Quantum and Artificial Intelligence in Drugs and Pharmaceutics
Abstract
1. Introduction
2. Methodology and Synthesis
| Aspect | Classical AI/Classical Physical | Quantum-Accelerated AI/Quantum Physical |
|---|---|---|
| Computational Power/Underlying Principles | AI: Relies on binary bits (0 s and 1 s) and classical logic. Exponentially slow and computationally intractable for simulating complex quantum states [24]. Physical: Based on classical physics (Newtonian mechanics, electromagnetism, thermodynamics). Measurements of the macroscopic averages of many particles [25]. | AI: Leverages qubits for superposition and entanglement [26,27]. Can theoretically handle exponentially large computational spaces, making it ideal for simulating quantum-level molecular interactions [28,29]. Physical: Based on quantum mechanics (superposition [30,31], entanglement [32,33,34], quantum tunneling [35,36]. Measurements can be performed at the single-particle or quantum-state level. Quantum states are highly sensitive to decoherence, requiring controlled environments [37]. |
| Typical Instruments/Methods | AI: Widely accessible via cloud computing (AWS, Google Cloud, Azure) [38] and open-source frameworks (Scikit-learn, PyTorch from Linux Foundation) [39]. The entry barriers are relatively low for standard analytical and modeling tasks. Physical: Classical Sensing: Mass spectrometers, NMR machines (traditional), optical microscopes, HPLC, calorimetry, fluorescence spectrometers. Classical Control: Automated liquid handling systems, temperature-controlled reaction vessels, robotic arms. | AI: Highly specialized and expensive. Access primarily via cloud-based services (IBM Q, IonQ) [40]. Requires expertise in quantum algorithms and hybrid computing. Physical: Quantum sensors [41,42,43,44,45,46,47] (e.g., NV diamond, quantum dots, SQUID, which extends the sensitivity and resolution of classical instruments. Quantum control [17,48] (e.g., Quantum Rabi Oscillations [49,50], Quantum Shaped Pulses [51,52,53], Adiabatic Evolution [54,55], Quantum Spin Echo [56]) for manipulating molecular spin states and reaction pathways, which augment conventional control methods. |
| Ideal Use Cases | AI: Pattern recognition [57,58], data-intensive prediction, interpreting large datasets, ADMET prediction [59], biomarker identification from structured data [60,61], high-throughput screening (HTS) [62]. Example: Training a model to identify active compounds from thousands of microscopy images or predicting drug toxicity from vast datasets. Physical: High-throughput screening, structural analysis of large molecules (e.g., traditional NMR) [63], bulk sample analysis, measuring macroscopic properties (temperature, concentration). Example: HPLC to separate large compound samples or traditional mass spectrometers to measure protein molecular weight. | AI: Simulating molecular dynamics at quantum level, modeling complex protein folding, predicting binding affinities with high accuracy, optimizing chemical reactions intractable for classical computers [29]. Example: Precisely modeling how a new drug binds a specific protein, accounting for quantum effects. Physical: High-resolution sensing, single-molecule detection, in vivo sensing, manipulating matter at quantum level [64,65,66,67]. Example: NV diamond sensor detecting single electron spin in a living cell or quantum dot as an ultra-sensitive biomarker. |
| Limitations | AI: Struggles with problems involving quantum phenomena or subtle, high-dimensional molecular interactions (e.g., electronic correlations [29,68]). Provides only an approximation of physical reality. Identifying systems with strong electronic correlations remains difficult with limited indicators. Physical: Limited by shot-noise limit (fundamental precision limit in classical measurements). Cannot detect single quantum states or surpass classical optics limits [37,69]. | AI: Limited by current quantum hardware: noisy, error-prone, and fault tolerant quantum computing for classically challenging molecules (e.g., FeMoco [70]) estimated at ~2000 logical qubits (~4 million physical qubits). Near-term intermediate quantum (NISQ) devices require highly controlled environments [71] and specialized expertise; measurements scale exponentially with circuit depth [72]. Quantum machine learning faces data representation challenges [73], some approaches are subject to de-quantization. Physical: Quantum decoherence; quantum states lose properties when interacting with environment [74]. Requires highly controlled conditions; technology not yet mature for robust clinical use. |
| Future Trajectory and New Possibilities | AI: Will excel at data-centric tasks. Focus on larger models Artificial General Intelligence (AGI), better data integration (Multimodal AI), and autonomous action (Agentic AI) [17,48,75,76]. Physical: Provides high-quality bulk data but limited to ensemble-averaged results. Cannot capture subtle, real-time molecular dynamics at quantum level. | AI: Transition from noisy NISQ to fault tolerant system [70]; true quantum advantage on biochemical problems. Eventually standard tool for computational chemistry alongside classical AI. Physical: Potential for sub-nanometer resolution, ultra-high sensitivity (single-molecule), real-time non-invasive in vivo diagnostics, quantum control of reactions, and quantum-enhanced medical imaging. |
3. Foundational Technologies: The AI and Quantum Toolkit
3.1. Defining the Dual Nature of Quantum and AI in Pharma
- The Physical Basis of Quantum Technologies
- The Dual Paradigms: Computational and Physical
- The Role of AI in Orchestrating Quantum Systems
3.2. AI Paradigms and the Biomedical Data Ecosystem
- The Biomedical Data Ecosystem
| Paradigm | Definition and Core Features | Pharma/Biomed Applications and Examples | Methodologies/Tools/Limitation |
|---|---|---|---|
| Symbolic AI (Good Old-Fashioned AI) (Stage 1–3) | Rule-based, logic-driven systems and explicit knowledge representation. | Applied to metabolic pathway mapping [112,113] and early Quantitative Structure-Activity Relationship models [114,115]. Worked in stable, clean contexts but failed in dynamic biological systems due to brittleness. Analogy: brittle “if–then” rules. | Rule-based inference engines and symbolic logic. Failures are due to lack of scalability and inability to handle noisy/unstructured data. |
| Machine Learning (ML) (Stage 1–5) | Algorithms that learn from structured data to make predictions/decisions without explicit rules [100,116]. | Predictive models for drug discovery, ADME/ADMET, retrosynthesis, multi-omics (e.g., identifying genetic markers linked to drug response), and ethics/interpretability) [18,59,117,118,119,120] Analogy: “pattern spotter” uncovering hidden correlations humans miss. | Regression, classification, clustering, dimensionality reduction [58]. Platforms: Scikit-learn, XGBoost, AutoML frameworks (Google AutoML, H2O.ai) for automated model selection/tuning [39,121]. Dependent on quality/quantity of structured data. |
| Deep Learning (DL) (Stage 2–5) | Subset of ML using neural networks with multiple deep layers to learn complex hierarchical patterns from raw data. | Protein folding prediction (e.g., AlphaFold) [122,123,124], structure-based drug design, de novo small-molecule generation [120,125], high-throughput image analysis. Analogy: stacked layers like visual cortex neurons or GNNs learn like networks of interacting proteins. | CNNs (Convolutional Neural Networks), RNNs (Recurrent Neural Networks), Transformers, reinforcement learning and Graph Neural Networks [58,121]. High data/computational requirements and black-box nature are limitations; it excels with unstructured data. |
| Generative AI (Stage 2–4) | Models creating new data resembling training data (GANs (Generative Adversarial Networks), VAEs (Variational Autoencoders), LLMs (Large Language Models) [120]. | Novel molecular design, scaffold optimization, synthetic data for scarce datasets, regulatory document drafting [120,125,126]. Moves drug design from library screening to invention. (e.g., LLMs for summarizing papers or drafting regulatory submissions). Analogy: “two dueling artists” (generator vs. discriminator). | GANs (Generative Adversarial Networks), VAEs (Variational Autoencoders), LLMs (Large Language Models) (GPT-style) [120], diffusion models. Risk of hallucination and need for domain constraints. |
| Agentic AI (Stage 1–5) | AI with autonomy, planning, and capacity to act in dynamic environments [93,94]. | Lab automation, self-optimizing robotic experimental workflows, autonomous design-synthesis-test loops [127,128,129,130]. (e.g., Eve, Insilico Medicine labs, LabGenius). Analogy: pneumonia workflow (detect, check history, order labs, notify nurses, update EHR). | Planning algorithms, reinforcement learning and robotics integration [129,131]. Significant regulatory and safety considerations; high complexity. |
| Collective AI (Stage 1–5) | Multiple specialized AI agents collaborating and sharing intelligence. | Collaborative drug discovery, federated learning for multi-hospital datasets, distributed problem-solving [132]. Analogy: “team of AIs” each with a specialized role. | Federated learning frameworks, agent-based modelling [132]. Coordination overhead and hard to predict emergent behaviors. |
| Multimodal AI (Stage 2–5) | Integrates diverse data types (omics, imaging, EHR, text) for holistic predictions. | Patient phenotype prediction, drug–disease interaction mapping, precision interventions [58,133,134] (e.g., DeepMind RETFound, IBM Watson Health). Analogy: doctor synthesizing labs + scans + history. | Transformers, multimodal fusion architectures [58,133,134]. Complex integration pipelines. Aligning modalities is challenging but improves understanding of complex biological phenomena. |
| AGI/ Superintelligence (Stage 1–6) | AI with general reasoning across domains, akin to human cognition (long-term vision). | Governance, complex system optimization, synergy with quantum for pharma R&D. Analogy: “human-like scientist partner,” but with unknown risks [75,76,135] | Theoretical models (still aspirational). Ethical, philosophical, and regulatory debates and currently speculative. |
- Key AI Methodologies and Quantum Integration
4. Computational and Physical Layers
4.1. Computational Design, Analysis and Biochemical Realism
| Quantum Aspect | Classical Physical Layer | Quantum Physical Layer | Classical AI Layer | Quantum AI Layer |
|---|---|---|---|---|
| Quantum Simulation | Scalable modeling via HPC (High-Performance Computing) clusters, NVIDIA DGX, Cryo-EM, X-ray crystallography, SAR (Structure–Activity Relationship), HTS (High-Throughput Screening), MD (Molecular Dynamics), and structure-based design. | High-fidelity quantum modeling with Qiskit Nature, PennyLane, Bloqade, Variational Quantum Eigensolver, Density Matrix Embedding Theory, Quantum Mechanics/Molecular Mechanics, NV centers, and quantum-enhanced spectroscopy—delivering near ab initio accuracy for physical and chemical systems. | AlphaFold for protein folding, generative design, Automated Machine Learning using KNIME and RDKit, multi-omics integration, Quantitative Structure–Activity Relationship, and iterative Design–Make–Test–Analyze cycles. | Hybrid workflows via Chemistry42 and AWS Braket, supporting mechanistic and micro-mechanistic toxicity analyses. Quantum Machine Learning for chemical space exploration, binding affinity prediction, quantum Variational Autoencoders, recursive learning, and multi-objective ADMET optimization. |
| Quantum Analysis | Fast hit prioritization via docking, HTS, MD, and ligand-based screening with interpretable pathway insights. | Stratification in small cohorts and subtle causal inference via quantum clustering, reverse docking, quantum kernel methods, and signal detection in noisy data. Quantum causal inference enhances pathway modeling. | NLP (Natural Language Processing)-based integration of heterogeneous datasets, anomaly detection, and pathway modeling. | Adaptive chemical space exploration beyond classical reach using hybrid Quantum Generative Adversarial Networks, recursive learning, quantum co-evolution of molecular architectures, and quantum-enhanced feedback loops for stratification. |
| Quantum Security (Digital + Physical) | Mature, regulation-ready, cross-jurisdiction compliant. Advanced Encryption Standard/Rivest–Shamir–Adleman encryption, Hardware Security Modules, audit trails, and secure lab infrastructure. | Immutable audit trails and cross-vendor secure exchange. Quantum-safe protocols including Quantum Key Distribution or Post-Quantum Cryptography such as CRYSTALS-Kyber and Dilithium. Quantum-optimized firewalls, and Quantum Random Number Generators. | Explainable AI for compliance, anomaly tracking, secure data exchange, and NLP-based monitoring. | Quantum explainability, provenance-aware Agentic AI, embedded safeguards, and quantum adversarial defense mechanisms with adaptive monitoring beyond classical AI. |
| Hardware Security | Standardized, robust containment and device-level security: secure instruments, Biosafety Level 3/4 labs, containment protocols, and sequencer safeguards. | Hardened assay/sensor capture appliances, quantum tamper-evident storage, biosensors, edge devices, archival systems, and quantum-safe firmware. | AI-driven monitoring of instruments and compliance systems. | Quantum simulations for pathogen evolution, agentic monitoring, ultra-sensitive hazard detection, and biosurveillance modeling with predictive risk anticipation beyond classical systems. |
| Quantum Sensors | Reliable, interpretable measurements: high-throughput phenotypic readouts via robotics, AFM, HTS readers, pipetting, and microscopy. | Single-molecule sensitivity and bedside/surgical micro-sensing with wearable diagnostics, enabled by ultra-sensitive detection using NV centers, quantum dots, optical frequency combs, magnetometers, and real-time systems. | High-content phenotypic screening using Convolutional Neural Networks, Long Short-Term Memory networks, and Vision Transformers for imaging, feedback, and anomaly detection. | Quantum-assisted interpretation, control optimization, and adaptive diagnostics using quantum algorithms and sensor fusion—enabling real-time adaptive experimentation. |
| Quantum Actuators | Mature and scalable robotic and optical manipulation for precise, reproducible perturbations. | Mode-selective bond editing, photo-triggered validation, metabolic/repair pathway nudging, and reflexive control via quantum-level photonic actuators, optogenetics, and therapeutic modulation—for precise molecular actuation and quantum-scale physico-chemical manipulation. | Reinforcement learning for multi-objective design, self-tuning optimization, and lead prioritization for fast, interpretable refinement. | AI-optimized quantum control for co-evolving molecular architectures and self-optimizing therapeutic interventions via hybrid quantum-classical systems—enabling interventions beyond classical capability. |
- Bridging the Gap: From Statistical Models to Physical Reality
- Classical AI as foundational pillar
- Quantum-Enhanced AI to bridge the Gap
- Advancing the Computational Workflow
4.2. Real-Time Control, Sensing, and Automation
- The Promise of Quantum Sensing
- The Promise of Quantum Actuation
- From Lab Automation to Self-Driving Labs
4.3. Optimizing Clinical Trials and Operations
| Stage | Classical Physical Layer | Quantum Physical Layer | Classical AI Layer | Quantum AI Layer |
|---|---|---|---|---|
| Stage 1: Target Discovery/Patient Stratification | Immunoassays, blood tests, traditional imaging (Magnetic Resonance Imaging, Computed Tomography), bulk sample analysis, flow cytometry, Enzyme-Linked Immunosorbent Assay (ELISA) panels for known biomarkers. | Quantum sensors (e.g., Superconducting Quantum Interference Devices (SQUIDs) [47,191,192,209,210], Nitrogen-Vacancy center [37,41,45,64]; and Quantum Dots [211]), allow high-resolution, real-time biomarker detection, continuous monitoring, and multiplexed sensing | Integrates multi-omics [57,58], Electronic Health Records, and wearable data for subtyping, digital twin modelling [212,213,214], and patient risk stratification. | Simulates mutation effects on protein function, metabolism, and signaling pathways at atomic resolution; quantum-enhanced biomarker discovery in high-dimensional combinatorial space [71]. |
| Stage 2: Lead Identification/Mechanism of Action Clarity | Traditional spectroscopy (Circular Dichroism, Fourier Transform Infrared Spectroscopy), bulk kinetics assays (Surface Plasmon Resonance), gel electrophoresis; fluorescence and absorbance assays for enzymatic activity; NMR for secondary structure. | Quantum spectroscopy (e.g., Quantum Cascade Lasers [42], SPR, interferometry), tracks conformational changes, reaction fingerprints, electron tunneling, and excited state dynamics [114]. | Uses graph neural networks (GNNs), causal inference, and network analysis to model drug actions [139,140], allosteric modulation, and off-target interactions. | Models binding, tunneling, and excited states ab initio; simulates electron density changes, transition states, and quantum-induced allosteric effects invisible to classical methods [114,164,215,216]. |
| Stage 3: Lead Optimization/Molecular Specificity | Rational drug design using classical molecular mechanics and docking [71,217]; traditional synthetic chemistry; high-throughput screening; chromatography and NMR for structure confirmation. | Coherent control (laser shaping, pulse sequences) enables enantioselective synthesis, bond activation, and monitoring of transient reactive intermediates [41,42,43,44,45,67] | Generative Adversarial Networks, Variational Autoencoders and Large Language Models [120], explore chemical space, predict synthesis routes, optimize ADMET properties [59,120,218,219], and design selective ligands or bifunctional molecules. | Simulates stereoselective pathways, electron-level interactions, reaction energetics, and molecular dynamics for stable, targeted designs; evaluates quantum tunneling in catalytic reactions [215,220,221,222]. |
| Stage 4: Preclinical Development/Manufacturability and Stability | High-Performance Liquid Chromatography for purity, rheometers for viscosity, stability chambers, traditional machine vision for Quality Control, spectrophotometry for degradation monitoring; thermal analysis (DSC, TGA). | Quantum sensors [200,201,223], detect nanoscale stability shifts, polymorph transitions, and excipient interactions; quantum electrodynamics could support light-activated carriers and precision-controlled reactions. | Predicts formulation compatibility, degradation kinetics, excipient selection, and process parameters; uses classical ML to anticipate stability issues under multiple conditions [59,224,225]. | Models’ polymorph transitions, solubility, chemical degradation pathways, and metabolism with quantum precision; simulates quantum-controlled excipient interactions for improved stability [114,222]. |
| Stage 5: Clinical Trials/ Toxicity and Safety Profiling | Traditional toxicology assays (MTT, LDH), animal models, clinical observations, ECG/biomarker monitoring; imaging for organ toxicity; hematology and biochemistry panels. | Quantum sensors detect early toxicity biomarkers, metabolic perturbations, and reactive intermediates, enabling closed-loop adaptive safety testing in preclinical and clinical phases. | Uses toxicogenomic, pharmacovigilance, and population health data to forecast adverse events; AI models predict dose–response relationships and rare events [100,111,226]. | Simulates reactive intermediates, metabolic byproducts, enzyme interactions, and off-target effects at quantum resolution; could enable atomic-level modeling of toxicity pathways. [6,117]. |
| Stage 6: Post-Market and Trial Optimization | Manual data collection, fixed trial designs, EHRs, patient registries; traditional remote monitoring; periodic lab tests and imaging. | Quantum sensors enable dynamic endpoint adjustment, decentralized trial monitoring, real-time physiological tracking, and high-resolution wearable integration [227]. | Designs adaptive trials, stratifies patients using real-world and longitudinal data, and predicts adherence and long-term outcomes [111,228]. | Models’ patient-drug interactions under uncertainty, evaluates population-level responses, predicts rare adverse events, and improves trial robustness with quantum-enhanced simulations [71,111]. |
5. Mechanistic Layer: Biomarkers, Targets, and Pathways
5.1. The Role of Classical Physics and AI in Mechanistic Discovery
| System/Application | Machines and Hardware | Classical Physical Capabilities | Quantum Machines and Hardware | Classical-AI Solutions | Quantum-AI Solutions |
|---|---|---|---|---|---|
| Target Discovery and Interaction (Stage 1–2) | High-Performance Computing clusters, Cryogenic Electron Microscopy; Quantum chemistry platforms | Traditional spectroscopy (Circular Dichroism, Fourier Transform Infrared Spectroscopy) [235], bulk kinetics assays (Surface Plasmon Resonance). | Quantum-enhanced spectroscopy enables high-resolution structural analysis [236,237]. | Target prioritization using network biology and causal inference [238]. | Atomic-level simulations for binding energy and drugability [114,163,216] |
| Generative Drug Design (Stage 2–4) | NVIDIA DGX systems, generative platforms; Quantum annealers. | Molecular docking [71,217,239,240], HTS, MD simulations; Structure-based drug design [241,242]; Rational drug design; Traditional synthetic chemistry. | Quantum spectroscopy and coherent control techniques (e.g., laser pulse shaping) enable high-resolution structural analysis and enantioselective synthesis [243]. | Graph Neural Networks, Generative Adversarial Networks, Variational Autoencoders and Large Language Models [244] for molecule design; AlphaFold predicts protein folding [122,123,124] | Quantum generative models (e.g., quantum Generative Adversarial Networks, quantum Variational Autoencoders) for chemical space exploration and optimize molecular generation [222,245] |
| Molecular Design and Optimization (Stage 2–4) | High-Performance Computing robotics, Mass Spectrometry, NMR, Cryo-EM; Quantum simulators. | Traditional spectroscopy (Circular Dichroism, Fourier Transform Infrared Spectroscopy), bulk kinetics assays (Surface Plasmon Resonance). | Quantum-enhanced spectroscopy [17,48,71,217] | Property prediction and hypothesis generation (e.g., Graph Neural Networks, AlphaFold) [122,123,124]; | Quantum deep learning for property prediction and chemical space navigation [17,48,92,245,246] |
| Protein-Ligand Binding (Stage 2–4) | Docking platforms; MD engines. | Bulk kinetics assays (SPR), gel electrophoresis [247]. | Quantum-enhanced spectroscopy for detailed binding dynamics [17] | ML scoring functions and absorption models for binding prediction [59,119,248]. | Hybrid quantum neural networks for FCI-level binding affinity prediction [245,249]. |
| Chirality and IDP Dynamics (Stage 2–4) | Circular Dichroism spectrometers, Vibrational Circular Dichroism systems; Quantum simulators. | Circular Dichroism and Vibrational Circular Dichroism spectroscopy. | Quantum-enhanced spectroscopy resolves chirality with precision [250,251]. | Docking, chirality, and protein dynamics modeling [71,217,252]. | Quantum simulations for tunneling and stereoselectivity; generative AI for Intrinsically Disordered Proteins conformations [253]. |
| Toxicity and Safety Profiling (Stage 3–5) | Toxicology assays (MTT), animal models | MTT assays; clinical observations | Quantum sensors detect early toxicity biomarkers, while closed-loop systems enable adaptive safety testing [254]. | Prediction of ADMET properties and adverse events [59,120,218]. | Quantum simulations for metabolic reactions and toxicity validation [255,256]. |
| Biosecurity and Dual-Use Risk (Stage 1–6) | Pathogen surveillance, genomic profiling. | Genomic profiling, traditional toxicology. | Ultra-sensitive detection of biohazards with quantum biosensors [254,257]. | NLP prediction of outbreak signals and dual-use risks [258] | Quantum models for protein-pathogen interactions and risk forecasting [22]. |
5.2. The Complementary Contributions of Quantum in Mechanistic Discovery
- Quantum Mechanics Provides Access to Electronic-Level Phenomena
- Quantum Mechanics For Accurate Description of Potential Energy Surfaces
- Quantum Mechanics Accounts for Unique Physical Phenomena
| Objective | Classical Methods | Benefits and Limitations | Quantum Methods | Benefits and Limitations |
|---|---|---|---|---|
| Molecular Structure and Dynamics (Stage 1–4) | NMR, X-ray Crystallography, Cryo-Electron Microscopy, Atomic Force Microscopy, Circular Dichroism (CD). | Benefits: High-resolution 3D structures (X-ray, Cryo-EM), non-destructive analysis (NMR), conformational insights (CD). Limitations: Requires large sample sizes (NMR), static snapshots (X-ray), complex image processing (Cryo-EM), limited spatial resolution (NMR), bulk averaging (CD). | Quantum-enhanced NMR [209,210]; Quantum tunneling Microscopy [274]; Nitrogen-Vacancy center (NV-center) nanoscopy [41,45,64]; Quantum embedding and hybrid QM/MM | Benefits: Enhanced sensitivity and resolution, nanoscale imaging of electronic and spin properties, single-molecule kinetics, label-free detection. QM/MM and Fragment Molecular Orbital (FMO) methods enable scalable modeling of large biomolecules. Limitations: Early-stage technologies, specialized instrumentation, potential imaging artifacts (QTM), complex data interpretation. |
| Reaction Kinetics (Stage 1–4) | Spectroscopy, Plate Readers, Liquid Chromatography-Mass Spectrometry/Mass Spectrometry | Benefits: Established protocols, high-throughput screening, detailed mass and concentration analysis. Limitations: Bulk measurements, limited temporal resolution, indirect detection, potential invasiveness. | Quantum-enhanced Spectroscopy [202,243], Pulse-shaped ultrafast lasers [275], Quantum Causal Reasoning [276]; Quantum optimal control | Benefits: Femtosecond-scale time resolution, sub-diffraction imaging, mechanistic inference, single-molecule sensitivity. Quantum optimal control enables steering of reaction pathways and lowering of activation barriers. Limitations: Technically demanding, high cost, limited accessibility, early-stage development. |
| Photophysical Properties (Stage 1–4) | Fluorescence/Absorbance Spectroscopy | Benefits: Cost-effective, widely used for quantum yield and absorption spectra. Limitations: Bulk averaging, indirect measurements, limited specificity, diffraction-limited resolution. | Quantum-enhanced Spectroscopy [243], Quantum plasmonic biosensing, entangled-photon Fluorescence Lifetime Imaging Microscopy (FLIM) [201,277]. | Benefits: Shot-noise-limited sensitivity, sub-diffraction resolution, enhanced lifetime and energy transfer measurements. Quantum plasmonics enables real-time monitoring of photophysical changes. Limitations: Requires stable squeezed light sources, homodyne detection, and specialized equipment. |
| Cellular and in vivo Imaging (Stage 2–5) | Confocal/Fluorescence Microscopy, Cryo-EM | Benefits: High-resolution imaging, applicable to live-cell studies. Limitations: Limited depth penetration, photo-toxicity, static imaging (Cryo-EM), diffraction-limited resolution. | NV-center nanoscopy [41,45,64]; Quantum Dot Microscopy [278]; Hyperpolarized quantum MRI/NMR [279]. | Benefits: Nanoscale resolution, single-molecule sensitivity, non-invasive imaging, deeper tissue penetration, metabolic imaging. Limitations: Requires specialized setups, susceptible to noise, quantum dot toxicity varies with coating and size. |
| Binding, Stability and Thermodynamics (Stage 2–5) | Surface Plasmon Resonance/Biolayer Interferometry, Dynamic Light Scattering, Differential Scanning Calorimetry. | Benefits: Real-time kinetics, label-free detection, particle sizing, thermal stability profiling. Limitations: Bulk measurements, low spatial resolution, slow acquisition, sensitivity to non-specific interactions. | Vibrational Strong Coupling [280,281], Quantum diamond magnetometry [282]. | Benefits: High sensitivity to subtle molecular changes, single-molecule binding detection, sub-Hz resolution of interactions. Limitations: Requires precise cavity design, strong coupling conditions, and specialized setups (still in early experimental stages). |
| Reaction Rates and Pathways (Stage 2–4) | Temperature/Pressure Controllers, Chiral Catalysts | Benefits: Scalable, well-established for bulk synthesis and control. Limitations: Non-specific control, thermally driven reactions, catalyst design constraints. | Pulse-shaped ultrafast lasers [275], Vibrational Strong Coupling [280,283], Radiofrequency (RF)/B-field Spin Control [284]; Quantum control via photonic reagents | Benefits: Selective bond excitation, manipulation of energy landscapes, spin-selective reactions, precise control over reaction outcomes. Photonic reagents enable targeted reaction control. Limitations: Complex setups, early-stage development, sensitive to environmental noise. |
| Catalysis and Selectivity (Stage 2–4) | Traditional Catalysts, Photochemistry | Benefits: Broad applicability, diverse reaction conditions. Limitations: Non-specific catalysis, bulk-level control, often requires harsh conditions. | Quantum-enhanced Catalysis [283,285], Vibrational Strong Coupling | Benefits: Enhanced reaction rates via quantum tunneling, energy landscape modulation, electrocatalysis via heteroatom doping and defect engineering. Limitations: Theoretical and early experimental stages, conductivity limitations in CQDs, scalability challenges. |
| Synthesis and Automation (Stage 2–4) | Microfluidics, Robotics | Benefits: Precise fluid handling, small sample volumes, automated workflows. Limitations: Limited scalability, specialized fabrication required. | Quantum-assisted Microfluidics or Robotics Synthesis [286]; Quantum-enhanced process optimization | Benefits: Potential for quantum-level precision in synthesis, optimization of synthetic routes using hybrid quantum-classical algorithms. Limitations: Largely conceptual, limited experimental validation, high complexity. |
- Solving Complex Biological Challenges with Quantum Computing
- Ultra-Sensitive Quantum Sensing in Mechanistic Discovery
6. Molecule and Modality Design Layer
6.1. Modality as Strategic Constraint and Design
| Biomarker | Challenges | Classical AI Layer | Quantum AI Layer | Quantum Physical Layer |
|---|---|---|---|---|
| Antigen | Subtle changes; specificity vs. similar proteins [307]. | Predict antigenicity, epitopes; Natural Language Processing (NLP) for associations [308,309]. | Quantum algorithms simulate antigen–antibody binding and explore conformational spaces for epitope prediction [114,213]. | Quantum-enhanced plasmonic spectroscopy, including Surface Plasmon Resonance [237,310] and NV centers [41,311], could enable ultra-sensitive detection of antigen–antibody kinetics; and conformational changes; quantum photonic sensors support real-time epitope mapping [82,312]. |
| Protein & Peptides | Folding diversity; Post-Translational Modifications (PTMs) hard to detect [307]. | Deep learning for 3D structure; aggregation motif prediction. | Quantum algorithms model protein folding and PTMs [114,213,245] outperform classical methods in conformational sampling [313]. | Quantum Chemical Mass Spectrometry (QC-MS) [196,197], quantum dots [65,66], and hyperpolarized NMR with magnetometry enable PTM detection and folding state analysis. [209,210]. |
| Enzyme | Complex reaction pathways; inhibitor design [307]. | Docking for inhibitors; generative AI for active sites. | Quantum simulations model transition states and reaction pathways; quantum annealing accelerates inhibitor design [114,118,264,314]. | Quantum Cascade Lasers [42] and quantum dot nanosensors detect enzymatic activity [17,81,315]; NV centers monitor catalytic sites with high spatial resolution [316]. |
| Gene, Nucleic Acids | Regulatory complexity; off-target edits; epigenetic modulation [307]. | Network analysis; multimodal AI for mutations and methylation [18,57,134]. | Quantum neural networks discover genetic biomarkers in large search spaces; classical machine learning struggles with multi-omics [79,317]. | Quantum microscopy and quantum-enhanced fluorescence enable visualization of gene expression [318] and DNA damage [319,320]. |
| Ion Channels | Rapid conformational changes; subtype specificity [307]. | Time-series gating prediction; allosteric site screening. | Quantum simulations model gating dynamics and ion transport with higher fidelity than classical Molecular Dynamics [79,245]. | Quantum sensors detect weak electric fields [321,322], NV centers [41,45,64] and quantum dots [83,323,324] track gating and conformational changes, or ion channel gating. |
| Transporter | Drug binding/release; drug–drug interactions [307]. | Substrate prediction; Graph Neural Networks for transporter networks. | Quantum simulations model dynamic drug-transporter interactions and substrate flux [17,79]. | Quantum-enhanced microscopy and tunneling sensors detect substrate flux and pH gradients in transporter dynamics [17,79,325]. |
| Receptor | Allosteric modulation complexity [307]. | Predictive AI for allosteric sites; generative molecule design. | Quantum simulations [17,79]. Capture allosteric modulation effects and ligand-induced conformational shifts. | Quantum-enhanced SPR and single-molecule FRET detect binding kinetics and ligand-induced shifts; quantum field sensors monitor conformational changes [17,79]. |
| Chemical Biomarkers | Trace levels; oxidation state; metabolite profiling [307]. | Predictive analytics for disease correlation [18,134]. | Quantum simulations detect redox states and trace metabolites with high sensitivity [326]. | Quantum sensors (e.g., SQUIDs [47,191,192], NV centers [195,311], Quantum Cascade Lasers [42]) detect trace metabolites and redox states with ultra-sensitivity [196,197]. |
| Toxin | High potency; rapid detection in mixtures [307]. | Toxicity prediction [59,218]; symptom correlation via multimodal AI. | Quantum simulations [114,117] resolve toxin binding mechanisms and enable sorting in complex mixtures. | Quantum biosensors [65,66,211], and quantum microfluidics [17] enable rapid screening and sorting of toxins. [327,328]; Quantum Mass Spectrometry [196,197]. |
| Cell & Organelle | Heterogeneity; system-level modeling; imaging complexity [307]. | Image analysis; signaling network mapping [18,134]. | Quantum simulations build digital twins of cells and model electron transport across organelles [17,166,329]. | Quantum dots and microscopy enable organelle imaging [65,66,211]; hyperpolarized MRI/NMR and magnetometry map metabolic flux [209,210]. |
| Biological Factor | Broad category (hormones, cytokines, microbiome); feedback loops [307]. | Predict downstream effects; correlate with disease [18,134]. | Quantum computing models nonlinear feedback loops and high-dimensional biological networks [317]. | Quantum-enhanced sensors and interferometry detect hormones, cytokines, and microbial metabolites [17]. |
| Digital Biomarkers | Noise, artifacts, variability; linking signals to clinical outcomes [307]. | Signal processing; predictive modeling from wearables/apps. | Quantum ML [17,166,329] improves pattern recognition in noisy wearable data; classical ML struggles with real-time inference. | Quantum sensors in wearables; quantum encryption for secure data transmission; quantum accelerometers and magnetometers for physiological signal fidelity [17]. |
6.2. Molecule Synthesis and Retrosynthesis
- Advancing Retrosynthesis with AI and Quantum Computing
| Molecule Type | Challenges | Classical AI Layer | Quantum AI Layer | Quantum Physical Layer |
|---|---|---|---|---|
| RNA-Based Therapeutics (Antisense, siRNA, miRNA, mRNA) | Degradation; off-target binding; intracellular delivery. | Predictive modeling for degradation/off targets; generative AI for sequence/mRNA vaccine design. | Quantum simulation of RNA folding and hybridization; quantum annealing for guide optimization [17,341]. | SQUID [47] or NV center-based electric field sensing [41,316] enable degradation profiling and RNA folding analysis; quantum dots [65,66] support intracellular tracking and Quantum Mass Spectrometry [196,197]. |
| Antibody-Based Therapeutics (Monoclonal, ADCs) | Immunogenicity; binding specificity; payload delivery; manufacturing. | Generative AI for low-immunogenicity design; predictive analytics for affinity and conjugation [117,213]. | Quantum chemistry models binding energy and simulates mutation and linker behavior in antibody-drug conjugates [114,245]. | Quantum-enhanced plasmonic spectroscopy, including Surface Plasmon Resonance [237,310], biosensing improve binding kinetics and specificity; NV centers could detect conformational changes [17,41,311]. |
| Small Molecule Therapeutics (includes PROTACs) | ADMET prediction [59,218]; polymorphism; synthesis; targeted degradation. | Generative AI for ADMET and PROT40AC design; GNNs for interaction modeling [59,218]. | Quantum simulation of metabolism, polymorph prediction, and degrader dynamics [245]. | Quantum Cascade Lasers [42] and quantum-enhanced NMR [45,67] enable vibrational fingerprinting and purity analysis; quantum hydration mapping supports ligand binding. |
| Synthetic Peptides | Rapid degradation; folding complexity; delivery. | Deep learning for 3D structure (e.g., AlphaFold); predictive AI for stability [122,123,124]. | Quantum simulation of folding and peptide-membrane interactions [245,342]. | Quantum-enhanced microscopy and quantum dot sensors [65,66] visualize folding and track degradation; single-photon detection reveals conformational states. |
| Polysaccharides | Branching complexity; immunogenicity. | Predictive analytics for immune response; generative AI for novel structures. | Quantum simulation of branching and delivery optimization. | Raman spectroscopies analyze branching and immune profiles, NV centers map charge distribution. Quantum Mass Spectrometry [196,197]. |
| Vaccines (including mRNA) | Immunogenicity; degradation; targeted delivery. | Generative AI for mRNA/epitope design; deep learning for immunogenicity. | Quantum simulation of vaccine-receptor interactions; adjuvant design [343]. | Quantum dot [65,66] tracking and NV center-based sensing monitor payload distribution and immune activation; quantum-enhanced cytokine assays improve profiling [17]. |
| Cell Therapy | Batch variability; targeting; patient-specific efficacy. | Multimodal AI for response prediction [134,344]; image analysis for QC. | Quantum digital twins [345] simulate cell behavior and optimize manufacturing. | Quantum microscopy and sensors monitor cell tracking and metabolic states; molecular quantum sensors assess membrane protein dynamics [17]. |
| Gene Therapy and Editing (CRISPR, TALENs) | Off-target integration; vector safety; immunogenicity. | Predictive AI for vector design; deep learning for off-target prediction. | Quantum annealing [17] for CRISPR guide optimization; simulation of editing mechanics. | Quantum-enhanced microscopy (e.g., NV centers and Quantum dot) detects gene expression and vector delivery [17]. |
| Fusion/Recombinant Proteins | Folding accuracy; aggregation; immunogenicity. | Deep learning for structure; predictive analytics for aggregation [59,124]. | Quantum simulation of folding and aggregation; quantum chemistry for binding energy [114,245]. | NV centers [41,316] and quantum fluorescence probe [65,66] assess folding states. Quantum Mass Spectrometry [196,197] detects aggregation. |
| Allergenics | Allergen identification; desensitization; batch consistency. | Predictive AI for epitope identification; multimodal AI for patient correlation. | Quantum simulation of allergen-receptor interactions and mixture modeling. | Quantum-enhanced assays [346] and Quantum-enhanced NMR [45,67] ensure batch consistency; biosensors detect hypersensitivity responses. |
| Biologics (proteins, antibodies, enzymes) | Structural complexity; stability; immunogenicity; scalability. | Generative AI for novel biologics; predictive analytics for stability or sequence [17,59,213]. | Quantum simulation [114,213,245] of folding, catalysis, and manufacturing optimization. | Quantum mass spectrometry and molecular sensors assess purity and stability; Quantum Cascade Lasers [42] and NV centers [41,316] and Quantum Dots [81,315]; monitor enzymatic activity and conformational dynamics. |
- The Computational Challenges for Accurate Molecular Prediction
- Applicability to High-Accuracy Methods
- Advanced Synthesis and Optimization
6.3. Formulation and Stability
7. Feedback Loops: From Broken Pipelines to a Learning System
8. Functional Landscape
8.1. Emerging AI Contributions
8.2. Emerging Quantum Contributions
- Quantum Computing landscape
- Quantum Sensing and Quantum Control Landscape
- Emerging Hybrid Quantum-AI Clinical Trials
8.3. Hybrid Quantum–AI Ecosystems
8.4. The Hybrid Future Quantum-AI Convergence in Pharmaceuticals
9. Operationalizing and Governing Pharmaceutical Innovation
9.1. Operational Complexities and Adoption Hurdles
- Infrastructure, Financial, and Logistical Hurdles
- Human Capital and Procedural Challenges
9.2. Socioethical, Regulatory, and Commercial Dynamics
- Ethical and Regulatory Challenges
- Commercialization Dynamics and Ethical Governance
9.3. Decentralizing and Democratizing Innovation for Global Health
- The Foundational Data Sovereignty Kit
- Building a Distributed Network
- Advanced Integration and Governance
10. Strategic Roadmaps and Policy Solutions
10.1. Strategic and Institutional Hurdles
10.2. Policy Solutions and Roadmaps
10.3. Roadmap by Timeline
- Immediate and Near-Term (0–3 years)
- Mid-Term (3–7 years)
- Long-Term (7–10+ years)
- Grand Outlook Challenge: Energy and Computational Scale
11. Conclusions
Supplementary Materials
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| VQE | Variational Quantum Eigensolver |
| QAOA | Quantum Approximate Optimization Algorithm |
| QITE | Quantum Imaginary Time Evolution |
| QKD | Quantum Key Distribution |
| FAIR | Findable, Accessible, Interoperable, and Reusable |
| FHE | Fully Homomorphic Encryption |
| NV | Nitrogen-Vacancy |
| SNSPD | Superconducting Nanowire Single-Photon Detector |
| QC-MS | Quantum Computing Mass Spectrometry |
References
- Khanna, I. Drug discovery in pharmaceutical industry: Productivity challenges and trends. Drug Discov. Today 2012, 17, 1088–1102. [Google Scholar] [CrossRef]
- Sun, D.; Gao, W.; Hu, H.; Zhou, S. Why 90% of clinical drug development fails and how to improve it? Acta Pharm. Sin. B 2022, 12, 3049–3062. [Google Scholar] [CrossRef]
- Villoutreix, B.O. Post-Pandemic Drug Discovery and Development: Facing Present and Future Challenges. Front. Drug Discov. 2021, 1, 728469. [Google Scholar] [CrossRef]
- Ioannidis, J.P.A.; Bossuyt, P.M.M. Waste, leaks, and failures in the biomarker pipeline. Clin. Chem. 2017, 63, 963–972. [Google Scholar] [CrossRef] [PubMed]
- Hassan Baig, M.; Ahmad, K.; Roy, S.; Ashraf, J.M.; Adil, M.; Siddiqui, M.H.; Khan, S.; Kamal, M.A.; Provazník, I.; Choi, I. Computer Aided Drug Design: Success and Limitations. Curr. Pharm. Des. 2016, 22, 572–581. [Google Scholar] [CrossRef]
- von Korff, M.; Sander, T. Limits of Prediction for Machine Learning in Drug Discovery. Front. Pharmacol. 2022, 13, 832120. [Google Scholar] [CrossRef]
- Simoens, S.; Huys, I. R&D Costs of New Medicines: A Landscape Analysis. Front. Med. 2021, 8, 760762. [Google Scholar] [CrossRef] [PubMed]
- Kalepu, S.; Nekkanti, V. Insoluble drug delivery strategies: Review of recent advances and business prospects. Acta Pharm. Sin. B 2015, 5, 442–453. [Google Scholar] [CrossRef]
- Censi, R.; Di Martino, P. Polymorph impact on the bioavailability and stability of poorly soluble drugs. Molecules 2015, 20, 18759–18776. [Google Scholar] [CrossRef] [PubMed]
- Rahban, M.; Ahmad, F.; Piatyszek, M.A.; Haertlé, T.; Saso, L.; Saboury, A.A. Stabilization challenges and aggregation in protein-based therapeutics in the pharmaceutical industry. RSC Adv. 2023, 13, 35947–35963. [Google Scholar] [CrossRef] [PubMed]
- Pushkaran, A.C.; Arabi, A.A. From understanding diseases to drug design: Can artificial intelligence bridge the gap? Artif. Intell. Rev. 2024, 57, 86. [Google Scholar] [CrossRef]
- Paul, D.; Sanap, G.; Shenoy, S.; Kalyane, D.; Kalia, K.; Tekade, R.K. Artificial intelligence in drug discovery and development. Drug Discov. Today 2021, 26, 80–93. [Google Scholar] [CrossRef]
- Miller, F.A.; Young, S.B.; Dobrow, M.; Shojania, K.G. Vulnerability of the medical product supply chain: The wake-up call of COVID-19. BMJ Qual. Saf. 2021, 30, 331–335. [Google Scholar] [CrossRef]
- Mulumba, M.; Oga, J.; Koomson, N.; Kara, T.A.; Cynthia, A.N.; Forman, L. Decolonizing global health: Africa’s pursuit of pharmaceutical sovereignty. BMC Health Serv. Res. 2025, 25, 1015. [Google Scholar] [CrossRef]
- Fadojutimi, B. Exploring the critical factors in pharmaceutical supply chains revealed during COVID-19 and addressing missing links. World J. Adv. Res. Rev. 2024, 24, 2407–2425. [Google Scholar] [CrossRef]
- Cova, T.; Vitorino, C.; Ferreira, M.; Nunes, S.; Rondon-Villarreal, P.; Pais, A. Artificial Intelligence and Quantum Computing as the Next Pharma Disruptors. In Methods in Molecular Biology; Humana: New York, NY, USA, 2022. [Google Scholar]
- Matarèse, B.F.E.; Purushotham, A. Quantum Oncology. Quantum Rep. 2025, 7, 9. [Google Scholar] [CrossRef]
- Narayanan, R.; Peker, E.; DeGroat, W.; Mendhe, D.; Zeeshan, S.; Ahmed, Z. 3D IntelliGenes: AI/ML application using multi-omics data for biomarker discovery and disease prediction with multi-dimensional visualization. BMC Med. Res. Methodol. 2025, 25, 193. [Google Scholar] [CrossRef]
- Maskara, N.; Ostermann, S.; Shee, J.; Kalinowski, M.; Gomez, A.M.; Bravo, R.A.; Wang, D.S.; Krylov, A.I.; Yao, N.Y.; Head-Gordon, M.; et al. Programmable simulations of molecules and materials with reconfigurable quantum processors. Nat. Phys. 2025, 21, 289–297. [Google Scholar] [CrossRef]
- Clinton, L.; Cubitt, T.; Flynn, B.; Gambetta, F.M.; Klassen, J.; Montanaro, A.; Piddock, S.; Santos, R.A.; Sheridan, E. Towards near-term quantum simulation of materials. Nat. Commun. 2024, 15, 211. [Google Scholar] [CrossRef] [PubMed]
- Kadowaki, T. Quantum Computing and AI: Perspectives on Advanced Automation in Science and Engineering. arXiv 2025, arXiv:2505.10012. [Google Scholar] [CrossRef]
- Braga, D.M.; Rawal, B. Harnessing AI and Quantum Computing for Revolutionizing Drug Discovery and Approval Processes: Case Example for Collagen Toxicity. JMIR Bioinform. Biotechnol. 2025, 6, e69800. [Google Scholar] [CrossRef] [PubMed]
- Plé, T.; Adjoua, O.; Benali, A.; Posenitskiy, E.; Villot, C.; Lagardère, L.; Piquemal, J.-P. A Foundation Model for Accurate Atomistic Simulations in Drug Design. ChemRxiv 2025. [Google Scholar] [CrossRef]
- Zhou, Y.; Stoudenmire, E.M.; Waintal, X. What Limits the Simulation of Quantum Computers? Phys. Rev. X 2020, 10, 041038. [Google Scholar] [CrossRef]
- Strocchi, F. Microscopic and macroscopic quantities in statistical mechanics. Il Nuovo C. B Ser. 10 1970, 65, 239–265. [Google Scholar] [CrossRef]
- Chae, E.; Choi, J.; Kim, J. An elementary review on basic principles and developments of qubits for quantum computing. Nano Converg. 2024, 11, 11. [Google Scholar] [CrossRef]
- Zhang, H. Quantum Entanglement and Qubit Interactions: The Key to Quantum Supremacy. Theor. Nat. Sci. 2024, 41, 112–118. [Google Scholar] [CrossRef]
- Shen, L.; Yang, W. Molecular Dynamics Simulations with Quantum Mechanics/Molecular Mechanics and Adaptive Neural Networks. J. Chem. Theory Comput. 2018, 14, 1442–1455. [Google Scholar] [CrossRef]
- Kulkarni, P.U.; Shah, H.; Vyas, V.K. Hybrid Quantum Mechanics/Molecular Mechanics (QM/MM) Simulation: A Tool for Structure-Based Drug Design and Discovery. Mini Rev. Med. Chem. 2021, 22, 1096–1107. [Google Scholar] [CrossRef] [PubMed]
- Schrödinger, E. Quantisierung als Eigenwertproblem. Ann. Phys. 1926, 384, 361–376. [Google Scholar] [CrossRef]
- Oszmaniec, M.; Grudka, A.; Horodecki, M.; Wójcik, A. Creating a Superposition of Unknown Quantum States. Phys. Rev. Lett. 2016, 116, 110403. [Google Scholar] [CrossRef]
- Freedman, S.J.; Clauser, J.F. Experimental test of local hidden-variable theories. Phys. Rev. Lett. 1972, 28, 938–941. [Google Scholar] [CrossRef]
- Aspect, A.; Dalibard, J.; Roger, G. Experimental test of Bell’s inequalities using time- varying analyzers. Phys. Rev. Lett. 1982, 49, 1804–1807. [Google Scholar] [CrossRef]
- Pan, J.W.; Bouwmeester, D.; Weinfurter, H.; Zeilinger, A. Experimental entanglement swapping: Entangling photons that never interacted. Phys. Rev. Lett. 1998, 80, 3891–3894. [Google Scholar] [CrossRef]
- Gamow, G. Zur Quantentheorie des Atomkernes. Z. Phys. 1928, 51, 204–212. [Google Scholar] [CrossRef]
- Gurney, R.W.; Condon, E.U. Quantum mechanics and radioactive disintegration. Phys. Rev. 1929, 33, 127–140. [Google Scholar] [CrossRef]
- Wrachtrup, J.; Finkler, A. Single spin magnetic resonance. J. Magn. Reson. 2016, 269, 225–236. [Google Scholar] [CrossRef]
- Kaushik, P.; Rao, A.M.; Singh, D.P.; Vashisht, S.; Gupta, S. Cloud Computing and Comparison based on Service and Performance between Amazon AWS, Microsoft Azure, and Google Cloud. In Proceedings of the 2021 International Conference on Technological Advancements and Innovations (ICTAI), Tashkent, Uzbekistan, 10–12 November 2021; IEEE: Piscataway, NJ, USA, 2021; pp. 268–273. [Google Scholar]
- Pajankar, A.; Joshi, A. Hands-on Machine Learning with Python: Implement Neural Network Solutions with Scikit-Learn and PyTorch; Apress: New York, NY, USA, 2022. [Google Scholar]
- Liliopoulos, I.; Varsamis, G.D.; Karamanidou, T.; Papalitsas, C.; Koulouras, G.; Pantazopoulos, V.; Stavropoulos, T.G.; Karafyllidis, I.G. Quantum algorithm for protein-ligand docking sites identification in the interaction space. J. Comput. Aided Mol. Des. 2025, 39, 40. [Google Scholar] [CrossRef]
- Du, J.; Shi, F.; Kong, X.; Jelezko, F.; Wrachtrup, J. Single-molecule scale magnetic resonance spectroscopy using quantum diamond sensors. Rev. Mod. Phys. 2024, 96, 025001. [Google Scholar] [CrossRef]
- Bird, B.; Baker, M.J. Quantum Cascade Lasers in Biomedical Infrared Imaging. Trends Biotechnol. 2015, 33, 557–558. [Google Scholar] [CrossRef]
- Chemla, Y.R.; Grossman, H.L.; Poon, Y.; McDermott, R.; Stevens, R.; Alper, M.D.; Clarke, J. Ultrasensitive magnetic biosensor for homogeneous immunoassay. Proc. Natl. Acad. Sci. USA 2000, 97, 14268–14272. [Google Scholar] [CrossRef]
- Meinel, J.; Kwon, M.; Maier, R.; Dasari, D.; Sumiya, H.; Onoda, S.; Isoya, J.; Vorobyov, V.; Wrachtrup, J. High-resolution nanoscale NMR for arbitrary magnetic fields. Commun. Phys. 2023, 6, 302. [Google Scholar] [CrossRef]
- Bucher, D.B.; Aude Craik, D.P.L.; Backlund, M.P.; Turner, M.J.; Dor, O.B.; Glenn, D.R.; Walsworth, R.L. Quantum diamond spectrometer for nanoscale NMR and ESR spectroscopy. Nat. Protoc. 2019, 14, 2707–2747. [Google Scholar] [CrossRef]
- Zwick, A.; Álvarez, G.A. Quantum sensing tools to characterize physical, chemical and biological processes with magnetic resonance. J. Magn. Reson. Open 2023, 16–17, 100113. [Google Scholar] [CrossRef]
- Wu, S.; Gong, C.; Li, S.; Ni, R.; Zhu, J. Weak Radio Frequency Signal Reception Sensitivity Based on DC Superconducting Quantum Interference Devices. IEEE Trans. Appl. Supercond. 2023, 33, 1–8. [Google Scholar] [CrossRef]
- Matarèse, B.F.E.; Purushotham, A. Unlocking clinical quantum oncology through quantum control. Eur. J. Cancer 2025, 226, 115632. [Google Scholar] [CrossRef]
- Chang, B.Y.; Sola, I.R.; Malinovsky, V.S. Anomalous Rabi Oscillations in Multilevel Quantum Systems. Phys. Rev. Lett. 2018, 120, 133201. [Google Scholar] [CrossRef] [PubMed]
- Wigger, D.; Schneider, C.; Gerhardt, S.; Kamp, M.; Höfling, S.; Kuhn, T.; Kasprzak, J. Rabi oscillations of a quantum dot exciton coupled to acoustic phonons: Coherence and population readout. Optica 2018, 5, 1442–1450. [Google Scholar] [CrossRef]
- Daems, D.; Ruschhaupt, A.; Sugny, D.; Guérin, S. Robust quantum control by a single-shot shaped pulse. Phys. Rev. Lett. 2013, 111, 050404. [Google Scholar] [CrossRef] [PubMed]
- Torosov, B.T.; Vitanov, N.V. Smooth composite pulses for high-fidelity quantum information processing. Phys. Rev. A 2011, 83, 053420. [Google Scholar] [CrossRef]
- Ndong, M.; Djotyan, G.; Ruschhaupt, A.; Guérin, S. Robust coherent superposition of states by single-shot shaped pulse. J. Phys. B At. Mol. Opt. Phys. 2015, 48, 174007. [Google Scholar] [CrossRef]
- Yin, Z.; Li, C.; Allcock, J.; Zheng, Y.; Gu, X.; Dai, M.; Zhang, S.; An, S. Shortcuts to adiabaticity for open systems in circuit quantum electrodynamics. Nat. Commun. 2022, 13, 188. [Google Scholar] [CrossRef]
- Rangelov, A.A.; Vitanov, N.V.; Yatsenko, L.P.; Shore, B.W.; Halfmann, T.; Bergmann, K. Stark-shift-chirped rapid-adiabatic-passage technique among three states. Phys. Rev. A 2005, 72, 053403. [Google Scholar] [CrossRef]
- Gregefalk, A.; Sjöqvist, E. Transitionless Quantum Driving in Spin Echo. Phys. Rev. Appl. 2022, 17, 024012. [Google Scholar] [CrossRef]
- Lin, M.; Guo, J.; Gu, Z.; Tang, W.; Tao, H.; You, S.; Jia, D.; Sun, Y.; Jia, P. Machine learning and multi-omics integration: Advancing cardiovascular translational research and clinical practice. J. Transl. Med. 2025, 23, 388. [Google Scholar] [CrossRef] [PubMed]
- Chandra, A.; Tünnermann, L.; Löfstedt, T.; Gratz, R. Transformer-based deep learning for predicting protein properties in the life sciences. eLife 2023, 12, e82819. [Google Scholar] [CrossRef]
- de Sá, A.G.C.; Ascher, D.B. Auto-ADMET: An Effective and Interpretable AutoML Method for Chemical ADMET Property Prediction. arXiv 2025, arXiv:2502.16378. [Google Scholar]
- Thiele, M.; Villesen, I.F.; Niu, L.; Johansen, S.; Sulek, K.; Nishijima, S.; Espen, L.V.; Keller, M.; Israelsen, M.; Suvitaival, T.; et al. Opportunities and barriers in omics-based biomarker discovery for steatotic liver diseases. J. Hepatol. 2024, 81, 345–359. [Google Scholar] [CrossRef] [PubMed]
- Paananen, J.; Fortino, V. An omics perspective on drug target discovery platforms. Brief. Bioinform. 2020, 21, 1937–1953. [Google Scholar] [CrossRef] [PubMed]
- Raniga, K.; Stebbeds, W.; Shivalingam, A.; Pembertond, M.; Denning, C. Design of experiments for the automated development of a multicellular cardiac model for high-throughput screening. SLAS Discov. 2023, 28, 410–417. [Google Scholar] [CrossRef]
- Yang, Z.; Ren, Z.; Cheng, Y.; Sun, W.; Xi, Z.; Jia, W.; Li, G.; Wang, Y.; Guo, M.; Li, D. Review and prospect on portable mass spectrometer for recent applications. Vacuum 2022, 199, 110889. [Google Scholar] [CrossRef]
- Leung, H.M.; Chu, H.C.; Mao, Z.W.; Lo, P.K. Versatile nanodiamond-based tools for therapeutics and bioimaging. Chem. Commun. 2023, 59, 2039–2055. [Google Scholar] [CrossRef]
- Truskewycz, A.; Yin, H.; Halberg, N.; Lai, D.T.H.; Ball, A.S.; Truong, V.K.; Rybicka, A.M.; Cole, I. Carbon Dot Therapeutic Platforms: Administration, Distribution, Metabolism, Excretion, Toxicity, and Therapeutic Potential. Small 2022, 18, 2106342. [Google Scholar] [CrossRef] [PubMed]
- Zrazhevskiy, P.; Sena, M.; Gao, X. Designing multifunctional quantum dots for bioimaging, detection, and drug delivery. Chem. Soc. Rev. 2010, 39, 4326–4354. [Google Scholar] [CrossRef]
- Liu, K.S.; Henning, A.; Heindl, M.W.; Bucher, D.B. Surface NMR using quantum sensors in diamond. Proc. Natl. Acad. Sci. USA 2022, 119, e2111607119. [Google Scholar] [CrossRef]
- Zhou, C.; Hermes, M.R.; Wu, D.; Bao, J.J.; Pandharkar, R.; King, D.S.; Zhang, D.; Scott, T.R.; Lykhin, A.O.; Gagliardi, L.; et al. Electronic structure of strongly correlated systems: Recent developments in multiconfiguration pair-density functional theory and multiconfiguration nonclassical-energy functional theory. Chem. Sci. 2022, 13, 7685–7706. [Google Scholar] [CrossRef]
- Schmidt-Böcking, H.; Eckart, S.; Lüdde, H.J.; Gruber, G.; Jahnke, T. The Precision Limits in a Single-Event Quantum Measurement of Electron Momentum and Position. In Molecular Beams in Physics and Chemistry: From Otto Stern’s Pioneering Exploits to Present-Day Feat; Springer: Berlin/Heidelberg, Germany, 2021. [Google Scholar]
- Lee, J.; Berry, D.W.; Gidney, C.; Huggins, W.J.; McClean, J.R.; Wiebe, N.; Babbush, R.B. Even More Efficient Quantum Computations of Chemistry through Tensor Hypercontraction. PRX Quantum 2021, 2, 030305. [Google Scholar] [CrossRef]
- Yu, S.; Zhong, Z.-P.; Fang, Y.; Patel, R.B.; Li, Q.-P.; Liu, W.; Li, Z.; Xu, L.; Sagona-Stophel, S.; Mer, E.; et al. A universal programmable Gaussian boson sampler for drug discovery. Nat. Comput. Sci. 2023, 3, 839–848. [Google Scholar] [CrossRef]
- Lau, J.W.Z.; Lim, K.H.; Shrotriya, H.; Kwek, L.C. NISQ computing: Where are we and where do we go? AAPPS Bull. 2022, 32, 27. [Google Scholar] [CrossRef]
- Schuld, M.; Killoran, N. Quantum Machine Learning in Feature Hilbert Spaces. Phys. Rev. Lett. 2019, 122, 040504. [Google Scholar] [CrossRef] [PubMed]
- Fellous-Asiani, M.; Chai, J.H.; Whitney, R.S.; Auffèves, A.; Ng, H.K. Limitations in Quantum Computing from Resource Constraints. PRX Quantum 2021, 2, 040335. [Google Scholar] [CrossRef]
- Westerberg, H. The Superintelligence That Cares About Us; GitHub: San Francisco, CA, USA, 2025. [Google Scholar]
- Talati, D. Quantum Minds: Merging Quantum Computing with Next-Gen AI. SSRN 2025. [Google Scholar] [CrossRef]
- Holdgate, G.; Embrey, K.; Milbradt, A.; Davies, G. Biophysical methods in early drug discovery. ADMET DMPK 2019, 7, 222–241. [Google Scholar] [CrossRef]
- Iyengar, R. Computational biochemistry: Systems biology minireview series. J. Biol. Chem. 2009, 284, 5425–5426. [Google Scholar] [CrossRef]
- Matarèse, B.F.E.; Rusin, A.; Seymour, C.; Mothersill, C. Quantum Biology and the Potential Role of Entanglement and Tunneling in Non-Targeted Effects of Ionizing Radiation: A Review and Proposed Model. Int. J. Mol. Sci. 2023, 24, 16464. [Google Scholar] [CrossRef] [PubMed]
- Iyer, G.; Michalet, X.; Chang, Y.P.; Weiss, S. Tracking Single Proteins in Live Cells Using Single-Chain Antibody Fragment-Fluorescent Quantum Dot Affinity Pair. Methods Enzymol. 2010, 475, 61–79. [Google Scholar]
- Knudsen, B.R.; Jepsen, M.L.; Ho, Y.P. Quantum dot-based nanosensors for diagnosis via enzyme activity measurement. Expert Rev. Mol. Diagn. 2013, 13, 367–375. [Google Scholar] [CrossRef] [PubMed]
- Mousavi, S.M.; Hashemi, S.A.; Yari Kalashgrani, M.; Omidifar, N.; Lai, C.W.; Vijayakameswara Rao, N.; Gholami, A.; Chiang, W.H. The Pivotal Role of Quantum Dots-Based Biomarkers Integrated with Ultra-Sensitive Probes for Multiplex Detection of Human Viral Infections. Pharmaceuticals 2022, 15, 880. [Google Scholar] [CrossRef] [PubMed]
- Sotoma, S.; Okita, H.; Chuma, S.; Harada, Y. Quantum nanodiamonds for sensing of biological quantities: Angle, temperature, and thermal conductivity. Biophys. Physicobiol. 2022, 19, e190034. [Google Scholar] [CrossRef]
- Luo, S.; Sun, Y. Quantum coherence versus quantum uncertainty. Phys. Rev. A 2017, 96, 022130. [Google Scholar] [CrossRef]
- Bischof, F.; Kampermann, H.; Bru, D. Quantifying coherence with respect to general quantum measurements. Phys. Rev. A 2021, 103, 032429. [Google Scholar] [CrossRef]
- Pullman, A.; Pullman, B. Electronic Structure and Carcinogenic Activity of Aromatic Molecules New Developments. Adv. Cancer Res. 1955, 3, 117–169. [Google Scholar] [CrossRef]
- Kar, R.K. Benefits of hybrid QM/MM over traditional classical mechanics in pharmaceutical systems. Drug Discov. Today 2023, 28, 103374. [Google Scholar]
- Cheng, H.P.; Deumens, E.; Freericks, J.K.; Li, C.; Sanders, B.A. Application of Quantum Computing to Biochemical Systems: A Look to the Future. Front. Chem. 2020, 8, 587143. [Google Scholar] [CrossRef]
- Rice, S.A.; Tannor, D.J.; Kosloff, R. Coherent pulse sequence induced control of selectivity of reactions. Exact quantum-mechanical calculations. J. Chem. Soc. Faraday Trans. 2 1986, 82, 2423. [Google Scholar] [CrossRef]
- Wang, J.; Li, D.; Betzholz, R.; Cai, J. Real-Time Adaptive Sensing of Nuclear Spins by a Single-Spin Quantum Sensor. Phys. Rev. Appl. 2022, 18, 024040. [Google Scholar] [CrossRef]
- Li, H.; Gong, X.; Ni, H.; Lu, P.; Luo, X.; Wen, J.; Yang, Y.; Qian, X.; Sun, Z.; Wu, J. Light-Induced Ultrafast Molecular Dynamics: From Photochemistry to Optochemistry. J. Phys. Chem. Lett. 2022, 13, 5881–5893. [Google Scholar] [CrossRef] [PubMed]
- Sultanow, E.; Tehrani, M.; Dutta, S.; Buchanan, W.J.; Khan, M.S. Quantum Agents. arXiv 2025, arXiv:2506.01536. [Google Scholar] [PubMed]
- Mione, F.M.; Kaspersetz, L.; Luna, M.F.; Aizpuru, J.; Scholz, R.; Borisyak, M.; Kemmer, A.; Schermeyer, M.T.; Martinez, E.C.; Neubauer, P.; et al. A workflow management system for reproducible and interoperable high-throughput self-driving experiments. Comput. Chem. Eng. 2024, 187, 108720. [Google Scholar] [CrossRef]
- Shahin, M.H.; Goswami, S.; Lobentanzer, S.; Corrigan, B.W. Agents for Change: Artificial Intelligent Workflows for Quantitative Clinical Pharmacology and Translational Sciences. Clin. Transl. Sci. 2025, 18, e70188. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Huang, K.; Yang, D.; Zhao, W.; Zhou, X. Biomedical Big Data Technologies, Applications, and Challenges for Precision Medicine: A Review. Glob. Chall. 2024, 8, 2300163. [Google Scholar]
- Swift, B.; Jain, L.; White, C.; Chandrasekaran, V.; Bhandari, A.; Hughes, D.A.; Jadhav, P.R. Innovation at the Intersection of Clinical Trials and Real-World Data Science to Advance Patient Care. Clin. Transl. Sci. 2018, 11, 450–460. [Google Scholar] [CrossRef] [PubMed]
- Chen, K.Y.; Borglund, E.M.; Postema, E.C.; Dunn, A.G.; Bourgeois, F.T. Reporting of clinical trial safety results in ClinicalTrials.gov for FDA-approved drugs: A cross-sectional analysis. Clin. Trials 2022, 19, 442–451. [Google Scholar] [CrossRef]
- Ding, H.; Wu, F. Image guided biodistribution and pharmacokinetic studies of theranostics. Theranostics 2012, 2, 1040–1053. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.; Anwar, S.; Choudhury, A.; Mohammad, T.; Hassan, M.I. Validation of Drug Targets Using Molecular Methodologies and Enzymatic Activity Assays for Validation of Inhibitory Potential. In Bacterial Enzymes as Targets for Drug Discovery; Elsevier: Amsterdam, The Netherlands, 2025; pp. 91–112. [Google Scholar]
- Wilson, L.J.; Kiffer, F.C.; Berrios, D.C.; Bryce-Atkinson, A.; Costes, S.V.; Gevaert, O.; Matarèse, B.F.E.; Miller, J.; Mukherjee, P.; Peach, K.; et al. Machine intelligence for radiation science: Summary of the Radiation Research Society 67th annual meeting symposium. Int. J. Radiat. Biol. 2023, 99, 1291–1300. [Google Scholar] [CrossRef]
- Lavertu, A.; Vora, B.; Giacomini, K.M.; Altman, R.; Rensi, S. A New Era in Pharmacovigilance: Toward Real-World Data and Digital Monitoring. Clin. Pharmacol. Ther. 2021, 109, 1197–1202. [Google Scholar] [CrossRef]
- Ng, D.Q.; Dang, E.; Chen, L.; Nguyen, M.T.; Nguyen, M.K.N.; Samman, S.; Nguyen, T.M.T.; Cadiz, C.L.; Nguyen, L.; Chan, A. Current and recommended practices for evaluating adverse drug events using electronic health records: A systematic review. JACCP J. Am. Coll. Clin. Pharm. 2021, 4, 1457–1468. [Google Scholar] [CrossRef]
- Agarwal, P.; Searls, D.B. Can literature analysis identify innovation drivers in drug discovery? Nat. Rev. Drug Discov. 2009, 8, 865–878. [Google Scholar] [CrossRef]
- Gadiya, Y.; Gribbon, P.; Hofmann-Apitius, M.; Zaliani, A. Pharmaceutical patent landscaping: A novel approach to understand patents from the drug discovery perspective. Artif. Intell. Life Sci. 2023, 3, 100069. [Google Scholar] [CrossRef]
- Long, R.A.; Ballard, S.; Shah, S.; Bianchi, O.; Jones, L.; Koretsky, M.J.; Kuznetsov, N.; Marsan, E.; Jen, B.; Chiang, P.; et al. A new AI-assisted data standard accelerates interoperability in biomedical research. medRxiv 2024. [Google Scholar] [CrossRef] [PubMed]
- Tabari, P.; Costagliola, G.; De Rosa, M.; Boeker, M. State-of-the-Art Fast Healthcare Interoperability Resources (FHIR)-Based Data Model and Structure Implementations: Systematic Scoping Review. JMIR Med. Inform. 2024, 12, e58445. [Google Scholar] [CrossRef]
- Babuc, D.; Fortiş, T.-F. Federated Learning Platforms for Privacy-Preserving Histopathological Image Classification; Springer: Cham, Switzerland, 2025; pp. 317–328. [Google Scholar]
- Sinaci, A.A.; Gencturk, M.; Teoman, H.A.; Laleci Erturkmen, G.B.; Alvarez-Romero, C.; Martinez-Garcia, A.; Poblador-Plou, B.; Carmona-Pírez, J.; Löbe, M.; Parra-Calderon, C.L. A Data Transformation Methodology to Create Findable, Accessible, Interoperable, and Reusable Health Data: Software Design, Development, and Evaluation Study. J. Med. Internet Res. 2023, 25, e42822. [Google Scholar] [CrossRef] [PubMed]
- Zheng, L.; Cao, Y.; Yoshikawa, M.; Shen, Y.; Rashed, E.A.; Taura, K.; Hanaoka, S.; Zhang, T. Sensitivity-Aware Differential Privacy for Federated Medical Imaging. Sensors 2025, 25, 2847. [Google Scholar] [CrossRef] [PubMed]
- Cross, J.L.; Choma, M.A.; Onofrey, J.A. Bias in medical AI: Implications for clinical decision-making. PLoS Digit. Health 2024, 3, e0000651. [Google Scholar] [CrossRef]
- Hanna, M.G.; Pantanowitz, L.; Jackson, B.; Palmer, O.; Visweswaran, S.; Pantanowitz, J.; Deebajah, M.; Rashidi, H.H. Ethical and Bias Considerations in Artificial Intelligence/Machine Learning. Mod. Pathol. 2025, 38, 100686. [Google Scholar] [CrossRef]
- Shah, H.A.; Liu, J.; Yang, Z.; Feng, J. Review of Machine Learning Methods for the Prediction and Reconstruction of Metabolic Pathways. Front. Mol. Biosci. 2021, 8, 634141. [Google Scholar] [CrossRef]
- Cit, A.; Elly, B.; Alam, D. AI-Driven Optimization of Drug Synthesis Pathways. Preprints 2025, 2025020911. [Google Scholar] [CrossRef]
- Molani, F.; Cho, A.E. Accurate protein-ligand binding free energy estimation using QM/MM on multi-conformers predicted from classical mining minima. Commun. Chem. 2024, 7, 247. [Google Scholar] [CrossRef]
- Neves, B.J.; Braga, R.C.; Melo-Filho, C.C.; Moreira-Filho, J.T.; Muratov, E.N.; Andrade, C.H. QSAR-Based Virtual Screening: Advances and Applications in Drug Discovery. Front. Pharmacol. 2018, 9, 1275. [Google Scholar] [CrossRef]
- Brnabic, A.; Hess, L.M. Systematic literature review of machine learning methods used in the analysis of real-world data for patient-provider decision making. BMC Med. Inform. Decis. Mak. 2021, 21, 54. [Google Scholar] [CrossRef]
- Bhatia, A.S.; Saggi, M.K.; Kais, S. Quantum Machine Learning Predicting ADME-Tox Properties in Drug Discovery. J. Chem. Inf. Model. 2023, 63, 6476–6486. [Google Scholar] [CrossRef]
- Wei, Y.; Shan, L.; Qiu, T.; Lu, D.; Liu, Z. Machine learning-assisted retrosynthesis planning: Current status and future prospects. Chin. J. Chem. Eng. 2025, 77, 273–292. [Google Scholar] [CrossRef]
- Raschka, S.; Kaufman, B. Machine learning and AI-based approaches for bioactive ligand discovery and GPCR-ligand recognition. Methods 2020, 180, 89–110. [Google Scholar] [CrossRef] [PubMed]
- Das, U. Generative AI for drug discovery and protein design: The next frontier in AI-driven molecular science. Med. Drug Discov. 2025, 27, 100213. [Google Scholar] [CrossRef]
- Kapoor, A.; Gulli, A.; Pal, S.; Chollet, F. Build and Deploy Supervised, Unsupervised, Deep, and Reinforcement Learning Models. In Deep Learning with TensorFlow and Keras, 3rd ed.; Packt Publishing Ltd.: Birmingham, UK, 2022. [Google Scholar]
- Nussinov, R.; Zhang, M.; Liu, Y.; Jang, H. AlphaFold, Artificial Intelligence (AI), and Allostery. J. Phys. Chem. B 2022, 126, 6372–6383. [Google Scholar] [CrossRef] [PubMed]
- Terwilliger, T.C.; Poon, B.K.; Afonine, P.V.; Schlicksup, C.J.; Croll, T.I.; Millán, C.; Richardson, J.S.; Read, R.J.; Adams, P.D. Improved AlphaFold modeling with implicit experimental information. Nat. Methods 2022, 19, 1376–1382. [Google Scholar] [CrossRef]
- Pak, M.A.; Markhieva, K.A.; Novikova, M.S.; Petrov, D.S.; Vorobyev, I.S.; Maksimova, E.S.; Kondrashov, F.A.; Ivankov, D.N. Using AlphaFold to predict the impact of single mutations on protein stability and function. PLoS ONE 2023, 18, e0282689. [Google Scholar] [CrossRef]
- Korshunova, M.; Huang, N.; Capuzzi, S.; Radchenko, D.S.; Savych, O.; Moroz, Y.S.; Wells, C.I.; Willson, T.M.; Tropsha, A.; Isayev, O. Generative and reinforcement learning approaches for the automated de novo design of bioactive compounds. Commun. Chem. 2022, 5, 129. [Google Scholar] [CrossRef]
- Gangwal, A.; Lavecchia, A. Unleashing the power of generative AI in drug discovery. Drug Discov. Today 2024, 29, 103992. [Google Scholar] [CrossRef]
- Bayley, O.; Savino, E.; Slattery, A.; Noël, T. Autonomous chemistry: Navigating self-driving labs in chemical and material sciences. Matter 2024, 7, 2382–2398. [Google Scholar] [CrossRef]
- Tom, G.; Schmid, S.P.; Baird, S.G.; Cao, Y.; Darvish, K.; Hao, H.; Lo, S.; Pablo-García, S.; Rajaonson, E.M.; Skreta, M. Self-Driving Laboratories for Chemistry and Materials Science. Chem. Rev. 2024, 124, 9633–9732. [Google Scholar] [CrossRef]
- Burger, B.; Maffettone, P.M.; Gusev, V.V.; Aitchison, C.M.; Bai, Y.; Wang, X.; Li, X.; Alston, B.M.; Li, B.; Clowes, R.; et al. A mobile robotic chemist. Nature 2020, 583, 237–241. [Google Scholar] [CrossRef]
- Dai, T.; Vijayakrishnan, S.; Szczypiński, F.T.; Ayme, J.-F.; Simaei, E.; Fellowes, T.; Clowes, R.; Kotopanov, L.; Shields, C.E.; Zhou, Z. Autonomous mobile robots for exploratory synthetic chemistry. Nature 2024, 635, 890–897. [Google Scholar] [CrossRef]
- Yang, Y.; Juntao, L.; Lingling, P. Multi-robot path planning based on a deep reinforcement learning DQN algorithm. CAAI Trans. Intell. Technol. 2020, 5, 177–183. [Google Scholar] [CrossRef]
- Huang, D.; Ye, X.; Zhang, Y.; Sakurai, T. Collaborative analysis for drug discovery by federated learning on non-IID data. Methods 2023, 219, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Schouten, D.; Nicoletti, G.; Dille, B.; Chia, C.; Vendittelli, P.; Schuurmans, M.; Litjens, G.; Khalili, N. Navigating the landscape of multimodal AI in medicine: A scoping review on technical challenges and clinical applications. Med. Image Anal. 2025, 105, 103621. [Google Scholar] [CrossRef]
- DeGroat, W.; Abdelhalim, H.; Peker, E.; Sheth, N.; Narayanan, R.; Zeeshan, S.; Liang, B.T.; Ahmed, Z. Multimodal AI/ML for discovering novel biomarkers and predicting disease using multi-omics profiles of patients with cardiovascular diseases. Sci. Rep. 2024, 14, 26503. [Google Scholar] [CrossRef]
- Russell, S. Artificial intelligence: The future is superintelligent. Nature 2017, 548, 520–521. [Google Scholar] [CrossRef]
- Tang, X.; Dai, H.; Knight, E.; Wu, F.; Li, Y.; Li, T.; Gerstein, M. A survey of generative AI for de novo drug design: New frontiers in molecule and protein generation. Brief. Bioinform. 2024, 25, bbae338. [Google Scholar] [CrossRef]
- Withers, C.A.; Rufai, A.M.; Venkatesan, A.; Tirunagari, S.; Lobentanzer, S.; Harrison, M.; Zdrazil, B. Natural language processing in drug discovery: Bridging the gap between text and therapeutics with artificial intelligence. Expert Opin. Drug Discov. 2025, 20, 765–783. [Google Scholar] [CrossRef]
- Guellil, I.; Wu, J.; Pradipta Gema, A.; Francis, F.; Berrachedi, Y.; Chenni, N.; Tobin, R.; Llewellyn, C.; Arakelyan, S.; Wu, H.; et al. Natural language processing for detecting adverse drug events: A systematic review protocol. NIHR Open Res. 2024, 3, 67. [Google Scholar] [CrossRef]
- Roche, R.; Moussad, B.; Shuvo, M.H.; Bhattacharya, D. E(3) equivariant graph neural networks for robust and accurate protein-protein interaction site prediction. PLoS Comput. Biol. 2023, 19, e1011435. [Google Scholar] [CrossRef]
- Xu, M.; Qian, P.; Zhao, Z.; Zeng, Z.; Chen, J.; Liu, W.; Yang, X. Graph Neural Networks for Protein-Protein Interactions—A Short Survey. arXiv 2024, arXiv:2404.10450. [Google Scholar]
- Nerella, S.; Bandyopadhyay, S.; Zhang, J.; Contreras, M.; Siegel, S.; Bumin, A.; Silva, B.; Sena, J.; Shickel, B.; Bihorac, A.; et al. Transformers and large language models in healthcare: A review. Artif. Intell. Med. 2024, 154, 102900. [Google Scholar] [CrossRef]
- Chen, J.-Y.; Wang, J.-F.; Hu, Y.; Li, X.-H.; Qian, Y.-R.; Song, C.-L. Evaluating the advancements in protein language models for encoding strategies in protein function prediction: A comprehensive review. Front. Bioeng. Biotechnol. 2025, 13, 1506508. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.-H.; Yin, T.; Zhang, D.-B. Hybrid quantum-classical algorithms for solving quantum chemistry in Hamiltonian–wave-function space. Phys. Rev. A 2021, 103, 012413. [Google Scholar] [CrossRef]
- Braga, D.M.; Rawal, B.S. Harnessing AI and Quantum Computing for Accelerated Drug Discovery: Regulatory Frameworks for In Silico to In Vivo Validation. J. Pharm. BioTech Ind. 2025, 2, 11. [Google Scholar] [CrossRef]
- Cui, M.; Chang, L.; Chau, A.; Mekuria, H.; Adwankar, L.; Pendyala, S. Efficient and Optimized Small Organic Molecular Graph Generation Pathway Using a Quantum Generative Adversarial Network. In Proceedings of the 2024 IEEE International Conference on Quantum Computing and Engineering (QCE), Montreal, QC, Canada, 15–20 September 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 1565–1570. [Google Scholar]
- Bauer, N.; Siopsis, G. Post-Variational Ground State Estimation via QPE-Based Quantum Imaginary Time Evolution. arXiv 2025, arXiv:2504.11549. [Google Scholar]
- Ullah, S.; Shah, M.H.; Anjum, A. Quantum Enhanced Federated Learning with Differential Privacy. In Proceedings of the 2024 International Conference on Frontiers of Information Technology (FIT), Islamabad, Pakistan, 9–10 December 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 1–6. [Google Scholar]
- Bender, A.; Cortés-Ciriano, I. Artificial intelligence in drug discovery: What is realistic, what are illusions? Part 1: Ways to make an impact, and why we are not there yet. Drug Discov. Today 2021, 26, 511–524. [Google Scholar] [CrossRef]
- Bender, A.; Cortes-Ciriano, I. Artificial intelligence in drug discovery: What is realistic, what are illusions? Part 2: A discussion of chemical and biological data. Drug Discov. Today 2021, 26, 1040–1052. [Google Scholar] [CrossRef]
- Price, S.S.L. Computed Crystal Energy Landscapes for Understanding and Predicting Organic Crystal Structures and Polymorphism. Acc. Chem. Res. 2009, 42, 117–126. [Google Scholar] [CrossRef]
- Davis, B.; Mcloughlin, K.; Allen, J.; Ellingson, S.R. Quantifying Overfitting Potential in Drug Binding Datasets. In Proceedings of the Computational Science—ICCS 2020, Amsterdam, The Netherlands, 15 June 2020; pp. 585–598. [Google Scholar]
- Fan, Z.; Yu, J.; Zhang, X.; Chen, Y.; Sun, S.; Zhang, Y.; Chen, M.; Xiao, F.; Wu, W.; Li, X.; et al. Reducing overconfident errors in molecular property classification using Posterior Network. Patterns 2024, 5, 100991. [Google Scholar] [CrossRef]
- Nyman, J.; Day, G.M. Static and lattice vibrational energy differences between polymorphs. CrystEngComm 2015, 17, 5154–5165. [Google Scholar] [CrossRef]
- Ding, Q.-M.; Huang, Y.-M.; Yuan, X. Molecular docking via quantum approximate optimization algorithm. Phys. Rev. Appl. 2024, 21, 034036. [Google Scholar] [CrossRef]
- Al-Dahlaki, M.H.; Mohammed, A.H. Quantum mechanics/molecular mechanics (QM/MM) methods in drug design: A comprehensive review of development and applications. Int. J. Adv. Chem. 2024, 12, 86–92. [Google Scholar] [CrossRef]
- Hayashi, H.; Maeda, S.; Mita, T. Quantum chemical calculations for reaction prediction in the development of synthetic methodologies. Chem. Sci. 2023, 14, 11601–11616. [Google Scholar] [CrossRef]
- Han, C.; Zhang, D.; Xia, S.; Zhang, Y. Accurate Prediction of NMR Chemical Shifts: Integrating DFT Calculations with Three-Dimensional Graph Neural Networks. J. Chem. Theory Comput. 2024, 20, 5250–5258. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Wu, X.; Brooks, B.R.; Wang, J. Accurate Free Energy Calculation via Multiscale Simulations Driven by Hybrid Machine Learning and Molecular Mechanics Potentials. J. Chem. Theory Comput. 2024, 21, 6979–6987. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Day, G.M. Global analysis of the energy landscapes of molecular crystal structures by applying the threshold algorithm. Commun. Chem. 2022, 5, 86. [Google Scholar] [CrossRef] [PubMed]
- Asano, D.; Takakusa, H.; Nakai, D. Oral Absorption of Middle-to-Large Molecules and Its Improvement, with a Focus on New Modality Drugs. Pharmaceutics 2023, 16, 47. [Google Scholar] [CrossRef]
- Kokudeva, M.; Vichev, M.; Naseva, E.; Miteva, D.G.; Velikova, T. Artificial intelligence as a tool in drug discovery and development. World J. Exp. Med. 2024, 14, 96042. [Google Scholar] [CrossRef]
- Mustafa, H.; Morapakula, S.N.; Jain, P.; Ganguly, S. Variational Quantum Algorithms for Chemical Simulation and Drug Discovery. In Proceedings of the 2022 International Conference on Trends in Quantum Computing and Emerging Business Technologies, TQCEBT 2022, Pune, India, 13–15 October 2022. [Google Scholar]
- Li, W.; Yin, Z.; Li, X.; Ma, D.; Yi, S.; Zhang, Z.; Zou, C.; Bu, K.; Dai, M.; Yue, J.; et al. A hybrid quantum computing pipeline for real world drug discovery. Sci. Rep. 2024, 14, 16942. [Google Scholar] [CrossRef]
- Zubatyuk, R.; Biczysko, M.; Ranasinghe, K.; Moriarty, N.W.; Gokcan, H.; Kruse, H.; Poon, B.K.; Adams, P.D.; Waller, M.P.; Roitberg, A.E.; et al. AQuaRef: Machine learning accelerated quantum refinement of protein structures. Nat. Commun. 2024, 16, 9224. [Google Scholar]
- Pasupuleti, M.K. Hybrid Quantum-Classical Algorithms for Drug Discovery and Molecular Simulation. Int. J. Acad. Ind. Res. Innov. 2025, 5, 303–314. [Google Scholar] [CrossRef]
- Upama, P.B.; Kolli, A.; Kolli, H.; Alam, S.; Syam, M.; Shahriar, H.; Ahamed, S.I. Quantum Machine Learning in Disease Detection and Prediction: A survey of Applications and Future Possibilities. In Proceedings of the International Computer Software and Applications Conference, Torino, Italy, 26–30 June 2023. [Google Scholar]
- Ghazi Vakili, M.; Gorgulla, C.; Snider, J.; Nigam, A.; Bezrukov, D.; Varoli, D.; Aliper, A.; Polykovsky, D.; Das, K.M.P.; Cox, H.; et al. Quantum-computing-enhanced algorithm unveils potential KRAS inhibitors. Nat. Biotechnol. 2025, 43, 1954–1959. [Google Scholar] [CrossRef]
- Pyrkov, A.; Aliper, A.; Bezrukov, D.; Lin, Y.-C.; Polykovskiy, D.; Kamya, P.; Ren, F.; Zhavoronkov, A. Quantum computing for near-term applications in generative chemistry and drug discovery. Drug Discov. Today 2023, 28, 103675. [Google Scholar] [CrossRef]
- Liu, J.-G.; Wang, L. Differentiable learning of quantum circuit Born machines. Phys. Rev. A 2018, 98, 062324. [Google Scholar] [CrossRef]
- Pfau, D.; Axelrod, S.; Sutterud, H.; von Glehn, I.; Spencer, J.S. Accurate computation of quantum excited states with neural networks. Science 2024, 385, eadn0137. [Google Scholar] [CrossRef] [PubMed]
- Sudarsanan Ragini, N.; Sazim, S. Higher-order incompatibility improves distinguishability of causal quantum networks. New J. Phys. 2024, 26, 123003. [Google Scholar] [CrossRef]
- Zheng, K.; Fu, T.; Liang, Z. QCS-ADME: Quantum Circuit Search for Drug Property Prediction with Imbalanced Data and Regression Adaptation. arXiv 2025, arXiv:2503.01927. [Google Scholar] [CrossRef]
- Rath, M.; Date, H. Quantum Inspired Encoding Strategies for Machine Learning Models: Proposing and Evaluating Instance Level, Global Discrete, and Class Conditional Representations. arXiv 2025, arXiv:2507.00019. [Google Scholar]
- Alizadehsani, R.; Oyelere, S.S.; Hussain, S.; Jagatheesaperumal, S.K.; Calixto, R.R.; Rahouti, M. Explainable Artificial Intelligence for Drug Discovery and Development: A Comprehensive Survey. IEEE Access 2024, 12, 35796–35812. [Google Scholar] [CrossRef]
- Salih, A.M.; Raisi-Estabragh, Z.; Galazzo, I.B.; Radeva, P.; Petersen, S.E.; Lekadir, K.; Menegaz, G. A Perspective on Explainable Artificial Intelligence Methods: SHAP and LIME. Adv. Intell. Syst. 2025, 7, 2400304. [Google Scholar] [CrossRef]
- Wang, Q.; He, M.; Guo, L.; Chai, H. AFEI: Adaptive optimized vertical federated learning for heterogeneous multi-omics data integration. Brief. Bioinform. 2023, 24, bbad269. [Google Scholar] [CrossRef] [PubMed]
- Fehlis, Y.; Mandel, P.; Crain, C.; Liu, B.; Fuller, D. Accelerating drug discovery with Artificial: A whole-lab orchestration and scheduling system for self-driving labs. arXiv 2025, arXiv:2504.00986. [Google Scholar]
- Häse, F.; Roch, L.M.; Aspuru-Guzik, A. Next-Generation Experimentation with Self-Driving Laboratories. Trends Chem. 2019, 1, 282–291. [Google Scholar] [CrossRef]
- El Arab, R.A.; Abu-Mahfouz, M.S.; Abuadas, F.H.; Alzghoul, H.; Almari, M.; Ghannam, A.; Seweid, M.M. Bridging the Gap: From AI Success in Clinical Trials to Real-World Healthcare Implementation-A Narrative Review. Healthcare 2025, 13, 701. [Google Scholar] [CrossRef]
- Babbush, R.; Huggins, W.J.; Berry, D.W.; Ung, S.F.; Zhao, A.; Reichman, D.R.; Neven, H.; Baczewski, A.D.; Lee, J. Quantum simulation of exact electron dynamics can be more efficient than classical mean-field methods. Nat. Commun. 2023, 14, 4058. [Google Scholar] [CrossRef]
- Lappala, A. The next revolution in computational simulations: Harnessing AI and quantum computing in molecular dynamics. Curr. Opin. Struct. Biol. 2024, 89, 102919. [Google Scholar] [CrossRef]
- Ahmed, M.; Maldonado, A.M.; Durrant, J.D. From byte to bench to bedside: Molecular dynamics simulations and drug discovery. BMC Biol. 2023, 21, 299. [Google Scholar] [CrossRef]
- Mahdi, G.; Alamdary, A.; Kaghazian, H.; Ahmadi, N.; Mardani, R.; Gholami, A. Biopharmaceutical Analysis by HPLC: Practices and Challenges. Chonnam Med. J. 2025, 61, 19. [Google Scholar] [CrossRef] [PubMed]
- Pienaar, E.; Theron, M.; Nelson, M.; Viljoen, H.J. A quantitative model of error accumulation during PCR amplification. Comput. Biol. Chem. 2006, 30, 102–111. [Google Scholar] [CrossRef]
- Moinard, S.; Piau, D.; Laporte, F.; Rioux, D.; Taberlet, P.; Gonindard-Melodelima, C.; Coissac, E. Towards quantitative DNA Metabarcoding: A method to overcome PCR amplification bias. bioRxiv 2023. [Google Scholar] [CrossRef]
- Kiepas, A.; Voorand, E.; Mubaid, F.; Siegel, P.M.; Brown, C.M. Optimizing live-cell fluorescence imaging conditions to minimize phototoxicity. J. Cell Sci. 2020, 133, jcs242834. [Google Scholar] [CrossRef]
- Godin, A.G.; Lounis, B.; Cognet, L. Super-resolution microscopy approaches for live cell imaging. Biophys. J. 2014, 107, 1777–1784. [Google Scholar] [CrossRef] [PubMed]
- Mohs, R.C.; Greig, N.H. Drug discovery and development: Role of basic biological research. Alzheimer’s Dement. 2017, 3, 651–657. [Google Scholar] [CrossRef] [PubMed]
- Farid, S.S.; Baron, M.; Stamatis, C.; Nie, W.; Coffman, J. Benchmarking biopharmaceutical process development and manufacturing cost contributions to R&D. MAbs 2020, 12, 1754999. [Google Scholar] [CrossRef] [PubMed]
- Pérez, I.; Krueger, A.; Wrachtrup, J.; Jelezko, F.; Börsch, M. Single NV Center in Nanodiamond for Quantum Sensing of Protein Dynamics in AN ABEL Trap. In Proceedings of the Single Molecule Spectroscopy and Superresolution Imaging XVII, San Francisco, CA, USA, 27 January–1 February 2024. [Google Scholar]
- Noh, T.; Kindseth, A.; Chandrasekhar, V. Nonlocal superconducting quantum interference device. Phys. Rev. B 2021, 104, 064503. [Google Scholar] [CrossRef]
- Bao, B.; Hua, Y.; Wang, R.; Li, D. Quantum-Based Magnetic Field Sensors for Biosensing. Adv. Quantum Technol. 2023, 6, 2200146. [Google Scholar] [CrossRef]
- Modicano, P.; Trutschel, M.; Phan-Xuan, T.; Matarèse, B.F.E.; Urbano, L.; Green, M.; Mäder, K.; Dailey, L.A. Does Encapsulation of π-Conjugated Polymer Nanoparticles within Biodegradable PEG–PLGA Matrices Mitigate Photoinduced Free Radical Production and Phototoxicity? Adv. Ther. 2025, 8, 2400190. [Google Scholar] [CrossRef]
- Feyen, P.L.C.; Matarèse, B.F.E.; Urbano, L.; Abelha, T.F.; Rahmoune, H.; Green, M.; Dailey, L.A.; de Mello, J.C.; Benfenati, F. Photosensitized and Photothermal Stimulation of Cellular Membranes by Organic Thin Films and Nanoparticles. Front. Bioeng. Biotechnol. 2022, 10, 932877. [Google Scholar] [CrossRef]
- Rondin, L.; Tetienne, J.P.; Hingant, T.; Roch, J.-F.; Maletinsky, P.; Jacques, V. Magnetometry with nitrogen-vacancy defects in diamond. Rep. Prog. Phys. 2014, 77, 056503. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez, D. A quantum sensor for high-performance mass spectrometry. Appl. Phys. B 2012, 107, 1031–1042. [Google Scholar] [CrossRef]
- Cautereels, J.; Blockhuys, F. Quantum Chemical Mass Spectrometry: Verification and Extension of the Mobile Proton Model for Histidine. J. Am. Soc. Mass Spectrom. 2017, 28, 1227–1235. [Google Scholar] [CrossRef]
- Sarkar, A.; Jones, Z.R.; Parashar, M.; Druga, E.; Akkiraju, A.; Conti, S.; Krishnamoorthi, P.; Nachuri, S.; Aman, P.; Hashemi, M.; et al. High-precision chemical quantum sensing in flowing monodisperse microdroplets. Sci. Adv. 2024, 10, eadp4033. [Google Scholar] [CrossRef]
- Fujiwara, M. Diamond quantum sensors in microfluidics technology. Biomicrofluidics 2023, 17, 054107. [Google Scholar] [CrossRef]
- Lucivero, V.G.; Jiménez-Martínez, R.; Kong, J.; Mitchell, M.W. Squeezed-light spin noise spectroscopy. Phys. Rev. A 2016, 93, 053802. [Google Scholar] [CrossRef]
- Tsao, C.; Ling, H.; Hinkle, A.; Chen, Y.; Jha, K.K.; Yan, Z.-L.; Utzat, H. Enhancing spectroscopy and microscopy with emerging methods in photon correlation and quantum illumination. Nat. Nanotechnol. 2025, 20, 1001–1016. [Google Scholar] [CrossRef] [PubMed]
- Prajapati, N.; Niu, Z.; Novikova, I. Quantum-enhanced two-photon spectroscopy using two-mode squeezed light. Opt. Lett. 2021, 46, 1800. [Google Scholar] [CrossRef]
- Zafar, A.; Takeda, C.; Manzoor, A.; Tanaka, D.; Kobayashi, M.; Wadayama, Y.; Nakane, D.; Majeed, A.; Iqbal, M.A.; Akitsu, T. Towards Industrially Important Applications of Enhanced Organic Reactions by Microfluidic Systems. Molecules 2024, 29, 398. [Google Scholar] [CrossRef] [PubMed]
- Goutham, R.; Gopinath, K.P.; Ramprasath, A.; Srikanth, B.; Narayan, R.B. High-Performance Photocatalysts for Organic Reactions; Springer: Cham, Switzerland, 2019; pp. 219–270. [Google Scholar]
- Levin, L.; Skomorowski, W.; Rybak, L.; Kosloff, R.; Koch, C.P.; Amitay, Z. Coherent Control of Bond Making. Phys. Rev. Lett. 2015, 114, 233003. [Google Scholar] [CrossRef]
- Leibscher, M.; Pozzoli, E.; Pérez, C.; Schnell, M.; Sigalotti, M.; Boscain, U.; Koch, C.P. Full quantum control of enantiomer-selective state transfer in chiral molecules despite degeneracy. Commun. Phys. 2022, 5, 110. [Google Scholar] [CrossRef]
- Furusho, Y.; Yagi, R.; Suda, M.; Maruhashi, T.; Takagi, I.; Kayanuma, Y.; Minami, F.; Nakamura, K.G. Ultrafast quantum-path interferometry of photo-absorption involving excitons in a GaAs multiple-quantum-well structure. Solid State Commun. 2023, 360, 115056. [Google Scholar] [CrossRef]
- Camlik, G.; Bilakaya, B.; Güven, G.K.; Akkol, E.K.; Degim, Z.; Sobarzo-Sánchez, E.; Degim, I.T. Quantum Drugs (Q-Drugs): A New Discovery and Taboo Breaking Approach; Producing Carbon Quantum Dots from Drug Molecules. Pharmaceuticals 2025, 18, 767. [Google Scholar] [CrossRef]
- Hartwig, S.; Albrecht, H.H.; Scheer, H.J.; Burghoff, M.; Trahms, L. A Superconducting Quantum Interference Device Measurement System for Ultra Low-Field Nuclear Magnetic Resonance. Appl. Magn. Reson. 2013, 44, 9–22. [Google Scholar] [CrossRef]
- Vasyukov, D.; Anahory, Y.; Embon, L.; Halbertal, D.; Cuppens, J.; Neeman, L.; Finkler, A.; Segev, Y.; Myasoedov, Y.; Rappaport, M.L.; et al. A scanning superconducting quantum interference device with single electron spin sensitivity. Nat. Nanotechnol. 2013, 8, 639–644. [Google Scholar] [CrossRef] [PubMed]
- Yong, K.T.; Wang, Y.; Roy, I.; Rui, H.; Swihart, M.T.; Law, W.-C.; Kwak, S.K.; Ye, L.; Liu, J.; Mahajan, S.D.; et al. Preparation of quantum dot/drug nanoparticle formulations for traceable targeted delivery and therapy. Theranostics 2012, 2, 681–694. [Google Scholar] [CrossRef] [PubMed]
- Magann, A.B.; Grace, M.D.; Rabitz, H.A.; Sarovar, M. Digital quantum simulation of molecular dynamics and control. Phys. Rev. Res. 2021, 3, 023165. [Google Scholar] [CrossRef]
- Gircha, A.I.; Boev, A.S.; Avchaciov, K.; Fedichev, P.O.; Fedorov, A.K. Hybrid quantum-classical machine learning for generative chemistry and drug design. Sci. Rep. 2023, 13, 8250. [Google Scholar] [CrossRef]
- Wang, L.; Zhou, Z.; Yang, X.; Shi, S.; Zeng, X.; Cao, D. The present state and challenges of active learning in drug discovery. Drug Discov. Today 2024, 29, 103985. [Google Scholar] [CrossRef]
- Kadam, E.A.J.J.; Kharmate, G.M.; Jadhav, M.R.; Howal, S.A.; Pandit, V.U. Quantum Machine Learning Technique for Automatic Retrosynthetic Reaction Pathway Search Method. Int. J. Recent Innov. Trends Comput. Commun. 2023, 11, 2111–2122. [Google Scholar] [CrossRef]
- Zhou, Y.; Chen, J.; Cheng, J.; Cao, X.; Zhang, Y.; Karemore, G.; Zitnik, M.; Chong, F.T.; Liu, J.; Fu, T.; et al. Quantum-Machine-Assisted Drug Discovery: Survey and Perspective. arXiv 2025, arXiv:2408.13479. [Google Scholar]
- Banchi, L.; Fingerhuth, M.; Babej, T.; Ing, C.; Arrazola, J.M. Molecular docking with Gaussian Boson Sampling. Sci. Adv. 2020, 6, eaax1950. [Google Scholar] [CrossRef]
- Niu, Z.; Xiao, X.; Wu, W.; Cai, Q.; Jiang, Y.; Jin, W.; Wang, M.; Yang, G.; Kong, L.; Jin, X.; et al. PharmaBench: Enhancing ADMET benchmarks with large language models. Sci. Data 2024, 11, 985. [Google Scholar] [CrossRef]
- Tran, T.T.; Van Tayara, H.; Chong, K.T. Recent Studies of Artificial Intelligence on In Silico Drug Absorption. J. Chem. Inf. Model. 2023, 63, 6198–6211. [Google Scholar] [CrossRef]
- Speybroeck, V.; Van Gani, R.; Meier, R.J. The calculation of thermodynamic properties of molecules. Chem. Soc. Rev. 2010, 39, 1764–1779. [Google Scholar] [CrossRef] [PubMed]
- Ali, R.S.A.E.; Meng, J.; Khan, M.E.I.; Jiang, X. Machine learning advancements in organic synthesis: A focused exploration of artificial intelligence applications in chemistry. Artif. Intell. Chem. 2024, 2, 100049. [Google Scholar] [CrossRef]
- Bogojeski, M.; Vogt-Maranto, L.; Tuckerman, M.E.; Müller, K.-R.; Burke, K. Quantum chemical accuracy from density functional approximations via machine learning. Nat. Commun. 2020, 11, 5223. [Google Scholar] [CrossRef]
- Zhuang, M.; Chen, S.; Huang, J.; Lee, C. Quantum lock-in measurement of weak alternating signals. Quantum Front. 2024, 3, 4. [Google Scholar] [CrossRef]
- Qi, J.; Ko, T.W.; Wood, B.C.; Pham, T.A.; Ong, S.P. Robust training of machine learning interatomic potentials with dimensionality reduction and stratified sampling. npj Comput. Mater. 2024, 10, 43. [Google Scholar] [CrossRef]
- Bannigan, P.; Aldeghi, M.; Bao, Z.; Häse, F.; Aspuru-Guzik, A.; Allen, C. Machine learning directed drug formulation development. Adv. Drug Deliv. Rev. 2021, 175, 113806. [Google Scholar] [CrossRef]
- Wang, Y.; Hu, Z.; Chang, J.; Yu, B. Thinking on the Use of Artificial Intelligence in Drug Discovery. J. Med. Chem. 2025, 68, 4996–4999. [Google Scholar] [CrossRef]
- Zhang, X.; Deng, H.; Wu, R.; Ren, J.; Ren, Y. PQSF: Post-quantum secure privacy-preserving federated learning. Sci. Rep. 2024, 14, 23553. [Google Scholar] [CrossRef]
- Morin, L.; Weber, V.; Meijer, G.I.; Yu, F.; Staar, P.W.J. PatCID: An open-access dataset of chemical structures in patent documents. Nat. Commun. 2024, 15, 6532. [Google Scholar] [CrossRef]
- Saratkar, S.Y.; Langote, M.; Kumar, P.; Gote, P.; Weerarathna, I.; Mishra, G. Digital twin for personalized medicine development. Front. Digit. Health 2025, 7, 1583466. [Google Scholar] [CrossRef]
- Khater, T.; Alkhatib, S.A.; AlShehhi, A.; Pitsalidis, C.; Pappa, A.; Ngo, S.; Chan, V.; Truong, V. Generative artificial intelligence based models optimization towards molecule design enhancement. J. Cheminform. 2025, 17, 116. [Google Scholar] [CrossRef] [PubMed]
- Udrescu, S.M.; Tegmark, M. AI Feynman: A physics-inspired method for symbolic regression. Sci. Adv. 2020, 6, eaay2631. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Zhao, J.; Xiong, M. Graphical Learning and Causal Inference for Drug Repurposing. medRxiv 2023. [Google Scholar] [CrossRef] [PubMed]
- Wang, G.; Wang, C.; Zhang, X.; Li, Z.; Zhou, J.; Sun, Z. Machine learning interatomic potential: Bridge the gap between small-scale models and realistic device-scale simulations. iScience 2024, 27, 109673. [Google Scholar] [CrossRef]
- Della Pia, F.; Shi, B.X.; Kapil, V.; Zen, A.; Alfè, D.; Michaelides, A. Accurate and efficient machine learning interatomic potentials for finite temperature modelling of molecular crystals. Chem. Sci. 2025, 16, 11419–11433. [Google Scholar] [CrossRef]
- Lin, J.C.; Glover, Z.K.; Sreedhara, A. Assessing the Utility of Circular Dichroism and FTIR Spectroscopy in Monoclonal-Antibody Comparability Studies. J. Pharm. Sci. 2015, 104, 4459–4466. [Google Scholar] [CrossRef]
- Schlawin, F.; Gessner, M. Theory of Quantum-Enhanced Stimulated Raman Scattering. arXiv 2025, arXiv:2502.19344. [Google Scholar] [CrossRef]
- Kalashnikov, D.A.; Pan, Z.; Kuznetsov, A.I.; Krivitsky, L.A. Quantum Spectroscopy of Plasmonic Nanostructures. Phys. Rev. X 2014, 4, 011049. [Google Scholar] [CrossRef]
- Michoel, T.; Zhang, J.D. Causal inference in drug discovery and development. Drug Discov. Today 2023, 28, 103737. [Google Scholar] [CrossRef]
- Deepasree, K.; Venugopal, S. Molecular docking and molecular dynamic simulation studies to identify potential terpenes against Internalin A protein of Listeria monocytogenes. Front. Bioinform. 2024, 4, 1463750. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; An, J.; Zhao, J.; Zhao, S.; Lv, H.; Wang, S. Drug-Target Interaction Prediction Based on Multisource Information Weighted Fusion. Contrast Media Mol. Imaging 2021, 2021, 6044256. [Google Scholar] [CrossRef]
- Heifetz, A. Accelerating COVID-19 Drug Discovery with High-Performance Computing; Humana: New York, NY, USA, 2024; pp. 405–411. [Google Scholar]
- Švajger, U.; Horvat, Ž.; Knez, D.; Rožman, P.; Turk, S.; Gobec, S. New antagonists of toll-like receptor 7 discovered through 3D ligand-based virtual screening. Med. Chem. Res. 2015, 24, 362–371. [Google Scholar] [CrossRef]
- Belsley, A. Quantum-Enhanced Absorption Spectroscopy with Bright Squeezed Frequency Combs. Phys. Rev. Lett. 2023, 130, 133602. [Google Scholar] [CrossRef] [PubMed]
- Brad, S. Enhancing Creativity in Deep Learning Models with SAVE-Inspired Activation Functions. In IFIP Advances in Information and Communication Technology; Springer: Cham, Switzerland, 2023. [Google Scholar]
- Pal, S.; Bhattacharya, M.; Lee, S.S.; Chakraborty, C. Quantum Computing in the Next-Generation Computational Biology Landscape: From Protein Folding to Molecular Dynamics. Mol. Biotechnol. 2024, 66, 163–178. [Google Scholar] [CrossRef]
- Miessen, A.; Ollitrault, P.J.; Tacchino, F.; Tavernelli, I. Quantum algorithms for quantum dynamics. Nat. Comput. Sci. 2022, 3, 25–37. [Google Scholar] [CrossRef]
- Day, E.C.; Chittari, S.S.; Bogen, M.P.; Knight, A.S. Navigating the Expansive Landscapes of Soft Materials: A User Guide for High-Throughput Workflows. ACS Polym. Au 2023, 3, 406–427. [Google Scholar] [CrossRef]
- Shapira, L.; Lerner, S.; Assayag, G.; Vardi, A.; Haham, D.; Bar, G.; Kozokaro, V.; Robicsek, M.; Lerner, I.; Michaeli, A. Discovery of novel spike/ACE2 inhibitory macrocycles using in silico reinforcement learning. Front. Drug Discov. 2022, 2, 1085701. [Google Scholar] [CrossRef]
- Lee, T.-S.; Hu, Y.; Sherborne, B.; Guo, Z.; York, D.M. Toward Fast and Accurate Binding Affinity Prediction with pmemdGTI: An Efficient Implementation of GPU-Accelerated Thermodynamic Integration. J. Chem. Theory Comput. 2017, 13, 3077–3084. [Google Scholar] [CrossRef]
- Ye, C.; Sun, Y.; Zhang, X. Entanglement-Assisted Quantum Chiral Spectroscopy. J. Phys. Chem. Lett. 2021, 12, 8591–8597. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; Miao, T.; Qian, Y.; Zhang, Z.; Zhang, W.; Zhu, X. Supramolecular Chirality in Azobenzene-Containing Polymer System: Traditional Postpolymerization Self-Assembly Versus In Situ Supramolecular Self-Assembly Strategy. Int. J. Mol. Sci. 2020, 21, 6186. [Google Scholar] [CrossRef]
- Gupta, A.; Dey, S.; Hicks, A.; Zhou, H.-X. Artificial intelligence guided conformational mining of intrinsically disordered proteins. Commun. Biol. 2022, 5, 610. [Google Scholar] [CrossRef]
- Richardson, J.O. Nonadiabatic Tunneling in Chemical Reactions. J. Phys. Chem. Lett. 2024, 15, 7387–7397. [Google Scholar] [CrossRef] [PubMed]
- Das, S.; Mazumdar, H.; Khondakar, K.R.; Mishra, Y.; Kaushik, A. Review—Quantum Biosensors: Principles and Applications in Medical Diagnostics. ECS Sens. Plus 2024, 3, 025001. [Google Scholar] [CrossRef]
- Jinich, A.; Rappoport, D.; Dunn, I.; Sanchez-Lengeling, B.; Olivares-Amaya, R.; Noor, E.; Bar Even, A.; Aspuru-Guzik, A. Quantum Chemical Approach to Estimating the Thermodynamics of Metabolic Reactions. Sci. Rep. 2014, 4, 7022. [Google Scholar] [CrossRef] [PubMed]
- Vidossich, P.; De Vivo, M. The role of first principles simulations in studying (bio)catalytic processes. Chem. Catal. 2021, 1, 69–87. [Google Scholar] [CrossRef]
- Sagar Shrikrishna, N.; Sharma, R.; Sahoo, J.; Kaushik, A.; Gandhi, S. Navigating the landscape of optical biosensors. Chem. Eng. J. 2024, 490, 151661. [Google Scholar] [CrossRef]
- Aly, S.M.; Fathi, M. Advancing aquaculture biosecurity: A scientometric analysis and future outlook for disease prevention and environmental sustainability. Aquac. Int. 2024, 32, 8763–8789. [Google Scholar] [CrossRef]
- Yu, T.; Guo, H. Understanding Enzyme Catalysis Mechanism Using QM/MM Simulation Methods. Mech. Enzymol. Bridg. Struct. Funct. 2020, 1357, 121–137. [Google Scholar]
- Hernandez, A.B.; Cortés-Arriagada, D.; García, H.C.; Anota, E.; Villanueva, M. Quantum molecular study on doping effect in titanium and vanadium clusters: Their application to remove some chemical species. Appl. Nanosci. 2020, 10, 37–49. [Google Scholar] [CrossRef]
- Wang, T.; He, X.; Li, M.; Li, Y.; Bi, R.; Wang, Y.; Cheng, C.; Shen, X.; Meng, J.; Zhang, H.; et al. Ab initio characterization of protein molecular dynamics with AI2BMD. Nature 2024, 635, 1019–1027. [Google Scholar] [CrossRef]
- Li, X.; Huo, P. Investigating Tunneling-Controlled Chemical Reactions through Ab Initio Ring Polymer Molecular Dynamics. J. Phys. Chem. Lett. 2021, 12, 6714–6721. [Google Scholar] [CrossRef]
- Wen, J.; Mai, S.; González, L. Excited-State Dynamics Simulations of a Light-Driven Molecular Motor in Solution. J. Phys. Chem. A 2023, 127, 9520–9529. [Google Scholar] [CrossRef]
- Thürlemann, M.; Riniker, S. Hybrid classical/machine-learning force fields for the accurate description of molecular condensed-phase systems. Chem. Sci. 2023, 14, 12661–12675. [Google Scholar] [CrossRef] [PubMed]
- Lin, F.-Y.; MacKerell, A.D. Force Fields for Small Molecules. Methods Mol. Biol. 2019, 2022, 21–54. [Google Scholar] [CrossRef]
- Skogh, M.; Dobrautz, W.; Lolur, P.; Warren, C.; Biznárová, J.; Osman, A.; Tancredi, G.; Bylander, J.; Rahm, M. The electron density: A fidelity witness for quantum computation. Chem. Sci. 2024, 15, 2257–2265. [Google Scholar] [CrossRef] [PubMed]
- Niazi, S.K. The Quantum Paradox in Pharmaceutical Science: Understanding Without Comprehending—A Centennial Reflection. Int. J. Mol. Sci. 2025, 26, 4658. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-L.; Li, B. Reliability of Computing van der Waals Bond Lengths of Some Rare Gas Diatomics. Int. J. Mol. Sci. 2022, 23, 13944. [Google Scholar] [CrossRef] [PubMed]
- Świderek, K.; Bertran, J.; Zinovjev, K.; Tuñón, I.; Moliner, V. Advances in the Simulations of Enzyme Reactivity in the Dawn of the Artificial Intelligence Age. WIREs Comput. Mol. Sci. 2025, 15, e70003. [Google Scholar] [CrossRef]
- Tiwari, A.; Honingh, C.; Ensing, B. Accurate calculation of zero point energy from molecular dynamics simulations of liquids and their mixtures. J. Chem. Phys. 2019, 151, 244124. [Google Scholar] [CrossRef] [PubMed]
- Bakowies, D.; Von Lilienfeld, O.A. Density Functional Geometries and Zero-Point Energies in Ab Initio Thermochemical Treatments of Compounds with First-Row Atoms (H, C, N, O, F). J. Chem. Theory Comput. 2021, 17, 4872–4890. [Google Scholar] [CrossRef] [PubMed]
- Karandashev, K.; Xu, Z.-H.; Meuwly, M.; Vaníček, J.; Richardson, J. Kinetic isotope effects and how to describe them. Struct. Dyn. 2017, 4, 061501. [Google Scholar] [CrossRef]
- Dale, H.J.A.; Leach, A.G.; Lloyd-Jones, G.C. Heavy-Atom Kinetic Isotope Effects: Primary Interest or Zero Point? J. Am. Chem. Soc. 2021, 143, 21079–21099. [Google Scholar] [CrossRef]
- Choi, D.-J.; Phark, S.; Heinrich, A.J.; Lorente, N. Electron spin resonance with scanning tunneling microscopy: A tool for an on-surface quantum platform of identical qubits. Nanoscale Adv. 2025, 7, 4551–4558. [Google Scholar] [CrossRef]
- Hishikawa, A.; Matsuda, A.; Fushitani, M. Ultrafast reaction imaging and control by ultrashort intense laser pulses. Bull. Chem. Soc. Jpn. 2020, 93, 1293–1304. [Google Scholar] [CrossRef]
- Lorenz, R. Quantum causal models: The merits of the spirit of Reichenbach’s principle for understanding quantum causal structure. Synthese 2022, 200, 424. [Google Scholar] [CrossRef]
- Alvarez-Mendoza, R.; Uboldi, L.; Lyons, A.; Cogdell, R.; Cerullo, G.; Faccio, D. Entangled-photon time- and frequency-resolved optical spectroscopy. arXiv 2025, arXiv:2505.02940. [Google Scholar]
- Walling, M.A.; Novak, J.A.; Shepard, J.R.E. Quantum Dots for Live Cell and In Vivo Imaging. Int. J. Mol. Sci. 2009, 10, 441–491. [Google Scholar] [CrossRef] [PubMed]
- Yukawa, H.; Kono, H.; Ishiwata, H.; Igarashi, R.; Takakusagi, Y.; Arai, S.; Hirano, Y.; Suhara, T.; Baba, Y. Quantum life science: Biological nano quantum sensors, quantum technology-based hyperpolarized MRI/NMR, quantum biology, and quantum biotechnology. Chem. Soc. Rev. 2025, 54, 3293–3322. [Google Scholar] [CrossRef] [PubMed]
- Nagarajan, K.; Thomas, A.; Ebbesen, T.W. Chemistry under Vibrational Strong Coupling. J. Am. Chem. Soc. 2021, 143, 16877–16889. [Google Scholar] [CrossRef]
- Lindoy, L.P.; Mandal, A.; Reichman, D.R. Quantum dynamical effects of vibrational strong coupling in chemical reactivity. Nat. Commun. 2023, 14, 2733. [Google Scholar] [CrossRef] [PubMed]
- Shen, Y.-Y.; Wang, B.; Ke, D.-Q.; Zheng, D.-D.; Li, Z.-H.; Wen, H.-F.; Guo, H.; Li, X.; Tang, J.; Ma, Z.-M.; et al. High-frequency resolution diamond nitrogen-vacancy center wide-spectrum imaging technology. Acta Phys. Sin. 2024, 73, 067601. [Google Scholar] [CrossRef]
- Hiura, H.; Shalabney, A.; George, J. Cavity Catalysis—Accelerating Reactions under Vibrational Strong Coupling. ChemRxiv 2018. Available online: https://chemrxiv.org/engage/chemrxiv/article-details/60c759bd0f50db605439873c (accessed on 6 January 2026).
- Burd, S.C.; Bagheri, N.; Ingaramo, M.; Condon, A.F.; Mondal, S.; Dowlatshahi, D.P.; Summers, J.A.; Mukherjee, S.; York, A.G.; Wakatsuki, S.; et al. Magnetic resonance control of reaction yields through genetically-encoded protein:flavin spin-correlated radicals in a live animal. bioRxiv 2025. [Google Scholar] [CrossRef]
- von Burg, V.; Low, G.H.; Häner, T.; Steiger, D.S.; Reiher, M.; Roetteler, M.; Troyer, M. Quantum computing enhanced computational catalysis. Phys. Rev. Res. 2021, 3, 033055. [Google Scholar] [CrossRef]
- Cheng, Y.; Ling, S.; Geng, Y.; Wang, Y.; Xu, J. Microfluidic synthesis of quantum dots and their applications in bio-sensing and bio-imaging. Nanoscale Adv. 2021, 3, 2180–2195. [Google Scholar] [CrossRef]
- Freddolino, L.; Harrison, C.B.; Liu, Y.; Schulten, K. Challenges in protein folding simulations: Timescale, representation, and analysis. Nat. Phys. 2010, 6, 751–758. [Google Scholar] [CrossRef]
- Samantray, S.; Schumann, W.; Illig, A.M.; Carballo-Pacheco, M.; Paul, A.; Barz, B.; Strodel, B. Molecular Dynamics Simulations of Protein Aggregation: Protocols for Simulation Setup and Analysis with Markov State Models and Transition Networks. In Methods in Molecular Biology; Humana: New York, NY, USA, 2022. [Google Scholar]
- Wang, W.; Singh, S.K.; Li, N.; Toler, M.R.; King, K.R.; Nema, S. Immunogenicity of protein aggregates—Concerns and realities. Int. J. Pharm. 2012, 431, 1–11. [Google Scholar] [CrossRef]
- Gillet, J.N. From molecular dynamics to quantum mechanics of misfolded proteins and amyloid-like macroaggregates applied to neurodegenerative diseases. J. Mol. Graph. Model. 2022, 110, 108046. [Google Scholar] [CrossRef] [PubMed]
- Wong, R.; Chang, W.L. Fast quantum algorithm for protein structure prediction in hydrophobic-hydrophilic model. J. Parallel Distrib. Comput. 2022, 164, 178–190. [Google Scholar] [CrossRef]
- Noshay, J.M.; Walker, T.; Alexander, W.G.; Klingeman, D.M.; Romero, J.; Walker, A.M.; Prates, E.; Eckert, C.; Irle, S.; Kainer, D.; et al. Quantum biological insights into CRISPR-Cas9 sgRNA efficiency from explainable-AI driven feature engineering. Nucleic Acids Res. 2023, 51, 10147–10161. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhou, X. Efficient quantum algorithm for lattice protein folding. Quantum Sci. Technol. 2025, 10, 015056. [Google Scholar] [CrossRef]
- Sarkar, M.; Roy, L.; Gutal, A.; Kumar, A.; Paranjothy, M. Quantum Simulations of Chemical Reactions: Achieving Accuracy with NISQ Devices. arXiv 2025, arXiv:2503.12084. [Google Scholar] [CrossRef]
- Guan, H.; Sun, H.; Zhao, X. Application of Density Functional Theory to Molecular Engineering of Pharmaceutical Formulations. Int. J. Mol. Sci. 2025, 26, 3262. [Google Scholar] [CrossRef]
- Bartlett, R.J.; Musiał, M. Coupled-cluster theory in quantum chemistry. Rev. Mod. Phys. 2007, 79, 291–352. [Google Scholar] [CrossRef]
- Vimal, D. PIMA Diabetes Prediction Using Machine Learning and Quantum Machine Learning Techniques. In Proceedings of the 2024 ITU Kaleidoscope: Innovation and Digital Transformation for a Sustainable World (ITU K), New Delhi, India, 21–23 October 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 1–7. [Google Scholar]
- Senthil, G.A.; Monica, K.M.; Prabha, R.; Prinslin, L.; Elavarasi, R. Quantum AI: A Cognitive Machine Learning Technique based on Nurturing Food Security Sustainability Predictive Analysis for Life Science—Bioengineering in Healthcare. BIO Web Conf. 2025, 172, 02002. [Google Scholar] [CrossRef]
- Neuling, N.R.; Allert, R.D.; Bucher, D.B. Prospects of single-cell nuclear magnetic resonance spectroscopy with quantum sensors. Curr. Opin. Biotechnol. 2023, 83, 102975. [Google Scholar] [CrossRef]
- Zaporski, L.; Liu, Q.; Velez, G.; Radzihovsky, M.; Li, Z.; Colombo, S.; Pedrozo-Peñafiel, E.; Vuletić, V. Quantum-amplified global-phase spectroscopy on an optical clock transition. Nature 2025, 646, 309–314. [Google Scholar] [CrossRef]
- Wu, W.; Davis, E.J.; Hughes, L.B.; Ye, B.; Wang, Z.; Kufel, D.; Ono, T.; Meynell, S.A.; Block, M.; Liu, C.; et al. Spin squeezing in an ensemble of nitrogen–vacancy centres in diamond. Nature 2025, 646, 74–80. [Google Scholar] [CrossRef]
- Dietze, K.; Pelzer, L.; Krinner, L.; Dawel, F.; Kramer, J.; Spethmann, N.C.H.; Kielinski, T.; Hammerer, K.; Stahl, K.; Klose, J.; et al. Entanglement-Enhanced Optical Ion Clock. arXiv 2025, arXiv:2506.11810. [Google Scholar] [CrossRef]
- Marques, J.M.C.; Prudente, F.V.; Pirani, F. Intermolecular Forces: From Atoms and Molecules to Nanostructures. Molecules 2022, 27, 3072. [Google Scholar] [CrossRef] [PubMed]
- Ritter, S.K. The art of the chemical bond. ACS Cent. Sci. 2016, 2, 769–772. [Google Scholar] [CrossRef]
- Korchef, A. Chemical Bonding and Molecular Geometry. In Understanding General Chemistry; CRC Press: Boca Raton, FL, USA, 2022. [Google Scholar]
- Akman, F.; Demirpolat, A.; Kazachenko, A.S.; Kazachenko, A.S.; Issaoui, N.; Al-Dossary, O. Molecular Structure, Electronic Properties, Reactivity (ELF, LOL, and Fukui), and NCI-RDG Studies of the Binary Mixture of Water and Essential Oil of Phlomis bruguieri. Molecules 2023, 28, 2684. [Google Scholar] [CrossRef]
- Ng, S.; Masarone, S.; Watson, D.; Barnes, M.R. The benefits and pitfalls of machine learning for biomarker discovery. Cell Tissue Res. 2023, 394, 17–31. [Google Scholar] [CrossRef] [PubMed]
- Liu, C.; Denzler, L.; Chen, Y.; Martin, A.; Paige, B. AsEP: Benchmarking Deep Learning Methods for Antibody-specific Epitope Prediction. arXiv 2024, arXiv:2407.18184. [Google Scholar]
- Zeng, X.; Bai, G.; Sun, C.; Ma, B. Recent Progress in Antibody Epitope Prediction. Antibodies 2023, 12, 52. [Google Scholar] [CrossRef]
- Hearty, S.; Leonard, P.; O’Kennedy, R. Measuring Antibody-Antigen Binding Kinetics Using Surface Plasmon Resonance. In Antibody Engineering; Humana Press: Totowa, NJ, USA, 2018; pp. 421–455. [Google Scholar] [CrossRef]
- Wu, K.; Vedelaar, T.A.; Damle, V.G.; Morita, A.; Mougnaud, J.; Martin, C.R.S.; Zhang, Y.; van der Pol, D.P.I.; Ende-Metselaar, H.; Rodenhuis-Zybert, I.; et al. Applying NV center-based quantum sensing to study intracellular free radical response upon viral infections. Redox Biol. 2022, 52, 102279. [Google Scholar] [CrossRef]
- Zheng, Y.; Song, K.; Cai, K.; Liu, L.; Tang, D.; Long, W.; Zhai, B.; Chen, J.; Tao, Y.; Zhao, Y.; et al. B-Cell-Epitope-Based Fluorescent Quantum Dot Biosensors for SARS-CoV-2 Enable Highly Sensitive COVID-19 Antibody Detection. Viruses 2022, 14, 1031. [Google Scholar] [CrossRef]
- Robert, A.; Barkoutsos, P.K.; Woerner, S.; Tavernelli, I. Resource-efficient quantum algorithm for protein folding. npj Quantum Inf. 2021, 7, 38. [Google Scholar] [CrossRef]
- Náray-Szabó, G.; Oláh, J.; Krámos, B. Quantum Mechanical Modeling: A Tool for the Understanding of Enzyme Reactions. Biomolecules 2013, 3, 662–702. [Google Scholar] [CrossRef]
- Tong, X.; Shi, S.; Tong, C.; Iftikhar, A.; Long, R.; Zhu, Y. Quantum/carbon dots-based fluorescent assays for enzyme activity. TrAC Trends Anal. Chem. 2020, 131, 116008. [Google Scholar] [CrossRef]
- Mzyk, A.; Sigaeva, A.; Schirhagl, R. Relaxometry with Nitrogen Vacancy (NV) Centers in Diamond. Acc. Chem. Res. 2022, 55, 3572–3580. [Google Scholar] [CrossRef] [PubMed]
- Roman-Vicharra, C.; Cai, J.J. Quantum gene regulatory networks. npj Quantum Inf. 2023, 9, 67. [Google Scholar] [CrossRef]
- Green, C.M.; Spangler, J.; Susumu, K.; Stenger, D.A.; Medintz, I.L.; Díaz, S.A. Quantum Dot-Based Molecular Beacons for Quantitative Detection of Nucleic Acids with CRISPR/Cas(N) Nucleases. ACS Nano 2022, 16, 20693–20704. [Google Scholar] [CrossRef]
- Giovannetti, V.; Lloyd, S.; Maccone, L. Quantum-enhanced measurements: Beating the standard quantum limit. Science 2004, 306, 1330–1336. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.-H.; Zhang, R.-Z.; Xu, Y.; Chen, X.-Q.; Zheng, H.; Li, Q.; Liu, R.-B.; Pan, X.-Y.; Budker, D.; Liu, G.-Q. Optically detected magnetic resonance of nitrogen-vacancy centers in diamond under weak laser excitation. Phys. Rev. Appl. 2024, 21, 044051. [Google Scholar] [CrossRef]
- Hansen, N.W.; Webb, J.L.; Troise, L.; Olsson, C.; Tomasevic, L.; Brinza, O.; Achard, J.; Staacke, R.; Kieschnick, M.; Meijer, J.; et al. Microscopic-scale magnetic recording of brain neuronal electrical activity using a diamond quantum sensor. Sci. Rep. 2023, 13, 12407. [Google Scholar] [CrossRef]
- Tan, Y.; Hu, X.; Hou, Y.; Chu, Z. Emerging Diamond Quantum Sensing in Bio-Membranes. Membranes 2022, 12, 957. [Google Scholar] [CrossRef]
- Crane, J.M.; Haggie, P.M.; Verkman, A.S. Quantum Dot Single Molecule Tracking Reveals a Wide Range of Diffusive Motions of Membrane Transport Proteins. In Proceedings of the Colloidal Quantum Dots for Biomedical Applications IV, San Jose, CA, USA, 24–26 January 2009. [Google Scholar]
- Hall, L.T.; Hill, C.D.; Cole, J.H.; Städler, B.; Caruso, F.; Mulvaney, P.; Wrachtrup, J.; Hollenberg, L.C.L. Monitoring ion-channel function in real time through quantum decoherence. Proc. Natl. Acad. Sci. USA 2010, 107, 18777–18782. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.J.; Von Kugelgen, S.; Laorenza, D.W.; Freedman, D.E. A Molecular Approach to Quantum Sensing. ACS Cent. Sci. 2021, 7, 712–723. [Google Scholar] [CrossRef]
- Li, X.; Zhao, W.; Liu, J.; Li, R.; Jia, S.; Li, C.; Bao, D.; Zhu, S.; Wang, P.; Huang, L.; et al. Harnessing non-equilibrium hot electrons in a quantum-engineered ternary heterostructure for sub-ppb C9 biomarker detection. Chem. Sci. 2025, 16, 17200–17206. [Google Scholar] [CrossRef]
- Pogorzelski, J.; Horsthemke, L.; Homrighausen, J.; Stiegekötter, D.; Gregor, M.; Glösekötter, P. Compact and Fully Integrated LED Quantum Sensor Based on NV Centers in Diamond. Sensors 2024, 24, 743. [Google Scholar] [CrossRef]
- Ajoy, A. High-Throughput in-Flow Quantum Sensing Based on Droplet Microfluidics. In Proceedings of the Quantum Effects and Measurement Techniques in Biology and Biophotonics, San Francisco, CA, USA, 27–30 January 2024. [Google Scholar]
- Al Shammri, F.K.; Al-Shareeda, M.A.; Abbood, A.A.J.; Almaiah, M.A.; AlAli, R. Quantum-Enhanced AI and Machine Learning: Transforming Predictive Analytics. In Recent Advances in Electrical & Electronic Engineering (Formerly Recent Patents on Electrical & Electronic Engineering); Bentham Science: Sharjah, United Arab Emirates, 2025; p. 18. [Google Scholar] [CrossRef]
- Kim, J.; McFee, M.; Fang, Q.; Abdin, O.; Kim, P.M. Computational and artificial intelligence-based methods for antibody development. Trends Pharmacol. Sci. 2023, 44, 175–189. [Google Scholar] [CrossRef]
- Basu, S.; Born, J.; Bose, A.; Capponi, S.; Chalkia, D.; Chan, T.A.; Doga, H.; Flother, F.F.; Getz, G.; Goldsmith, M.; et al. Towards Quantum-Enabled Cell-Centric Therapeutics. arXiv 2023, arXiv:2307.05734. [Google Scholar]
- Utro, F.; Tolunay, M.; Rhrissorrakrai, K.; Gujarati, T.P.; Shi, J.; Capponi, S.; Amico, M.; Earnest-Noble, N.; Parida, L. Enhanced Prediction of CAR T-Cell Cytotoxicity with Quantum-Kernel Methods. arXiv 2025, arXiv:2507.22710. [Google Scholar]
- Reymond, J.-L.; van Deursen, R.; Blum, L.C.; Ruddigkeit, L. Chemical space as a source for new drugs. Medchemcomm 2010, 1, 30. [Google Scholar] [CrossRef]
- Han, P.; Zhao, P.; Lu, C.; Huang, J.; Wu, J.; Shang, S.; Yao, B.; Zhang, X. GNN-Retro: Retrosynthetic Planning with Graph Neural Networks. In Proceedings of the 36th AAAI Conference on Artificial Intelligence, AAAI 2022, Vancouver, BC, Canada, 22 February–8 March 2022. [Google Scholar]
- Zhao, P.-C.; Wei, X.-X.; Wang, Q.; Wang, Q.-H.; Li, J.-N.; Shang, J.; Lu, C.; Shi, J.-Y. Single-step retrosynthesis prediction via multitask graph representation learning. Nat. Commun. 2025, 16, 814. [Google Scholar] [CrossRef] [PubMed]
- Koch, M.; Duigou, T.; Faulon, J.L. Reinforcement learning for bioretrosynthesis. ACS Synth. Biol. 2020, 9, 157–168. [Google Scholar] [CrossRef]
- Lu, J.-M.; Pan, J.-Z.; Mo, Y.-M.; Fang, Q. Automated Intelligent Platforms for High-Throughput Chemical Synthesis. Artif. Intell. Chem. 2024, 2, 100057. [Google Scholar] [CrossRef]
- Cavasotto, C.N.; Di Filippo, J.I. Artificial intelligence in the early stages of drug discovery. Arch. Biochem. Biophys. 2021, 698, 108730. [Google Scholar] [CrossRef]
- Wang, Y.; Pang, C.; Wang, Y.; Jin, J.; Zhang, J.; Zeng, X.; Su, R.; Zou, Q.; Wei, L. Retrosynthesis prediction with an interpretable deep-learning framework based on molecular assembly tasks. Nat. Commun. 2023, 14, 6155. [Google Scholar] [CrossRef]
- Reiher, M.; Wiebe, N.; Svore, K.M.; Wecker, D.; Troyer, M. Elucidating reaction mechanisms on quantum computers. Proc. Natl. Acad. Sci. USA 2017, 114, 7555–7560. [Google Scholar] [CrossRef]
- Fox, D.M.; MacDermaid, C.M.; Schreij, A.M.A.; Zwierzyna, M.; Walker, R.C. RNA folding using quantum computers. PLoS Comput. Biol. 2022, 18, e1010032. [Google Scholar] [CrossRef]
- Conde-Torres, D.; Mussa-Juane, M.; Faílde, D.; Gómez, A.; García-Fandiño, R.; Piñeiro, Á. Interface-Driven Peptide Folding: Quantum Computations on Simulated Membrane Surfaces. arXiv 2024, arXiv:2401.05075. [Google Scholar] [CrossRef]
- de la Fuente, J.; Contreras, M. Quantum vaccinomics platforms to advance in vaccinology. Front. Immunol. 2023, 14, 1172734. [Google Scholar] [CrossRef]
- Liu, J.; Guan, S.; Zou, Q.; Wu, H.; Tiwari, P.; Ding, Y. AMDGT: Attention aware multi-modal fusion using a dual graph transformer for drug–disease associations prediction. Knowl. Based Syst. 2024, 284, 111329. [Google Scholar] [CrossRef]
- Shen, S.; Qi, W.; Liu, X.; Zeng, J.; Li, S.; Zhu, X.; Dong, C.; Wang, B.; Shi, Y.; Yao, J.; et al. From virtual to reality: Innovative practices of digital twins in tumor therapy. J. Transl. Med. 2025, 23, 348. [Google Scholar] [CrossRef] [PubMed]
- Mohkam, M.; Sadraeian, M.; Lauto, A.; Gholami, A.; Nabavizadeh, S.H.; Esmaeilzadeh, H.; Alyasin, S. Exploring the potential and safety of quantum dots in allergy diagnostics. Microsyst. Nanoeng. 2023, 9, 145. [Google Scholar] [CrossRef]
- Papalitsas, C.; Guan, Y.; Waghe, S.; Liakos, A.; Balatsos, I.; Pantazopoulos, V. Quantum Approximate Optimization Algorithms for Molecular Docking. arXiv 2025, arXiv:2503.04239. [Google Scholar] [CrossRef]
- Jameel, F.; Stein, M. Chemical accuracy for ligand-receptor binding Gibbs energies through multi-level SQM/QM calculations. Phys. Chem. Chem. Phys. 2024, 26, 21197–21203. [Google Scholar] [CrossRef] [PubMed]
- Hoja, J.; Tkatchenko, A. First-principles stability ranking of molecular crystal polymorphs with the DFT+MBD approach. Faraday Discuss. 2018, 211, 253–274. [Google Scholar] [CrossRef]
- Hoja, J.; Ko, H.Y.; Neumann, M.A.; Car, R.; DiStasio, R.A.; Tkatchenko, A. Reliable and practical computational description of molecular crystal polymorphs. Acta Hortic. Sin. 2019, 45, eaau3338. [Google Scholar] [CrossRef]
- Feller, D. Climbing mount PSI: The level of theory needed to obtain 1 KCAL:/MOL accuracy in thermochemical properties. In Abstracts of Papers of the American Chemical Society; American Chemical Society: Washington, DC, USA, 1999; Volume 44. [Google Scholar]
- Baker, T.E.; Burke, K.; White, S.R. Accurate correlation energies in one-dimensional systems from small system-adapted basis functions. Phys. Rev. B 2018, 97, 085139. [Google Scholar] [CrossRef]
- Behera, S.; Hahn, D.F.; Wilson, C.J.; Marsili, S.; Tresadern, G.; Gapsys, V.; de Groot, B.L. Quantification of the Impact of Structure Quality on Predicted Binding Free Energy Accuracy. J. Chem. Inf. Model. 2025, 65, 6927–6938. [Google Scholar] [CrossRef] [PubMed]
- Salo-Ahen, O.M.H.; Alanko, I.; Bhadane, R.; Bonvin, A.M.J.J.; Honorato, R.V.; Hossain, S.; Juffer, A.H.; Kabedev, A.; Lahtela-Kakkonen, M.; Larsen, A.S.; et al. Molecular dynamics simulations in drug discovery and pharmaceutical development. Processes 2021, 9, 71. [Google Scholar] [CrossRef]
- Kállay, M.; Horváth, R.A.; Gyevi-Nagy, L.; Nagy, P.R. Basis Set Limit CCSD(T) Energies for Extended Molecules via a Reduced-Cost Explicitly Correlated Approach. J. Chem. Theory Comput. 2023, 19, 174–189. [Google Scholar] [CrossRef]
- Lesiuk, M. Quintic-scaling rank-reduced coupled cluster theory with single and double excitations. J. Chem. Phys. 2022, 156, 064103. [Google Scholar] [CrossRef]
- Nandi, A.; Pandey, P.; Houston, P.L.; Qu, C.; Yu, Q.; Conte, R.; Tkatchenko, A.; Bowman, J.M. Δ-Machine Learning to Elevate DFT-based Potentials and a Force Field to the CCSD(T) Level Illustrated for Ethanol. J. Chem. Theory Comput. 2024, 20, 8807–8819. [Google Scholar] [CrossRef]
- Nandi, A.; Qu, C.; Houston, P.L.; Conte, R.; Bowman, J.M. Δ-machine learning for potential energy surfaces: A PIP approach to bring a DFT-based PES to CCSD(T) level of theory. J. Chem. Phys. 2021, 154, 051102. [Google Scholar] [CrossRef]
- Abdolmaleki, S.; Aliabadi, A.; Khaksar, S. Riding the metal wave: A review of the latest developments in metal-based anticancer agents. Coord. Chem. Rev. 2024, 501, 215579. [Google Scholar] [CrossRef]
- Sheng, X.; Kazemi, M.; Planas, F.; Himo, F. Modeling Enzymatic Enantioselectivity using Quantum Chemical Methodology. ACS Catal. 2020, 10, 6430–6449. [Google Scholar] [CrossRef]
- Robo, M.T.; Hayes, R.L.; Ding, X.; Pulawski, B.; Vilseck, J.Z. Fast free energy estimates from λ-dynamics with bias-updated Gibbs sampling. Nat. Commun. 2023, 14, 8515. [Google Scholar] [CrossRef]
- Champion, C.; Gall, R.; Ries, B.; Rieder, S.R.; Barros, E.P.; Riniker, S. Accelerating Alchemical Free Energy Prediction Using a Multistate Method: Application to Multiple Kinases. J. Chem. Inf. Model. 2023, 63, 7133–7147. [Google Scholar] [CrossRef]
- Bueno, P.R. Quantum electroanalysis in drug discovery. Chem. Commun. 2025, 61, 8632–8642. [Google Scholar] [CrossRef] [PubMed]
- Santos, J.; Pujols, J.; Pallarès, I.; Iglesias, V.; Ventura, S. Computational prediction of protein aggregation: Advances in proteomics, conformation-specific algorithms and biotechnological applications. Comput. Struct. Biotechnol. J. 2020, 18, 1403–1413. [Google Scholar] [CrossRef] [PubMed]
- Hu, H. Quantum Computer Simulation of Protein Protonation. J. Chem. Theory Comput. 2023, 19, 5671–5676. [Google Scholar] [CrossRef]
- Takaya, D.; Ohno, S.; Miyagishi, T.; Tanaka, S.; Okuwaki, K.; Watanabe, C.; Kato, K.; Tian, Y.-S.; Fukuzawa, K. Quantum chemical calculation dataset for representative protein folds by the fragment molecular orbital method. Sci. Data 2024, 11, 1164. [Google Scholar] [CrossRef]
- Danhof, M.; Klein, K.; Stolk, P.; Aitken, M.; Leufkens, H. The future of drug development: The paradigm shift towards systems therapeutics. Drug Discov. Today 2018, 23, 1990–1995. [Google Scholar] [CrossRef]
- Posey Norris, S.M.; Pankevich, D.E.; Davis, M.; Altevogt, B.M. Improving and Accelerating Therapeutic Development for Nervous System Disorders: Workshop Summary; National Academies Press: Washington, DC, USA, 2014. [Google Scholar]
- Ocana, A.; Pandiella, A.; Privat, C.; Bravo, I.; Luengo-Oroz, M.; Amir, E.; Gyorffy, B. Integrating artificial intelligence in drug discovery and early drug development: A transformative approach. Biomark. Res. 2025, 13, 45. [Google Scholar] [CrossRef]
- Malheiro, V.; Santos, B.; Figueiras, A.; Mascarenhas-Melo, F. The Potential of Artificial Intelligence in Pharmaceutical Innovation: From Drug Discovery to Clinical Trials. Pharmaceuticals 2025, 18, 788. [Google Scholar] [CrossRef] [PubMed]
- Woo, H.-M.; Qian, X.; Tan, L.; Jha, S.; Alexander, F.J.; Dougherty, E.R.; Yoon, B.-J. Optimal decision-making in high-throughput virtual screening pipelines. Patterns 2023, 4, 100875. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Hu, J.; Zhang, X.; Jin, S.; Xu, X. Drug target affinity prediction based on multi-scale gated power graph and multi-head linear attention mechanism. PLoS ONE 2025, 20, e0315718. [Google Scholar] [CrossRef]
- Fromer, J.C.; Graff, D.E.; Coley, C.W. Pareto optimization to accelerate multi-objective virtual screening. Digit. Discov. 2024, 3, 467–481. [Google Scholar] [CrossRef]
- Wang, W.; Liu, Y.; Wang, Z.; Hao, G.; Song, B. The way to AI-controlled synthesis: How far do we need to go? Chem. Sci. 2022, 13, 12604–12615. [Google Scholar] [CrossRef] [PubMed]
- Philippidis, A. Pipeline Pruning: Exscientia Cuts Programs to Sharpen Precision Oncology Focus. GEN Edge 2023, 5, 889–897. [Google Scholar] [CrossRef]
- Philippidis, A. In the Pipeline: Recursion’s Approach to AI and Machine Learning. GEN Biotechnol. 2024, 3, 269–273. [Google Scholar] [CrossRef]
- Redl, I.; Fisicaro, C.; Dutton, O.; Hoffmann, F.; Henderson, L.; Owens, B.M.J.; Heberling, M.; Paci, E.; Tamiola, K. ADOPT: Intrinsic protein disorder prediction through deep bidirectional transformers. NAR Genom. Bioinform. 2023, 5, lqad041. [Google Scholar] [CrossRef]
- Thomas, D.A.; Fleming, T.A.; Bittner, M.-I. Abstract 295: Implementing fully automated kinase inhibitor characterization using a robotic system. Cancer Res. 2021, 81, 295. [Google Scholar] [CrossRef]
- Segal, M. An operating system for the biology lab. Nature 2019, 573, S112–S113. [Google Scholar] [CrossRef]
- YWong, G.K.; Costa, K.D.; Fermini, B.; Li, R.A. Modeling the Heart with Novoheart’s MyheartTM Platform. Future Drug Discov. 2020, 2, FDD32. [Google Scholar] [CrossRef]
- Hough, S.H.; Ajetunmobi, A.; Brody, L.; Humphryes-Kirilov, N.; Perello, E. Desktop genetics. Pers. Med. 2016, 13, 517–521. [Google Scholar] [CrossRef] [PubMed]
- Kumar, K.; Chupakhin, V.; Vos, A.; Morrison, D.; Rassokhin, D.; Dellwo, M.J.; McCormick, K.; Paternoster, E.; Ceulemans, H.; DesJarlais, R.L. Development and implementation of an enterprise-wide predictive model for early absorption, distribution, metabolism and excretion properties. Future Med. Chem. 2021, 13, 1639–1654. [Google Scholar] [CrossRef]
- Ghiandoni, G.M.; Evertsson, E.; Riley, D.J.; Tyrchan, C.; Rathi, P.C. Augmenting DMTA using predictive AI modelling at AstraZeneca. Drug Discov. Today 2024, 29, 103945. [Google Scholar] [CrossRef] [PubMed]
- Benson, B.M.; Ingman, V.M.; Agarwal, A.; Keinan, S. A CQM-Based Approach to Solving a Combinatorial Problem with Applications in Drug Design. arXiv 2023, arXiv:2303.15419. [Google Scholar] [CrossRef]
- Chandran, A. Biopharma foresees a ‘quantum advantage’: They could be right. Nat. Biotechnol. 2024, 42, 690–692. [Google Scholar] [CrossRef]
- Feniou, C.; Hassan, M.; Claudon, B.; Courtat, A.; Adjoua, O.; Maday, Y.; Piquemal, J.-P. Greedy gradient-free adaptive variational quantum algorithms on a noisy intermediate scale quantum computer. Sci. Rep. 2025, 15, 18689. [Google Scholar] [CrossRef]
- Petch, J.; Di, S.; Nelson, W. Opening the Black Box: The Promise and Limitations of Explainable Machine Learning in Cardiology. Can. J. Cardiol. 2022, 38, 204–213. [Google Scholar] [CrossRef]
- Liu, Y. Superconducting quantum computing optimization based on multi-objective deep reinforcement learning. Sci. Rep. 2025, 15, 3828. [Google Scholar] [CrossRef]
- Zhong, H.; Ju, K.; Shen, J.; Zhang, X.; Qin, X.; Ohtsuki, T.; Pan, M.; Han, Z. Differential Privacy Preserving Distributed Quantum Computing. arXiv 2025, arXiv:2412.12387. [Google Scholar]
- Kannan, E.; MJ, C.M.B.; Ravikumar, S.; Alex, D.S.; Sriram, K.; Vijay, K. Quantum-Safe Federated Learning: Enhancing Data Privacy and Security. In Proceedings of the 2024 International Conference on Emerging Research in Computational Science (ICERCS), Coimbatore, India, 12–14 December 2024; IEEE: Piscataway, NJ, USA, 2024; pp. 1–6. [Google Scholar]
- Gao, X.; Zhang, Z.Y.; Duan, L.M. A quantum machine learning algorithm based on generative models. Sci. Adv. 2018, 4, eaat9004. [Google Scholar] [CrossRef] [PubMed]
- Dang, T.L.P.; Sadreddin, A.; Ahuja, S. Readily available technologies in low-resource communities: A review and synthesis. Inf. Technol. Dev. 2024, 30, 132–172. [Google Scholar] [CrossRef]
- Knapik, E.; Brandimarte, L.; Usher, M. Maintenance in sustainable stormwater management: Issues, barriers and challenges. J. Environ. Plan. Manag. 2025, 68, 2769–2795. [Google Scholar] [CrossRef]
- Whalen, C.J.; Donnell, D.; Tartakovsky, M. Supporting research sites in resource-limited settings: Challenges in implementing information technology infrastructure. J. Acquir. Immune Defic. Syndr. 2014, 65, S44–S49. [Google Scholar] [CrossRef]
- Jindal, M. How Can Hybrid Algorithms Combining Quantum Machine Learning and Classical Machine Learning be Optimised for Performance and Accuracy? Int. J. Multidiscip. Res. 2024, 6, 33629. [Google Scholar] [CrossRef]
- Zhang, X.-M.; Li, T.; Yuan, X. Quantum State Preparation with Optimal Circuit Depth: Implementations and Applications. Phys. Rev. Lett. 2022, 129, 230504. [Google Scholar] [CrossRef]
- Aiello, C.D.; Awschalom, D.D.; Bernien, H.; Brower, T.; Brown, K.R.; Brun, T.A.; Caram, J.R.; Chitambar, E.; Di Felice, R.; Edmonds, K.M. Achieving a quantum smart workforce. Quantum Sci. Technol. 2021, 6, 030501. [Google Scholar] [CrossRef]
- De Michielis, M.; Ferraro, E. Energy and power scaling in quantum computers based on rotated surface codes with silicon flip-flop qubits. EPJ Quantum Technol. 2025, 12, 64. [Google Scholar] [CrossRef]
- Ennab, M.; Mcheick, H. Enhancing interpretability and accuracy of AI models in healthcare: A comprehensive review on challenges and future directions. Front. Robot. AI 2024, 11, 1444763. [Google Scholar] [CrossRef]
- Lellouch, S.; Holynski, M. Integration of a high-fidelity model of quantum sensors with a map-matching filter for quantum-enhanced navigation. Quantum Sci. Technol. 2025, 10, 045007. [Google Scholar] [CrossRef]
- Christensen, M.; Yunker, L.P.E.; Shiri, P.; Zepel, T.; Prieto, P.L.; Grunert, S.; Borka, F.; Hein, J.E. Automation isn’t automatic. Chem. Sci. 2021, 12, 15473–15490. [Google Scholar] [CrossRef]
- Hosseinzadehtaher, M.; D’silva, S.; Baker, M.; Kumar, R.; Hein, N.T.; Shadmand, M.B.; Jagadish, S.V.K.; Ghanbarian, B. On Design Challenges of Portable Nuclear Magnetic Resonance System. J. Nucl. Eng. 2023, 4, 323–337. [Google Scholar] [CrossRef]
- Kennedy, I.G.; Whitehead, D.; Ferdinand-James, D. Serendipity: A way of stimulating researchers’ creativity. J. Creat. 2022, 32, 100014. [Google Scholar] [CrossRef]
- Kothari, A.R.; Bickford, J.J.; Edwards, N.; Dobbins, M.J.; Meyer, M. Uncovering tacit knowledge: A pilot study to broaden the concept of knowledge in knowledge translation. BMC Health Serv. Res. 2011, 11, 198. [Google Scholar] [CrossRef]
- Guedes, M.; de la Serna Bazan, A.; Rubio-Martín, E.; Pulido, L.B.; Palomo, V.; Piljić, A.; Leclerc, Q.J.; Aris, E.; Vella, V.; Dambrauskienė, A.; et al. How to: Share and reuse data—Challenges and solutions from predicting the impact of monoclonal antibodies & vaccines on antimicrobial resistance project. Clin. Microbiol. Infect. 2025, 31, 753–760. [Google Scholar] [CrossRef] [PubMed]
- Ballantyne, A. How should we think about clinical data ownership? J. Med. Ethics 2020, 46, 289–294. [Google Scholar] [CrossRef]
- Patino, R.M. Moving research to patient applications through commercialization: Understanding and evaluating the role of intellectual property. J. Am. Assoc. Lab. Anim. Sci. 2010, 49, 147–154. [Google Scholar]
- Pratt, B.; Hyder, A.A. Governance of global health research consortia: Sharing sovereignty and resources within Future Health Systems. Soc. Sci. Med. 2017, 174, 113–121. [Google Scholar] [CrossRef]
- Dutta, M.; Ramasubramanian, S.; Barrett, M.; Elers, C.; Sarwatay, D.; Raghunath, P.; Kaur, S.; Dutta, D.; Jayan, P.; Rahman, M.; et al. Decolonizing Open Science: Southern Interventions. J. Commun. 2021, 71, 803–826. [Google Scholar] [CrossRef]
- Wilkinson, B.D. The Limits of Neuroscience in Counseling: A Humanistic Perspective and Proposed Model. J. Humanist. Couns. 2018, 57, 70–78. [Google Scholar] [CrossRef]
- Frank, A.; Gleiser, M.; Thompson, E. The Blind Spot: Why Science Cannot Ignore Human Experience; The MIT Press: Cambridge, MA, USA, 2024. [Google Scholar]
- Ricci, G. Pharmacological Human Enhancement: An Overview of the Looming Bioethical and Regulatory Challenges. Front. Psychiatry 2020, 11, 53. [Google Scholar] [CrossRef]
- Barra, M.E.; Solt, K.; Yu, X.; Edlow, B.L. Restoring consciousness with pharmacologic therapy: Mechanisms, targets, and future directions. Neurotherapeutics 2024, 21, e00374. [Google Scholar] [CrossRef]
- Wenzel, T. Open hardware: From DIY trend to global transformation in access to laboratory equipment. PLoS Biol. 2023, 21, e3001931. [Google Scholar] [CrossRef]
- Sarkar, S.; Mateus, S. Doing more with less—How frugal innovations can contribute to improving healthcare systems. Soc. Sci. Med. 2022, 306, 115127. [Google Scholar] [CrossRef] [PubMed]
- Mugambi, M.L.; Peter, T.; Martins, S.F.; Giachetti, C. How to implement new diagnostic products in low-resource settings: An end-to-end framework. BMJ Glob. Health 2018, 3, e000914. [Google Scholar] [CrossRef] [PubMed]
- Pearce, J.M. Economic savings for scientific free and open source technology: A review. HardwareX 2020, 8, e00139. [Google Scholar] [CrossRef] [PubMed]
- D’Alton, L.; Carrara, S.; Barbante, G.J.; Hoxley, D.; Hayne, D.J.; Francis, P.S.; Hogan, C.F. A simple, low-cost instrument for electrochemiluminescence immunoassays based on a Raspberry Pi and screen-printed electrodes. Bioelectrochemistry 2022, 146, 108107. [Google Scholar] [CrossRef]
- Rujano, M.A.; Boiten, J.-W.; Ohmann, C.; Canham, S.; Contrino, S.; David, R.; Ewbank, J.; Filippone, C.; Connellan, C.; Custers, I.; et al. Sharing sensitive data in life sciences: An overview of centralized and federated approaches. Brief. Bioinform. 2024, 25, bbae262. [Google Scholar] [CrossRef]
- Kotev, M.; Diaz Gonzalez, C. Molecular Dynamics and Other HPC Simulations for Drug Discovery. Methods Mol. Biol. 2024, 2716, 265–291. [Google Scholar] [CrossRef]
- Kushwaha, P.; Srivastava, N.; Kushwaha, S.P. Enhancing clinical drug trial monitoring with blockchain technology. Contemp. Clin. Trials 2024, 146, 107684. [Google Scholar] [CrossRef] [PubMed]
- Keršič, V.; Karakatič, S.; Turkanović, M. On-chain zero-knowledge machine learning: An overview and comparison. J. King Saud Univ. Comput. Inf. Sci. 2024, 36, 102207. [Google Scholar] [CrossRef]
- Cha, Y.; Erez, T.; Reynolds, I.J.; Kumar, D.; Ross, J.; Koytiger, G.; Kusko, R.; Zeskind, B.; Risso, S.; Kagan, E.; et al. Drug repurposing from the perspective of pharmaceutical companies. Br. J. Pharmacol. 2018, 175, 168–180. [Google Scholar] [PubMed]
- Stevens, W. Good clinical laboratory practice (GCLP): The need for a hybrid of good laboratory practice and good clinical practice guidelines/standards for medical testing laboratories conducting clinical trials in developing countries. Qual. Assur. 2003, 10, 83–89. [Google Scholar] [CrossRef]
- Satterthwaite, E.V.; Bax, N.J.; Miloslavich, P.; Ratnarajah, L.; Canonico, G.; Dunn, D.; Simmons, S.E.; Carini, R.J.; Evans, K.; Allain, V.; et al. Establishing the Foundation for the Global Observing System for Marine Life. Front. Mar. Sci. 2021, 8, 737416. [Google Scholar] [CrossRef]
- Matarése, B.F.E.; Desai, R.; Oughton, D.H.; Motherisill, C. EGO to ECO: Tracing the History of Radioecology from the 1950s to the Present Day. Radiat. Res. 2024, 202, 273–288. [Google Scholar] [CrossRef]
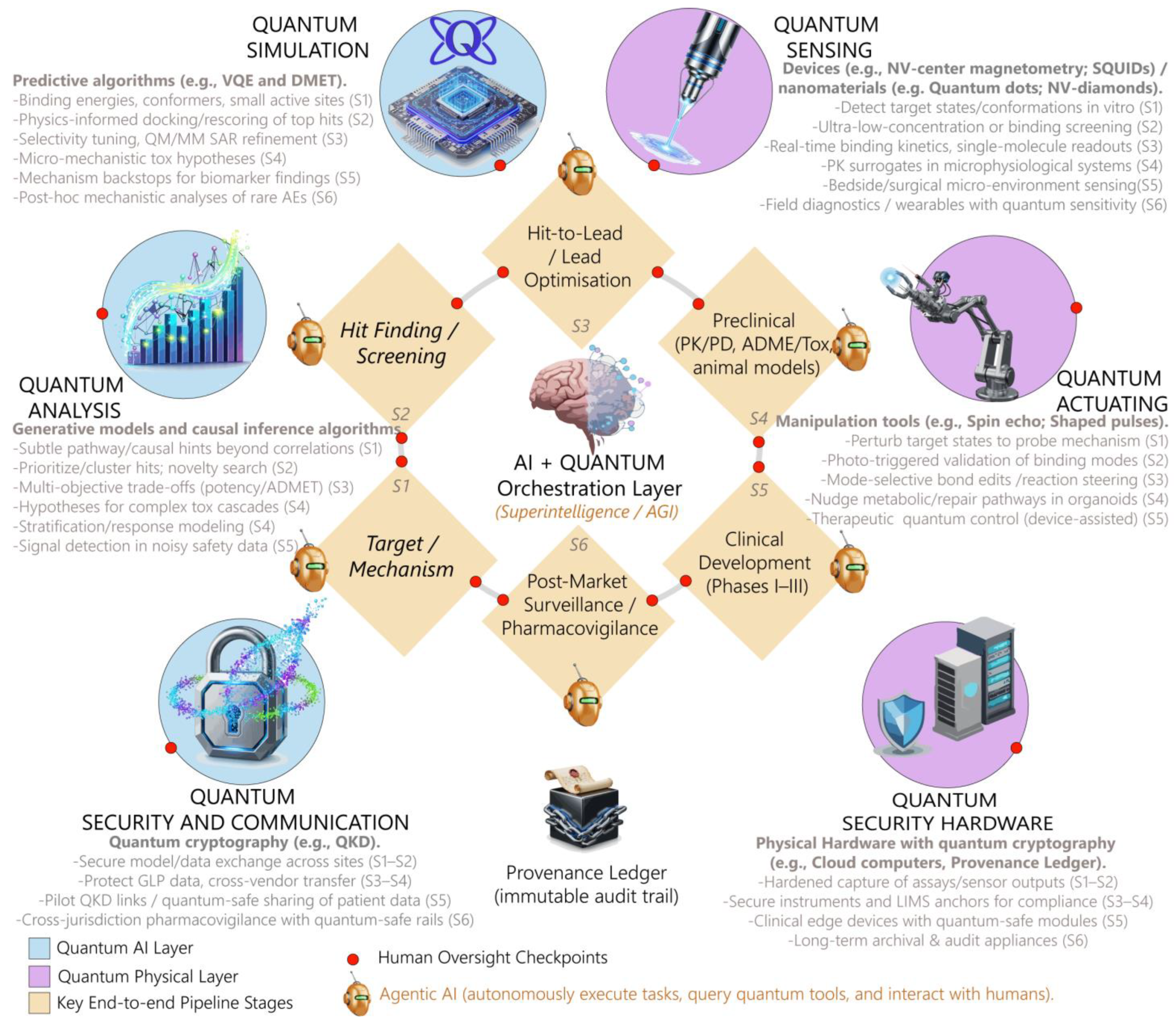


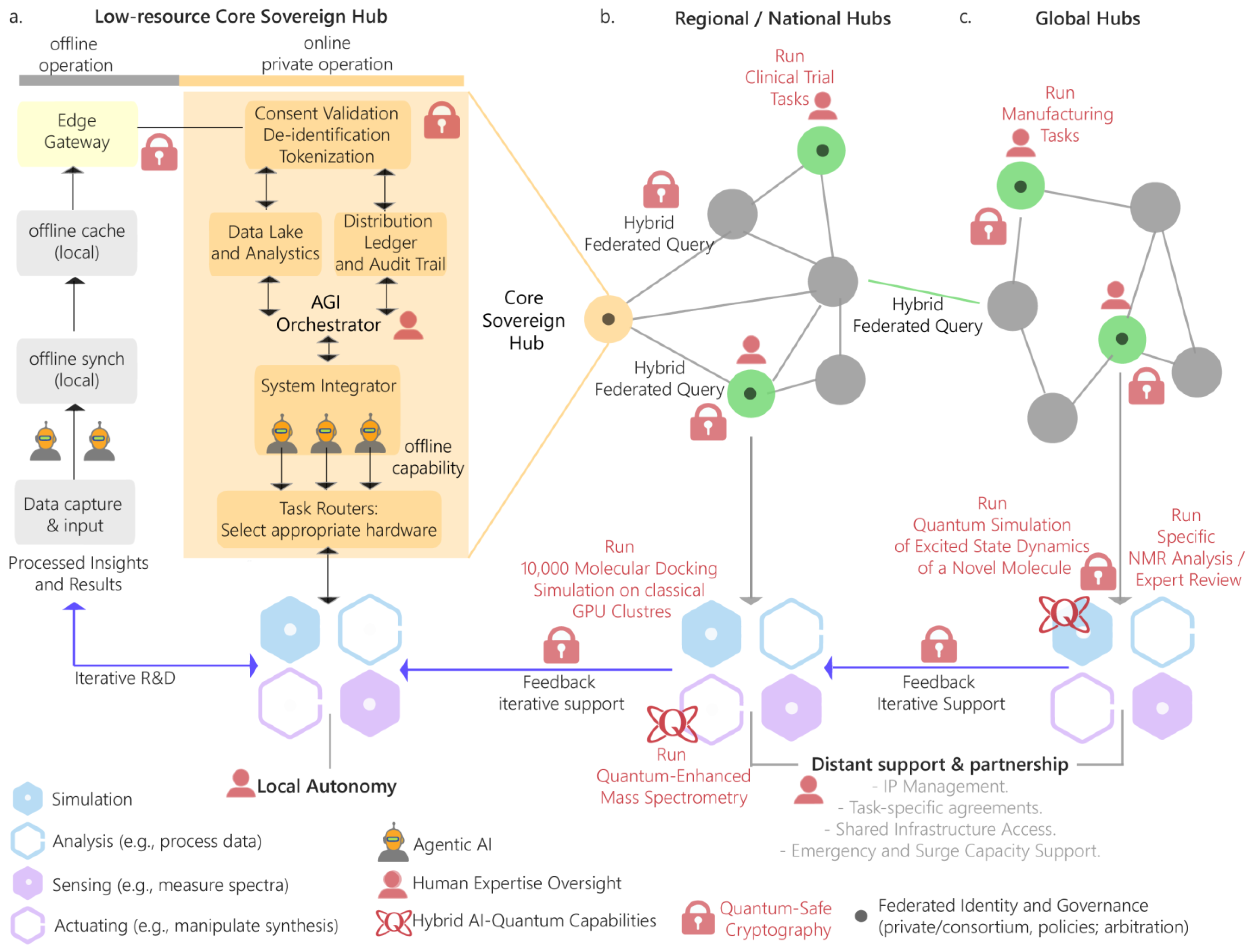
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Matarèse, B.F.E. Quantum and Artificial Intelligence in Drugs and Pharmaceutics. BioChem 2026, 6, 2. https://doi.org/10.3390/biochem6010002
Matarèse BFE. Quantum and Artificial Intelligence in Drugs and Pharmaceutics. BioChem. 2026; 6(1):2. https://doi.org/10.3390/biochem6010002
Chicago/Turabian StyleMatarèse, Bruno F. E. 2026. "Quantum and Artificial Intelligence in Drugs and Pharmaceutics" BioChem 6, no. 1: 2. https://doi.org/10.3390/biochem6010002
APA StyleMatarèse, B. F. E. (2026). Quantum and Artificial Intelligence in Drugs and Pharmaceutics. BioChem, 6(1), 2. https://doi.org/10.3390/biochem6010002






