An Automated Medical Diagnosis System for Neoplasm Medical Image Classification Using Supervised and Unsupervised Techniques †
Abstract
1. Introduction
2. Proposed NMICS
2.1. Data Collection Stage
2.2. Data Preprocessing Stage
2.2.1. Resizing
2.2.2. Noise Reduction
2.3. Feature Extraction Stage
| Algorithm 1: Feature Extraction |
Output: The set of images with extracted features. Begin
|
2.4. Training Stage
| Algorithm 2: Training Data Model using Clustering |
| Input: The set of images with extracted features. Output: The two distinct clusters (tumor) are obtained. Begin
|
2.5. Testing and Classification Stage
| Algorithm 3: Classification |
| Input: Trained data set with two clusters and feature vector of test sample features Output: KNN classification result either tum as tumor or no tumor. Begin
|
3. Results and Discussion
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Guy-Fernand, K.N.; Zhao, J.J.; Sabuni, F.M.; Wang, J. Classification of Brain Tumor Leveraging Goal-Driven Visual Attention with the Support of Transfer Learning. In Proceedings of the Information Communication Technologies Conference, Kigali, Rwanda, 6–8 May 2020; pp. 328–332. [Google Scholar]
- Jayade, S.; Ingole, D.T.; Ingole, M.D. Review of Brain Tumor Detection Concept using MRI Images. In Proceedings of the International Conference on Innovative Trends and Advances in Engineering and Technology, Shegoaon, India, 27–28 December 2019; pp. 206–209. [Google Scholar]
- Patil, D.O.; Hamde, S.T. Monogenic Wavelet Phase Encoded Descriptors for Brain Tumor Image Detection. In Proceedings of the IEEE Third International Conference on Multimedia Processing, Communication & Information Technology, Shivamogga, India, 11–12 December 2020; pp. 40–44. [Google Scholar]
- Gouskir, M.; Zyad, M.A.; Boutalline, M. Automatic Analysis of Brain Tumor from Magnetic Resonance Images based on Geometric Median Shift. In Proceedings of the 2nd International Conference on Knowledge-Based Engineering and Innovation, Beni Mellal, Morocco, 20–21 April 2020; Volume 8, pp. 11–17. [Google Scholar]
- Grampurohit, S.; Shalavadi, V.; Dhotargavi, V.R.; Kudari, M. Brain Tumor Detection Using Deep Learning Models. In Proceedings of the IEEE India Council International Subsections Conference, Visakhapatnam, India, 3–4 October 2020; pp. 129–134. [Google Scholar]
- Kabir, A.; Gyorfi, A.; Kovacs, L.; Szilagyi, L.O. Early-Stage Brain Tumor Detection on MRI Image Using a Hybrid Technique. In Proceedings of the IEEE Region 10 Symposium, Dhaka, Bangladesh, 5–7 June 2020; pp. 1828–1831. [Google Scholar]
- Gyorfi, A.; Sudirman, R.; Wei, S.T.C. A feature ranking and selection algorithm for brain tumor segmentation in multi spectral magnetic resonance image data. In Proceedings of the 2019 41st Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Berlin, Germany, 23–27 July 2019; pp. 804–807. [Google Scholar]
- Chew, K.M.; Yong, C.Y.; Sudirman, R.; Wei, S.T.C. Human Brain Modeling Tumor Detection in 2D and 3D Representation Using Microwave Signal Analysis. In Proceedings of the 2018 IEEE Symposium on Computer Applications & Industrial Electronics (ISCAIE), Penang, Malaysia, 28–29 April 2018; pp. 310–316. [Google Scholar]
- Polly, F.P.; Aziz, A.I.; Rana, M.; Islam, A.; Inum, R. Detection and Classification of HGG and LGG Brain Tumor Using Machine Learning, Expert Systems with Applications. In Proceedings of the 2018 International Conference on Information Networking (ICOIN), Chiang Mai, Thailand, 10–12 January 2018; pp. 813–817. [Google Scholar]
- Kumar, S.; Ahmed, S.T.; Sandeep, S.; Madheswaran, M.; Basha, S.M. Unstructured Oncological Image Cluster Identification Using Improved Unsupervised Clustering Techniques. Comput. Mater. Contin. 2022, 72, 281–298. [Google Scholar] [CrossRef]
- Sreedhar Kumar, S.; Ahmed, S.T.; Fathima, A.S.; Mathivanan, S.K.; Jayagopal, P.; Saif, A.; Gupta, S.K.; Sinha, G. iLIAC: An approach of identifying dissimilar groups on unstructured numerical image dataset using improved agglomerative clustering technique. Multimed. Tools Appl. 2024, 83, 86359–86381. [Google Scholar]
- Ahmed, S.T.; Kumar, S.; Guptha, N.S.; Basha, S.M.; Fathima, A.S. Improving medical image pixel quality using micq unsupervised machine learning technique. Malays. J. Comput. Sci. 2022, 2022, 53–64. [Google Scholar] [CrossRef]
- (Msoud Nickparvar). Available online: https://www.kaggle.com/datasets/masoudnickparvar/brain-tumor-mri-dataset (accessed on 10 August 2025).





| Test MRI Image Sample | Improved MRI Samples | Feature of MRI Samples | Classification Result | |
|---|---|---|---|---|
| Cluster 1 (Normal) | Cluster 2 (Tumor) | |||
 |  |  |  | |
 | 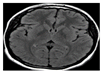 |  |  | |
 |  |  |  | |
 |  |  |  | |
 |  |  |  | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Seetharaman, S.K.; Kumar, B.; Rajanna, M.C.; Ahmed, S.T. An Automated Medical Diagnosis System for Neoplasm Medical Image Classification Using Supervised and Unsupervised Techniques. Eng. Proc. 2026, 124, 49. https://doi.org/10.3390/engproc2026124049
Seetharaman SK, Kumar B, Rajanna MC, Ahmed ST. An Automated Medical Diagnosis System for Neoplasm Medical Image Classification Using Supervised and Unsupervised Techniques. Engineering Proceedings. 2026; 124(1):49. https://doi.org/10.3390/engproc2026124049
Chicago/Turabian StyleSeetharaman, Sreedhar Kumar, Basant Kumar, Manjunath Chikkanjinappa Rajanna, and Syed Thouheed Ahmed. 2026. "An Automated Medical Diagnosis System for Neoplasm Medical Image Classification Using Supervised and Unsupervised Techniques" Engineering Proceedings 124, no. 1: 49. https://doi.org/10.3390/engproc2026124049
APA StyleSeetharaman, S. K., Kumar, B., Rajanna, M. C., & Ahmed, S. T. (2026). An Automated Medical Diagnosis System for Neoplasm Medical Image Classification Using Supervised and Unsupervised Techniques. Engineering Proceedings, 124(1), 49. https://doi.org/10.3390/engproc2026124049






