubMed]
- Adamatzky, A.; Holley, J.; Bull, L.; De Lacy Costello, B. On computing in fine-grained compartmentalised Belousov–Zhabotinsky medium. Chaos Solitons Fractals 2011, 44, 779–790. [Google Scholar] [CrossRef]
- Holley, J.; Adamatzky, A.; Bull, L.; De Lacy Costello, B.; Jahan, I. Computational modalities of Belousov–Zhabotinsky encapsulated vesicles. Nano Commun. Netw. 2011, 2, 50–61. [Google Scholar] [CrossRef]
- Adamatzky, A.; Holley, J.; Dittrich, P.; Gorecki, J.; De Lacy Costello, B.; Zauner, K.-P.; Bull, L. On architectures of circuits implemented in simulated Belousov–Zhabotinsky droplets. Biosystems 2012, 109, 72–77. [Google Scholar] [CrossRef]
- Gizynski, K.; Gorecki, J. Cancer classification with a network of chemical oscillators. Phys. Chem. Chem. Phys. 2017, 19, 28808–28819. [Google Scholar] [CrossRef]
- Gizynski, K.; Gruenert, G.; Dittrich, P.; Gorecki, J. Evolutionary design of classifiers made of oscillators containing a nonlinear chemical medium. MIT Evol. Comput. 2017, 25, 643–671. [Google Scholar] [CrossRef]
- Gorecki, J.; Bose, A. How Does a Simple Network of Chemical Oscillators See the Japaneseese Flag? Front. Chem. 2020, 8, 580703. [Google Scholar] [CrossRef]
- Duenas-Diez, M.; Perez-Mercader, J. How Chemistry Computes: Language Recognition by Non-Biochemical Chemical Automata. From Finite Automata to Turing Machines. iScience 2019, 19, 514–526. [Google Scholar] [CrossRef]
- Bose, A.; Gorecki, J. Computing With Networks of Chemical Oscillators and its Application for Schizophrenia Diagnosis. Front. Chem. 2022, 10, 848685. [Google Scholar] [CrossRef]
- Field, R.J.; Noyes, R.M. Oscillations in chemical systems. IV. Limit cycle behavior in a model of a real chemical reaction. J. Chem. Phys. 1974, 60, 1877–1884. [Google Scholar] [CrossRef]
- Sutthiopad, M.; Luengviriya, J.; Porjai, P.; Tomapatanaget, B.; Müller, S.C.; Luengviriya, C. Unpinning of spiral waves by electrical forcing in excitable chemical media. Phys. Rev. E 2014, 89, 052902. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, M.; Nagahara, H.; Kitahata, H.; Krinsky, V.; Agladze, K.; Yoshikawa, K. Survival versus collapse: Abrupt drop of excitability kills the traveling pulse, while gradual change results in adaptation. Phys. Rev. E 2007, 76, 016205. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gorecki, J.; Gorecka, J.N.; Adamatzky, A. Information coding with frequency of oscillations in Belousov–Zhabotinsky encapsulated disks. Phys. Rev. E 2014, 89, 042910. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Gizynski, K.; Gorecki, J. Chemical memory with states coded in light controlled oscillations of interacting Belousov–Zhabotinsky droplets. Phys. Chem. Chem. Phys. 2017, 19, 6519–6531. [Google Scholar] [CrossRef]
- Press, W.H.; Teukolsky, S.A.; Vetterling, W.T.; Flannery, B.P. Numerical Recipes: The Art of Scientific Computing; Cambridge University Press: New York, NY, USA, 2007; ISBN 978-0-521-88068-8. [Google Scholar]
- MacKay, D.J.C. Information Theory, Inference and Learning Algorithms; Cambridge University Press: Cambridge, UK, 2003; ISBN 978-0-521-64298-9. [Google Scholar]
- Cover, T.M.; Thomas, J.A. Elements of Information Theory; Wiley InterScience: New York, NY, USA, 2006; ISBN 978-0-471-24195-9. [Google Scholar]
- Shannon, C.E. A Mathematical Theory of Communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Gorecki, J. Applications of Information Theory Methods for Evolutionary Optimization of Chemical Computers. Entropy 2020, 22, 313. [Google Scholar] [CrossRef]
- Goldberg, D.E. Genetic Algorithms in Search, Optimization and Machine Learning; Addison-Wesley Longman Publishing Co., Inc.: Boston, MA, USA, 1989. [Google Scholar]


| Oregonator | Method | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Model I | activator maxima | 34.4 | 11.6 | 19.9 | 10.1 | 0.87 | 0.72 | 0.16 | 0.43 | 0.29 |
| Model II | activator maxima | 8.45 | 3.77 | 8.03 | 5.42 | 0.96 | 0.46 | 0.53 | 0.38 | 0.42 |
| Model II | u-integral | 8.43 | 3.77 | 7.41 | 5.00 | 0.65 | 0.50 | 0.83 | 0.26 | 0.29 |
| Model II | v-integral | 9.06 | 3.77 | 7.56 | 5.71 | 0.75 | 0.44 | 0.60 | 0.29 | 0.33 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gorecki, J. Information Processing Using Networks of Chemical Oscillators. Entropy 2022, 24, 1054. https://doi.org/10.3390/e24081054
Gorecki J. Information Processing Using Networks of Chemical Oscillators. Entropy. 2022; 24(8):1054. https://doi.org/10.3390/e24081054
Chicago/Turabian StyleGorecki, Jerzy. 2022. "Information Processing Using Networks of Chemical Oscillators" Entropy 24, no. 8: 1054. https://doi.org/10.3390/e24081054
APA StyleGorecki, J. (2022). Information Processing Using Networks of Chemical Oscillators. Entropy, 24(8), 1054. https://doi.org/10.3390/e24081054






