Fourier Transform Infrared Spectroscopy-Based Detection of Amoxicillin and Ampicillin for Advancing Antibiotic Monitoring with Optical Techniques †
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Preparation
2.2. FTIR Spectra Acquisition and Exploration
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ATB | Antibiotic |
| ATR | Attenuated Total Reflection |
| FTIR | Fourier Transform Infrared Spectroscopy |
| PCA | Principal Component Analysis |
| LoD | Limit of Detection |
| AMX | Amoxicillin |
| AMP | Ampicillin |
References
- Chiumia, F.K.; Sinjani Muula, A.; Chimimba, F.; Nyirongo, H.M.; Kampira, E.; Khuluza, F. Substandard Antibiotics and Their Clinical Outcomes among Hospitalized Patients in Southern Malawi: A Pilot Study. Front. Pharmacol. 2025, 16, 1535501. [Google Scholar] [CrossRef] [PubMed]
- ‘t Hoen, E.F.M.; Hogerzeil, H.V.; Quick, J.D.; Sillo, H.B. A Quiet Revolution in Global Public Health: The World Health Organization’s Prequalification of Medicines Programme. J. Public Health Pol. 2014, 35, 137–161. [Google Scholar] [CrossRef] [PubMed]
- Medicines|WHO—Prequalification of Medical Products (IVDs, Medicines, Vaccines and Immunization Devices, Vector Control). Available online: https://extranet.who.int/prequal/medicines (accessed on 29 May 2025).
- World Health Organization. Quality Assurance of Pharmaceuticals: A Compendium of Guidelines and Related Materials: Vol. 2: Good Manufacturing Practices and Inspection. In Good Manufacturing Practices and Inspection; World Health Organization: Geneva, Switzerland, 2007. [Google Scholar]
- Dyar, O.J.; Huttner, B.; Schouten, J.; Pulcini, C. What Is Antimicrobial Stewardship? Clin. Microbiol. Infect. 2017, 23, 793–798. [Google Scholar] [CrossRef] [PubMed]
- Charani, E.; Holmes, A. Antibiotic Stewardship—Twenty Years in the Making. Antibiotics 2019, 8, 7. [Google Scholar] [CrossRef] [PubMed]
- CDC Core Elements of Hospital Antibiotic Stewardship Programs. Available online: https://www.cdc.gov/antibiotic-use/hcp/core-elements/hospital.html (accessed on 29 May 2025).
- Conte, D.; Palmeiro, J.K.; Bavaroski, A.A.; Rodrigues, L.S.; Cardozo, D.; Tomaz, A.P.; Camargo, J.O.; Dalla-Costa, L.M. Antimicrobial Resistance in Aeromonas Species Isolated from Aquatic Environments in Brazil. J. Appl. Microbiol. 2021, 131, 169–181. [Google Scholar] [CrossRef] [PubMed]
- Maranni, A.C. Identificação de Bactérias Utilizando Espectroscopia Raman ou FTIR e Análise Multivariada: Uma Revisão Sistemática; UFMS: Pioneiros, Brazil, 2023. [Google Scholar]
- Cielecka-Piontek, J.; Paczkowska, M.; Lewandowska, K.; Barszcz, B.; Zalewski, P.; Garbacki, P. Solid-State Stability Study of Meropenem—Solutions Based on Spectrophotometric Analysis. Chem. Cent. J. 2013, 7, 98. [Google Scholar] [CrossRef] [PubMed]
- Ataka, K.; Heberle, J. Biochemical Applications of Surface-Enhanced Infrared Absorption Spectroscopy. Anal. Bioanal. Chem. 2007, 388, 47–54. [Google Scholar] [CrossRef] [PubMed]
- Fanelli, S.; Zimmermann, A.; Totóli, E.G.; Salgado, H.R.N. FTIR Spectrophotometry as a Green Tool for Quantitative Analysis of Drugs: Practical Application to Amoxicillin. J. Chem. 2018, 2018, 3920810. [Google Scholar] [CrossRef]
- Hassall, J.; Coxon, C.; Patel, V.C.; Goldenberg, S.D.; Sergaki, C. Limitations of Current Techniques in Clinical Antimicrobial Resistance Diagnosis: Examples and Future Prospects. Npj Antimicrob. Resist. 2024, 2, 16. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Li, B.; Liu, M.; Mao, S. Demand, Status, and Prospect of Antibiotics Detection in the Environment. Sens. Actuators B Chem. 2022, 369, 132383. [Google Scholar] [CrossRef]
- Ribeiro da Cunha, B.; Fonseca, L.P.; Calado, C.R.C. Metabolic Fingerprinting with Fourier-Transform Infrared (FTIR) Spectroscopy: Towards a High-Throughput Screening Assay for Antibiotic Discovery and Mechanism-of-Action Elucidation. Metabolites 2020, 10, 145. [Google Scholar] [CrossRef] [PubMed]
- Verma, K.; Akhtar, M.; Anchliya, A. Combination of FTIR Spectroscopy and Chemometric Method on Quantitative Approach—A Review. Austin J. Anal. Pharm. Chem. 2021, 8, 1128. [Google Scholar]
- Eid, S.M.; Soliman, S.S.; Elghobashy, M.R.; Abdalla, O.M. ATR-FTIR Coupled with Chemometrics for Quantification of Vildagliptin and Metformin in Pharmaceutical Combinations Having Diverged Concentration Ranges. Vib. Spectrosc. 2020, 106, 102995. [Google Scholar] [CrossRef]
- Nägele, E.; Moritz, R. Structure Elucidation of Degradation Products of the Antibiotic Amoxicillin with Ion Trap MSn and Accurate Mass Determination by ESI TOF. J. Am. Soc. Mass Spectrom. 2005, 16, 1670–1676. [Google Scholar] [CrossRef] [PubMed]
- Currie, L.A. Nomenclature in Evaluation of Analytical Methods Including Detection and Quantification Capabilities (IUPAC Recommendations 1995). Pure Appl. Chem. 1995, 67, 1699–1723. [Google Scholar] [CrossRef]
- Baraldi, C.; Tinti, A.; Ottani, S.; Gamberini, M.C. Characterization of Polymorphic Ampicillin Forms. J. Pharm. Biomed. Anal. 2014, 100, 329–340. [Google Scholar] [CrossRef] [PubMed]
- Bebu, A.; Szabó, L.; Leopold, N.; Berindean, C.; David, L. IR, Raman, SERS and DFT Study of Amoxicillin. J. Mol. Struct. 2011, 993, 52–56. [Google Scholar] [CrossRef]
- Müller, A.L.H.; Flores, É.M.M.; Müller, E.I.; Silva, F.E.B.; Ferrão, M.F. Attenuated Total Reflectance with Fourier Transform Infrared Spectroscopy (ATR/FTIR) and Different PLS Algorithms for Simultaneous Determination of Clavulanic Acid and Amoxicillin in Powder Pharmaceutical Formulation. J. Braz. Chem. Soc. 2011, 22, 1903–1912. [Google Scholar] [CrossRef]
- Nairi, V.; Medda, L.; Monduzzi, M.; Salis, A. Adsorption and Release of Ampicillin Antibiotic from Ordered Mesoporous Silica. J. Colloid Interface Sci. 2017, 497, 217–225. [Google Scholar] [CrossRef] [PubMed]
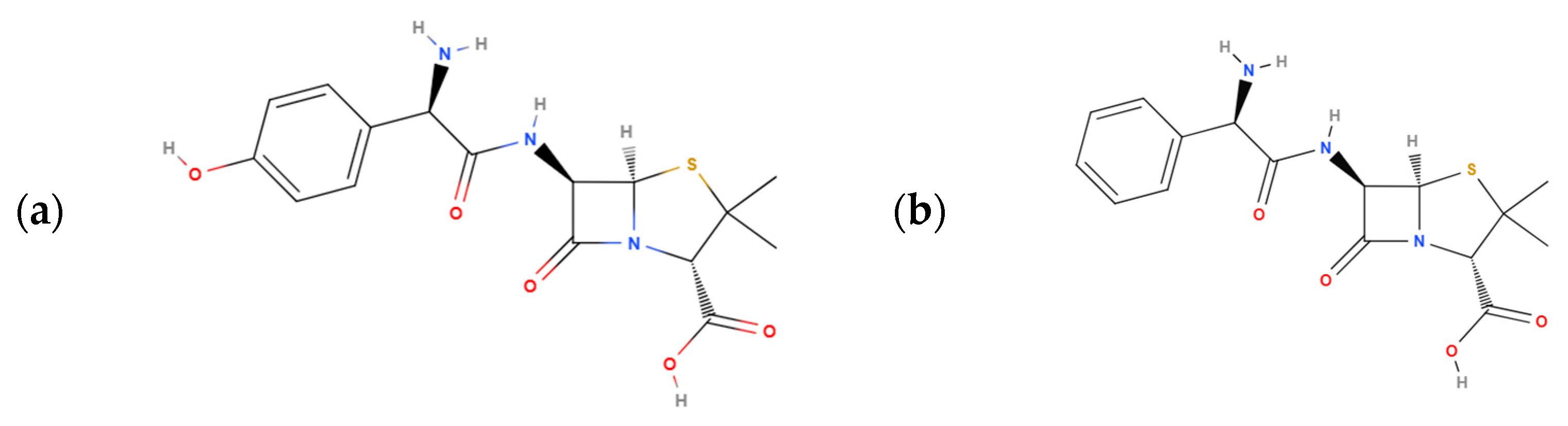

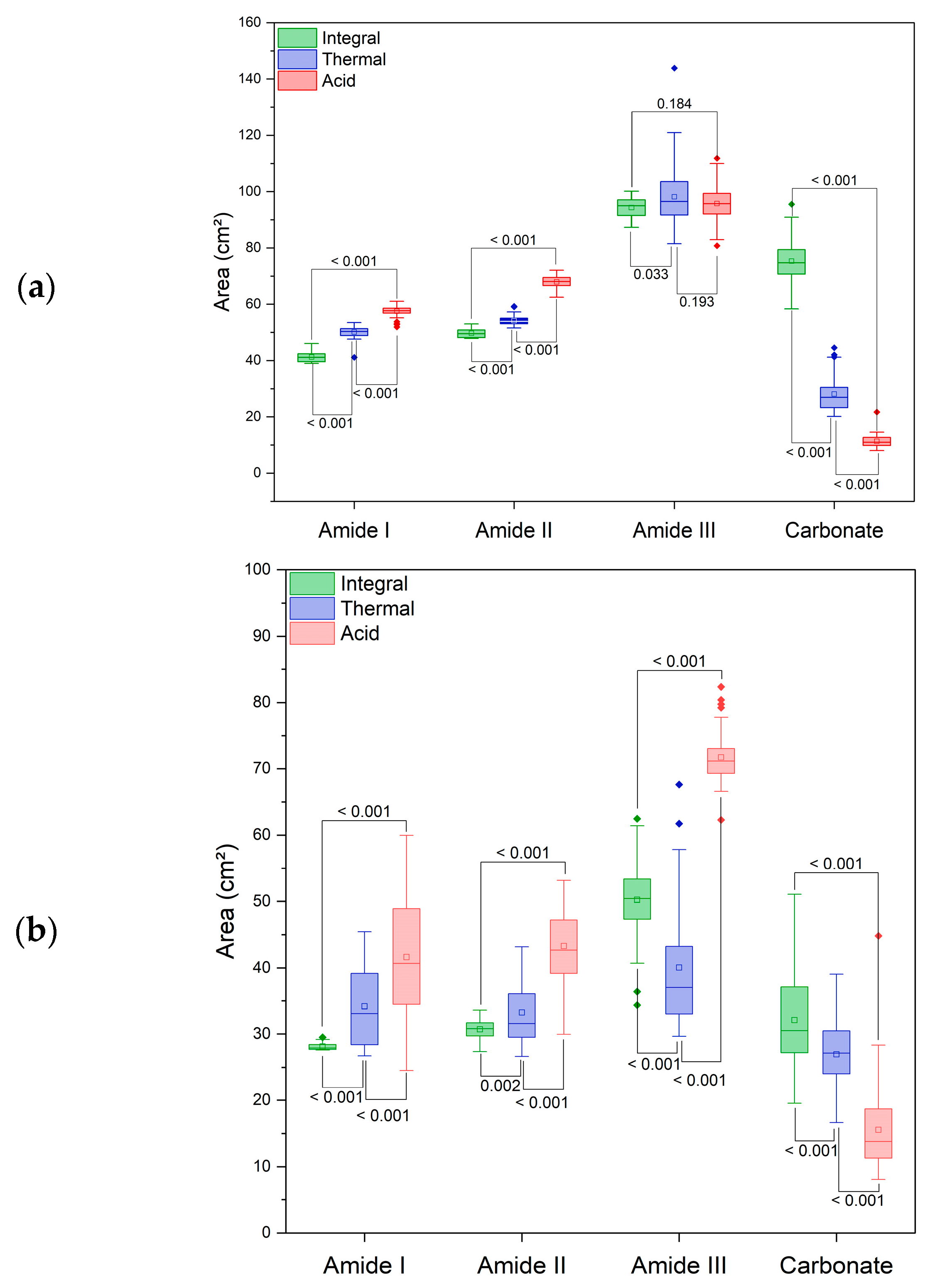



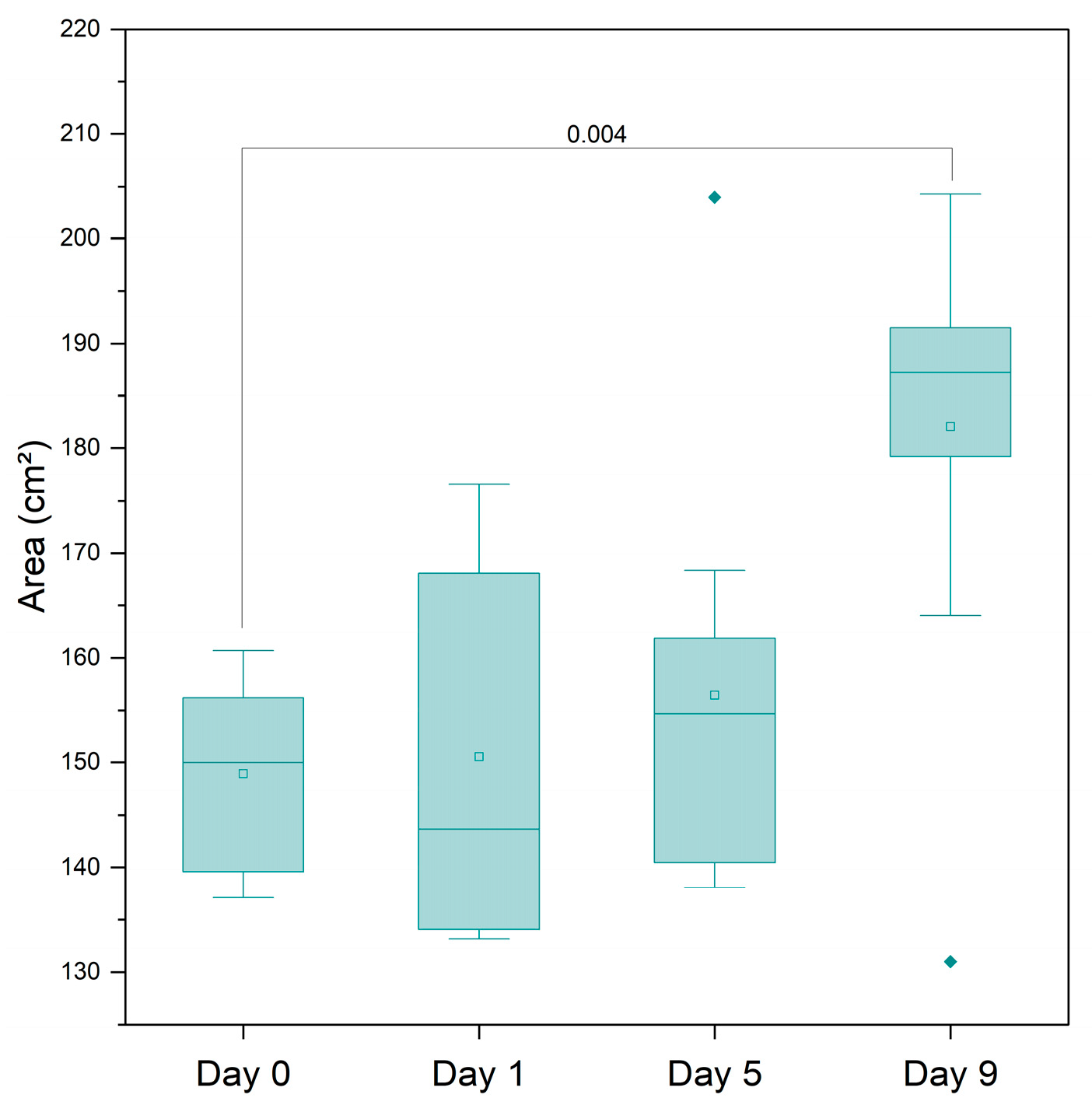



| AMP | AMX | ||||
|---|---|---|---|---|---|
| Experimental Wavenumbers (cm−1) | Theoretical Wavenumbers (cm−1) | Assignment | Experimental Wavenumbers (cm−1) | Theoretical Wavenumbers (cm−1) | Assignment |
| 1776 | 1760–1700 | C=O carboxylate ion | 1774 | 1793 | C8O14 + C3H + O13H |
| 1689 | 1680–1630 | C=O amides | 1685 | 1709 | N10H + C15O17 + C16H |
| 1600 | 1650–1550 | COO− carboxylate ion | 1616 | 1610 | C=C benzene ring + N19H |
| 1600; 1504 | 1600–1450 | C=C aromatic ring | 1574 | 1585 | C=C benzene ring + CH benzene ring + O25H |
| 1417 | 1400 | COO− carboxylate ion | 1396 | 1398 | CH3 |
| 1298; 1084 | 1350–1000 | C–N | 1282 | 1278 | N1C3 + N10H + N19H2 + CH + O13H |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anjos, V.P.; Freire, M.R.V.B.; Stasi, R.; Silva, D.F.T.; Zezell, D.M. Fourier Transform Infrared Spectroscopy-Based Detection of Amoxicillin and Ampicillin for Advancing Antibiotic Monitoring with Optical Techniques. Med. Sci. Forum 2025, 35, 7. https://doi.org/10.3390/msf2025035007
Anjos VP, Freire MRVB, Stasi R, Silva DFT, Zezell DM. Fourier Transform Infrared Spectroscopy-Based Detection of Amoxicillin and Ampicillin for Advancing Antibiotic Monitoring with Optical Techniques. Medical Sciences Forum. 2025; 35(1):7. https://doi.org/10.3390/msf2025035007
Chicago/Turabian StyleAnjos, Vinicius Pereira, Maria Renata Valente Brandão Freire, Raffaele Stasi, Daniela Fátima Teixeira Silva, and Denise Maria Zezell. 2025. "Fourier Transform Infrared Spectroscopy-Based Detection of Amoxicillin and Ampicillin for Advancing Antibiotic Monitoring with Optical Techniques" Medical Sciences Forum 35, no. 1: 7. https://doi.org/10.3390/msf2025035007
APA StyleAnjos, V. P., Freire, M. R. V. B., Stasi, R., Silva, D. F. T., & Zezell, D. M. (2025). Fourier Transform Infrared Spectroscopy-Based Detection of Amoxicillin and Ampicillin for Advancing Antibiotic Monitoring with Optical Techniques. Medical Sciences Forum, 35(1), 7. https://doi.org/10.3390/msf2025035007








