- Article
Whole-Genome Sequencing in Premature Coronary Artery Disease in South Asians: A Pilot Case–Control Study
- Iftikhar Ali Ch,
- Azhar Chaudhry and
- Khurram Nasir
- + 11 authors
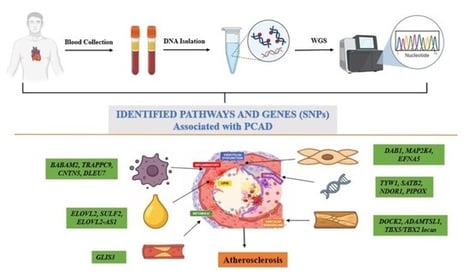
Background/Objectives: Coronary artery disease (CAD) remains the leading cause of mortality worldwide, with South Asia bearing a disproportionately high and rising burden, particularly at younger ages. The present study aimed to investigate genetic variants associated with premature coronary artery disease (PCAD) using whole-genome sequencing (WGS). Methods: WGS was conducted on 12 people (five PCAD cases, seven matched controls) to assess feasibility and methodology for future large-scale research. High-quality genomic DNA was sequenced at a minimum read depth of 10× with a quality threshold of Q30. Variant calling with stringent quality control identified single-nucleotide polymorphisms (SNPs), followed by annotation against gnomAD for allele frequencies and ClinVar for pathogenicity. Protein-coding variants were filtered, and candidate genes were prioritized for comparative analysis between cases and controls. Results: An average of over 8.8 million SNPs per individual was identified, with comparable overall variant distributions between cases and controls. Initial analyses revealed 120 SNPs exclusively present in PCAD cases. All protein-coding variants were rare (allele frequency < 0.0001), and none were previously classified as pathogenic in ClinVar. After filtration, 87 candidate genes were prioritized. Enriched or unique variants in PCAD cases are mapped to genes involved in lipid metabolism, endothelial dysfunction, inflammatory signaling, immune regulation, thrombosis, vascular remodeling, and metabolic processes. Additional variants were identified in genes related to smooth muscle proliferation, oxidative stress, and other biological pathways. Conclusions: This WGS pilot study provides an initial overview of the genomic landscape of PCAD in a South Asian cohort, highlighting rare variants across multiple biological pathways implicated in atherosclerosis that need validation in a large-scale study.
29 April 2026