RNA World Modeling: A Comparison of Two Complementary Approaches
Abstract
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. RP Model
2.2. Modeling with Partial Differential Equations (PDE)
2.2.1. Replication
2.2.2. Decay
2.2.3. Resource Formation
2.2.4. Diffusion
2.2.5. PDE Model and Its Assumptions
2.2.6. The Solutions for a PDE Model
Well-Mixed Solution
- : replicases and parasites can coexist in any proportion, but only if ;
- The denominator in Equation (22) is equal to 0. Unless the previous special case is also fulfilled, there is no solution.
Linear Stability
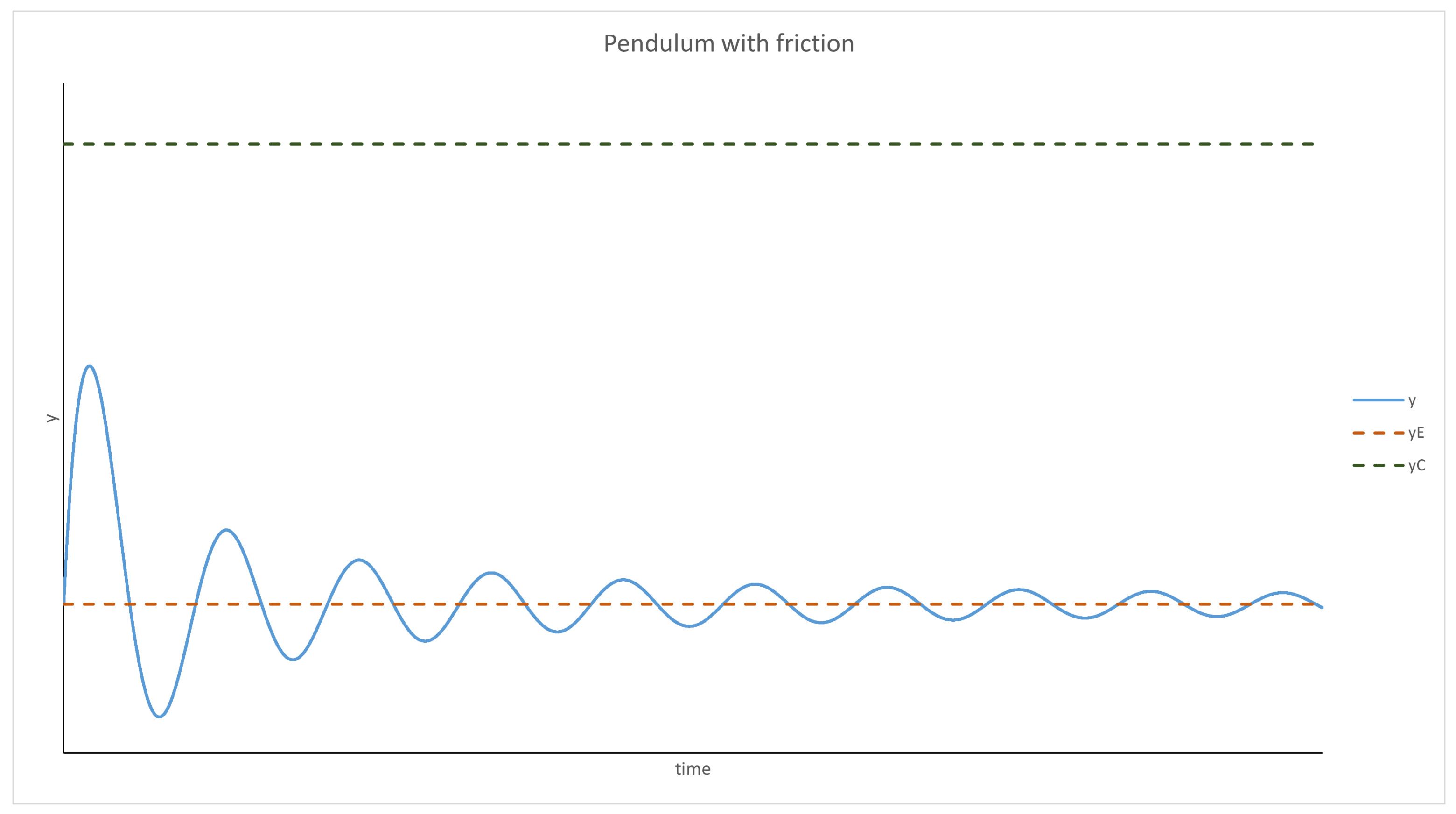
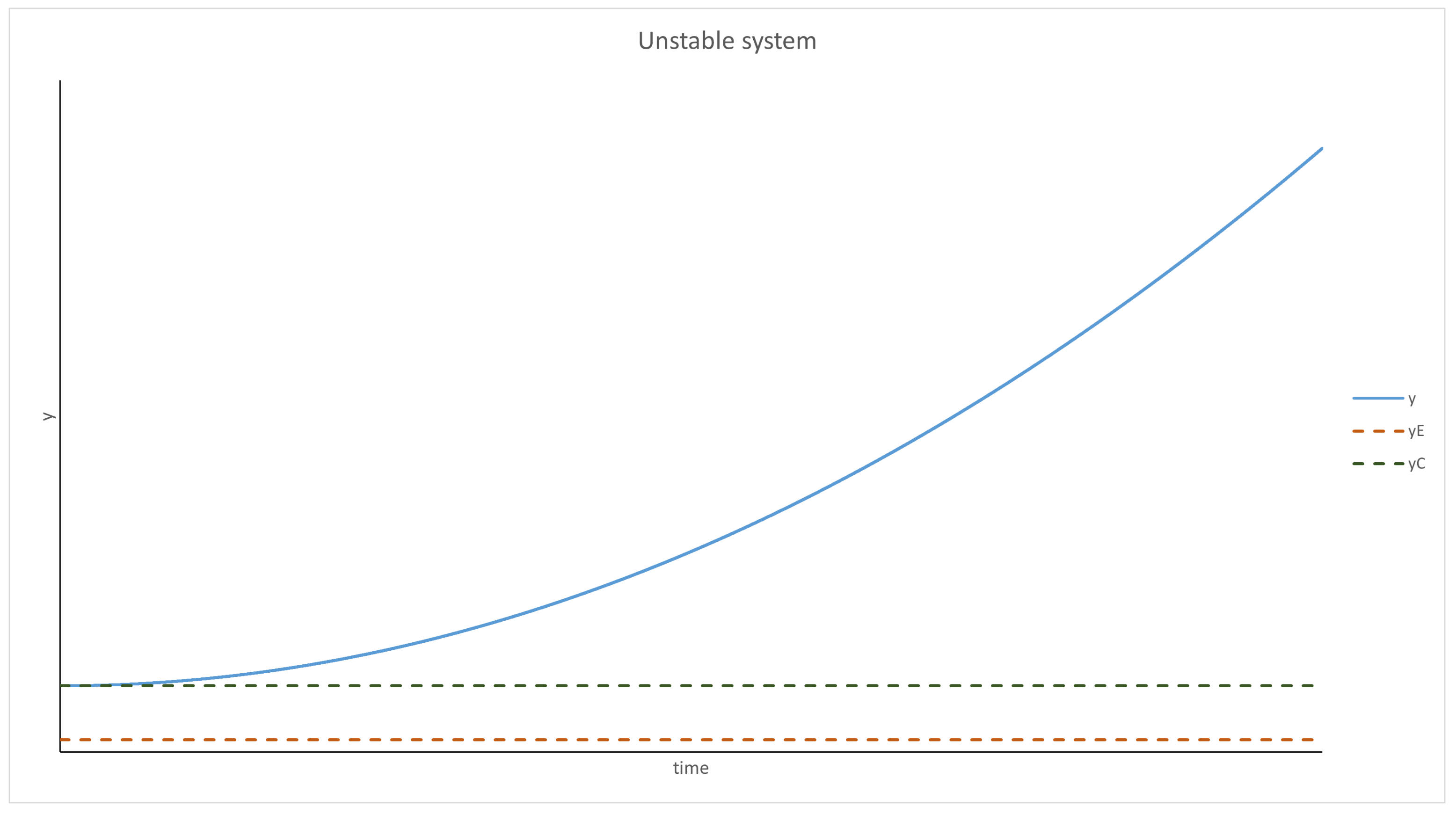
Evolution of a Well-Mixed Solution
- has to have a root—an equilibrium point. Moreover, it has to be negative for any y bigger than it and positive for anything smaller;
- cannot be positive.
Mutation
Partial Differential Equations Model Robustness
- Diffusion: The presence of diffusion allows spatial interactions to occur. If there exists a stable, local equilibrium, this will be reached at every point and the system will become homogenous. However, for more complicated scenarios (such as the presence of mutation and the lack of a stable, local equilibrium), more complex structures can emerge and lead to the system’s survival. These cases are analyzed using computer simulations in the latter part of this article;
- Complex formation: In this article, the replication of RNA molecules is treated as an instantaneous reaction, whereas, in reality, it takes time. In order to account for this, a new type of molecule can be added to the model—a complex of replicase with the template representing the ongoing replication—just as in [41]. However, the results obtained in the aforementioned work and this article show very similar behavior; hence, the addition of complexes does not seem to have much influence;
- Different chemical reaction dynamics: reactions are assumed to follow the law of mass action. Due to the very complicated nature of complex molecules, especially in biology, replicases can behave differently depending on the density of resources and potential templates. The simplest case was assumed due to the fact that no functional replicase has been created in vitro as of yet; thus, their exact properties remain unknown. Changing the way the chemical reactions in the model work can radically change the outcome; for instance, replicases “programmed” to ignore parasites and only copy other replicases (but only if there are plenty of resources present) would be able to ensure the system’s survival without any additional mechanisms.
2.3. Description of the Partial Differential Equation Simulation Algorithm
- p: the density of parasites;
- r: the density of replicases;
- a: the average of parasites;
- n: the density of resources necessary for replication.
| Algorithm 1 Differential equation simulation |
| Initialize two 2D arrays— and |
| for all field f of do |
| Set , , and to 1. |
| end for |
| for all simulation step do |
| for all field f of do |
| if and then |
| end if |
| Compute average values for f’s adjacent neighbors: , , and . |
| Find the corresponing field of f in - . |
| if then |
| else |
| end if |
| Randomly change according to mutation probability distribution. |
| Set all negative variables of to 0. |
| end for |
| Save results of the current simulation step. |
| Swap and . |
| end for |
2.4. Description of the Multi-Agent Approach and Simulation Algorithm (MAS)
2.4.1. MAS Parameters
- Position;
- Type (replicase or parasite);
- (parasites only): the probability of being replicated.
2.4.2. First-Order Reactions
2.4.3. Second Order Reactions
- Two parasites: no reaction;
- Parasite and replicase: parasite becomes a template for replication and its becomes the probability of the reaction;
- Two replicases: the reaction occurs with the probability of (global parameter).
2.4.4. Mutation
2.4.5. Diffusion
| Algorithm 2 Multi-agent algorithm |
| Initialize the agents’ positions randomly. |
| Initialize all parasites with equal values. |
| while (simulation time < time limit) and (there are both parasites and replicases present) do |
| Decrease the RLT time for each agent. |
| for all agents with the RLT = 0 do |
| Remove the agent. |
| end for |
| Randomize the order of all agents. |
| for all agent do |
| Initialize an empty set of neighbors N. |
| for all agent that overlaps do |
| Add it to the set of neighbors N |
| end for |
| if then |
| Remove agent from the simulation. |
| end if |
| Randomize the order of N. |
| Move by a random vector (Gaussian distribution with variance ). |
| for all do |
| if ( or is a replicase) and reaction occurred (probability /) then |
| Create a copy of the template and place it in the same position. |
| Initialize new agents’ RLT value. |
| if New agent is a parasite then |
| Mutate new agent’s . |
| end if |
| end if |
| end for |
| end for |
| end while |
3. Results
3.1. Scenarios 1–8
| Diffusion Constant | Mutation Rate | Result |
|---|---|---|
| 5 | 0.1 | Alive (Figure 4) |
| 5 | 0.2 | Alive (Figure 5) |
| 10 | 0.1 | Alive (Figure 6) |
| 10 | 0.2 | Extinction |
| 15 | 0.1 | Extinction |
| 15 | 0.2 | Extinction |
| 20 | 0.1 | Extinction |
| 20 | 0.2 | Extinction |
3.2. Scenario 9
3.3. Scenario 10
3.4. Scenario 11
3.5. Scenarios 12 and 13
3.6. The Summary of Scenarios 1–13
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Szostak, N.; Wasik, S.; Blazewicz, J. Understanding Life: A Bioinformatics Perspective. Eur. Rev. 2017, 25, 231–245. [Google Scholar] [CrossRef]
- Szostak, J. The Narrow Road to the Deep Past: In Search of the Chemistry of the Origin of Life. Angew. Chem. Int. Ed. Engl. 2017, 56, 11037–11043. [Google Scholar] [CrossRef] [PubMed]
- Totani, T. Emergence of life in an inflationary universe. Sci. Rep. 2020, 10, 1671. [Google Scholar] [CrossRef] [PubMed]
- Schneider, T. A brief review of molecular information theory. Nano Commun. Netw. 2010, 1, 173–180. [Google Scholar] [CrossRef] [PubMed]
- Adami, C. The use of information theory in evolutionary biology. Ann. N. Y. Acad. Sci. 2012, 1256, 49–65. [Google Scholar] [CrossRef]
- Landauer, R. Information is Physical. Physics Today 1991, 44, 23–29. [Google Scholar] [CrossRef]
- Shannon, C. A mathematical theory of communication. Bell. Syst. Technol. J. 1948, 27, 623–656. [Google Scholar] [CrossRef]
- Yockey, H. Information theory, evolution and the origin of life. Inf. Sci. 2002, 141, 219–225. [Google Scholar] [CrossRef]
- MacKay, D. Information, Mechanism and Meaning; MIT Press: Cambridge, MA, USA, 1969. [Google Scholar]
- Kauffman, S. Investigations; Oxford University Press: Oxford, UK, 2000. [Google Scholar]
- Gilbert, W. Origin of life: The RNA world. Nature 1986, 319, 618. [Google Scholar] [CrossRef]
- Cech, T. Structural biology. The ribosome is a ribozyme. Science 2000, 289, 878–879. [Google Scholar] [CrossRef]
- Weinberg, C.; Weinberg, Z.; Hammann, C. Novel ribozymes: Discovery, catalytic mechanisms, and the quest to understand biological function. Nucleic Acids Res. 2019, 47, 9480–9494. [Google Scholar] [CrossRef] [PubMed]
- Kun, Á.; Szilágyi, A.; Könnyu, B.; Boza, G.; Zachar, I.; Szathmáry, E. The dynamics of the RNA world: Insights and challenges. Ann. N. Y. Acad. Sci. 2015, 1341, 75–95. [Google Scholar] [CrossRef] [PubMed]
- De Farias, S.; dos Santos Junior, A.; Rego, T.; Jose, M. Origin and evolution of RNA-dependent RNA polymerase. Front. Genet. 2017, 28, 125. [Google Scholar] [CrossRef] [PubMed]
- Dos Santos Junior, A.; Jose, M.; de Farias, S. From RNA to DNA: Insights about the transition of informational molecule in the biological systems based on the structural proximity between the polymerases. Biosystems 2021, 206, 104442. [Google Scholar] [CrossRef]
- Robertson, M.; Joyce, G. The origins of the RNA world. Cold Spring Harb. Perspect. Biol. 2012, 4, a003608. [Google Scholar] [CrossRef]
- Wachowius, F.; Holliger, P. Non-Enzymatic Assembly of a Minimized RNA Polymerase Ribozyme. ChemSystemsChem 2019, 1, 1–4. [Google Scholar] [CrossRef]
- Wochner, A.; Attwater, J.; Coulson, A.; Holliger, P. Ribozyme-Catalyzed Transcription of an Active Ribozyme. Science 2011, 332, 209–212. [Google Scholar] [CrossRef]
- Attwater, J.; Raguram, A.; Morgunov, A.; Gianni, E.; Holliger, P. Ribozyme-catalysed RNA synthesis using triplet building blocks. eLife 2018, 7, e35255. [Google Scholar] [CrossRef]
- Oliver, C.; Reinharz, V.; Waldispühl, J. On the Emergence of Structural Complexity in RNA Replicators. RNA 2019, 25, 1579–1591. [Google Scholar] [CrossRef]
- Doudna, J.; Couture, S.; Szostak, J. A multisubunit ribozyme that is a catalyst of and template for complementary strand RNA synthesis. Science 1991, 251, 1605–1608. [Google Scholar] [CrossRef]
- Mutschler, H.; Wochner, A.; Holliger, P. Freeze-thaw cycles as drivers of complex ribozyme assembly. Science 2015, 7, 502–508. [Google Scholar] [CrossRef] [PubMed]
- Eigen, M.; Schuster, P. The Hypercycle: A Principle of Natural Self-Organization; Springer: New York, NY, USA, 1979. [Google Scholar] [CrossRef]
- Szilágyi, A.; Zachar, I.; Scheuring, I.; Kun, A.; Könnyu, B.; Czárán, T. Ecology and Evolution in the RNA World Dynamics and Stability of Prebiotic Replicator Systems. Life 2017, 7, 48. [Google Scholar] [CrossRef] [PubMed]
- Shah, V.; de Bouter, J.; Pauli, Q.; Tupper, A.; Higgs, P. Survival of RNA Replicators is much Easier in Protocells than in Surface-Based, Spatial Systems. Life 2019, 9, 65. [Google Scholar] [CrossRef] [PubMed]
- Smith, J. Models of evolution. Proc. R. Soc. Lond. B 1983, 219, 315–325. [Google Scholar] [CrossRef]
- Eigen, M. Selforganization of matter and the evolution of biological macromolecules. Die Naturwissenschaften 1971, 58, 465–523. [Google Scholar] [CrossRef]
- Eigen, M.; Schuster, P. The hypercycle. A principle of natural self-organization. Part A: Emergence of the hypercycle. Die Naturwissenschaften 1977, 64, 541–565. [Google Scholar] [CrossRef]
- Smith, J.; Szathmáry, E. The Major Transitions in Evolution; Oxford University Press: Oxford, UK, 1995. [Google Scholar]
- Kun, A.; Szathmáry, E. Real ribozymes suggest a relaxed error threshold. Nat. Genet. 2005, 37, 1008–1011. [Google Scholar] [CrossRef]
- Szostak, N.; Wasik, S.; Blazewicz, J. Hypercycle. PLoS Comput. Biol. 2016, 12, e1004853. [Google Scholar] [CrossRef]
- Smith, J. Hypercycles and the origin of life. Nature 1979, 280, 445–446. [Google Scholar] [CrossRef]
- Hogeweg, P.; Takeuchi, N. Multilevel selection in models of prebiotic evolution: Compartments and spatial self-organization. Orig. Life Evol. Biosph. 2003, 33, 375–403. [Google Scholar] [CrossRef]
- Boerlijst, M.; Hogeweg, P. Spiral wave structure in pre-biotic evolution: Hypercycles stable against parasite. Phys. D Nonlinear Phenom. 1991, 48, 17–28. [Google Scholar] [CrossRef]
- Cronhjort, M.; Blomberg, C. Hypercycles versus parasites in a two dimensional partial differential equations model. J. Theor. Biol. 1994, 169, 31–49. [Google Scholar] [CrossRef]
- Takeuchi, N.; Hogeweg, P. Multilevel Selection in Models of Prebiotic Evolution II: A Direct Comparison of Compartmentalization and Spatial Self-Organization. PLoS Comput. Biol. 2009, 5, e1000542. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, N.; Hogeweg, P. Evolutionary dynamics of RNA-like replicator systems: A bioinformatic approach to the origin of life. Phys. Life Rev. 2012, 9, 219–263. [Google Scholar] [CrossRef]
- Boerlijst, M.; Hogeweg, P. Spatial gradients enhance persistence of hypercycles. Phys. D Nonlinear Phenom. 1988, 88, 29–39. [Google Scholar] [CrossRef]
- Takeuchi, N.; Hogeweg, P. Evolution of complexity in RNA-like replicator systems. Biol. Direct 2008, 3, 11. [Google Scholar] [CrossRef]
- Szostak, N.; Synak, J.; Borowski, M.; Wasik, S.; Blazewicz, J. Simulating the origins of life: The dual role of RNA replicases as an obstacle to evolution. PLoS ONE 2017, 12, e0180827. [Google Scholar] [CrossRef]
- Synak, J.; Rybarczyk, A.; Blazewicz, J. Multi-agent approach to sequence structure simulation in the RNA World hypothesis. PLoS ONE 2020, 15, e0238253. [Google Scholar] [CrossRef]
- Pearce, B.; Pudritz, R.; Semenov, D.; Henning, T. Fate of nucleobases in warm little ponds. Proc. Natl. Acad. Sci. USA 2017, 114, 11327–11332. [Google Scholar] [CrossRef]
- Szilágyi, A.; Könnyu, B.; Czárán, T. Dynamics and stability in prebiotic information integration: An RNA World model from first principles. Sci. Rep. 2020, 10, 51. [Google Scholar] [CrossRef]
- Takeuchi, N.; Hogeweg, P. The Role of Complex Formation and Deleterious Mutations for the Stability of RNA-Like Replicator Systems. J. Mol. Evol. 2007, 65, 668–686. [Google Scholar] [CrossRef] [PubMed]
- Berezovskaya, F.; Karev, G.; Katsnelson, M.; Wolf, Y.; Koonin, E. Stable coevolutionary regimes for genetic parasites and their hosts: You must differ to coevolve. Biol. Direct 2018, 13, 27. [Google Scholar] [CrossRef] [PubMed]
- Hofbauer, J.; Sigmund, K. The Theory of Evolution and Dynamical Systems: Mathematical Aspects of Selection; Cambridge University Press: Cambridge, UK, 1988. [Google Scholar]
- McCaskill, J.; Bauer, G. Images of evolution: Origin of spontaneous RNA replication waves. Proc. Natl. Acad. Sci. USA 1993, 90, 4191–4195. [Google Scholar] [CrossRef] [PubMed]
- Tupper, A.; Higgs, P. Error thresholds for RNA replication in the presence of both point mutations and premature termination errors. J. Theor. Biol. 2017, 428, 34–42. [Google Scholar] [CrossRef]
- Duim, H.; Otto, S. Towards open-ended evolution in self-replicating molecular systems. Beilstein J. Org. Chem. 2017, 13, 1189–1203. [Google Scholar] [CrossRef]
- Le Vay, K.; Mutschler, H. The difficult case of an RNA-only origin of life. Emerg. Top. Life Sci. 2019, 3, 469–475. [Google Scholar] [CrossRef]
- Furubayashi, T.; Ueda, K.; Bansho, Y.; Motooka, D.; Nakamura, S.; Mizuuchi, R.; Ichihashi, N. Emergence and diversification of a host-parasite RNA ecosystem through Darwinian evolution. eLife 2020, 9, e56038. [Google Scholar] [CrossRef]
- Ellinger, T.; Ehricht, R.; McCaskill, J. In vitro evolution of molecular cooperation in CATCH, a cooperatively coupled amplification system. Chem. Biol. 1998, 5, 729–741. [Google Scholar] [CrossRef][Green Version]
- Mizuuchi, R.; Furubayashi, T.; Ichihashi, N. Evolutionary transition from a single RNA replicator to a multiple replicator network. Nat. Commun. 2022, 13, 1460. [Google Scholar] [CrossRef]
- Bansho, Y.; Furubayashi, T.; Ichihashi, N.; Yomo, T. Host-parasite oscillation dynamics and evolution in a compartmentalized RNA replication system. Proc. Natl. Acad. Sci. USA 2016, 113, 4045–4050. [Google Scholar] [CrossRef]




| Parameter Name | Default Value | Description |
|---|---|---|
| , | 1000 | Simulation area size |
| d | 0.01 | Decay rate |
| 1.0 | Probability of replicases reaction with replicases | |
| 0.1 | Single step length ( in equations) | |
| D | 5.0 | Diffusion constant |
| 10.0 | Resources diffusion constant | |
| 1.0 | Resources production rate | |
| m | 0.0 | Mutation of a replicase into a parasite probability |
| Parameter Name | Default Value | Description |
|---|---|---|
| , | 800 | Simulation area size |
| 3.0 | Radius of the circle representing an agent | |
| 4 | Maximum number of neighbors | |
| d | 0.1 | Decay rate |
| 1.0 | Affinity of replicases towards replicases | |
| 1.0 | Single step length | |
| D | 15.0 | Diffusion constant |
| 0.1 | Parasite mutation speed | |
| 0.1 | Parasite mutation probability |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Synak, J.; Rybarczyk, A.; Blazewicz, J. RNA World Modeling: A Comparison of Two Complementary Approaches. Entropy 2022, 24, 536. https://doi.org/10.3390/e24040536
Synak J, Rybarczyk A, Blazewicz J. RNA World Modeling: A Comparison of Two Complementary Approaches. Entropy. 2022; 24(4):536. https://doi.org/10.3390/e24040536
Chicago/Turabian StyleSynak, Jaroslaw, Agnieszka Rybarczyk, and Jacek Blazewicz. 2022. "RNA World Modeling: A Comparison of Two Complementary Approaches" Entropy 24, no. 4: 536. https://doi.org/10.3390/e24040536
APA StyleSynak, J., Rybarczyk, A., & Blazewicz, J. (2022). RNA World Modeling: A Comparison of Two Complementary Approaches. Entropy, 24(4), 536. https://doi.org/10.3390/e24040536






