Hotspots of Somatic Genetic Variation in Pituitary Neuroendocrine Tumors
Abstract
Simple Summary
Abstract
1. Introduction
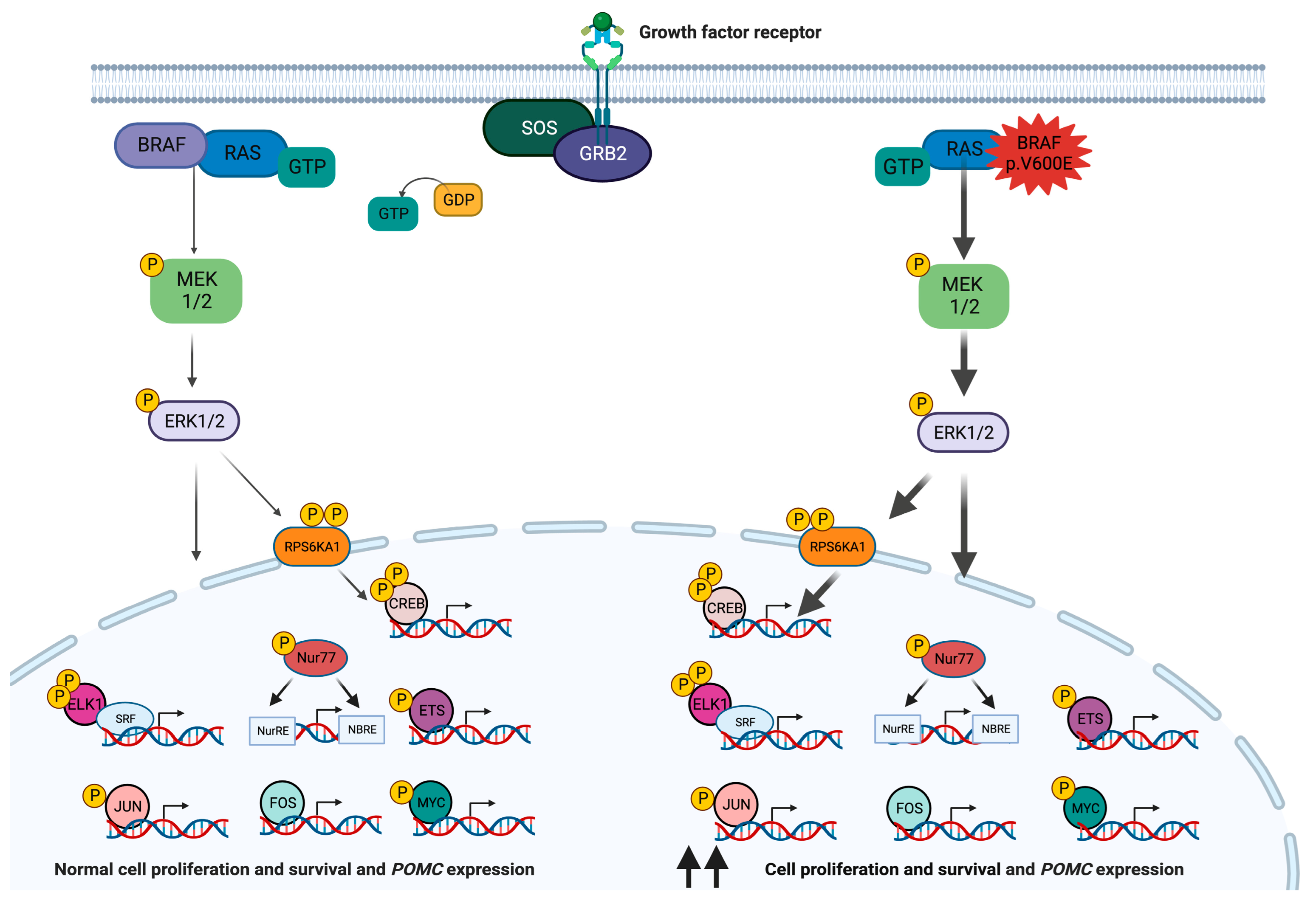
2. BRAF
3. GNAS

4. DICER1

5. SF3B1
6. USP8

7. USP48
8. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
References
- Juul, R.I.; Nielsen, M.M.; Juul, M.; Feuerbach, L.; Pedersen, J.S. The landscape and driver potential of site-specific hotspots across cancer genomes. NPJ Genom. Med. 2021, 6, 33. [Google Scholar] [CrossRef] [PubMed]
- Cooper, D.N.; Mort, M.; Stenson, P.D.; Ball, E.V.; Chuzhanova, N.A. Methylation-mediated deamination of 5-methylcytosine appears to give rise to mutations causing human inherited disease in CpNpG trinucleotides, as well as in CpG dinucleotides. Hum. Genom. 2010, 4, 406–410. [Google Scholar] [CrossRef] [PubMed]
- Marais, G. Biased gene conversion: Implications for genome and sex evolution. Trends Genet. 2003, 19, 330–338. [Google Scholar] [CrossRef] [PubMed]
- Nesta, A.V.; Tafur, D.; Beck, C.R. Hotspots of human mutation. Trends Genet. 2021, 37, 717–729. [Google Scholar] [CrossRef] [PubMed]
- Kato, T.; Kurahashi, H.; Emanuel, B.S. Chromosomal translocations and palindromic AT-rich repeats. Curr. Opin. Genet. Dev. 2012, 22, 221–228. [Google Scholar] [CrossRef] [PubMed]
- ICGC/TCGA Pan-Cancer Analysis of Whole Genomes Consortium. Pan-cancer analysis of whole genomes. Nature 2020, 578, 82–93. [Google Scholar] [CrossRef] [PubMed]
- Martincorena, I.; Campbell, P.J. Somatic mutation in cancer and normal cells. Science 2015, 349, 1483–1489. [Google Scholar] [CrossRef]
- Zhang, F.; Zhang, Q.; Zhu, J.; Yao, B.; Ma, C.; Qiao, N.; He, S.; Ye, Z.; Wang, Y.; Han, R.; et al. Integrated proteogenomic characterization across major histological types of pituitary neuroendocrine tumors. Cell Res. 2022, 32, 1047–1067. [Google Scholar] [CrossRef]
- Villa, C.; Baussart, B.; Assie, G.; Raverot, G.; Roncaroli, F. The World Health Organization classifications of pituitary neuroendocrine tumours: A clinico-pathological appraisal. Endocr. Relat. Cancer 2023, 30, e230021. [Google Scholar] [CrossRef]
- Landis, C.A.; Masters, S.B.; Spada, A.; Pace, A.M.; Bourne, H.R.; Vallar, L. GTPase inhibiting mutations activate the alpha chain of Gs and stimulate adenylyl cyclase in human pituitary tumours. Nature 1989, 340, 692–696. [Google Scholar] [CrossRef]
- Ma, Z.Y.; Song, Z.J.; Chen, J.H.; Wang, Y.F.; Li, S.Q.; Zhou, L.F.; Mao, Y.; Li, Y.M.; Hu, R.G.; Zhang, Z.Y.; et al. Recurrent gain-of-function USP8 mutations in Cushing’s disease. Cell Res. 2015, 25, 306–317. [Google Scholar] [CrossRef] [PubMed]
- Reincke, M.; Sbiera, S.; Hayakawa, A.; Theodoropoulou, M.; Osswald, A.; Beuschlein, F.; Meitinger, T.; Mizuno-Yamasaki, E.; Kawaguchi, K.; Saeki, Y.; et al. Mutations in the deubiquitinase gene USP8 cause Cushing’s disease. Nat. Genet. 2015, 47, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Greenman, C.; Stephens, P.; Smith, R.; Dalgliesh, G.L.; Hunter, C.; Bignell, G.; Davies, H.; Teague, J.; Butler, A.; Stevens, C.; et al. Patterns of somatic mutation in human cancer genomes. Nature 2007, 446, 153–158. [Google Scholar] [CrossRef] [PubMed]
- Bonner, T.I.; Kerby, S.B.; Sutrave, P.; Gunnell, M.A.; Mark, G.; Rapp, U.R. Structure and biological activity of human homologs of the raf/mil oncogene. Mol. Cell Biol. 1985, 5, 1400–1407. [Google Scholar] [CrossRef] [PubMed]
- Huebner, K.; ar-Rushdi, A.; Griffin, C.A.; Isobe, M.; Kozak, C.; Emanuel, B.S.; Nagarajan, L.; Cleveland, J.L.; Bonner, T.I.; Goldsborough, M.D.; et al. Actively transcribed genes in the raf oncogene group, located on the X chromosome in mouse and human. Proc. Natl. Acad. Sci. USA 1986, 83, 3934–3938. [Google Scholar] [CrossRef] [PubMed]
- Ikawa, S.; Fukui, M.; Ueyama, Y.; Tamaoki, N.; Yamamoto, T.; Toyoshima, K. B-raf, a new member of the raf family, is activated by DNA rearrangement. Mol. Cell Biol. 1988, 8, 2651–2654. [Google Scholar] [CrossRef]
- Sithanandam, G.; Druck, T.; Cannizzaro, L.A.; Leuzzi, G.; Huebner, K.; Rapp, U.R. B-raf and a B-raf pseudogene are located on 7q in man. Oncogene 1992, 7, 795–799. [Google Scholar]
- Wellbrock, C.; Karasarides, M.; Marais, R. The RAF proteins take centre stage. Nat. Rev. Mol. Cell Biol. 2004, 5, 875–885. [Google Scholar] [CrossRef]
- Gualtieri, A.; Kyprianou, N.; Gregory, L.C.; Vignola, M.L.; Nicholson, J.G.; Tan, R.; Inoue, S.I.; Scagliotti, V.; Casado, P.; Blackburn, J.; et al. Activating mutations in BRAF disrupt the hypothalamo-pituitary axis leading to hypopituitarism in mice and humans. Nat. Commun. 2021, 12, 2028. [Google Scholar] [CrossRef]
- Hebron, K.E.; Hernandez, E.R.; Yohe, M.E. The RASopathies: From pathogenetics to therapeutics. Dis. Model Mech. 2022, 15, dmm049107. [Google Scholar] [CrossRef]
- Hoshino, R.; Chatani, Y.; Yamori, T.; Tsuruo, T.; Oka, H.; Yoshida, O.; Shimada, Y.; Ari-i, S.; Wada, H.; Fujimoto, J.; et al. Constitutive activation of the 41-/43-kDa mitogen-activated protein kinase signaling pathway in human tumors. Oncogene 1999, 18, 813–822. [Google Scholar] [CrossRef]
- Davies, H.; Bignell, G.R.; Cox, C.; Stephens, P.; Edkins, S.; Clegg, S.; Teague, J.; Woffendin, H.; Garnett, M.J.; Bottomley, W.; et al. Mutations of the BRAF gene in human cancer. Nature 2002, 417, 949–954. [Google Scholar] [CrossRef] [PubMed]
- Garnett, M.J.; Marais, R. Guilty as charged: B-RAF is a human oncogene. Cancer Cell 2004, 6, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Xing, M. BRAF mutation in thyroid cancer. Endocr. Relat. Cancer 2005, 12, 245–262. [Google Scholar] [CrossRef] [PubMed]
- Wan, P.T.; Garnett, M.J.; Roe, S.M.; Lee, S.; Niculescu-Duvaz, D.; Good, V.M.; Jones, C.M.; Marshall, C.J.; Springer, C.J.; Barford, D.; et al. Mechanism of activation of the RAF-ERK signaling pathway by oncogenic mutations of B-RAF. Cell 2004, 116, 855–867. [Google Scholar] [CrossRef] [PubMed]
- Ikenoue, T.; Hikiba, Y.; Kanai, F.; Tanaka, Y.; Imamura, J.; Imamura, T.; Ohta, M.; Ijichi, H.; Tateishi, K.; Kawakami, T.; et al. Functional analysis of mutations within the kinase activation segment of B-Raf in human colorectal tumors. Cancer Res. 2003, 63, 8132–8137. [Google Scholar]
- Wellbrock, C.; Ogilvie, L.; Hedley, D.; Karasarides, M.; Martin, J.; Niculescu-Duvaz, D.; Springer, C.J.; Marais, R. V599EB-RAF is an oncogene in melanocytes. Cancer Res. 2004, 64, 2338–2342. [Google Scholar] [CrossRef]
- Hingorani, S.R.; Jacobetz, M.A.; Robertson, G.P.; Herlyn, M.; Tuveson, D.A. Suppression of BRAF(V599E) in human melanoma abrogates transformation. Cancer Res. 2003, 63, 5198–5202. [Google Scholar]
- Grothey, A.; Fakih, M.; Tabernero, J. Management of BRAF-mutant metastatic colorectal cancer: A review of treatment options and evidence-based guidelines. Ann. Oncol. 2021, 32, 959–967. [Google Scholar] [CrossRef]
- Broccoli, A.; Terragna, C.; Nanni, L.; Martello, M.; Armuzzi, S.; Agostinelli, C.; Morigi, A.; Casadei, B.; Pellegrini, C.; Stefoni, V.; et al. Droplet digital polymerase chain reaction for the assessment of disease burden in hairy cell leukemia. Hematol. Oncol. 2022, 40, 57–62. [Google Scholar] [CrossRef]
- Haston, S.; Pozzi, S.; Carreno, G.; Manshaei, S.; Panousopoulos, L.; Gonzalez-Meljem, J.M.; Apps, J.R.; Virasami, A.; Thavaraj, S.; Gutteridge, A.; et al. MAPK pathway control of stem cell proliferation and differentiation in the embryonic pituitary provides insights into the pathogenesis of papillary craniopharyngioma. Development 2017, 144, 2141–2152. [Google Scholar] [CrossRef] [PubMed]
- Prieto, R.; Pascual, J.M. Craniopharyngiomas with a mixed histological pattern: The missing link to the intriguing pathogenesis of adamantinomatous and squamous-papillary varieties? Neuropathology 2013, 33, 682–686. [Google Scholar] [CrossRef]
- Brastianos, P.K.; Taylor-Weiner, A.; Manley, P.E.; Jones, R.T.; Dias-Santagata, D.; Thorner, A.R.; Lawrence, M.S.; Rodriguez, F.J.; Bernardo, L.A.; Schubert, L.; et al. Exome sequencing identifies BRAF mutations in papillary craniopharyngiomas. Nat. Genet. 2014, 46, 161–165. [Google Scholar] [CrossRef] [PubMed]
- Larkin, S.J.; Preda, V.; Karavitaki, N.; Grossman, A.; Ansorge, O. BRAF V600E mutations are characteristic for papillary craniopharyngioma and may coexist with CTNNB1-mutated adamantinomatous craniopharyngioma. Acta Neuropathol. 2014, 127, 927–929. [Google Scholar] [CrossRef] [PubMed]
- Holsken, A.; Sill, M.; Merkle, J.; Schweizer, L.; Buchfelder, M.; Flitsch, J.; Fahlbusch, R.; Metzler, M.; Kool, M.; Pfister, S.M.; et al. Adamantinomatous and papillary craniopharyngiomas are characterized by distinct epigenomic as well as mutational and transcriptomic profiles. Acta Neuropathol. Commun. 2016, 4, 20. [Google Scholar] [CrossRef] [PubMed]
- Goschzik, T.; Gessi, M.; Dreschmann, V.; Gebhardt, U.; Wang, L.; Yamaguchi, S.; Wheeler, D.A.; Lauriola, L.; Lau, C.C.; Muller, H.L.; et al. Genomic alterations of adamantinomatous and papillary craniopharyngioma. J. Neuropathol. Exp. Neurol. 2017, 76, 126–134. [Google Scholar] [CrossRef]
- Yoshimoto, K.; Hatae, R.; Suzuki, S.O.; Hata, N.; Kuga, D.; Akagi, Y.; Amemiya, T.; Sangatsuda, Y.; Mukae, N.; Mizoguchi, M.; et al. High-resolution melting and immunohistochemical analysis efficiently detects mutually exclusive genetic alterations of adamantinomatous and papillary craniopharyngiomas. Neuropathology 2018, 38, 3–10. [Google Scholar] [CrossRef]
- Omay, S.B.; Chen, Y.N.; Almeida, J.P.; Ruiz-Trevino, A.S.; Boockvar, J.A.; Stieg, P.E.; Greenfield, J.P.; Souweidane, M.M.; Kacker, A.; Pisapia, D.J.; et al. Do craniopharyngioma molecular signatures correlate with clinical characteristics? J. Neurosurg. 2018, 128, 1473–1478. [Google Scholar] [CrossRef]
- La Corte, E.; Younus, I.; Pivari, F.; Selimi, A.; Ottenhausen, M.; Forbes, J.A.; Pisapia, D.J.; Dobri, G.A.; Anand, V.K.; Schwartz, T.H. BRAF V600E mutant papillary craniopharyngiomas: A single-institutional case series. Pituitary 2018, 21, 571–583. [Google Scholar] [CrossRef]
- Chen, J.; Jian, X.; Deng, S.; Ma, Z.; Shou, X.; Shen, Y.; Zhang, Q.; Song, Z.; Li, Z.; Peng, H.; et al. Identification of recurrent USP48 and BRAF mutations in Cushing’s disease. Nat. Commun. 2018, 9, 3171. [Google Scholar] [CrossRef]
- Sbiera, S.; Pérez-Rivas, L.G.; Taranets, L.; Weigand, I.; Flitsch, J.; Graf, E.; Monoranu, C.M.; Saeger, W.; Hagel, C.; Honegger, J.; et al. Driver mutations in USP8 wild-type Cushing’s disease. Neuro Oncol. 2019, 21, 1273–1283. [Google Scholar] [CrossRef] [PubMed]
- Uzilov, A.V.; Taik, P.; Cheesman, K.C.; Javanmard, P.; Ying, K.; Roehnelt, A.; Wang, H.; Fink, M.Y.; Lau, C.Y.; Moe, A.S.; et al. USP8 and TP53 drivers are associated with CNV in a corticotroph adenoma cohort enriched for aggressive tumors. J. Clin. Endocrinol. Metab. 2021, 106, 826–842. [Google Scholar] [CrossRef] [PubMed]
- Abraham, A.P.; Pai, R.; Beno, D.L.; Chacko, G.; Asha, H.S.; Rajaratnam, S.; Kapoor, N.; Thomas, N.; Chacko, A.G. USP8, USP48, and BRAF mutations differ in their genotype-phenotype correlation in Asian Indian patients with Cushing’s disease. Endocrine 2022, 75, 549–559. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ramírez, L.C.; Pankratz, N.; Lane, J.; Faucz, F.R.; Chittiboina, P.; Kay, D.M.; Beethem, Z.; Mills, J.L.; Stratakis, C.A. Genetic drivers of Cushing’s disease: Frequency and associated phenotypes. Genet. Med. 2022, 24, 2516–2525. [Google Scholar] [CrossRef] [PubMed]
- Schmid, S.; Solomon, D.A.; Perez, E.; Thieme, A.; Kleinschmidt-DeMasters, B.K.; Giannini, C.; Reinhardt, A.; Asa, S.L.; Mete, O.; Stichel, D.; et al. Genetic and epigenetic characterization of posterior pituitary tumors. Acta Neuropathol. 2021, 142, 1025–1043. [Google Scholar] [CrossRef] [PubMed]
- Hatzivassiliou, G.; Song, K.; Yen, I.; Brandhuber, B.J.; Anderson, D.J.; Alvarado, R.; Ludlam, M.J.; Stokoe, D.; Gloor, S.L.; Vigers, G.; et al. RAF inhibitors prime wild-type RAF to activate the MAPK pathway and enhance growth. Nature 2010, 464, 431–435. [Google Scholar] [CrossRef]
- Su, F.; Viros, A.; Milagre, C.; Trunzer, K.; Bollag, G.; Spleiss, O.; Reis-Filho, J.S.; Kong, X.; Koya, R.C.; Flaherty, K.T.; et al. RAS mutations in cutaneous squamous-cell carcinomas in patients treated with BRAF inhibitors. N. Engl. J. Med. 2012, 366, 207–215. [Google Scholar] [CrossRef]
- Aziz, S.N.; Proano, L.; Cruz, C.; Tenemaza, M.G.; Monteros, G.; Hassen, G.; Baskar, A.; Argudo, J.M.; Duenas, J.B.; Fabara, S.P. Vemurafenib in the Treatment of Erdheim Chester Disease: A Systematic Review. Cureus 2022, 14, e25935. [Google Scholar] [CrossRef]
- Maitre, E.; Paillassa, J.; Troussard, X. Novel targeted treatments in hairy cell leukemia and other hairy cell-like disorders. Front. Oncol. 2022, 12, 1068981. [Google Scholar] [CrossRef]
- Fernandez, M.F.; Choi, J.; Sosman, J. New approaches to targeted therapy in melanoma. Cancers 2023, 15, 3224. [Google Scholar] [CrossRef]
- Guaitoli, G.; Zullo, L.; Tiseo, M.; Dankner, M.; Rose, A.A.; Facchinetti, F. Non-small-cell lung cancer: How to manage. Drugs Context 2023, 12, 2022-11-3. [Google Scholar] [CrossRef] [PubMed]
- Hamidi, S.; Hofmann, M.C.; Iyer, P.C.; Cabanillas, M.E.; Hu, M.I.; Busaidy, N.L.; Dadu, R. Review article: New treatments for advanced differentiated thyroid cancers and potential mechanisms of drug resistance. Front. Endocrinol. 2023, 14, 1176731. [Google Scholar] [CrossRef] [PubMed]
- Juratli, T.A.; Jones, P.S.; Wang, N.; Subramanian, M.; Aylwin, S.J.B.; Odia, Y.; Rostami, E.; Gudjonsson, O.; Shaw, B.L.; Cahill, D.P.; et al. Targeted treatment of papillary craniopharyngiomas harboring BRAF V600E mutations. Cancer 2019, 125, 2910–2914. [Google Scholar] [CrossRef] [PubMed]
- Brastianos, P.K.; Twohy, E.; Geyer, S.; Gerstner, E.R.; Kaufmann, T.J.; Tabrizi, S.; Kabat, B.; Thierauf, J.; Ruff, M.W.; Bota, D.A.; et al. BRAF-MEK inhibition in newly diagnosed papillary craniopharyngiomas. N. Engl. J. Med. 2023, 389, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Grenier-Chartrand, F.; Barrit, S.; Racu, M.L.; Luce, S.; Spitaels, J.; Sadeghi-Meibodi, N.; Lebrun, L.; Salmon, I.; Lefranc, F.; De Witte, O. Dabrafenib monotherapy for a recurrent BRAFV600E-mutated TTF-1-positive posterior pituitary tumor. Acta. Neurochir. 2022, 164, 737–742. [Google Scholar] [CrossRef] [PubMed]
- Sollfrank, L.; Lettmaier, S.; Erdmann, M.; Uslu, U. Panniculitis under successful targeted inhibition of the MAPK/ERK signaling pathway in a patient with BRAF V600E-mutated spindle cell oncocytoma of the pituitary gland. Anticancer Res. 2019, 39, 3955–3959. [Google Scholar] [CrossRef] [PubMed]
- Dawoud, F.M.; Naylor, R.M.; Giannini, C.; Swanson, A.A.; Meyer, F.B.; Uhm, J.H. TTF-1 positive posterior pituitary tumor: Limitations of current treatment and potential new hope in BRAF V600E mutation variants. Clin. Neurol. Neurosurg. 2020, 196, 106059. [Google Scholar] [CrossRef]
- Chang, F.; Steelman, L.S.; Lee, J.T.; Shelton, J.G.; Navolanic, P.M.; Blalock, W.L.; Franklin, R.A.; McCubrey, J.A. Signal transduction mediated by the Ras/Raf/MEK/ERK pathway from cytokine receptors to transcription factors: Potential targeting for therapeutic intervention. Leukemia 2003, 17, 1263–1293. [Google Scholar] [CrossRef]
- Derwich, A.; Sykutera, M.; Brominska, B.; Rubis, B.; Ruchala, M.; Sawicka-Gutaj, N. The role of activation of PI3K/AKT/mTOR and RAF/MEK/ERK pathways in aggressive pituitary adenomas-new potential therapeutic approach—A systematic review. Int. J. Mol. Sci. 2023, 24, 10952. [Google Scholar] [CrossRef]
- Ishida, M.; Moore, G.E. The role of imprinted genes in humans. Mol. Aspects Med. 2013, 34, 826–840. [Google Scholar] [CrossRef]
- Hayward, B.E.; Moran, V.; Strain, L.; Bonthron, D.T. Bidirectional imprinting of a single gene: GNAS1 encodes maternally, paternally, and biallelically derived proteins. Proc. Natl. Acad. Sci. USA 1998, 95, 15475–15480. [Google Scholar] [CrossRef] [PubMed]
- Hayward, B.E.; Bonthron, D.T. An imprinted antisense transcript at the human GNAS1 locus. Hum. Mol. Genet 2000, 9, 835–841. [Google Scholar] [CrossRef] [PubMed]
- The UniProt Consortium. UniProt: The universal protein knowledgebase in 2023. Nucleic. Acids Res. 2023, 51, D523–D531. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, G.; Ballare, E.; Giammona, E.; Beck-Peccoz, P.; Spada, A. The gsalpha gene: Predominant maternal origin of transcription in human thyroid gland and gonads. J. Clin. Endocrinol. Metab. 2002, 87, 4736–4740. [Google Scholar] [CrossRef]
- Syrovatkina, V.; Alegre, K.O.; Dey, R.; Huang, X.Y. Regulation, signaling, and physiological functions of G-proteins. J. Mol. Biol. 2016, 428, 3850–3868. [Google Scholar] [CrossRef]
- Peverelli, E.; Mantovani, G.; Lania, A.G.; Spada, A. cAMP in the pituitary: An old messenger for multiple signals. J. Mol. Endocrinol. 2014, 52, R67–R77. [Google Scholar] [CrossRef]
- Duc, N.M.; Kim, H.R.; Chung, K.Y. Structural mechanism of G protein activation by G protein-coupled receptor. Eur. J. Pharmacol. 2015, 763, 214–222. [Google Scholar] [CrossRef]
- Lambright, D.G.; Noel, J.P.; Hamm, H.E.; Sigler, P.B. Structural determinants for activation of the alpha-subunit of a heterotrimeric G protein. Nature 1994, 369, 621–628. [Google Scholar] [CrossRef]
- Pierce, K.L.; Premont, R.T.; Lefkowitz, R.J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell. Biol. 2002, 3, 639–650. [Google Scholar] [CrossRef]
- Hernandez-Ramirez, L.C.; Trivellin, G.; Stratakis, C.A. Cyclic 3’,5’-adenosine monophosphate (cAMP) signaling in the anterior pituitary gland in health and disease. Mol. Cell Endocrinol. 2018, 463, 72–86. [Google Scholar] [CrossRef]
- Landis, C.A.; Harsh, G.; Lyons, J.; Davis, R.L.; McCormick, F.; Bourne, H.R. Clinical characteristics of acromegalic patients whose pituitary tumors contain mutant Gs protein. J. Clin. Endocrinol. Metab. 1990, 71, 1416–1420. [Google Scholar] [CrossRef] [PubMed]
- Lyons, J.; Landis, C.A.; Harsh, G.; Vallar, L.; Grunewald, K.; Feichtinger, H.; Duh, Q.Y.; Clark, O.H.; Kawasaki, E.; Bourne, H.R. Two G protein oncogenes in human endocrine tumors. Science 1990, 249, 655–659. [Google Scholar] [CrossRef] [PubMed]
- Suarez, H.G.; du Villard, J.A.; Caillou, B.; Schlumberger, M.; Parmentier, C.; Monier, R. gsp mutations in human thyroid tumours. Oncogene 1991, 6, 677–679. [Google Scholar] [PubMed]
- Spada, A.; Vallar, L. G-protein oncogenes in acromegaly. Horm. Res. 1992, 38, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Hosoi, E.; Yokogoshi, Y.; Hosoi, E.; Horie, H.; Sano, T.; Yamada, S.; Saito, S. Analysis of the Gs alpha gene in growth hormone-secreting pituitary adenomas by the polymerase chain reaction-direct sequencing method using paraffin-embedded tissues. Acta Endocrinol. 1993, 129, 301–306. [Google Scholar] [CrossRef]
- Parma, J.; Duprez, L.; Van Sande, J.; Cochaux, P.; Gervy, C.; Mockel, J.; Dumont, J.; Vassart, G. Somatic mutations in the thyrotropin receptor gene cause hyperfunctioning thyroid adenomas. Nature 1993, 365, 649–651. [Google Scholar] [CrossRef]
- Tordjman, K.; Stern, N.; Ouaknine, G.; Yossiphov, Y.; Razon, N.; Nordenskjold, M.; Friedman, E. Activating mutations of the Gs alpha-gene in nonfunctioning pituitary tumors. J. Clin. Endocrinol. Metab. 1993, 77, 765–769. [Google Scholar] [CrossRef]
- Yoshimoto, K.; Iwahana, H.; Fukuda, A.; Sano, T.; Itakura, M. Rare mutations of the Gs alpha subunit gene in human endocrine tumors. Mutation detection by polymerase chain reaction-primer-introduced restriction analysis. Cancer 1993, 72, 1386–1393. [Google Scholar] [CrossRef]
- Michiels, F.M.; Caillou, B.; Talbot, M.; Dessarps-Freichey, F.; Maunoury, M.T.; Schlumberger, M.; Mercken, L.; Monier, R.; Feunteun, J. Oncogenic potential of guanine nucleotide stimulatory factor alpha subunit in thyroid glands of transgenic mice. Proc. Natl. Acad. Sci. USA 1994, 91, 10488–10492. [Google Scholar] [CrossRef]
- Williamson, E.A.; Ince, P.G.; Harrison, D.; Kendall-Taylor, P.; Harris, P.E. G-protein mutations in human pituitary adrenocorticotrophic hormone-secreting adenomas. Eur. J. Clin. Invest 1995, 25, 128–131. [Google Scholar] [CrossRef]
- Williamson, E.A.; Johnson, S.J.; Foster, S.; Kendall-Taylor, P.; Harris, P.E. G protein gene mutations in patients with multiple endocrinopathies. J. Clin. Endocrinol. Metab. 1995, 80, 1702–1705. [Google Scholar] [CrossRef] [PubMed]
- Yang, I.M.; Woo, J.T.; Kim, S.W.; Kim, J.W.; Kim, Y.S.; Choi, Y.K. Characteristics of acromegalic patients with a good response to octreotide, a somatostatin analogue. Clin. Endocrinol. 1995, 42, 295–301. [Google Scholar] [CrossRef] [PubMed]
- Fragoso, M.C.B.V.; Latronico, C.; Carvalho, F.M.; Zerbini, M.C.N.; Marcondes, J.A.M.; Araujo, L.M.B.; Lando, V.S.; Frazzatto, E.T.; Mendonca, B.B.; Villares, S.M.F. Activating mutation of the stimulatory G protein (gsp) as a putative cause of ovarian and testicular human stromal Leydig cell tumors. J. Clin. Endocr. Metab. 1998, 83, 2074–2078. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Shi, Y.; Tang, D.; Deng, J.; Su, C. Detection of gsp oncogene in growth hormone-secreting pituitary adenomas and the study of clinical characteristics of acromegalic patients with gsp-positive pituitary tumors. Chin. Med. J. 1998, 111, 891–894. [Google Scholar] [PubMed]
- Buchfelder, M.; Fahlbusch, R.; Merz, T.; Symowski, H.; Adams, E.F. Clinical correlates in acromegalic patients with pituitary tumors expressing GSP oncogenes. Pituitary 1999, 1, 181–185. [Google Scholar] [CrossRef] [PubMed]
- Persani, L.; Lania, A.; Alberti, L.; Romoli, R.; Mantovani, G.; Filetti, S.; Spada, A.; Conti, M. Induction of specific phosphodiesterase isoforms by constitutive activation of the cAMP pathway in autonomous thyroid adenomas. J. Clin. Endocrinol. Metab. 2000, 85, 2872–2878. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Krohn, K.; Paschke, R. Somatic mutations in thyroid nodular disease. Mol. Genet. Metab. 2002, 75, 202–208. [Google Scholar] [CrossRef]
- Mendoza, V.; Sosa, E.; Espinosa-de-los-Monteros, A.L.; Salcedo, M.; Guinto, G.; Cheng, S.; Sandoval, C.; Mercado, M. GSPalpha mutations in Mexican patients with acromegaly: Potential impact on long term prognosis. Growth Horm. IGF. Res. 2005, 15, 28–32. [Google Scholar] [CrossRef]
- Freda, P.U.; Chung, W.K.; Matsuoka, N.; Walsh, J.E.; Kanibir, M.N.; Kleinman, G.; Wang, Y.; Bruce, J.N.; Post, K.D. Analysis of GNAS mutations in 60 growth hormone secreting pituitary tumors: Correlation with clinical and pathological characteristics and surgical outcome based on highly sensitive GH and IGF-I criteria for remission. Pituitary 2007, 10, 275–282. [Google Scholar] [CrossRef]
- Taboada, G.F.; Tabet, A.L.; Naves, L.A.; de Carvalho, D.P.; Gadelha, M.R. Prevalence of gsp oncogene in somatotropinomas and clinically non-functioning pituitary adenomas: Our experience. Pituitary 2009, 12, 165–169. [Google Scholar] [CrossRef]
- Lu, J.Y.; Hung, P.J.; Chen, P.L.; Yen, R.F.; Kuo, K.T.; Yang, T.L.; Wang, C.Y.; Chang, T.C.; Huang, T.S.; Chang, C.C. Follicular thyroid carcinoma with NRAS Q61K and GNAS R201H mutations that had a good (131)I treatment response. Endocrinol. Diabetes Metab. Case Rep. 2016, 2016, 150067. [Google Scholar] [CrossRef] [PubMed]
- Ronchi, C.L.; Peverelli, E.; Herterich, S.; Weigand, I.; Mantovani, G.; Schwarzmayr, T.; Sbiera, S.; Allolio, B.; Honegger, J.; Appenzeller, S.; et al. Landscape of somatic mutations in sporadic GH-secreting pituitary adenomas. Eur. J. Endocrinol. 2016, 174, 363–372. [Google Scholar] [CrossRef] [PubMed]
- Ritvonen, E.; Pitkanen, E.; Karppinen, A.; Vehkavaara, S.; Demir, H.; Paetau, A.; Schalin-Jantti, C.; Karhu, A. Impact of AIP and inhibitory G protein alpha 2 proteins on clinical features of sporadic GH-secreting pituitary adenomas. Eur. J. Endocrinol. 2017, 176, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Puig-Domingo, M.; Gil, J.; Sampedro-Nunez, M.; Jorda, M.; Webb, S.M.; Serra, G.; Pons, L.; Salinas, I.; Blanco, A.; Marques-Pamies, M.; et al. Molecular profiling for acromegaly treatment: A validation study. Endocr. Relat. Cancer 2020, 27, 375–389. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, A.; Jin, D.X.; De Marco, L.; Laitman, Y.; Friedman, E. Activating genomic alterations in the Gs alpha gene (GNAS) in 274 694 tumors. Genes Chromosomes Cancer 2020, 59, 503–516. [Google Scholar] [CrossRef] [PubMed]
- Jung, H.; Kim, K.; Kim, D.; Moon, J.H.; Kim, E.H.; Kim, S.H.; Ku, C.R.; Lee, E.J. Associations of GNAS mutations with surgical outcomes in patients with growth hormone-secreting pituitary adenoma. Endocrinol. Metab. 2021, 36, 342–350. [Google Scholar] [CrossRef] [PubMed]
- Romanet, P.; Galluso, J.; Kamenicky, P.; Hage, M.; Theodoropoulou, M.; Roche, C.; Graillon, T.; Etchevers, H.C.; De Murat, D.; Mougel, G.; et al. Somatotroph tumors and the epigenetic status of the GNAS locus. Int. J. Mol. Sci. 2021, 22, 7570. [Google Scholar] [CrossRef] [PubMed]
- Yamato, A.; Nagano, H.; Gao, Y.; Matsuda, T.; Hashimoto, N.; Nakayama, A.; Yamagata, K.; Yokoyama, M.; Gong, Y.; Shi, X.; et al. Proteogenomic landscape and clinical characterization of GH-producing pituitary adenomas/somatotroph pituitary neuroendocrine tumors. Commun. Biol. 2022, 5, 1304. [Google Scholar] [CrossRef]
- O’Hayre, M.; Vazquez-Prado, J.; Kufareva, I.; Stawiski, E.W.; Handel, T.M.; Seshagiri, S.; Gutkind, J.S. The emerging mutational landscape of G proteins and G-protein-coupled receptors in cancer. Nat. Rev. Cancer 2013, 13, 412–424. [Google Scholar] [CrossRef]
- Turan, S.; Bastepe, M. GNAS Spectrum of Disorders. Curr. Osteoporos. Rep. 2015, 13, 146–158. [Google Scholar] [CrossRef]
- Ritterhouse, L.L.; Vivero, M.; Mino-Kenudson, M.; Sholl, L.M.; Iafrate, A.J.; Nardi, V.; Dong, F. GNAS mutations in primary mucinous and non-mucinous lung adenocarcinomas. Mod. Pathol. 2017, 30, 1720–1727. [Google Scholar] [CrossRef] [PubMed]
- Faias, S.; Duarte, M.; Pereira, L.; Chaves, P.; Cravo, M.; Dias Pereira, A.; Albuquerque, C. Methylation changes at the GNAS imprinted locus in pancreatic cystic neoplasms are important for the diagnosis of malignant cysts. World J. Gastrointest. Oncol. 2020, 12, 1056–1064. [Google Scholar] [CrossRef] [PubMed]
- McCune, D.J. Osteitis fibrosa cystica: The case of a nine year old girl who also exhibits precocious puberty, multiple pigmentation of the skin and hyperthyroidism. Am. J. Dis. Child 1936, 52, 743–744. [Google Scholar]
- Weinstein, L.S.; Shenker, A.; Gejman, P.V.; Merino, M.J.; Friedman, E.; Spiegel, A.M. Activating mutations of the stimulatory G protein in the McCune-Albright syndrome. N. Engl. J. Med. 1991, 325, 1688–1695. [Google Scholar] [CrossRef] [PubMed]
- Boyce, A.M.; Florenzano, P.; de Castro, L.F.; Collins, M.T. Fibrous dysplasia/McCune-Albright syndrome. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2019. [Google Scholar]
- Lumbroso, S.; Paris, F.; Sultan, C. Activating Gsalpha mutations: Analysis of 113 patients with signs of McCune-Albright syndrome—A European Collaborative Study. J. Clin. Endocrinol. Metab. 2004, 89, 2107–2113. [Google Scholar] [CrossRef]
- Idowu, B.D.; Al-Adnani, M.; O’Donnell, P.; Yu, L.; Odell, E.; Diss, T.; Gale, R.E.; Flanagan, A.M. A sensitive mutation-specific screening technique for GNAS1 mutations in cases of fibrous dysplasia: The first report of a codon 227 mutation in bone. Histopathology 2007, 50, 691–704. [Google Scholar] [CrossRef] [PubMed]
- Spencer, T.; Pan, K.S.; Collins, M.T.; Boyce, A.M. The clinical spectrum of McCune-Albright syndrome and its management. Horm. Res. Paediatr. 2019, 92, 347–356. [Google Scholar] [CrossRef]
- Boyce, A.M.; Collins, M.T. Fibrous dysplasia/McCune-Albright syndrome: A rare, mosaic disease of Galpha s activation. Endocr. Rev. 2020, 41, 345–370. [Google Scholar] [CrossRef]
- Spada, A.; Vallar, L.; Faglia, G. G-proteins and hormonal signalling in human pituitary tumors: Genetic mutations and functional alterations. Front. Neuroendocrinol. 1993, 14, 214–232. [Google Scholar] [CrossRef]
- Weinstein, L.S. The stimulatory G protein alpha-subunit gene: Mutations and imprinting lead to complex phenotypes. J. Clin. Endocrinol. Metab. 2001, 86, 4622–4626. [Google Scholar] [CrossRef]
- Adams, E.F.; Lei, T.; Buchfelder, M.; Petersen, B.; Fahlbusch, R. Biochemical characteristics of human pituitary somatotropinomas with and without gsp mutations: In vitro cell culture studies. J. Clin. Endocrinol. Metab. 1995, 80, 2077–2081. [Google Scholar] [CrossRef] [PubMed]
- Rymuza, J.; Kober, P.; Rusetska, N.; Mossakowska, B.J.; Maksymowicz, M.; Nyc, A.; Baluszek, S.; Zielinski, G.; Kunicki, J.; Bujko, M. Transcriptomic classification of pituitary neuroendocrine tumors causing acromegaly. Cells 2022, 11, 3846. [Google Scholar] [CrossRef] [PubMed]
- Spada, A.; Arosio, M.; Bochicchio, D.; Bazzoni, N.; Vallar, L.; Bassetti, M.; Faglia, G. Clinical, biochemical, and morphological correlates in patients bearing growth hormone-secreting pituitary-tumors with or without constitutively active adenylyl cyclase. J. Clin. Endocr. Metab. 1990, 71, 1421–1426. [Google Scholar] [CrossRef] [PubMed]
- Larkin, S.; Reddy, R.; Karavitaki, N.; Cudlip, S.; Wass, J.; Ansorge, O. Granulation pattern, but not GSP or GHR mutation, is associated with clinical characteristics in somatostatin-naive patients with somatotroph adenomas. Eur. J. Endocrinol. 2013, 168, 491–499. [Google Scholar] [CrossRef] [PubMed]
- Lasolle, H.; Elsensohn, M.H.; Wierinckx, A.; Alix, E.; Bonnefille, C.; Vasiljevic, A.; Cortet, C.; Decoudier, B.; Sturm, N.; Gaillard, S.; et al. Chromosomal instability in the prediction of pituitary neuroendocrine tumors prognosis. Acta Neuropathol. Commun. 2020, 8, 190. [Google Scholar] [CrossRef] [PubMed]
- Neou, M.; Villa, C.; Armignacco, R.; Jouinot, A.; Raffin-Sanson, M.L.; Septier, A.; Letourneur, F.; Diry, S.; Diedisheim, M.; Izac, B.; et al. Pangenomic classification of pituitary neuroendocrine tumors. Cancer Cell 2020, 37, 123–134.e125. [Google Scholar] [CrossRef] [PubMed]
- Mantovani, G.; Bondioni, S.; Lania, A.G.; Corbetta, S.; de Sanctis, L.; Cappa, M.; Di Battista, E.; Chanson, P.; Beck-Peccoz, P.; Spada, A. Parental origin of Gsalpha mutations in the McCune-Albright syndrome and in isolated endocrine tumors. J. Clin. Endocrinol. Metab. 2004, 89, 3007–3009. [Google Scholar] [CrossRef][Green Version]
- Hayward, B.E.; Barlier, A.; Korbonits, M.; Grossman, A.B.; Jacquet, P.; Enjalbert, A.; Bonthron, D.T. Imprinting of the G(s)alpha gene GNAS1 in the pathogenesis of acromegaly. J. Clin. Investig. 2001, 107, R31–R36. [Google Scholar] [CrossRef]
- Picard, C.; Silvy, M.; Gerard, C.; Buffat, C.; Lavaque, E.; Figarella-Branger, D.; Dufour, H.; Gabert, J.; Beckers, A.; Brue, T.; et al. Gs alpha overexpression and loss of Gs alpha imprinting in human somatotroph adenomas: Association with tumor size and response to pharmacologic treatment. Int. J. Cancer 2007, 121, 1245–1252. [Google Scholar] [CrossRef]
- Altarejos, J.Y.; Montminy, M. CREB and the CRTC co-activators: Sensors for hormonal and metabolic signals. Nat. Rev. Mol. Cell Biol. 2011, 12, 141–151. [Google Scholar] [CrossRef]
- Schultz, K.A.P.; Stewart, D.R.; Kamihara, J.; Bauer, A.J.; Merideth, M.A.; Stratton, P.; Huryn, L.A.; Harris, A.K.; Doros, L.; Field, A.; et al. DICER1 tumor predisposition. In GeneReviews®; Adam, M.P., Mirzaa, G.M., Pagon, R.A., Wallace, S.E., Bean, L.J.H., Gripp, K.W., Amemiya, A., Eds.; University of Washington: Seattle, WA, USA, 2020. [Google Scholar]
- Hill, D.A.; Ivanovich, J.; Priest, J.R.; Gurnett, C.A.; Dehner, L.P.; Desruisseau, D.; Jarzembowski, J.A.; Wikenheiser-Brokamp, K.A.; Suarez, B.K.; Whelan, A.J.; et al. DICER1 mutations in familial pleuropulmonary blastoma. Science 2009, 325, 965. [Google Scholar] [CrossRef] [PubMed]
- Slade, I.; Bacchelli, C.; Davies, H.; Murray, A.; Abbaszadeh, F.; Hanks, S.; Barfoot, R.; Burke, A.; Chisholm, J.; Hewitt, M.; et al. DICER1 syndrome: Clarifying the diagnosis, clinical features and management implications of a pleiotropic tumour predisposition syndrome. J. Med. Genet. 2011, 48, 273–278. [Google Scholar] [CrossRef] [PubMed]
- de Kock, L.; Wang, Y.C.; Revil, T.; Badescu, D.; Rivera, B.; Sabbaghian, N.; Wu, M.; Weber, E.; Sandoval, C.; Hopman, S.M.; et al. High-sensitivity sequencing reveals multi-organ somatic mosaicism causing DICER1 syndrome. J. Med. Genet. 2016, 53, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Schultz, K.A.P.; Williams, G.M.; Kamihara, J.; Stewart, D.R.; Harris, A.K.; Bauer, A.J.; Turner, J.; Shah, R.; Schneider, K.; Schneider, K.W.; et al. DICER1 and associated conditions: Identification of at-risk individuals and recommended surveillance strategies. Clin. Cancer Res. 2018, 24, 2251–2261. [Google Scholar] [CrossRef] [PubMed]
- Klein, S.; Lee, H.; Ghahremani, S.; Kempert, P.; Ischander, M.; Teitell, M.A.; Nelson, S.F.; Martinez-Agosto, J.A. Expanding the phenotype of mutations in DICER1: Mosaic missense mutations in the RNase IIIb domain of DICER1 cause GLOW syndrome. J. Med. Genet. 2014, 51, 294–302. [Google Scholar] [CrossRef] [PubMed]
- Brenneman, M.; Field, A.; Yang, J.; Williams, G.; Doros, L.; Rossi, C.; Schultz, K.A.; Rosenberg, A.; Ivanovich, J.; Turner, J.; et al. Temporal order of RNase IIIb and loss-of-function mutations during development determines phenotype in pleuropulmonary blastoma/DICER1 syndrome: A unique variant of the two-hit tumor suppression model. F1000Res 2015, 4, 214. [Google Scholar] [CrossRef] [PubMed]
- MacRae, I.J.; Zhou, K.; Li, F.; Repic, A.; Brooks, A.N.; Cande, W.Z.; Adams, P.D.; Doudna, J.A. Structural basis for double-stranded RNA processing by Dicer. Science 2006, 311, 195–198. [Google Scholar] [CrossRef]
- Bernstein, E.; Caudy, A.A.; Hammond, S.M.; Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001, 409, 363–366. [Google Scholar] [CrossRef]
- Foulkes, W.D.; Priest, J.R.; Duchaine, T.F. DICER1: Mutations, microRNAs and mechanisms. Nat. Rev. Cancer 2014, 14, 662–672. [Google Scholar] [CrossRef]
- Chendrimada, T.P.; Gregory, R.I.; Kumaraswamy, E.; Norman, J.; Cooch, N.; Nishikura, K.; Shiekhattar, R. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 2005, 436, 740–744. [Google Scholar] [CrossRef]
- Svoboda, P. Key mechanistic principles and considerations concerning RNA interference. Front. Plant Sci. 2020, 11, 1237. [Google Scholar] [CrossRef] [PubMed]
- de Boer, C.M.; Eini, R.; Gillis, A.M.; Stoop, H.; Looijenga, L.H.; White, S.J. DICER1 RNase IIIb domain mutations are infrequent in testicular germ cell tumours. BMC. Res. Notes 2012, 5, 569. [Google Scholar] [CrossRef] [PubMed]
- Heravi-Moussavi, A.; Anglesio, M.S.; Cheng, S.W.; Senz, J.; Yang, W.; Prentice, L.; Fejes, A.P.; Chow, C.; Tone, A.; Kalloger, S.E.; et al. Recurrent somatic DICER1 mutations in nonepithelial ovarian cancers. N. Engl. J. Med. 2012, 366, 234–242. [Google Scholar] [CrossRef] [PubMed]
- de Kock, L.; Plourde, F.; Carter, M.T.; Hamel, N.; Srivastava, A.; Meyn, M.S.; Arseneau, J.; Bouron-Dal, S.D.; Foulkes, W.D. Germ-line and somatic DICER1 mutations in a pleuropulmonary blastoma. Pediatr. Blood Cancer 2013, 60, 2091–2092. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.K.; Sabbaghian, N.; Xu, B.; Addidou-Kalucki, S.; Bernard, C.; Zou, D.; Reeve, A.E.; Eccles, M.R.; Cole, C.; Choong, C.S.; et al. Biallelic DICER1 mutations occur in Wilms tumours. J. Pathol. 2013, 230, 154–164. [Google Scholar] [CrossRef] [PubMed]
- Tomiak, E.; de, K.L.; Grynspan, D.; Ramphal, R.; Foulkes, W.D. DICER1 mutations in an adolescent with cervical embryonal rhabdomyosarcoma (cERMS). Pediatr. Blood Cancer 2014, 61, 568–569. [Google Scholar] [CrossRef] [PubMed]
- de Kock, L.; Sabbaghian, N.; Soglio, D.B.D.; Guillerman, R.P.; Park, B.K.; Chami, R.; Deal, C.L.; Priest, J.R.; Foulkes, W.D. Exploring the association between DICER1 mutations and differentiated thyroid carcinoma. J. Clin. Endocr. Metab. 2014, 99, E1072–E1077. [Google Scholar] [CrossRef]
- de Kock, L.; Sabbaghian, N.; Druker, H.; Weber, E.; Hamel, N.; Miller, S.; Choong, C.S.; Gottardo, N.G.; Kees, U.R.; Rednam, S.P.; et al. Germ-line and somatic DICER1 mutations in pineoblastoma. Acta. Neuropathol. 2014, 128, 583–595. [Google Scholar] [CrossRef]
- de Kock, L.; Sabbaghian, N.; Plourde, F.; Srivastava, A.; Weber, E.; Bouron-Dal Soglio, D.; Hamel, N.; Choi, J.H.; Park, S.H.; Deal, C.L.; et al. Pituitary blastoma: A pathognomonic feature of germ-line DICER1 mutations. Acta. Neuropathol. 2014, 128, 111–122. [Google Scholar] [CrossRef]
- Doros, L.A.; Rossi, C.T.; Yang, J.; Field, A.; Williams, G.M.; Messinger, Y.; Cajaiba, M.M.; Perlman, E.J.; Schultz, A.; Cathro, H.P.; et al. DICER1 mutations in childhood cystic nephroma and its relationship to DICER1-renal sarcoma. Mod. Pathol. 2014, 27, 1267–1280. [Google Scholar] [CrossRef]
- Murray, M.J.; Bailey, S.; Raby, K.L.; Saini, H.K.; de, K.L.; Burke, G.A.; Foulkes, W.D.; Enright, A.J.; Coleman, N.; Tischkowitz, M. Serum levels of mature microRNAs in DICER1-mutated pleuropulmonary blastoma. Oncogenesis 2014, 3, e87. [Google Scholar] [CrossRef] [PubMed]
- Pugh, T.J.; Yu, W.; Yang, J.; Field, A.L.; Ambrogio, L.; Carter, S.L.; Cibulskis, K.; Giannikopoulos, P.; Kiezun, A.; Kim, J.; et al. Exome sequencing of pleuropulmonary blastoma reveals frequent biallelic loss of TP53 and two hits in DICER1 resulting in retention of 5p-derived miRNA hairpin loop sequences. Oncogene 2014, 33, 5295–5302. [Google Scholar] [CrossRef] [PubMed]
- Anglesio, M.S.; Wang, Y.; Yang, W.; Senz, J.; Wan, A.; Heravi-Moussavi, A.; Salamanca, C.; Maines-Bandiera, S.; Huntsman, D.G.; Morin, G.B. Cancer-associated somatic DICER1 hotspot mutations cause defective miRNA processing and reverse-strand expression bias to predominantly mature 3p strands through loss of 5p strand cleavage. J. Pathol. 2013, 229, 400–409. [Google Scholar] [CrossRef]
- Chong, A.S.; Han, H.; Albrecht, S.; Weon, Y.C.; Park, S.K.; Foulkes, W.D. DICER1 syndrome in a young adult with pituitary blastoma. Acta. Neuropathol. 2021, 142, 1071–1076. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.P.Y.; Kelsey, M.M.; Sabbaghian, N.; Park, S.H.; Deal, C.L.; Esbenshade, A.J.; Ploner, O.; Peet, A.; Traunecker, H.; Ahmed, Y.H.E.; et al. Clinical outcomes and complications of pituitary blastoma. J. Clin. Endocrinol. Metab. 2021, 106, 351–363. [Google Scholar] [CrossRef] [PubMed]
- Liu, A.P.; Li, K.K.; Chow, C.; Chan, S.; Leung, A.W.; Shing, M.M.; To, K.F.; Chan, D.T.; Chan, G.C.; Ng, H.K. Expanding the clinical and molecular spectrum of pituitary blastoma. Acta. Neuropathol. 2022, 143, 415–417. [Google Scholar] [CrossRef] [PubMed]
- Scheithauer, B.W.; Kovacs, K.; Horvath, E.; Kim, D.S.; Osamura, R.Y.; Ketterling, R.P.; Lloyd, R.V.; Kim, O.L. Pituitary blastoma. Acta. Neuropathol. 2008, 116, 657–666. [Google Scholar] [CrossRef]
- Nadaf, J.; de Kock, L.; Chong, A.S.; Korbonits, M.; Thorner, P.; Benlimame, N.; Fu, L.; Peet, A.; Warner, J.; Ploner, O.; et al. Molecular characterization of DICER1-mutated pituitary blastoma. Acta. Neuropathol. 2021, 141, 929–944. [Google Scholar] [CrossRef]
- Thorner, P.S.; Chong, A.S.; Nadaf, J.; Benlimame, N.; Marrano, P.; Chami, R.; Fu, L.; Foulkes, W.D. PRAME protein expression in DICER1-related tumours. J. Pathol. Clin. Res. 2022, 8, 579–581. [Google Scholar] [CrossRef]
- Xu, Y.; Zou, R.; Wang, J.; Wang, Z.W.; Zhu, X. The role of the cancer testis antigen PRAME in tumorigenesis and immunotherapy in human cancer. Cell Prolif. 2020, 53, e12770. [Google Scholar] [CrossRef]
- Asada, K.; Canestrari, E.; Fu, X.; Li, Z.; Makowski, E.; Wu, Y.C.; Mito, J.K.; Kirsch, D.G.; Baraban, J.; Paroo, Z. Rescuing dicer defects via inhibition of an anti-dicing nuclease. Cell Rep. 2014, 9, 1471–1481. [Google Scholar] [CrossRef] [PubMed]
- Asada, K.; Canestrari, E.; Paroo, Z. A druggable target for rescuing microRNA defects. Bioorg Med. Chem. Lett. 2016, 26, 4942–4946. [Google Scholar] [CrossRef] [PubMed]
- Muller, M.; Fazi, F.; Ciaudo, C. Argonaute proteins: From structure to function in development and pathological cell fate determination. Front Cell Dev. Biol. 2019, 7, 360. [Google Scholar] [CrossRef] [PubMed]
- Thunders, M.; Delahunt, B. Gene of the month: DICER1: Ruler and controller. J. Clin. Pathol. 2021, 74, 69–72. [Google Scholar] [CrossRef]
- Meiklejohn, K.M.; Darbinyan, A.; Barbieri, A.L. A review of DICER1: Structure, function and contribution to disease. Diagnostic Histopathology 2022, 28, 329–336. [Google Scholar] [CrossRef]
- Cretu, C.; Schmitzova, J.; Ponce-Salvatierra, A.; Dybkov, O.; De Laurentiis, E.I.; Sharma, K.; Will, C.L.; Urlaub, H.; Luhrmann, R.; Pena, V. Molecular architecture of SF3b and structural consequences of its cancer-related mutations. Mol. Cell 2016, 64, 307–319. [Google Scholar] [CrossRef]
- Alsafadi, S.; Houy, A.; Battistella, A.; Popova, T.; Wassef, M.; Henry, E.; Tirode, F.; Constantinou, A.; Piperno-Neumann, S.; Roman-Roman, S.; et al. Cancer-associated SF3B1 mutations affect alternative splicing by promoting alternative branchpoint usage. Nat. Commun. 2016, 7, 10615. [Google Scholar] [CrossRef]
- Darman, R.B.; Seiler, M.; Agrawal, A.A.; Lim, K.H.; Peng, S.; Aird, D.; Bailey, S.L.; Bhavsar, E.B.; Chan, B.; Colla, S.; et al. Cancer-associated SF3B1 hotspot mutations induce cryptic 3’ splice site selection through use of a different branch point. Cell Rep. 2015, 13, 1033–1045. [Google Scholar] [CrossRef]
- DeBoever, C.; Ghia, E.M.; Shepard, P.J.; Rassenti, L.; Barrett, C.L.; Jepsen, K.; Jamieson, C.H.; Carson, D.; Kipps, T.J.; Frazer, K.A. Transcriptome sequencing reveals potential mechanism of cryptic 3’ splice site selection in SF3B1-mutated cancers. PLoS Comput. Biol. 2015, 11, e1004105. [Google Scholar] [CrossRef]
- Will, C.L.; Lührmann, R. Spliceosome structure and function. Cold Spring Harb. Perspect. Biol. 2011, 3, a003707. [Google Scholar] [CrossRef]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Z.; Gong, Q.; Wang, Y.; Li, M.; Wang, L.; Ding, H.; Li, P. The biological function and clinical significance of SF3B1 mutations in cancer. Biomark Res. 2020, 8, 38. [Google Scholar] [CrossRef]
- Li, C.; Xie, W.; Rosenblum, J.S.; Zhou, J.; Guo, J.; Miao, Y.; Shen, Y.; Wang, H.; Gong, L.; Li, M.; et al. Somatic SF3B1 hotspot mutation in prolactinomas. Nat. Commun. 2020, 11, 2506. [Google Scholar] [CrossRef] [PubMed]
- Simon, J.; Perez-Rivas, L.G.; Zhao, Y.; Chasseloup, F.; Lasolle, H.; Cortet, C.; Descotes, F.; Villa, C.; Baussart, B.; Burman, P.; et al. Prevalence and clinical correlations of SF3B1 variants in lactotroph tumours. Eur. J. Endocrinol. 2023, 189, 372–378. [Google Scholar] [CrossRef] [PubMed]
- Cass, D.M.; Berglund, J.A. The SF3b155 N-terminal domain is a scaffold important for splicing. Biochemistry 2006, 45, 10092–10101. [Google Scholar] [CrossRef] [PubMed]
- Papaemmanuil, E.; Cazzola, M.; Boultwood, J.; Malcovati, L.; Vyas, P.; Bowen, D.; Pellagatti, A.; Wainscoat, J.S.; Hellstrom-Lindberg, E.; Gambacorti-Passerini, C.; et al. Somatic SF3B1 mutation in myelodysplasia with ring sideroblasts. N. Engl. J. Med. 2011, 365, 1384–1395. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Lawrence, M.S.; Wan, Y.; Stojanov, P.; Sougnez, C.; Stevenson, K.; Werner, L.; Sivachenko, A.; DeLuca, D.S.; Zhang, L.; et al. SF3B1 and other novel cancer genes in chronic lymphocytic leukemia. N. Engl. J. Med. 2011, 365, 2497–2506. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, K.; Sanada, M.; Shiraishi, Y.; Nowak, D.; Nagata, Y.; Yamamoto, R.; Sato, Y.; Sato-Otsubo, A.; Kon, A.; Nagasaki, M.; et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 2011, 478, 64–69. [Google Scholar] [CrossRef]
- Biankin, A.V.; Waddell, N.; Kassahn, K.S.; Gingras, M.C.; Muthuswamy, L.B.; Johns, A.L.; Miller, D.K.; Wilson, P.J.; Patch, A.M.; Wu, J.; et al. Pancreatic cancer genomes reveal aberrations in axon guidance pathway genes. Nature 2012, 491, 399–405. [Google Scholar] [CrossRef]
- Stephens, P.J.; Tarpey, P.S.; Davies, H.; Van Loo, P.; Greenman, C.; Wedge, D.C.; Nik-Zainal, S.; Martin, S.; Varela, I.; Bignell, G.R.; et al. The landscape of cancer genes and mutational processes in breast cancer. Nature 2012, 486, 400–404. [Google Scholar] [CrossRef]
- Harbour, J.W.; Roberson, E.D.; Anbunathan, H.; Onken, M.D.; Worley, L.A.; Bowcock, A.M. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat. Genet. 2013, 45, 133–135. [Google Scholar] [CrossRef] [PubMed]
- Kong, Y.; Krauthammer, M.; Halaban, R. Rare SF3B1 R625 mutations in cutaneous melanoma. Melanoma Res. 2014, 24, 332–334. [Google Scholar] [CrossRef]
- Yang, H.M.; Hsiao, S.J.; Schaeffer, D.F.; Lai, C.; Remotti, H.E.; Horst, D.; Mansukhani, M.M.; Horst, B.A. Identification of recurrent mutational events in anorectal melanoma. Mod. Pathol. 2017, 30, 286–296. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Li, C.; Fang, Q.; Liu, Y.; Wang, D.; Chen, Y.; Xie, W.; Zhang, Y. The SF3B1(R625H) mutation promotes prolactinoma tumor progression through aberrant splicing of DLG1. J. Exp. Clin. Cancer Res. 2022, 41, 26. [Google Scholar] [CrossRef] [PubMed]
- Vazquez-Borrego, M.C.; Fuentes-Fayos, A.C.; Venegas-Moreno, E.; Rivero-Cortes, E.; Dios, E.; Moreno-Moreno, P.; Madrazo-Atutxa, A.; Remon, P.; Solivera, J.; Wildemberg, L.E.; et al. Splicing machinery is dysregulated in pituitary neuroendocrine tumors and is associated with aggressiveness features. Cancers 2019, 11, 1439. [Google Scholar] [CrossRef]
- Zhang, L.; Zhang, X.; Zhang, H.; Liu, F.; Bi, Y.; Zhang, Y.; Cheng, C.; Liu, J. Knockdown of SF3B1 inhibits cell proliferation, invasion and migration triggering apoptosis in breast cancer via aberrant splicing. Breast Cancer 2020, 27, 464–476. [Google Scholar] [CrossRef] [PubMed]
- Effenberger, K.A.; Urabe, V.K.; Jurica, M.S. Modulating splicing with small molecular inhibitors of the spliceosome. Wiley Interdiscip. Rev. RNA 2017, 8, e1381. [Google Scholar] [CrossRef]
- Zhang, D.; Meng, F. A comprehensive overview of structure-activity relationships of small-molecule splicing modulators targeting SF3B1 as anticancer agents. ChemMedChem 2020, 15, 2098–2120. [Google Scholar] [CrossRef]
- Steensma, D.P.; Wermke, M.; Klimek, V.M.; Greenberg, P.L.; Font, P.; Komrokji, R.S.; Yang, J.; Brunner, A.M.; Carraway, H.E.; Ades, L.; et al. Phase I first-in-human dose escalation study of the oral SF3B1 modulator H3B-8800 in myeloid neoplasms. Leukemia 2021, 35, 3542–3550. [Google Scholar] [CrossRef]
- Pérez-Rivas, L.G.; Theodoropoulou, M.; Ferrau, F.; Nusser, C.; Kawaguchi, K.; Stratakis, C.A.; Faucz, F.R.; Wildemberg, L.E.; Assie, G.; Beschorner, R.; et al. The gene of the ubiquitin-specific protease 8 is frequently mutated in adenomas causing Cushing’s disease. J. Clin. Endocrinol. Metab. 2015, 100, E997–E1004. [Google Scholar] [CrossRef]
- Hayashi, K.; Inoshita, N.; Kawaguchi, K.; Ibrahim Ardisasmita, A.; Suzuki, H.; Fukuhara, N.; Okada, M.; Nishioka, H.; Takeuchi, Y.; Komada, M.; et al. The USP8 mutational status may predict drug susceptibility in corticotroph adenomas of Cushing’s disease. Eur. J. Endocrinol. 2016, 174, 213–226. [Google Scholar] [CrossRef] [PubMed]
- de Araujo, L.J.; Lerario, A.M.; de Castro, M.; Martins, C.S.; Bronstein, M.D.; Machado, M.C.; Trarbach, E.B.; Villares Fragoso, M.C. Transcriptome Analysis Showed a Differential Signature between Invasive and Non-invasive Corticotrophinomas. Front. Endocrinol. 2017, 8, 55. [Google Scholar] [CrossRef] [PubMed]
- Faucz, F.R.; Tirosh, A.; Tatsi, C.; Berthon, A.; Hernández-Ramírez, L.C.; Settas, N.; Angelousi, A.; Correa, R.; Papadakis, G.Z.; Chittiboina, P.; et al. Somatic USP8 gene mutations are a common cause of pediatric Cushing disease. J. Clin. Endocrinol. Metab. 2017, 102, 2836–2843. [Google Scholar] [CrossRef] [PubMed]
- Albani, A.; Pérez-Rivas, L.G.; Dimopoulou, C.; Zopp, S.; Colon-Bolea, P.; Roeber, S.; Honegger, J.; Flitsch, J.; Rachinger, W.; Buchfelder, M.; et al. The USP8 mutational status may predict long-term remission in patients with Cushing’s disease. Clin. Endocrinol. 2018, 89, 454–458. [Google Scholar] [CrossRef]
- Ballmann, C.; Thiel, A.; Korah, H.E.; Reis, A.C.; Saeger, W.; Stepanow, S.; Kohrer, K.; Reifenberger, G.; Knobbe-Thomsen, C.B.; Knappe, U.J.; et al. USP8 mutations in pituitary Cushing adenomas-targeted analysis by next-generation sequencing. J. Endocr. Soc. 2018, 2, 266–278. [Google Scholar] [CrossRef] [PubMed]
- Bujko, M.; Kober, P.; Boresowicz, J.; Rusetska, N.; Paziewska, A.; Dabrowska, M.; Piascik, A.; Pekul, M.; Zielinski, G.; Kunicki, J.; et al. USP8 mutations in corticotroph adenomas determine a distinct gene expression profile irrespective of functional tumour status. Eur. J. Endocrinol. 2019, 181, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Losa, M.; Mortini, P.; Pagnano, A.; Detomas, M.; Cassarino, M.F.; Pecori Giraldi, F. Clinical characteristics and surgical outcome in USP8-mutated human adrenocorticotropic hormone-secreting pituitary adenomas. Endocrine 2019, 63, 240–246. [Google Scholar] [CrossRef] [PubMed]
- Weigand, I.; Knobloch, L.; Flitsch, J.; Saeger, W.; Monoranu, C.M.; Hofner, K.; Herterich, S.; Rotermund, R.; Ronchi, C.L.; Buchfelder, M.; et al. Impact of USP8 gene mutations on protein deregulation in Cushing disease. J. Clin. Endocrinol. Metab. 2019, 104, 2535–2546. [Google Scholar] [CrossRef]
- Castellnou, S.; Vasiljevic, A.; Lapras, V.; Raverot, V.; Alix, E.; Borson-Chazot, F.; Jouanneau, E.; Raverot, G.; Lasolle, H. SST5 expression and USP8 mutation in functioning and silent corticotroph pituitary tumors. Endocr. Connect. 2020, 9, 243–253. [Google Scholar] [CrossRef]
- Martins, C.S.; Camargo, R.C.; Coeli-Lacchini, F.B.; Saggioro, F.P.; Moreira, A.C.; de Castro, M. USP8 mutations and cell cycle regulation in corticotroph adenomas. Horm. Metab. Res. 2020, 52, 117–123. [Google Scholar] [CrossRef]
- Sesta, A.; Cassarino, M.F.; Terreni, M.; Ambrogio, A.G.; Libera, L.; Bardelli, D.; Lasio, G.; Losa, M.; Pecori Giraldi, F. Ubiquitin-specific protease 8 mutant corticotrope adenomas present unique secretory and molecular features and shed light on the role of ubiquitylation on ACTH processing. Neuroendocrinology 2020, 110, 119–129. [Google Scholar] [CrossRef] [PubMed]
- Pasternak-Pietrzak, K.; Faucz, F.R.; Stratakis, C.A.; Moszczynska, E.; Roszkowski, M.; Grajkowska, W.; Pronicki, M.; Szalecki, M. Is there a common cause for paediatric Cushing’s disease? Endokrynol. Pol. 2021, 72, 104–107. [Google Scholar] [CrossRef] [PubMed]
- Treppiedi, D.; Barbieri, A.M.; Di Muro, G.; Marra, G.; Mangili, F.; Catalano, R.; Esposito, E.; Ferrante, E.; Serban, A.L.; Locatelli, M.; et al. Genetic Profiling of a Cohort of Italian Patients with ACTH-Secreting Pituitary Tumors and Characterization of a Novel USP8 Gene Variant. Cancers 2021, 13, 4022. [Google Scholar] [CrossRef] [PubMed]
- Rebollar-Vega, R.G.; Zuarth-Vázquez, J.M.; Hernández-Ramírez, L.C. Clinical spectrum of USP8 pathogenic variants in Cushing’s disease. Arch. Med. Res. 2023, 102899. [Google Scholar] [CrossRef] [PubMed]
- Clague, M.J.; Urbe, S.; Komander, D. Breaking the chains: Deubiquitylating enzyme specificity begets function. Nat. Rev. Mol. Cell Biol. 2019, 20, 338–352. [Google Scholar] [CrossRef] [PubMed]
- Mizuno, E.; Iura, T.; Mukai, A.; Yoshimori, T.; Kitamura, N.; Komada, M. Regulation of epidermal growth factor receptor down-regulation by UBPY-mediated deubiquitination at endosomes. Mol. Biol. Cell 2005, 16, 5163–5174. [Google Scholar] [CrossRef]
- Alwan, H.A.; van Leeuwen, J.E. UBPY-mediated epidermal growth factor receptor (EGFR) de-ubiquitination promotes EGFR degradation. J. Biol. Chem. 2007, 282, 1658–1669. [Google Scholar] [CrossRef]
- Berlin, I.; Schwartz, H.; Nash, P.D. Regulation of epidermal growth factor receptor ubiquitination and trafficking by the USP8.STAM complex. J. Biol. Chem. 2010, 285, 34909–34921. [Google Scholar] [CrossRef]
- Crespo-Yàñez, X.; Aguilar-Gurrieri, C.; Jacomin, A.C.; Journet, A.; Mortier, M.; Taillebourg, E.; Soleilhac, E.; Weissenhorn, W.; Fauvarque, M.O. CHMP1B is a target of USP8/UBPY regulated by ubiquitin during endocytosis. PLoS Genet. 2018, 14, e1007456. [Google Scholar] [CrossRef]
- Theodoropoulou, M.; Reincke, M.; Fassnacht, M.; Komada, M. Decoding the genetic basis of Cushing’s disease: USP8 in the spotlight. Eur. J. Endocrinol. 2015, 173, M73–M83. [Google Scholar] [CrossRef]
- Islam, M.T.; Chen, F.; Chen, H. The oncogenic role of ubiquitin specific peptidase (USP8) and its signaling pathways targeting for cancer therapeutics. Arch. Biochem. Biophys. 2021, 701, 108811. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Rivas, L.G.; Theodoropoulou, M.; Puar, T.H.; Fazel, J.; Stieg, M.R.; Ferrau, F.; Assie, G.; Gadelha, M.R.; Deutschbein, T.; Fragoso, M.C.; et al. Somatic USP8 mutations are frequent events in corticotroph tumor progression causing Nelson’s tumor. Eur. J. Endocrinol. 2018, 178, 59–65. [Google Scholar] [CrossRef] [PubMed]
- Treppiedi, D.; Marra, G.; Di Muro, G.; Esposito, E.; Barbieri, A.M.; Catalano, R.; Mangili, F.; Bravi, F.; Locatelli, M.; Lania, A.G.; et al. P720R USP8 Mutation Is Associated with a Better Responsiveness to Pasireotide in ACTH-Secreting PitNETs. Cancers 2022, 14, 2455. [Google Scholar] [CrossRef]
- Jian, F.F.; Li, Y.F.; Chen, Y.F.; Jiang, H.; Chen, X.; Zheng, L.L.; Zhao, Y.; Wang, W.Q.; Ning, G.; Bian, L.G.; et al. Inhibition of Ubiquitin-specific Peptidase 8 Suppresses Adrenocorticotropic Hormone Production and Tumorous Corticotroph Cell Growth in AtT20 Cells. Chin. Med. J. 2016, 129, 2102–2108. [Google Scholar] [CrossRef] [PubMed]
- Shichi, H.; Fukuoka, H.; Kanzawa, M.; Yamamoto, M.; Yamamoto, N.; Suzuki, M.; Urai, S.; Matsumoto, R.; Kanie, K.; Fujita, Y.; et al. Responsiveness to DDAVP in Cushing’s disease is associated with USP8 mutations through enhancing AVPR1B promoter activity. Pituitary 2022, 25, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Cohen, M.; Persky, R.; Stegemann, R.; Hernández-Ramírez, L.C.; Zeltser, D.; Lodish, M.B.; Chen, A.; Keil, M.F.; Tatsi, C.; Faucz, F.R.; et al. Germline USP8 mutation associated with pediatric Cushing disease and other clinical features: A new syndrome. J. Clin. Endocrinol. Metab. 2019, 104, 4676–4682. [Google Scholar] [CrossRef]
- Fukuoka, H.; Cooper, O.; Ben-Shlomo, A.; Mamelak, A.; Ren, S.G.; Bruyette, D.; Melmed, S. EGFR as a therapeutic target for human, canine, and mouse ACTH-secreting pituitary adenomas. J. Clin. Investig. 2011, 121, 4712–4721. [Google Scholar] [CrossRef]
- Kageyama, K.; Asari, Y.; Sugimoto, Y.; Niioka, K.; Daimon, M. Ubiquitin-specific protease 8 inhibitor suppresses adrenocorticotropic hormone production and corticotroph tumor cell proliferation. Endocr. J. 2020, 67, 177–184. [Google Scholar] [CrossRef]
- Treppiedi, D.; Di Muro, G.; Marra, G.; Barbieri, A.M.; Mangili, F.; Catalano, R.; Serban, A.; Ferrante, E.; Locatelli, M.; Lania, A.G.; et al. USP8 inhibitor RA-9 reduces ACTH release and cell growth in tumor corticotrophs. Endocr. Relat. Cancer 2021, 28, 573–582. [Google Scholar] [CrossRef]
- Clague, M.J.; Urbe, S. Ubiquitin: Same molecule, different degradation pathways. Cell 2010, 143, 682–685. [Google Scholar] [CrossRef]
- Mossakowska, B.J.; Rusetska, N.; Konopinski, R.; Kober, P.; Maksymowicz, M.; Pekul, M.; Zielinski, G.; Styk, A.; Kunicki, J.; Bujko, M. The expression of cell cycle-related genes in USP8-mutated corticotroph neuroendocrine pituitary tumors and their possible role in cell cycle-targeting treatment. Cancers 2022, 14, 5594. [Google Scholar] [CrossRef] [PubMed]
- Xia, R.; Jia, H.; Fan, J.; Liu, Y.; Jia, J. USP8 promotes smoothened signaling by preventing its ubiquitination and changing its subcellular localization. PLoS Biol. 2012, 10, e1001238. [Google Scholar] [CrossRef] [PubMed]
- Colombo, M.; Vallese, S.; Peretto, I.; Jacq, X.; Rain, J.C.; Colland, F.; Guedat, P. Synthesis and biological evaluation of 9-oxo-9H-indeno[1,2-b]pyrazine-2,3-dicarbonitrile analogues as potential inhibitors of deubiquitinating enzymes. ChemMedChem 2010, 5, 552–558. [Google Scholar] [CrossRef] [PubMed]
| Gene | Hotspot Defect | Effect on Protein | PitNET Subtypes Affected | Clinical Phenotype | Pharmacological Agents Targeting the Hotspot |
|---|---|---|---|---|---|
| BRAF | Somatic p.V600E | Destabilization of the inactive conformation of BRAF, promoting its activation | Corticotropinomas | Unclear (very infrequent genetic defect in PitNETs) | Vemurafenib, dabrafenib, and encorafenib, as monotherapy or combined with MEK inhibitors (in clinical use for other tumor types) |
| GNAS | Somatic missense variants affecting residues 201 or 227 | Stabilization of Gsα in its active conformation and inhibition of its GTPase activity | Somatotropinomas. Less frequently: NF-PitNETs and corticotropinomas | Older patients. Controversial: low GH or IGF1, small tumors with slow growth rate, densely granulated somatotropinomas | None reported |
| DICER1 | Germline and (or?) * somatic variants affecting residues 1705, 1709, 1809, 1810, or 1813 | Loss of RNase IIIb activity | PitBs | CD in infants, neonates, or, less frequently, in childhood and young adulthood. Rarely: silent corticotropinomas. Large, aggressive, and poorly differentiated tumors with an oncofetal signature. Presentation as isolated tumors or as part of DICER1 syndrome | Not reported. Indirect approach: pharmacological inhibition of the endonuclease complex TSN-TSNAX (not available for clinical use) |
| SF3B1 | Somatic p.R625H and p.R625C | Impaired interaction with the BP and with other U2 snRNP components | Prolactinomas | High serum prolactin, large tumors with increased Ki67 index, increased progression-free survival and mortality, and requirement for multiple treatments. Occasionally: metastatic PitNETs | H3B-8800 and pladienolide B (not available for clinical use) |
| USP8 | Somatic variants affecting residues 718–720 | Impaired interaction with 14-3-3, proteolytic cleavage, and enhanced deubiquitinase activity | Corticotropinomas | CD in young adults and teenagers, most frequently women, increased SST5 and MGMT, rarely Crooke’s cell adenomas, not associated with Nelson’s syndrome. Controversial: effects on tumor size, hormone secretion, response to dynamic tests, and remission/recurrence rates | DUBs-IN-2 and RA-9 (not available for clinical use). Indirect: gefitinib (in clinical use for other tumor types, under research for corticotropinomas) |
| USP48 | Somatic p.M415I or p.M415V | Enhanced deubiquitinase activity | Corticotropinomas | Controversial: small tumors in one study, high rate of cavernous sinus invasion in a different study. Requires further evaluation | Potential, not yet tested: DUBs-IN (not available for clinical use) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Torres-Morán, M.; Franco-Álvarez, A.L.; Rebollar-Vega, R.G.; Hernández-Ramírez, L.C. Hotspots of Somatic Genetic Variation in Pituitary Neuroendocrine Tumors. Cancers 2023, 15, 5685. https://doi.org/10.3390/cancers15235685
Torres-Morán M, Franco-Álvarez AL, Rebollar-Vega RG, Hernández-Ramírez LC. Hotspots of Somatic Genetic Variation in Pituitary Neuroendocrine Tumors. Cancers. 2023; 15(23):5685. https://doi.org/10.3390/cancers15235685
Chicago/Turabian StyleTorres-Morán, Mariana, Alexa L. Franco-Álvarez, Rosa G. Rebollar-Vega, and Laura C. Hernández-Ramírez. 2023. "Hotspots of Somatic Genetic Variation in Pituitary Neuroendocrine Tumors" Cancers 15, no. 23: 5685. https://doi.org/10.3390/cancers15235685
APA StyleTorres-Morán, M., Franco-Álvarez, A. L., Rebollar-Vega, R. G., & Hernández-Ramírez, L. C. (2023). Hotspots of Somatic Genetic Variation in Pituitary Neuroendocrine Tumors. Cancers, 15(23), 5685. https://doi.org/10.3390/cancers15235685








