The validation of the proposed approach was conducted using a collection of benchmark datasets for classification and regression tasks, which are publicly available online through the following sources:
3.2. Experimental Results
The experimental software was implemented in C++ using the freely available Optimus framework [
104]. Each experimental configuration was executed 30 independent times, with a distinct random seed assigned to each run. To ensure a reliable evaluation of the results, the ten-fold cross-validation methodology was employed. All experiments were repeated 30 times, and performance was quantified using the mean classification error for the classification datasets and the mean regression error for the regression datasets. The classification error is calculated using the following equation:
Here the set
represents the associated test set of the objective problem. Similarly, the regression error for the test set is given from the following equation:
The values for every parameter of the proposed algorithm are given in
Table 1.
Moreover, the following notation is used in the tables that provide the experimental results:
The column DATASET provides the name of the dataset.
The column ADAM denotes the experimental results by the usage of the ADAM optimization method [
21] to train a neural network with
processing nodes.
The column BFGS denotes the usage of the BFGS method to train an artificial neural network with processing nodes.
The column GENETIC denotes the usage of a Genetic Algorithm to train a neural network with processing nodes.
The column RBF denotes the application of a Radial Basis Function (RBF) network [
105,
106] with
hidden nodes.
The column NEAT denotes the incorporation of the NEAT method (NeuroEvolution of Augmenting Topologies) [
107].
The column PRUNE denotes the application of the OBS pruning method [
108].
The column PROPOSED stands for the application of the current method.
The row AVERAGE stands for the the average classification or regression error.
The experimental results by the application of the previously mentioned machine learning methods to the classification datasets are depicted in
Table 2. Also the corresponding results for the regression datasets are shown in
Table 3.
Table 2 reports classification error rates (lower is better) across 34 datasets for seven learning/training approaches, with the last row providing the average error per method. At the aggregate level, the proposed approach (PROPOSED) is clearly the best performer, achieving an average error of 20.57%, while the second-best average is obtained by GENETIC at 26.55%. This corresponds to an absolute improvement of 5.97 percentage points, i.e., an approximately 22.5% relative error reduction compared to GENETIC. The advantage remains consistent against the remaining baselines: relative to PRUNE (27.44%), the improvement is 6.87 points (~25.0% relative reduction), relative to RBF (29.42%), it is 8.85 points (~30.1%), relative to NEAT (32.11%), it is 11.54 points (~35.9%), and relative to ADAM/BFGS (34.48%/34.34%), it is about 13.9 points (~40% relative reduction). In addition, when considering variability across heterogeneous datasets, PROPOSED exhibits the lowest dispersion of errors (standard deviation ≈ 14.31), which is consistent with more stable behavior across different classification tasks.
From a per-dataset perspective (minimum error per row), PROPOSED attains the best result on 23 out of 34 datasets (≈67.6%), indicating that its superiority is systematic rather than driven by a small subset of cases. Moreover, PROPOSED ranks within the top two methods on 31/34 datasets and within the top three on 33/34 datasets, highlighting strong rank consistency. In the datasets where PROPOSED is not the best, it is often very close to the winner (e.g., SPIRAL: 44.90% vs. 44.87%; ALCOHOL: 15.99% vs. 15.75%; POPFAILURES: 5.06% vs. 4.79%), with a limited number of more pronounced gaps such as ECOLI (49.67% vs. 43.44%) and DERMATOLOGY (14.83% vs. 9.02%). Conversely, there are datasets where PROPOSED achieves large margins over the second-best method, notably SEGMENT (38.85% vs. 49.75%), HEARTATTACK (18.97% vs. 29.00%), HEART (18.37% vs. 27.21%), and WINE (8.12% vs. 16.62%). Overall,
Table 2 supports that the proposed method delivers the best mean performance, the best average ranking, and the highest number of dataset-level wins, with only sporadic cases where alternative methods outperform it.
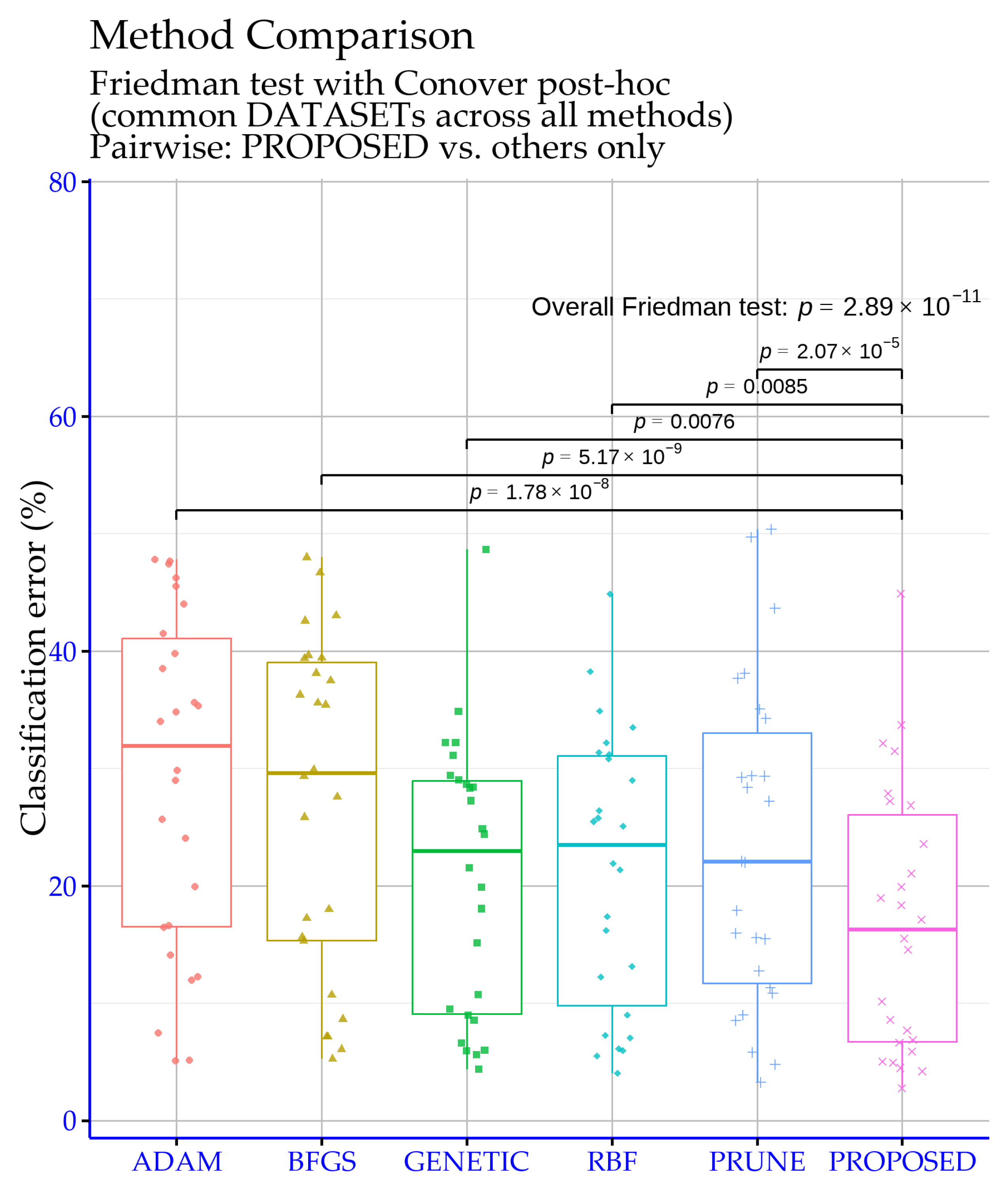
The significance levels shown in
Figure 2 were obtained via R-based analyses on the classification experiment tables, aiming to verify that the observed performance differences are statistically reliable rather than due to random variation. The overall Friedman test yields
p =
, which is extremely small and therefore strongly rejects the null hypothesis that all methods perform equivalently. This indicates that, across the set of datasets, genuine performance differences exist among the considered models and motivate post hoc pairwise comparisons against the proposed approach.
The pairwise results confirm that the proposed method differs significantly from each baseline. In particular, the comparisons ADAM vs. PROPOSED and BFGS vs. PROPOSED produce
p =
and
p =
, respectively, providing very strong evidence of a difference (well beyond the
p < 0.0001 threshold). The PRUNE vs. PROPOSED comparison is also highly decisive (
p =
, i.e.,
p < 0.0001). For GENETIC and RBF, the
p-values are larger but remain below 0.01 (
p = 0.0076 and
p = 0.0085), which corresponds to a “highly significant” difference. Overall,
Figure 2 supports that the proposed model’s superiority is not only reflected in the raw error rates, but is also substantiated by strong statistical significance against all competing baselines.
Also, a comparison between the Genetic Algorithm and the proposed one for different numbers of processing nodes is outlined in
Figure 3, which clearly demonstrates the potential of the current work. The numbers in the graph indicate the average classification error for all classification datasets that participated in the experiments.
In
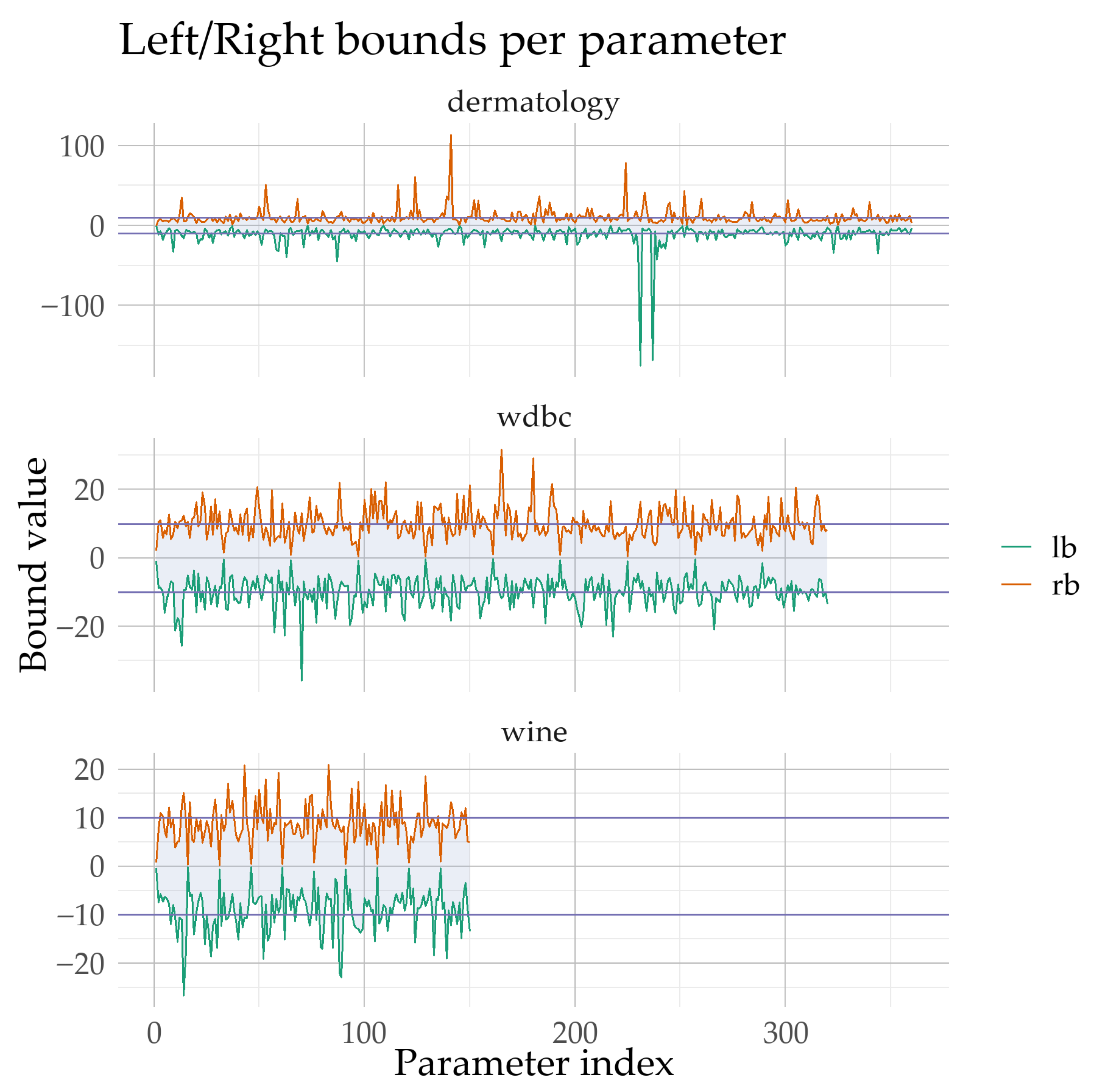
Figure 4, the computed parameter intervals of the neural network are reported as left and right bounds (L and R) obtained by the Simulated Annealing bound-optimization stage for three datasets with different dimensionalities (WINE: 150 parameters, WDBC: 320, DERMATOLOGY: 360). The visualization highlights that the bounds do not change uniformly; instead, they vary substantially across parameters and across datasets, indicating that the proposed stage does not enforce a fixed range but identifies problem-dependent search regions. Most intervals cross zero and remain centered close to 0, while their widths can become very large and exhibit occasional extreme expansions, particularly for DERMATOLOGY where a subset of parameters attains markedly wider intervals. As shown in
Figure 4, this variability supports the claim that the resulting bounds adapt to each dataset’s structure and scale, thereby providing a tailored search space for the subsequent training phase.
In the regression part of
Table 3, errors are reported in absolute units and span a very wide dynamic range (from approximately
up to hundreds), meaning that a few large-error cases can dominate average performance. Under this regime, the proposed approach (PROPOSED) delivers the most favorable aggregate outcome, with the lowest mean error of 5.33, compared to 9.31 for GENETIC and 10.02 for RBF. The reduction from 9.31 to 5.33 corresponds to a 3.98-unit absolute gain, i.e., about a 42.8% relative improvement over the best competing average (GENETIC). Importantly, PROPOSED also exhibits the strongest robustness against extreme values, showing the smallest variability across datasets (standard deviation ≈ 13.75). This point matters in regression benchmarking because heavy-tailed errors can materially affect the mean and may reveal instability. In contrast, ADAM and BFGS show very large worst-case outcomes (e.g., 180.89 and 302.43 on STOCK), which inflates their averages (22.46 and 30.29, respectively) and indicates sensitivity to high-scale or difficult regression settings.
Looking at dataset-level outcomes (row-wise minima), PROPOSED attains the best result or ties for best on 14 out of 21 regression datasets, including 11 outright wins and 3 top ties. Beyond wins, its rank consistency is strong: PROPOSED falls within the top two methods on 18/21 datasets and within the top three on 20/21 datasets, suggesting that its advantage is not driven by a small number of isolated successes. Several datasets show substantive margins rather than marginal differences, including HOUSING (26.76 vs. 43.26 for the runner-up), TREASURY (0.20 vs. 2.02), MORTGAGE (0.12 vs. 1.45), and BASEBALL (58.86 vs. 77.90). The few cases where PROPOSED is not the best do not overturn the overall pattern: concerning ABALONE, it is essentially tied with the best value (4.31 vs. 4.30), BL is dominated by RBF (0.013), and FY is the only dataset where PROPOSED falls outside the top three, yet the absolute gap remains small (0.057 vs. 0.038). Overall,
Table 3 indicates that the proposed method achieves the strongest balance of low mean error, frequent top performance, and reduced exposure to severe outliers, which is particularly relevant for regression evaluation across heterogeneous datasets.
Figure 5 reports statistical significance levels for the regression experiments, based on
p-values computed through R scripts. The overall Friedman test yields
p =
, which is well below 0.001, indicating that the compared methods do not behave equivalently across the regression datasets and that genuine performance differences exist. This justifies examining post hoc pairwise comparisons against the proposed method.
The pairwise outcomes reveal a more mixed pattern than in the classification case, which is expected in regression settings where absolute error scales can vary substantially across datasets. The strongest evidence of a difference in favor of the proposed approach is observed against BFGS (
), which falls in the extremely significant range (
). The comparison with GENETIC yields
, i.e., significant at the 0.05 level but not highly significant, implying a reliable yet weaker separation. In contrast, ADAM, RBF, and PRUNE produce
p-values of 0.0585, 0.1462, and 0.0898, respectively, all above 0.05, meaning that differences versus the proposed method are not statistically supported under the standard threshold. Overall,
Figure 5 suggests that, for regression, the proposed method exhibits statistically substantiated advantages primarily over BFGS and, to a lesser extent, over GENETIC, while differences relative to the remaining baselines are not strong enough to rule out random experimental variability.
Also in
Figure 6, an indicative plot is presented for the comparison of the training process between the Genetic Algorithm and the proposed method for the regression dataset MORTGAGE.
Figure 7 reports mean runtime as a function of the neural network’s number of nodes, comparing the Genetic Algorithm against the proposed method. Both runtimes increase with network size, but the proposed method consistently exhibits a much higher computational cost. Specifically, for two nodes, the Genetic Algorithm requires 11.34 on average versus 213.21 for the proposed method; for five nodes, the corresponding values are 29.18 versus 420.43; and for 10 nodes, they are 57.95 versus 790.35. The gap remains large in all cases, and the relative overhead decreases only slightly as network size grows, without changing the overall conclusion. Overall, the results document a clear trade-off: the proposed method is substantially more time-consuming, and its use is therefore justified when improvements in accuracy or generalization outweigh the additional computational burden.
3.3. Experiments with the Perturbation Parameter
An additional experiment was performed to determine the stability of the proposed technique to changes in the perturbation factor
. The experimental results for this experiment on the classification datasets are depicted in
Table 4 and for regression datasets in
Table 5.
Table 4 provides a targeted sensitivity study of the proposed classifier with respect to the crossover-related parameter
, reporting error rates (lower is better) on 34 classification datasets for three settings (
= 0.01, 0.02, 0.05). The averages at the bottom of the table are extremely close, yet they consistently favor
= 0.05: 20.33% versus 20.54% for
= 0.02 and 20.57% for
= 0.01. The absolute gain relative to
is only 0.24 percentage points (approximately a 1.2% relative reduction), which suggests that the method’s performance is not highly sensitive to moderate changes in
. This limited sensitivity is also reflected by the central tendency and spread: the medians are nearly identical (about 17.75%, 17.85%, and 17.90%), and the dispersion across datasets remains comparable for all three settings (standard deviation around 14%), indicating that
mainly shifts performance on particular datasets rather than reshaping the overall distribution.
A more informative view emerges from the dataset-wise minima. The setting
= 0.05 achieves the lowest error on 13 out of 34 datasets and ties for best on one additional dataset (STUDENT), i.e., 14/34 top outcomes. In comparison,
= 0.01 is best on 11 datasets, while
= 0.02 is best on nine datasets plus one top tie. Hence,
= 0.05 is the strongest default choice in terms of both average performance and frequency of best results, but the table also highlights clear dataset-specific preferences. For instance,
= 0.05 yields notable improvements on LIVERDISORDER (33.71% to 30.26%), APPENDICITIS (15.50% to 12.40%), DERMATOLOGY (14.83% to 11.92%), AUSTRALIAN (27.22% to 24.58%), and ECOLI (49.67% to 47.15%). Conversely,
= 0.01 is distinctly advantageous on SEGMENT (38.85% vs. 44.21–44.72), SPIRAL (44.90% vs. 45.29–47.77), and WINE (8.12% vs. 10.65–11.23). Overall,
Table 4 indicates that
acts as a fine-grained control parameter:
= 0.05 optimizes mean behavior and the count of best-case outcomes, while smaller values such as
= 0.01 can be preferable for specific datasets.
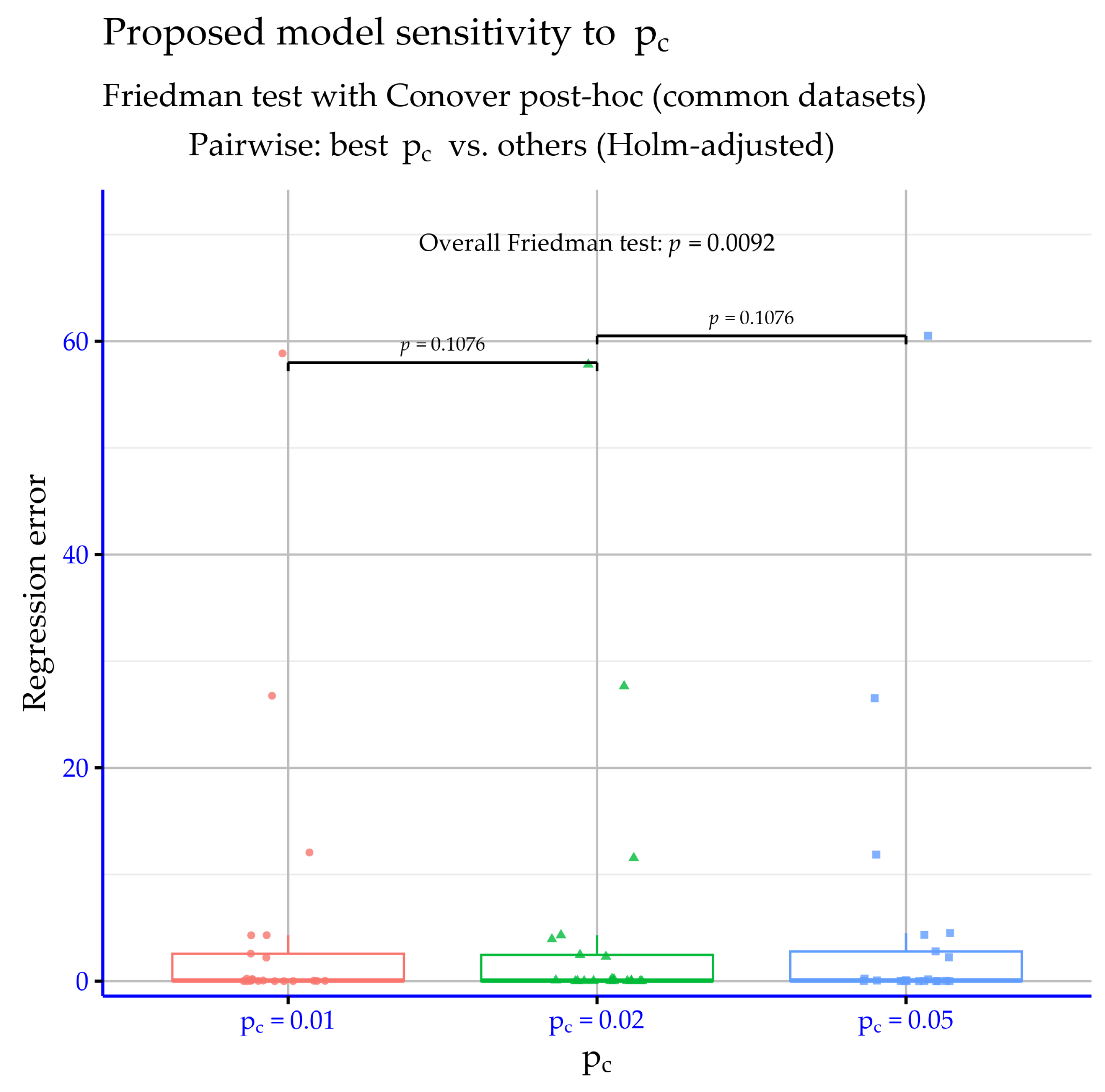
Figure 8 indicates that varying the pc parameter does not produce statistically detectable differences in the proposed model’s performance on the classification datasets. The overall Friedman test yields
p = 0.6556; hence, the null hypothesis of equivalent settings is not rejected. Moreover, the pairwise comparisons
= 0.01 vs.
= 0.05 and
= 0.02 vs.
= 0.05 both give
p = 1, confirming no evidence of separation. Therefore, within the tested range,
behaves as a practically neutral tuning choice, and the selection can be guided by secondary considerations (e.g., stability or computational cost). In summary, the three
settings are statistically indistinguishable for classification under the reported tests.
Table 5 evaluates the proposed method on regression tasks under three settings of the parameter
(0.01, 0.02, 0.05), reporting absolute errors per dataset. A key characteristic of these results is the pronounced scale heterogeneity: several datasets yield very small errors (on the order of
to
), whereas a small number of datasets produce much larger values (most notably BASEBALL and HOUSING). Consequently, the mean is strongly influenced by a few large-error cases, while the median better reflects the “typical” dataset behavior. Under the mean criterion,
= 0.02 provides the best overall outcome (average error ≈ 5.27), followed closely by
= 0.01 (≈5.33), where
= 0.05 is higher (≈5.40). This ordering is consistent with the medians as well (0.050 for
= 0.02, 0.057 for
= 0.01, 0.060 for
= 0.05), indicating that the 0.02 setting tends to reduce central errors slightly, without causing a major shift in the overall distribution.
At the dataset level,
= 0.02 emerges as the most consistently favorable choice. It achieves the lowest error with a strict advantage on 11 datasets and, when ties are included, it matches the best value on 17 out of 21 datasets. The clearest gains for
= 0.02 occur on datasets that also affect the mean, such as AUTO (11.55 vs. 12.07/11.87), BL (0.006 vs. 0.032/0.010), BASEBALL (57.83 vs. 58.86/60.52), MORTGAGE (0.079 vs. 0.12/0.085), QUAKE (0.007 vs. 0.04/0.011), and STOCK (3.93 vs. 4.30/4.51). In contrast,
= 0.05 provides a clear advantage only on a limited subset, primarily BK (0.019), HOUSING (26.53), and TREASURY (0.17), while
= 0.01 is strictly best only on PLASTIC (2.21). Additionally, several datasets are effectively insensitive to
(AIRFOIL, CONCRETE, HO, PL), where all settings yield identical outcomes. Overall, for regression performance as summarized in
Table 5,
= 0.02 is the most reliable default setting in terms of both average error and frequency of best results, with
= 0.05 being preferable in specific datasets and
= 0.01 offering isolated advantages.
Figure 9 shows no statistically significant differences among pc settings on the regression datasets. The Friedman test yields
p = 0.092 (>0.05), so equivalence is not rejected. The pairwise comparisons likewise provide no evidence of separation. Therefore, the tested
values can be treated as practically equivalent for regression.
3.4. Experiments with the Weight Parameter
In order to determine the stability of the proposed method under different initialization conditions, another experiment was conducted using it in which various values for the initialization parameter
were tested. The experimental results for the classification datasets are shown in
Table 6 and for regression datasets in
Table 7.
Table 6 examines the sensitivity of the proposed classifier to the parameter
using three settings (1, 10, 20) across 34 classification datasets, where lower error rates indicate better performance. The aggregate trend is monotonic and unfavorable as
increases: the average error rises from 19.45% at
= 1 to 20.57% at
= 10 and 21.15% at
= 20. In practical terms,
= 1 yields an absolute advantage of 1.12 percentage points over
= 10 and 1.70 points over
= 20, corresponding to approximately 5.5% and 8.0% relative error reductions, respectively. The same pattern is reflected in the distribution’s center, with medians of about 17.17% (
= 1), 17.75% (
= 10), and 18.49% (
= 20). Meanwhile, the spread across datasets remains comparable (standard deviation close to 14% for all three settings), suggesting that the degradation with higher
is not driven by a small number of extreme cases but by a consistent upward shift in errors.
A dataset-wise view further supports this conclusion. The setting
= 1 achieves the lowest error on 22 of the 34 datasets, whereas
= 10 and
= 20 each win on six datasets, with no ties for best. Importantly, larger
values are not uniformly detrimental: there are clear instances where
= 20 improves performance, such as CLEVELAND (43.20%), CIRCULAR (5.14%), PIMA (26.46%), and SPIRAL (43.86%). However, these gains are offset by pronounced losses on other datasets, including DERMATOLOGY, where
= 1 is markedly superior (6.03% vs. 13.97–14.83), SEGMENT (33.09% vs. 38.85–47.02), and WINE (6.82% vs. 8.12–12.00). Overall,
Table 6 indicates that
= 1 is the most reliable default for classification, while higher settings behave more like specialized adjustments that can benefit particular datasets but tend to reduce average performance across the benchmark suite.
Figure 10 suggests that the Iw parameter has a marginal yet detectable effect on the classification datasets, as the overall Friedman test yields
p = 0.0423 and slightly rejects the null of equivalent settings at the 0.05 level. However, the pairwise comparisons do not support strong separation:
= 1 vs.
= 10 gives
p = 0.2834 (not significant) and
= 1 vs.
= 20 gives
p = 0.069 (also not significant, but close to 0.05). This indicates that differences are modest and distributed across settings rather than producing a clear, strongly significant pairwise contrast. Practically, Iw behaves as a secondary tuning parameter with limited impact, where an overall difference is detectable but not pronounced in simple post hoc tests. Hence, selecting
can be guided by mean performance or stability, acknowledging that the statistical effects are weak.
Table 7 reports a regression sensitivity analysis of the proposed method with respect to
(1, 10, 20) using absolute error values. At the aggregate level, differences are modest: the average error is 6.10 for
= 1, 5.33 for
= 10, and 5.46 for
= 20. Thus,
= 10 is the best overall setting, improving the mean by 0.77 relative to
= 1 (≈12.6% reduction) and by 0.13 relative to
= 20 (≈2.4%). Given the heterogeneous error scales across datasets, these mean differences are influenced by a small number of high-magnitude cases, so it is also informative to consider the per-dataset pattern.
At the dataset level, no single setting dominates uniformly, rather,
behaves as a tuning parameter.
= 10 attains the minimum error on 8 out of 21 datasets and is close to the best on several others, while
= 20 is best on seven datasets and
= 1 on six datasets. The most visible gains favoring
= 10 occur on AUTO (12.07 vs. 16.88), BASEBALL (58.86 vs. 67.47), and TREASURY (0.20 vs. 1.98), whereas
= 20 is clearly preferable on ABALONE (4.22), AIRFOIL (0.001), FRIEDMAN (2.07), MORTGAGE (0.088), and STOCK (4.13). Conversely,
= 1 is best on BK (0.021), BL (0.003), FY (0.041), HO (0.008), HOUSING (23.12), LASER (0.005), and LW (0.012). Overall,
Table 7 suggests
= 10 as a reasonable default for regression under the mean criterion, while the optimal choice can vary by dataset without producing large shifts in the overall performance picture.
Figure 11 indicates that varying the
parameter does not produce statistically significant differences on the regression datasets. The overall Friedman test reports
; therefore, the null hypothesis of equivalent settings is not rejected under the standard 0.05 threshold. The pairwise comparisons are clearly non-significant as well, with
for
= 10 vs.
= 20 and
for
= 1 vs.
= 20. Such large
p-values imply practically indistinguishable behavior among the examined settings, with no evidence of a consistent winner. Hence, for regression,
can be selected based on secondary considerations because its statistical impact appears negligible.
3.5. Experiments with Initialization Methods
Additionally, three methods from the related literature were used as the initialization methods for the weights of the neural network:
The Xavier method [
49], which sets the variance of the weights based on both the number of input and output connections in the hidden layer.
The HE method [
50], which scales the weight variance according to the number of input connections.
The LeCun method [
109], which sets the weight variance inversely proportional to the number of input connections.
These weight initialization techniques were used by a Genetic Algorithm to train the artificial neural network and the results for the classification datasets are outlined in
Table 8 and for the regression datasets in
Table 9.
Table 8 isolates the role of the initialization distribution within a genetic training setting and compares it directly against the proposed machine learning model on the same set of classification datasets, using the error rate as the performance measure. In terms of averages, the four genetic variants differ in a noticeable yet bounded manner, ranging from 26.55% for the uniform initialization down to 24.69% for the LeCun initialization, a spread of about 1.86 percentage points. This indicates that initialization matters, but it does not, by itself, reshape the overall performance profile. The proposed machine learning model attains a substantially lower mean error of 20.57%, which is about 4.12 percentage points below even the strongest genetic initialization (LeCun), corresponding to a roughly 16.7% relative reduction in mean error.
The dataset-level pattern confirms that this advantage is not merely an artifact of aggregation. The proposed machine learning model achieves the lowest error on 26 out of 34 datasets, showing consistent superiority over all genetic initialization choices across most of the benchmark suite. The limited cases where genetic initialization is better occur on BALANCE, CIRCULAR, DERMATOLOGY, GLASS, HABERMAN, LIVERDISORDER, MAMMOGRAPHIC, and REGIONS2. Even among these, some gaps are marginal, while others are more pronounced, suggesting that dataset structure can selectively favor specific initialization schemes. At the same time, on several challenging datasets the proposed machine learning model yields clear reductions relative to every genetic variant, including APPENDICITIS, ALCOHOL, HEART, HEARTATTACK, SEGMENT, WDBC, WINE, and ZO_NF_S. Overall,
Table 8 supports that initialization choice within the genetic framework has a measurable but secondary effect, whereas the proposed machine learning model delivers a consistently stronger outcome both in average error and in the frequency of best per-dataset results.
Table 9 reports the regression results in absolute error units and compares four genetic training variants that differ only in the initialization distribution against the proposed machine learning model. The data indicate that initialization within the genetic framework has a tangible impact on aggregate performance, as the genetic mean error changes from 9.31 with UNIFORM to 7.24 with XAVIER and 7.32 with LECUN. Nevertheless, the proposed machine learning model achieves the lowest mean error, 5.33, which is 1.91 units below the best genetic mean, corresponding to an approximately 26% relative reduction. This suggests that the observed gains cannot be explained merely by a more favorable random initialization, but reflect a more effective overall training mechanism.
At the dataset level, the proposed machine learning model is dominant: it delivers the lowest error on 17 out of 21 datasets and ties for best on one additional dataset, while being outperformed on only three datasets. The advantage is particularly pronounced on problems where the genetic variants exhibit large errors or strong sensitivity to initialization, such as BL, HO, LW, SN, MORTGAGE, and HOUSING, where the proposed approach yields substantially smaller values than all genetic initializations. The few exceptions occur on AIRFOIL and STOCK, and also on FRIEDMAN where the genetic variants attain lower error. Overall,
Table 9 supports that while initialization choice improves genetic training to some extent, it does not match the consistently stronger regression performance of the proposed machine learning model across the benchmark suite.
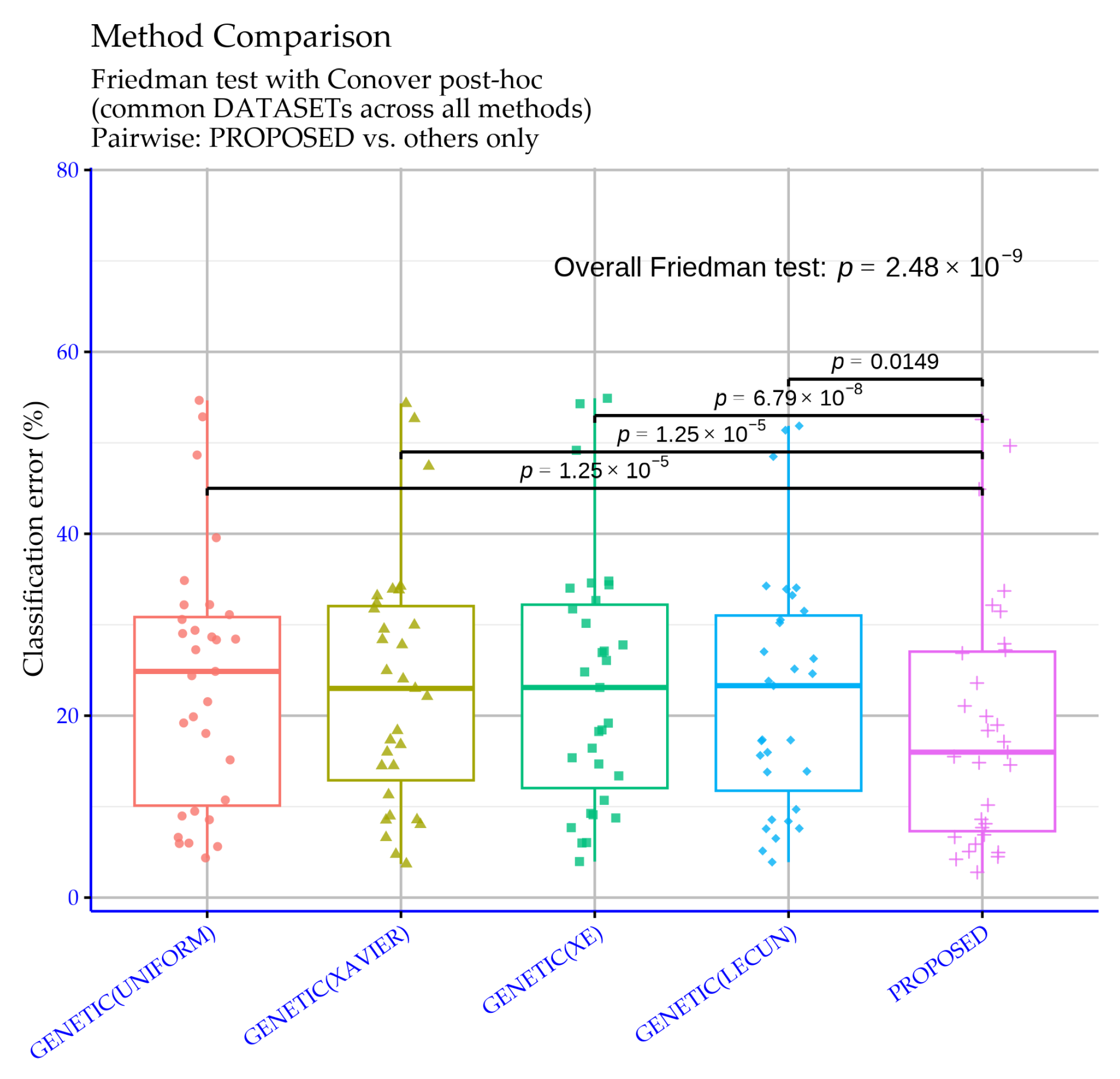
Figure 12 reports statistical significance levels for the classification experiments comparing the proposed machine learning model against four genetic training variants that differ by their initialization distribution. The overall Friedman test yields
, an extremely small value indicating that performance differences among the methods are real and cannot be attributed to random variability. In other words, across the dataset suite, the alternative genetic initializations and the proposed approach lead to measurably different performance behavior.
The pairwise comparisons against the proposed model show statistically supported differences in all cases. Specifically, GENETIC (UNIFORM) vs. the proposed model and GENETIC (XAVIER) vs. the proposed model both yield
, providing very strong evidence of a difference well below the
threshold. The GENETIC (XE) comparison is even more decisive with
, also far below
. The GENETIC (LECUN) comparison yields
, which remains significant at the 0.05 level but is clearly weaker, consistent with this variant being the most competitive among the genetic baselines. Overall,
Figure 12 confirms that the proposed method’s advantage over the genetic variants is not only reflected in mean error rates, but is also statistically substantiated across the full set of classification experiments.
Figure 13 reports R-based statistical significance levels for comparisons between the proposed machine learning model and four genetic variants that differ by their initialization distribution, using the regression experiment results. The overall Friedman test yields
, which strongly rejects the hypothesis that all methods behave equivalently across the regression datasets. Therefore, at the suite level, the genetic initialization choices and the proposed approach lead to performance differences that are unlikely to be explained by random variation.
The pairwise comparisons show that the strength of separation depends on the specific genetic variant. GENETIC(UNIFORM) vs. the proposed model and GENETIC(XE) vs. the proposed model yield
and
, respectively, both below
and thus extremely significant, providing strong evidence that the proposed method differs from these two initializations. In contrast, the comparisons against GENETIC(XAVIER) and GENETIC(LECUN) give
, which is above
, so a statistically significant difference is not supported under the standard threshold. Overall,
Figure 13 suggests that the proposed approach is statistically superior relative to the weaker genetic initializations, while against the more competitive genetic choices, the observed differences are not significant enough to rule out random experimental variability.
3.6. Experiments with the Cooling Strategy
An additional experiment was executed where the cooling strategy was altered, in order to verify the robustness of the proposed method. In this experiment, the following strategies were used:
Exponential decreasing (EXP), the temperature is decreased using the following equation:
where
k defines the current iteration.
Logarithmical multiplicative cooling (LOG), where the following formula is used:
Linear multiplicative cooling (LINEAR), using the following formula:
Quadratic multiplicative cooling (QUAD), with the following formula:
The experimental results for the classification datasets are reported in
Table 10 and for regression datasets in
Table 11.
In
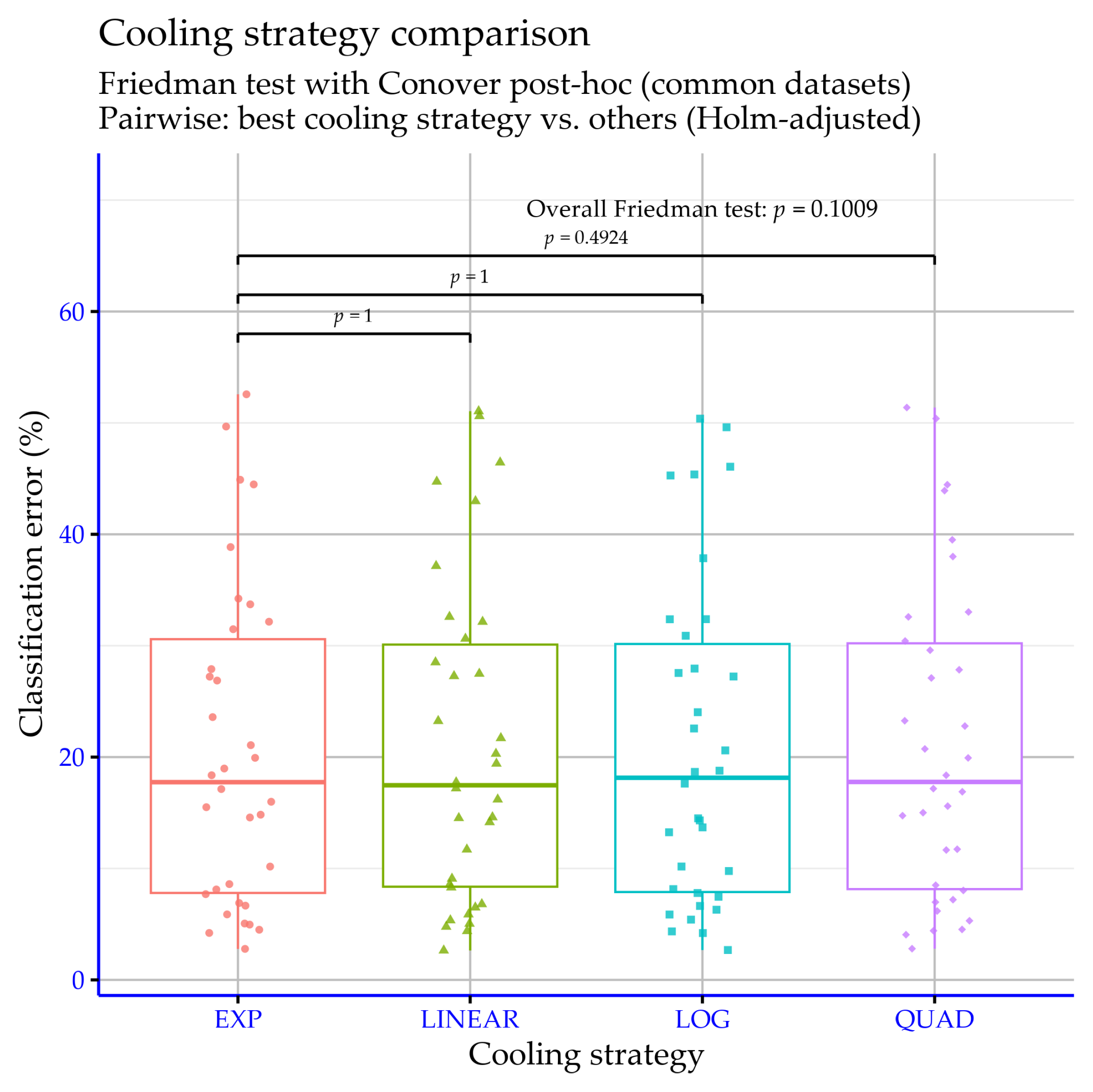
Table 10, the classification error rate (%) is reported for 34 datasets under four cooling schedules (EXP, LOG, LINEAR, QUAD) within the proposed model, where lower values indicate better performance. In terms of average error, EXP provides the best overall result (20.57%), while LOG (20.88%) and LINEAR (20.87%) are essentially tied and very close, and QUAD yields the highest average (21.01%). Although the average differences are small, the best schedule can be dataset-dependent, and some datasets exhibit higher sensitivity to the cooling choice (e.g., SEGMENT and WINE). As shown in
Figure 14, there is no statistical significance according to the
p-values.
In
Table 11, the reported values are the errors (lower is better) on 21 regression datasets, obtained by using four cooling schedules (EXP, LOG, LINEAR, QUAD) within the proposed model. If we focus on the overall picture, LOG has the lowest average error (5.16), EXP and LINEAR are essentially the same (5.33), and QUAD is slightly higher (5.42). In other words, the average differences are modest and do not suggest a single schedule that clearly dominates everywhere.
At the dataset level, however, the picture is less uniform. There are datasets where the cooling choice seems to matter more, such as BL, HOUSING, FRIEDMAN, and STOCK, because the gaps between schedules become noticeably larger. In contrast, datasets like CONCRETE, HO, and LASER remain almost unchanged across schedules, indicating that performance there is largely insensitive to how temperature is decreased. If a single, simple default must be used without per-dataset tuning, LOG is the most reasonable choice because it yields the best average.
Figure 15 indicates that the
p-values do not support statistically significant differences among the four cooling schedules.