Application of Nanotechnology in COVID-19 Infection: Findings and Limitations
Abstract
1. Introduction
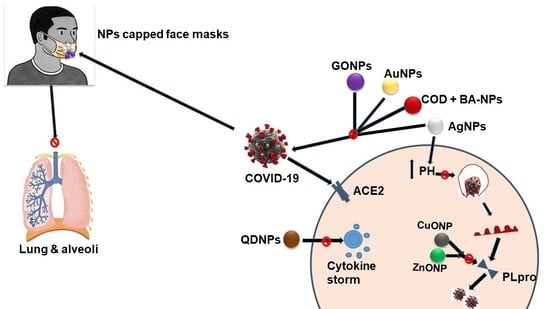
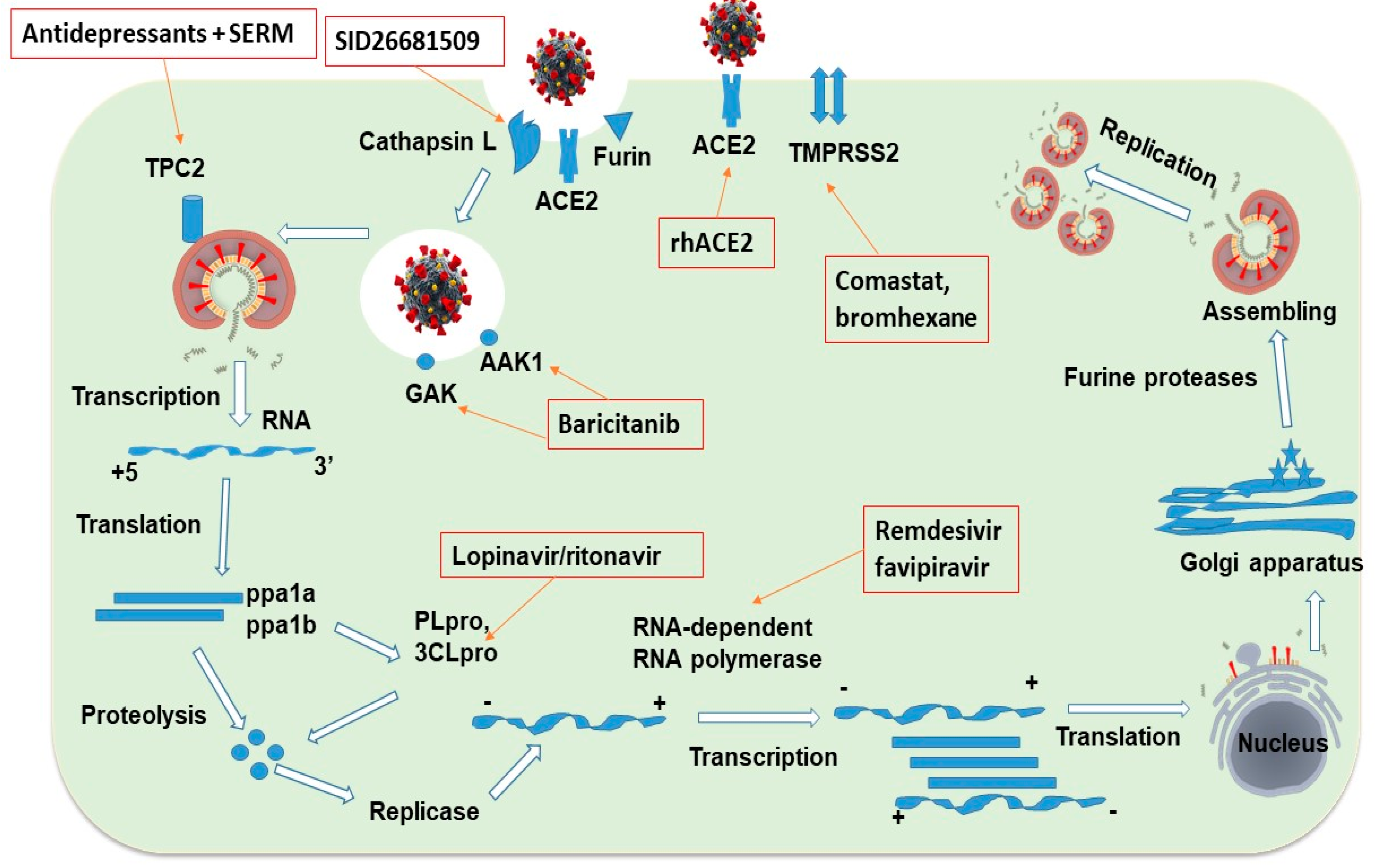
2. Possible Nano-Assisted Druggable Targets of COVID-19 Life Cycle
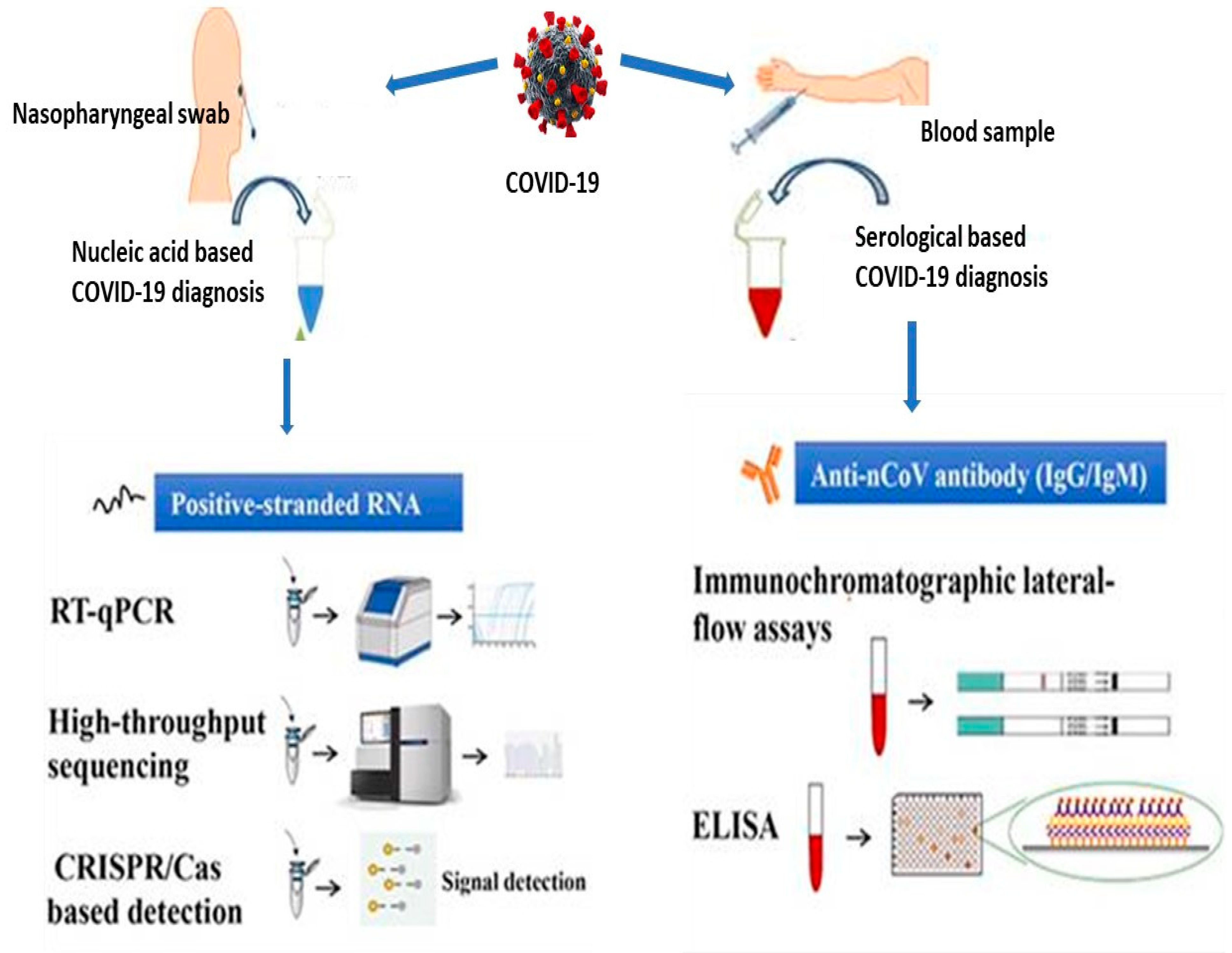
3. Nanotechnology for Efficient COVID-19 Diagnostics
3.1. Nucleic Acid-Based Sensing Mechanism
3.1.1. Antigen-Based Sensing
Reverse Transcription-Polymerase Chain Reaction (RT-PCR) Assays
Isothermal Amplification Techniques with Point-of-Care (POC) Potential
High-Throughput Sequencing Technology
Isothermal Temperature Nucleic Acid Amplification Technology and Recombinase Aided Amplification (RAA)
Nucleic Acid Mass Spectrometry
3.1.2. Antibody-Based Sensing
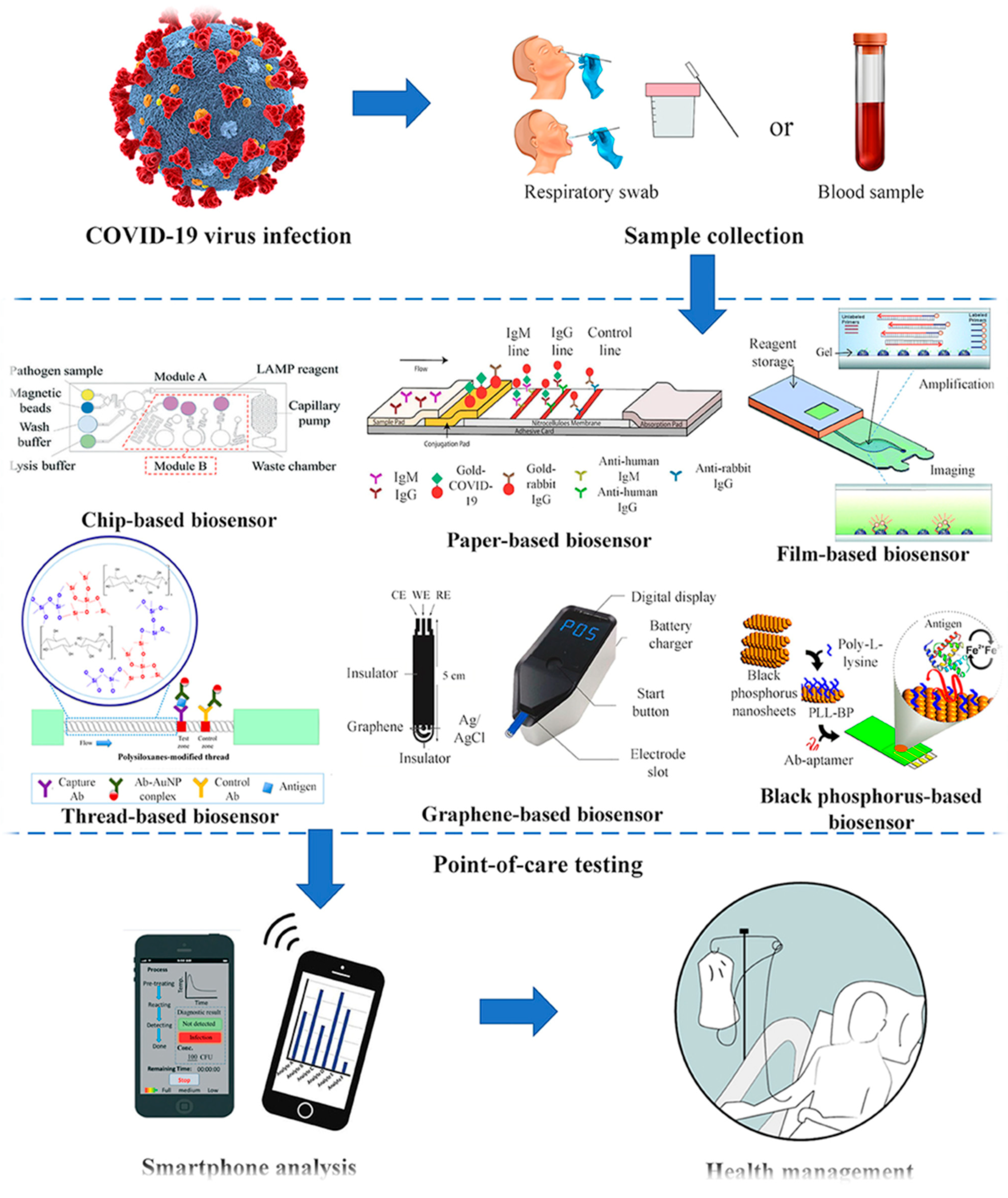
3.2. Biosensors and Signal Detection Methods for COVID-19 Diagnosis
3.2.1. Optical Biosensors
Lab-on-a-Chip Biosensors
Paper-Based Biosensors
Graphene-Based Biosensors
3.2.2. Electrochemical Nanobiosensors
Field-Effect Transistor (FET)
3.3. POC Diagnostic of COVID-19 Supported by AI and IoT
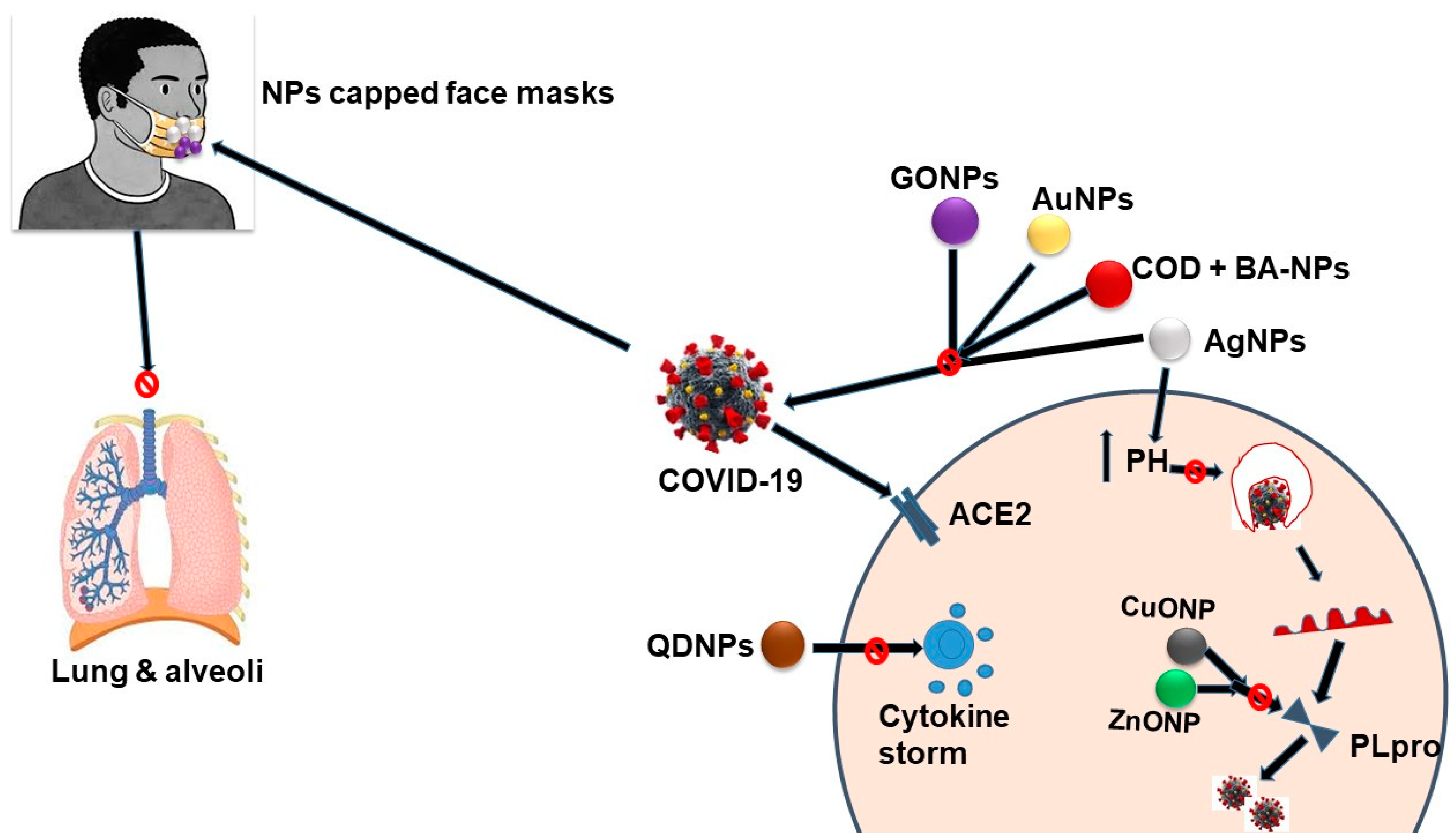
4. Inactivation of SARS-CoV-2 Transmission Using High-Performance Nanosystems
4.1. Blockage of Cellular Infiltration
4.2. Inhibition of Cellular Replication
4.3. Surface Sterilization/Personal Protection
4.4. Reusable Face Masks
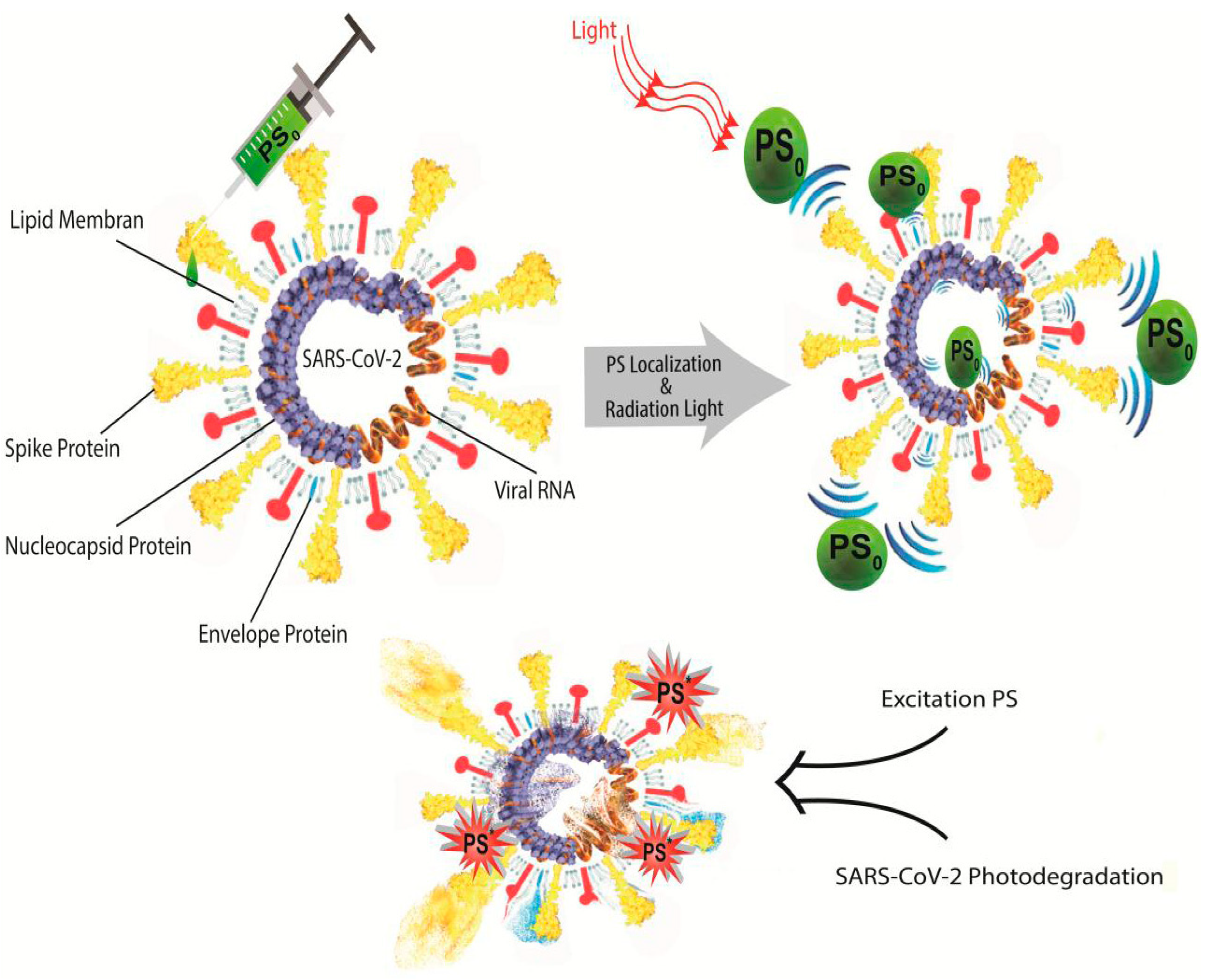
4.5. Photodynamic Approach to Eradicating SARS-CoV-2 Virus
5. Nanotechnology Based-Vaccines against COVID-19
6. Nano-Based Drug Delivery Systems for COVID-19 Management
7. Challenges and Perspectives
8. Conclusions and Viewpoint
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wu, A.; Peng, Y.; Huang, B.; Ding, X.; Wang, X.; Niu, P.; Meng, J.; Zhu, Z.; Zhang, Z.; Wang, J.; et al. Clinical features of patients infected with 2019 novel coronavirus in Wuhan, China. Cell Host Microbe 2020, 27, 325–328. [Google Scholar] [CrossRef]
- Channappanavar, R.; Perlman, S. Pathogenic Human Coronavirus Infections: Causes and Consequences of Cytokine Storm and Immunopathology. In Seminars in Immunopathology; Springer: Berlin/Heidelberg, Germany, 2017. [Google Scholar] [CrossRef]
- Petrosillo, N.; Viceconte, G.; Ergonul, O.; Ippolito, G.; Petersen, E. COVID-19, SARS and MERS: Are They Closely Related? Clin. Microbiol. Infect. 2020, 26, 729–734. [Google Scholar] [CrossRef] [PubMed]
- Chauhan, S. Comprehensive review of coronavirus disease 2019 (COVID-19). Biomed. J. 2020, 43, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Ge, Z.-Y.; Yang, L.-M.; Xia, J.-J.; Fu, X.-H.; Zhang, Y.-Z. Possible Aerosol Transmission of COVID-19 and Special Precautions in Dentistry. J. Zhejiang Univ. Sci. B 2020. [Google Scholar] [CrossRef] [PubMed]
- Hilgenfeld, R.; Peiris, M. From SARS to MERS: 10 Years of Research on Highly Pathogenic Human Coronaviruses. Antivir. Res. 2013, 100, 286–295. [Google Scholar] [CrossRef] [PubMed]
- Eckerle, I.; Müller, M.A.; Kallies, S.; Gotthardt, D.N.; Drosten, C. In-vitro renal epithelial cell infection reveals a viral kidney tropism as a potential mechanism for acute renal failure during Middle East Respiratory Syndrome (MERS) Coronavirus infection. Virol. 2013, 10, 1–5. [Google Scholar] [CrossRef]
- Zaki, A.M.; Van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Isolation of a Novel Coronavirus from a Man with Pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Cucinotta, D.; Vanelli, M. WHO Declares COVID-19 a Pandemic. Acta Biomed. 2020, 91, 157–160. [Google Scholar] [CrossRef]
- Weekly Epidemiological Update on COVID-19–7 September 2022. Available online: https://www.who.int/publications/m/item/weekly-epidemiological-update-on-covid-19---7-september-2022 (accessed on 3 October 2022).
- Weiner, D.L.; Balasubramaniam, V.; Shah, S.I.; Javier, J.R. COVID-19 Impact on Research, Lessons Learned from COVID-19 Research, Implications for Pediatric Research. Pediatric Res. 2020, 88, 148–150. [Google Scholar] [CrossRef]
- Liu, S.; Zheng, Q.; Wang, Z.; Elofsson, A. Potential covalent drugs targeting the main protease of the SARS-CoV-2 coronavirus. Bioinformatics 2020, 36, 3295–3298. [Google Scholar] [CrossRef]
- Li, L.; Fernández-Cruz, M.L.; Connolly, M.; Conde, E.; Fernández, M.; Schuster, M.; Navas, J.M. The potentiation effect makes the difference: Non-toxic concentrations of ZnO nanoparticles enhance Cu nanoparticle toxicity in vitro. Sci. Total Environ. 2015, 505, 253–260. [Google Scholar] [CrossRef] [PubMed]
- Moin, I.J. Tackling COVID-19 with Nanotechnology. Nanomed. Nanotechnol. 2020, 11, 10224. [Google Scholar] [CrossRef]
- Yalcin, H.C.; Kaushik, A. Support of intelligent emergent materials to combat COVID-19 pandemic. Emergent Mater. 2021, 4, 3–4. [Google Scholar] [CrossRef]
- Yang, D. Application of Nanotechnology in the COVID-19 Pandemic. Int. J. Nanomed. 2021, 16, 623–649. [Google Scholar] [CrossRef] [PubMed]
- Buschmann, M.D.; Carrasco, M.J.; Alishetty, S.; Paige, M.; Alameh, M.G.; Weissman, D. Nanomaterial Delivery Systems for mRNA Vaccines. Vaccines 2021, 9, 65. [Google Scholar] [CrossRef] [PubMed]
- Butkovich, N.; Li, E.; Ramirez, A.; Burkhardt, A.M.; Wang, S.W. Advancements in Protein Nanoparticle Vaccine Platforms to Combat Infectious Disease. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2020, 13, e1681. [Google Scholar] [CrossRef] [PubMed]
- Shehu, I.A.; Musa, M.K.; Alok, S.; Verma, A. Clinical features, treatment approaches and promising contributions of nanomedicines towards COVID-19. Int. J. Pharm. Sci. Res. 2021, 12, 1958–1965. [Google Scholar] [CrossRef]
- Kiplin Guy, R.; DiPaola, R.S.; Romanelli, F.; Dutch, R.E. Rapid repurposing of drugs for COVID-19. Science 2020, 368, 829–830. [Google Scholar] [CrossRef]
- Heinrich, M.A.; Martina, B.; Prakash, J. Nanomedicine Strategies to Target Coronavirus. Nano Today 2020, 35, 100961. [Google Scholar] [CrossRef]
- Gil, C.; Ginex, T.; Maestro, I.; Nozal, V.; Barrado-Gil, L.; Cuesta-Geijo, M.Á.; Urquiza, J.; Ramírez, D.; Alonso, C.; Campillo, N.E.; et al. COVID-19: Drug Targets and Potential Treatments. Med. Chem. 2020, 63, 12359–12386. [Google Scholar] [CrossRef]
- Iwata-Yoshikawa, N.; Okamura, T.; Shimizu, Y.; Hasegawa, H.; Takeda, M.; Nagata, N.J. TMPRSS2 Contributes to Virus Spread and Immunopathology in the Airways of Murine Models after Coronavirus Infection. J. Virol. 2019, 93, e01815–e01818. [Google Scholar] [CrossRef] [PubMed]
- Hasan, A.; Paray, B.A.; Hussain, A.; Qadir, F.A.; Attar, F.; Aziz, F.M.; Sharifi, M.; Derakhshankhah, H.; Rasti, B.; Mehrabi, M.; et al. A Review on the Cleavage Priming of the Spike Protein on Coronavirus by Angiotensin-Converting Enzyme-2 and Furin. J. Biomol. Struct. Dyn. 2021, 39, 3025–3033. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, A.O.; Severson, W.; Jonsson, C.; Singh, K.; Weiss, S.R.; Sarafianos, S.G.J. Novel Inhibitors of Severe Acute Respiratory Syndrome Coronavirus Entry That Act by Three Distinct Mechanisms. J. Virol. 2013, 87, 8017–8028. [Google Scholar] [CrossRef] [PubMed]
- Padmanabhan, P.; Desikan, R.; Dixit, N.M. Targeting TMPRSS2 and Cathepsin B/L together may be synergistic against SARS-CoV-2 infection. PLoS Comput. Biol. 2020, 16, e1008461. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, Y.; Kolokoltsov, A.A.; Chen, C.C.; Tidwell, M.W.; Bauta, W.E.; Klugbauer, N.; Grimm, C.; Wahl-Schott, C.; Biel, M.; Davey, R.A. Two-pore channels control Ebola virus host cell entry and are drug targets for disease treatment. Science 2015, 347, 995–998. [Google Scholar] [CrossRef]
- Penny, C.J.; Vassileva, K.; Jha, A.; Yuan, Y.; Chee, X.; Yates, E.; Mazzon, M.; Kilpatrick, B.S.; Muallem, S.; Marsh, M.; et al. Mining of Ebola virus entry inhibitors identifies approved drugs as two-pore channel pore blockers. Biochim. Biophys. Acta-Mol. Cell Res. 2019, 1866, 1151–1161. [Google Scholar] [CrossRef]
- Wang, C.; Wang, J.; Shuai, L.; Ma, X.; Zhang, H.; Liu, R.; Chen, W.; Wang, X.; Ge, J.; Wen, Z.; et al. The Serine/Threonine Kinase AP2-Associated Kinase 1 Plays an Important Role in Rabies Virus Entry. Viruses 2019, 12, 45. [Google Scholar] [CrossRef]
- Stebbing, J.; Phelan, A.; Griffin, I.; Tucker, C.; Oechsle, O.; Smith, D.; Richardson, P. COVID-19: Combining Antiviral and Anti-Inflammatory Treatments. Lancet Infect. Dis. 2020, 20, 400–402. [Google Scholar] [CrossRef]
- Favalli, E.G.; Biggioggero, M.; Maioli, G.; Caporali, R. Baricitinib for COVID-19: A Suitable Treatment? Lancet Infect. Dis. 2020, 20, 1012–1013. [Google Scholar] [CrossRef]
- Dong, S.; Sun, J.; Mao, Z.; Wang, L.; Lu, Y.L.; Li, J.J. A guideline for homology modeling of the proteins from newly discovered betacoronavirus, 2019 novel coronavirus (2019-nCoV). Med. Virol. 2020, 92, 1542–1548. [Google Scholar] [CrossRef]
- Kandeel, M.; Ibrahim, A.; Fayez, M.; Al-Nazawi, M.J. From SARS and MERS CoVs to SARS-CoV-2: Moving toward more biased codon usage in viral structural and nonstructural genes. Med. Virol. 2020, 92, 660–666. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Lin, D.; Sun, X.; Curth, U.; Drosten, C.; Sauerhering, L.; Becker, S.; Rox, K.; Hilgenfeld, R. Crystal structure of SARS-CoV-2 main protease provides a basis for design of improved α-ketoamide inhibitors. Science 2020, 368, 409–412. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Vedantham, P.; Lu, K.; Agudelo, J.; Carrion, R.; Nunneley, J.W.; Barnard, D.; Pöhlmann, S.; McKerrow, J.H.; Renslo, A.R.; et al. Protease inhibitors targeting coronavirus and filovirus entry. Antivir. Res. 2015, 116, 76–84. [Google Scholar] [CrossRef]
- Chan, J.F.W.; Kok, K.H.; Zhu, Z.; Chu, H.; To, K.K.W.; Yuan, S.; Yuen, K.Y. Genomic characterization of the 2019 novel human-pathogenic coronavirus isolated from a patient with atypical pneumonia after visiting Wuhan. Emerg. Microbes Infect. 2020, 9, 221–236. [Google Scholar] [CrossRef]
- Mcdonald, S.M. RNA synthetic mechanisms employed by diverse families of RNA viruses. Wiley Interdiscip. Rev. RNA 2013, 4, 351–367. [Google Scholar] [CrossRef] [PubMed]
- Kubik, T.; Bogunia-Kubik, K.; Sugisaka, M. Nanotechnology on Duty in Medical Applications. Curr. Pharm. Biotechnol. 2016, 16, 17–33. [Google Scholar] [CrossRef]
- Suleman, S.; Shukla, S.K.; Malhotra, N.; Bukkitgar, S.D.; Shetti, N.P.; Pilloton, R.; Narang, J.; Tan, Y.N.; Aminabhavi, T.M. Point of care detection of COVID-19: Advancement in biosensing and diagnostic methods. Chem. Eng. J. 2021, 414, 128759. [Google Scholar] [CrossRef]
- Bisht, A.; Mishra, A.; Bisht, H.; Tripathi, R.M. Nanomaterial based biosensors for detection of viruses including sars-cov-2: A review. J. Anal. Test. 2021, 5, 327–340. [Google Scholar] [CrossRef]
- Wanunu, M.; Dadosh, T.; Ray, V.; Jin, J.; McReynolds, L.; Drndić, M. Rapid electronic detection of probe-specific microRNAs using thin nanopore sensors. Nat. Nanotechnol. 2010, 5, 807–814. [Google Scholar] [CrossRef]
- Wang, X.; Liu, L.H.; Ramström, O.; Yan, M. Engineering Nanomaterial Surfaces for Biomedical Applications. Exp. Biol. Med. 2009, 234, 1128–1139. [Google Scholar] [CrossRef]
- Shi, L.; Sun, Q.; He, J.; Xu, H.; Liu, C.; Zhao, C.; Xu, Y.; Wu, C.; Xiang, J.; Gu, D.; et al. Development of SPR biosensor for simultaneous detection of multiplex respiratory viruses. Bio-Med. Mater. Eng. 2015, 26, S2207–S2216. [Google Scholar] [CrossRef] [PubMed]
- Cho, S.J.; Woo, H.M.; Kim, K.S.; Oh, J.W.; Jeong, Y.J.J. Novel system for detecting SARS coronavirus nucleocapsid protein using an ssDNA aptamer. Biosci. Bioeng. 2011, 112, 535–540. [Google Scholar] [CrossRef] [PubMed]
- Park, T.J.; Lee, S.K.; Yoo, S.M.; Yang, S.M.; Lee, S.Y. Development of Reflective Biosensor Using Fabrication of Functionalized Photonic Nanocrystals. J. Nanosci. Nanotechnol. 2011, 11, 632–637. [Google Scholar] [CrossRef]
- Ishikawa, F.N.; Curreli, M.; Olson, C.A.; Liao, H.I.; Sun, R.; Roberts, R.W.; Cote, R.J.; Thompson, M.E.; Zhou, C. Importance of Controlling Nanotube Density for Highly Sensitive and Reliable Biosensors Functional in Physiological Conditions. ACS Nano 2010, 11, 6914–6922. [Google Scholar] [CrossRef] [PubMed]
- Zhao, V.X.T.; Wong, T.I.; Zheng, X.T.; Tan, Y.N.; Zhou, X. Colorimetric biosensors for point-of-care virus detections. Mater. Sci. Energy Technol. 2020, 3, 237–249. [Google Scholar] [CrossRef]
- Aldewachi, H.; Chalati, T.; Woodroofe, M.N.; Bricklebank, N.; Sharrack, B.; Gardiner, P. Gold Nanoparticle-Based Colorimetric Biosensors. Nanoscale 2018, 10, 18–33. [Google Scholar] [CrossRef]
- Layqah, L.A.; Eissa, S. An electrochemical immunosensor for the corona virus associated with the Middle East respiratory syndrome using an array of gold nanoparticle-modified carbon electrodes. Microchim. Acta 2019, 186, 224. [Google Scholar] [CrossRef]
- Ahmed, M.K.; Afifi, M.; Uskoković, V.J. Protecting healthcare workers during COVID-19 pandemic with nanotechnology: A protocol for a new device from Egypt. Infect. Public Health 2020, 13, 1243–1246. [Google Scholar] [CrossRef]
- Weiss, C.; Carriere, M.; Fusco, L.; Fusco, L.; Capua, I.; Regla-Nava, J.A.; Pasquali, M.; Pasquali, M.; Pasquali, M.; Scott, J.A.; et al. Toward Nanotechnology-Enabled Approaches against the COVID-19 Pandemic. ACS Nano 2020, 14, 6383–6406. [Google Scholar] [CrossRef]
- Lin, Y.; Zhang, H.; Song, W.; Si, S.; Han, Y.; Jiang, J. Identification and characterization of Zika virus NS5 RNA-dependent RNA polymerase inhibitors. Int. J. Antimicrob. Agents 2019, 54, 502–506. [Google Scholar] [CrossRef]
- Hecel, A.; Ostrowska, M.; Stokowa-Sołtys, K.; Wątły, J.; Dudek, D.; Miller, A.; Potocki, S.; Matera-Witkiewicz, A.; Dominguez-Martin, A.; Kozłowski, H.; et al. Zinc(II)—The Overlooked Éminence Grise of Chloroquine’s Fight against COVID-19? Pharmaceuticals 2020, 13, 228. [Google Scholar] [CrossRef] [PubMed]
- Pokhrel, P.; Hu, C.; Mao, H. Detecting the Coronavirus (COVID-19). ACS Sens. 2020, 5, 2283–2296. [Google Scholar] [CrossRef] [PubMed]
- Kneller, D.W.; Phillips, G.; O’Neill, H.M.; Jedrzejczak, R.; Stols, L.; Langan, P.; Joachimiak, A.; Coates, L.; Kovalevsky, A. Structural plasticity of SARS-CoV-2 3CL Mpro active site cavity revealed by room temperature X-ray crystallography. Nat. Commun. 2020, 11, 3202. [Google Scholar] [CrossRef]
- Chittapragada, M.; Roberts, S.; Ham, Y.W. Aminoglycosides: Molecular Insights on the Recognition of RNA and Aminoglycoside Mimics. Perspect. Med. Chem. 2009, 3, 21–37. [Google Scholar] [CrossRef] [PubMed]
- Lin, K.; Gallay, P. Curing a Viral Infection by Targeting the Host: The Example of Cyclophilin Inhibitors. Antivir. Res. 2013, 99, 68–77. [Google Scholar] [CrossRef]
- Luo, Z.; Ang, M.J.Y.; Chan, S.Y.; Yi, Z.; Goh, Y.Y.; Yan, S.; Tao, J.; Liu, K.; Li, X.; Zhang, H.; et al. Combating the Coronavirus Pandemic: Early Detection, Medical Treatment, and a Concerted Effort by the Global Community. Research 2020, 2020. [Google Scholar] [CrossRef]
- Keni, R.; Alexander, A.; Nayak, P.G.; Mudgal, J.; Nandakumar, K. COVID-19: Emergence, spread, possible treatments, and global burden. Front. Public Health 2020, 8, 216. [Google Scholar] [CrossRef]
- Viljoen, G.J.; Romito, M.; Kara, P.D. Current and Future Developments in Nucleic Acid-Based Diagnostics. In Applications of Gene-Based Technologies for Improving Animal Production and Health in Developing Countries, Proceedings of the FAO/IAEA International Symposium on Applications of Gene-Based Technologies for Improving Animal Production and Health in Developing Countries, Vienna, Austria, 6–10 October 2003; Makkar, H.P.S., Viljoen, G.J., Eds.; Springer: Dordrecht, The Netherlands, 2005. [Google Scholar] [CrossRef]
- Feng, W.; Newbigging, A.M.; Le, C.; Pang, B.; Peng, H.; Cao, Y.; Wu, J.; Abbas, G.; Song, J.; Wang, D.B.; et al. Molecular Diagnosis of COVID-19: Challenges and Research Needs. Anal. Chem. 2020, 92, 10196–10209. [Google Scholar] [CrossRef]
- Lv, H.; Wu, N.C.; Tsang, O.T.Y.; Yuan, M.; Perera, R.A.P.M.; Leung, W.S.; So, R.T.Y.; Chan, J.M.C.; Yip, G.K.; Chik, T.S.H.; et al. Cross-reactive antibody response between SARS-CoV-2 and SARS-CoV infections. bioRxiv 2020, 31, 107725. [Google Scholar] [CrossRef]
- Ju, B.; Zhang, Q.; Ge, X.; Wang, R.; Yu, J.; Shan, S.; Zhou, B.; Song, S.; Tang, X.; Yu, J.; et al. Potent human neutralizing antibodies elicited by SARS-CoV-2 infection. bioRxiv 2020. [Google Scholar] [CrossRef]
- Islam, K.U.; Iqbal, J. An Update on Molecular Diagnostics for COVID-19. Front. Cell. Infect. Microbiol. 2020, 10, 560616. [Google Scholar] [CrossRef] [PubMed]
- Vahedifard, F.; Chakravarthy, K. Nanomedicine for COVID-19: The role of nanotechnology in the treatment and diagnosis of COVID-19. Emergent Mater. 2021, 4, 75–99. [Google Scholar] [CrossRef] [PubMed]
- Samson, R.; Navale, G.R.; Dharne, M.S. Biosensors: Frontiers in rapid detection of COVID-19. 3 Biotech 2020, 10, 1–9. [Google Scholar] [CrossRef] [PubMed]
- FDA Authorizes First Point-of-Care Antibody COVID-19 Test—COVID-19—Labmedica.com. Available online: https://www.labmedica.com/covid-19/articles/294784685/fda-authorizes-first-point-of-care-antibody-covid-19-test.html (accessed on 28 September 2020).
- Graphene Updates—The Graphene Council. Available online: https://www.thegraphenecouncil.org/blogpost/1501180/Graphene-Updates?tag=&DGPCrSrt=&DGPCrPg=12 (accessed on 28 September 2020).
- Zheng, G.; Xie, W.-Q.; Albarakati, S.; Algarni, M.; Tan, C.; Wang, Y.; Peng, J.; Partridge, J.; Farrar, L.; Yi, J.; et al. Gate-Tuned Interlayer Coupling in van der Waals Ferromagnet Fe3GeTe2 Nanoflakes. Phys. Rev. Lett. 2020, 125, 047202. [Google Scholar] [CrossRef] [PubMed]
- Jiang, C.; Li, X.; Ge, C.; Ding, Y.; Zhang, T.; Cao, S.; Meng, L.; Lu, S.J. Molecular detection of SARS-CoV-2 being challenged by virus variation and asymptomatic infection. Pharm. Anal. 2021, 11, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Li, N.; Wang, P.; Wang, X.; Geng, C.; Chen, J.; Gong, Y. Molecular diagnosis of COVID-19: Current situation and trend in China. Exp. Ther. Med. 2020, 20, 1. [Google Scholar] [CrossRef]
- Van Dijk, E.L.; Jaszczyszyn, Y.; Naquin, D.; Thermes, C. The Third Revolution in Sequencing Technology. Trends Genet. 2018, 34, 666–681. [Google Scholar] [CrossRef]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA Detection Using Recombination Proteins. PLoS Biol. 2006, 4, 1115–1121. [Google Scholar] [CrossRef]
- Summary of NMPA Approved Novel Coronavirus 2019-nCoV Test Kits. Available online: http://english.nmpa.gov.cn/2020-04/03/c_468570.htm (accessed on 7 June 2021).
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef]
- Broughton, J.P.; Deng, X.; Yu, G.; Fasching, C.L.; Servellita, V.; Singh, J.; Miao, X.; Streithorst, J.A.; Granados, A.; Sotomayor-Gonzalez, A.; et al. CRISPR–Cas12-based detection of SARS-CoV-2. Nat. Biotechnol. 2020, 38, 870–874. [Google Scholar] [CrossRef]
- Rashid, Z.Z.; Othman, S.N.; Samat, M.N.A.; Ali, U.K.; Wong, K.K. Diagnostic Performance of COVID-19 Serology Assays. Malays. J. Pathol. 2020, 42, 13–21. [Google Scholar]
- Tan, W.; Lu, Y.; Zhang, J.; Wang, J.; Dan, Y.; Tan, Z.; He, X.; Qian, C.; Sun, Q.; Hu, Q.; et al. Viral Kinetics and Antibody Responses in Patients with COVID-19. medRxiv 2020. [Google Scholar] [CrossRef]
- Saylan, Y.; Erdem, Ö.; Ünal, S.; Denizli, A. An Alternative Medical Diagnosis Method: Biosensors for Virus Detection. Biosensors 2019, 9, 65. [Google Scholar] [CrossRef] [PubMed]
- Ngom, B.; Guo, Y.; Wang, X.; Bi, D. Development and Application of Lateral Flow Test Strip Technology for Detection of Infectious Agents and Chemical Contaminants: A Review. Anal. Bioanal. Chem. 2010, 397, 1113–1135. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Wang, Y.; Wang, J.; Tang, Z.; Pounds, J.G.; Lin, Y. Rapid and Sensitive Detection of Protein Biomarker Using a Portable Fluorescence Biosensor Based on Quantum Dots and a Lateral Flow Test Strip. Anal. Chem. 2010, 82, 7008–7014. [Google Scholar] [CrossRef]
- Choi, J.R.; Yong, K.W.; Choi, J.Y.; Nilghaz, A.; Lin, Y.; Xu, J.; Lu, X. Black Phosphorus and Its Biomedical Applications. Theranostics 2018, 8, 1005–1026. [Google Scholar] [CrossRef]
- Ling, X.; Wang, H.; Huang, S.; Xia, F.; Dresselhaus, M.S. The Renaissance of Black Phosphorus. Proc. Natl. Acad. Sci. USA 2015, 112, 4523–4530. [Google Scholar] [CrossRef]
- Damborský, P.; Švitel, J.; Katrlík, J. Optical biosensors. Essays Biochem. 2016, 60, 91–100. [Google Scholar] [CrossRef]
- Huang, J.C.; Chang, Y.F.; Chen, K.H.; Su, L.C.; Lee, C.W.; Chen, C.C.; Chen, Y.M.A.; Chou, C. Detection of severe acute respiratory syndrome (SARS) coronavirus nucleocapsid protein in human serum using a localized surface plasmon coupled fluorescence fiber-optic biosensor. Biosens. Bioelectron. 2009, 25, 320–325. [Google Scholar] [CrossRef]
- Aoki, V.; Fukumori, L.M.I.; Freitas, E.L.; Sousa, J.X.; Périgo, A.M.; Oliveira, Z.N.P. Direct and Indirect Immunofluorescence. An. Bras. Dermatol. 2010, 85, 490–500. [Google Scholar] [CrossRef]
- Patolsky, F.; Zheng, G.; Lieber, C.M. Nanowire Sensors for Medicine and the Life Sciences. Nanomedicine 2006, 1, 51–65. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Wang, H.; Cao, Z.; Jin, H.; Chi, H.; Zhao, J.; Yu, B.; Yan, F.; Hu, X.; Wu, F.; et al. Rapid and Specific Assay for the Detection of MERS-CoV. Front. Microbiol. 2018, 9, 1101. [Google Scholar] [CrossRef] [PubMed]
- Pohanka, M.; Jun, D.; Kuca, K. Amperometric Biosensors for Real Time Assays of Organophosphates. Sensors 2008, 8, 5303–5312. [Google Scholar] [CrossRef] [PubMed]
- Skládal, P.; Dos Santos Riccardi, C.; Yamanaka, H.; Da Costa, P.I.J. Piezoelectric biosensors for real-time monitoring of hybridization and detection of hepatitis C virus. J. Virol. Methods 2004, 117, 145–151. [Google Scholar] [CrossRef] [PubMed]
- Dzyadevych, S.; Jaffrezic-Renault, N. Conductometric Biosensors. In Biological Identification: DNA Amplification and Sequencing, Optical Sensing, Lab-On-Chip and Portable Systems; Elsevier Ltd.: Amsterdam, The Netherlands, 2014; pp. 153–193. [Google Scholar] [CrossRef]
- Qiu, G.; Gai, Z.; Tao, Y.; Schmitt, J.; Kullak-Ublick, G.A.; Wang, J. Dual-Functional Plasmonic Photothermal Biosensors for Highly Accurate Severe Acute Respiratory Syndrome Coronavirus 2 Detection. ACS Nano 2020, 14, 5268–5277. [Google Scholar] [CrossRef]
- Chen, Z.; Zhang, Z.; Zhai, X.; Li, Y.; Lin, L.; Zhao, H.; Bian, L.; Li, P.; Yu, L.; Wu, Y.; et al. Rapid and Sensitive Detection of anti-SARS-CoV-2 IgG, Using Lanthanide-Doped Nanoparticles-Based Lateral Flow Immunoassay. Anal. Chem. 2020, 92, 7226–7231. [Google Scholar] [CrossRef]
- Wang, X.; Zhong, M.; Liu, Y.; Ma, P.; Dang, L.; Meng, Q.; Wan, W.; Ma, X.; Liu, J.; Yang, G.; et al. Rapid and sensitive detection of COVID-19 using CRISPR/Cas12a-based detection with naked eye readout, CRISPR/Cas12a-NER. Sci. Bull. 2020, 65, 1436–1439. [Google Scholar] [CrossRef]
- Seo, G.; Lee, G.; Kim, M.J.; Baek, S.H.; Choi, M.; Ku, K.B.; Lee, C.S.; Jun, S.; Park, D.; Kim, H.G.; et al. Rapid Detection of COVID-19 Causative Virus (SARS-CoV-2) in Human Nasopharyngeal Swab Specimens Using Field-Effect Transistor-Based Biosensor. ACS Nano 2020, 14, 5135–5142. [Google Scholar] [CrossRef]
- Yan, C.; Cui, J.; Huang, L.; Du, B.; Chen, L.; Xue, G.; Li, S.; Zhang, W.; Zhao, L.; Sun, Y.; et al. Rapid and visual detection of 2019 novel coronavirus (SARS-CoV-2) by a reverse transcription loop-mediated isothermal amplification assay. Clin. Microbiol. Infect. 2020, 26, 773–779. [Google Scholar] [CrossRef]
- Yu, F.; Yan, L.; Wang, N.; Yang, S.; Wang, L.; Tang, Y.; Gao, G.; Wang, S.; Ma, C.; Xie, R.; et al. Quantitative Detection and Viral Load Analysis of SARS-CoV-2 in Infected Patients. Clin. Infect. Dis. 2020, 71, 793–798. [Google Scholar] [CrossRef]
- Zhu, X.; Wang, X.; Han, L.; Chen, T.; Wang, L.; Li, H.; Li, S.; He, L.; Fu, X.; Chen, S.; et al. Multiplex reverse transcription loop-mediated isothermal amplification combined with nanoparticle-based lateral flow biosensor for the diagnosis of COVID-19. Biosens. Bioelectron. 2020, 166, 112437. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.L.; Zhou, R.; Zhao, X.S.J. Graphene-based materials as supercapacitor electrodes. J. Mater. Chem. 2010, 20, 5983–5992. [Google Scholar] [CrossRef]
- Ke, Q.; Wang, J. Graphene-Based Materials for Supercapacitor Electrodes—A Review. J. Mater. 2016, 2, 37–54. [Google Scholar] [CrossRef]
- Arduini, F.; Micheli, L.; Moscone, D.; Palleschi, G.; Piermarini, S.; Ricci, F.; Volpe, G. Electrochemical Biosensors Based on Nanomodified Screen-Printed Electrodes: Recent Applications in Clinical Analysis. TrAC—Trends Anal. Chem. 2016, 79, 114–126. [Google Scholar] [CrossRef]
- Ma, Y.D.; Li, K.H.; Chen, Y.H.; Lee, Y.M.; Chou, S.T.; Lai, Y.Y.; Huang, P.C.; Ma, H.P.; Lee, G. Bin. A sample-to-answer, portable platform for rapid detection of pathogens with a smartphone interface. Lab Chip 2019, 19, 3804–3814. [Google Scholar] [CrossRef]
- Li, Z.; Yi, Y.; Luo, X.; Xiong, N.; Liu, Y.; Li, S.; Sun, R.; Wang, Y.; Hu, B.; Chen, W.; et al. Development and clinical application of a rapid IgM-IgG combined antibody test for SARS-CoV-2 infection diagnosis. J. Med. Virol. 2020, 92, 1518–1524. [Google Scholar] [CrossRef]
- Linger, Y.; Knickerbocker, C.; Sipes, D.; Golova, J.; Franke, M.; Calderon, R.; Lecca, L.; Thakore, N.; Holmberg, R.; Qu, P.; et al. Genotyping Multidrug-Resistant Mycobacterium tuberculosis from Primary Sputum and Decontaminated Sediment with an Integrated Microfluidic Amplification Microarray Test. J. Clin. Microbiol. 2018, 56, e01652-17. [Google Scholar] [CrossRef]
- Nguyen, N.V.; Jen, C.P. Selective Detection of Human Lung Adenocarcinoma Cells Based on the Aptamer-Conjugated Self-Assembled Monolayer of Gold Nanoparticles. Micromachines 2019, 10, 195. [Google Scholar] [CrossRef]
- Kampeera, J.; Pasakon, P.; Karuwan, C.; Arunrut, N.; Sappat, A.; Sirithammajak, S.; Dechokiattawan, N.; Sumranwanich, T.; Chaivisuthangkura, P.; Ounjai, P.; et al. Point-of-care rapid detection of Vibrio parahaemolyticus in seafood using loop-mediated isothermal amplification and graphene-based screen-printed electrochemical sensor. Biosens. Bioelectron. 2019, 132, 271–278. [Google Scholar] [CrossRef]
- Choi, J.R. Development of Point-of-Care Biosensors for COVID-19. Front. Chem. 2020, 8, 517. [Google Scholar] [CrossRef]
- Sin, M.L.; Mach, K.E.; Wong, P.K.; Liao, J.C. Advances and Challenges in Biosensor-Based Diagnosis of Infectious Diseases. Expert Rev. Mol. Diagn. 2014, 14, 225–244. [Google Scholar] [CrossRef] [PubMed]
- Kobayashi, M.; Kikuchi, N.; Sato, A. Optical tomography of fluorophores in dense scattering media based on ultrasound-enhanced chemiluminescence. Appl. Phys. Lett. 2015, 106, 021103. [Google Scholar] [CrossRef]
- Tansi, F.L.; Rüger, R.; Rabenhold, M.; Steiniger, F.; Fahr, A.; Hilger, I.J. Fluorescence-quenching of a Liposomal-encapsulated Near-infrared Fluorophore as a Tool for In Vivo Optical Imaging. J. Vis. Exp. 2015, 5, e52136. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Guo, Q.; Cui, D. Recent Advances in Nanotechnology Applied to Biosensors. Sensors 2009, 9, 1033–1053. [Google Scholar] [CrossRef]
- Sharma, A.; Mishra, R.K.; Goud, K.Y.; Mohamed, M.A.; Kummari, S.; Tiwari, S.; Li, Z.; Narayan, R.; Stanciu, L.A.; Marty, J.L. Optical biosensors for diagnostics of infectious viral disease: A recent update. Diagnostics 2021, 11, 2083. [Google Scholar] [CrossRef]
- Alafeef, M.; Moitra, P.; Dighe, K.; Pan, D. RNA-extraction-free nano-amplified colorimetric test for point-of-care clinical diagnosis of COVID-19. Nat. Protoc. 2021, 16, 3141–3162. [Google Scholar] [CrossRef]
- Feng, T.; Nie, C.; Peng, P.; Liu, H.; Wang, T.; Li, P.; Huang, W.; Wang, J. Nanoagent-based theranostic strategies against human coronaviruses. Nano Res. 2022, 15, 3323–3337. [Google Scholar] [CrossRef]
- Yin, J.; Zou, Z.; Hu, Z.; Zhang, S.; Zhang, F.; Wang, B.; Lv, S.; Mu, Y. A “sample-in-multiplex-digital-answer-out” chip for fast detection of pathogens. Lab Chip 2020, 20, 979–986. [Google Scholar] [CrossRef]
- Zanoli, L.M.; Spoto, G. Isothermal amplification methods for the detection of nucleic acids in microfluidic devices. Biosensors 2012, 3, 18–43. [Google Scholar] [CrossRef]
- Jiao, J.; Duan, C.; Xue, L.; Liu, Y.; Sun, W.; Xiang, Y. DNA nanoscaffold-based SARS-CoV-2 detection for COVID-19 diagnosis. Biosens. Bioelectron. 2020, 167, 112479. [Google Scholar] [CrossRef]
- Cennamo, N.; D’Agostino, G.; Perri, C.; Arcadio, F.; Chiaretti, G.; Parisio, E.M.; Camarlinghi, G.; Vettori, C.; Di Marzo, F.; Cennamo, R.; et al. Proof of Concept for a Quick and Highly Sensitive On-Site Detection of SARS-CoV-2 by Plasmonic Optical Fibers and Molecularly Imprinted Polymers. Sensors 2021, 21, 1681. [Google Scholar] [CrossRef] [PubMed]
- Dincer, C.; Bruch, R.; Kling, A.; Dittrich, P.S.; Urban, G.A. Multiplexed Point-of-Care Testing—XPOCT. Trends Biotechnol. 2017, 35, 728–742. [Google Scholar] [CrossRef]
- Patel, S.; Nanda, R.; Sahoo, S.; Mohapatra, E. Biosensors in Health Care: The Milestones Achieved in Their Development towards Lab-on-Chip-Analysis. Biochem. Res. Int. 2016, 3130469. [Google Scholar] [CrossRef] [PubMed]
- Kukhtin, A.C.; Sebastian, T.; Golova, J.; Perov, A.; Knickerbocker, C.; Linger, Y.; Bueno, A.; Qu, P.; Villanueva, M.; Holmberg, R.C.; et al. Lab-on-a-Film disposable for genotyping multidrug-resistant Mycobacterium tuberculosis from sputum extracts. Lab Chip 2019, 19, 1217–1225. [Google Scholar] [CrossRef] [PubMed]
- Kukhtin, A.V.; Norville, R.; Bueno, A.; Qu, P.; Parrish, N.; Murray, M.; Chandler, D.P.; Holmberg, R.C.; Cooney, C.G. A Benchtop Automated Sputum-to-Genotype System Using a Lab-on-a-Film Assembly for Detection of Multidrug-Resistant Mycobacterium tuberculosis. Anal. Chem. 2020, 92, 5311–5318. [Google Scholar] [CrossRef]
- Darwish, N.T.; Sekaran, S.D.; Khor, S.M. Point-of-Care Tests: A Review of Advances in the Emerging Diagnostic Tools for Dengue Virus Infection. Sens. Actuators B Chem. 2018, 255, 3316–3331. [Google Scholar] [CrossRef]
- Böhm, A.; Trosien, S.; Avrutina, O.; Kolmar, H.; Biesalski, M. Covalent Attachment of Enzymes to Paper Fibers for Paper-Based Analytical Devices. Front. Chem. 2018, 6, 214. [Google Scholar] [CrossRef]
- Choi, J.R.; Yong, K.W.; Choi, J.Y.; Cowie, A.C. Emerging Point-of-Care Technologies for Food Safety Analysis. Sensors 2019, 19, 817. [Google Scholar] [CrossRef]
- Ni, L.; Ye, F.; Cheng, M.L.; Feng, Y.; Deng, Y.Q.; Zhao, H.; Wei, P.; Ge, J.; Gou, M.; Li, X.; et al. Detection of SARS-CoV-2-Specific Humoral and Cellular Immunity in COVID-19 Convalescent Individuals. Immunity 2020, 52, 971–977.e3. [Google Scholar] [CrossRef]
- Reboud, J.; Xu, G.; Garrett, A.; Adriko, M.; Yang, Z.; Tukahebwa, E.M.; Rowell, C.; Cooper, J.M. Paper-based microfluidics for DNA diagnostics of malaria in low resource underserved rural communities. Proc. Natl. Acad. Sci. USA 2019, 116, 4834–4842. [Google Scholar] [CrossRef]
- Park, B.H.; Oh, S.J.; Jung, J.H.; Choi, G.; Seo, J.H.; Kim, D.H.; Lee, E.Y.; Seo, T.S. An integrated rotary microfluidic system with DNA extraction, loop-mediated isothermal amplification, and lateral flow strip based detection for point-of-care pathogen diagnostics. Biosens. Bioelectron. 2016, 91, 334–340. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Choi, J.R.; Wang, S.; Gong, Y.; Feng, S.; Pingguan-Murphy, B.; Lu, T.J.; Xu, F. Multiple test zones for improved detection performance in lateral flow assays. Sens. Actuators B Chem. 2016, 243, 484–488. [Google Scholar] [CrossRef]
- Santiago, I. Trends and Innovations in Biosensors for COVID-19 Mass Testing. ChemBioChem. 2020, 21, 2880–2889. [Google Scholar] [CrossRef] [PubMed]
- Singhal, C.; Dubey, A.; Mathur, A.; Pundir, C.S.; Narang, J. Paper based DNA biosensor for detection of chikungunya virus using gold shells coated magnetic nanocubes. Process Biochem. 2018, 74, 35–42. [Google Scholar] [CrossRef]
- Singhal, C.; Khanuja, M.; Chaudhary, N.; Pundir, C.S.; Narang, J. Detection of chikungunya virus DNA using two-dimensional MoS2 nanosheets based disposable biosensor. Sci. Rep. 2018, 8, 7734. [Google Scholar] [CrossRef]
- Rao, C.N.R.; Sood, A.K.; Subrahmanyam, K.S.; Govindaraj, A. Graphen, das neue zweidimensionale Nanomaterial. Angew. Chem. 2009, 121, 7890–7916. [Google Scholar] [CrossRef]
- Ge, X.; Xia, Z.; Guo, S. Recent Advances on Black Phosphorus for Biomedicine and Biosensing. Adv. Funct. Mater. 2019, 29, 1900318. [Google Scholar] [CrossRef]
- Luo, M.; Fan, T.; Zhou, Y.; Zhang, H.; Mei, L. 2D Black Phosphorus–Based Biomedical Applications. Adv. Funct. Mater. 2019, 29, 1808306. [Google Scholar] [CrossRef]
- Kumar, V.; Brent, J.R.; Shorie, M.; Kaur, H.; Chadha, G.; Thomas, A.G.; Lewis, E.A.; Rooney, A.P.; Nguyen, L.; Zhong, X.L.; et al. Nanostructured Aptamer-Functionalized Black Phosphorus Sensing Platform for Label-Free Detection of Myoglobin, a Cardiovascular Disease Biomarker. ACS Appl. Mater. Interfaces 2016, 8, 22860–22868. [Google Scholar] [CrossRef]
- Le, L.T.; Ervin, M.H.; Qiu, H.; Fuchs, B.E.; Lee, W.Y. Graphene supercapacitor electrodes fabricated by inkjet printing and thermal reduction of graphene oxide. Electrochem. Commun. 2011, 13, 355–358. [Google Scholar] [CrossRef]
- Lin, L.P.; Tan, M.T.T.; Loh, H.S. Electrochemical DNA Sensor Based on Graphene/Zirconia Nanocomposite for Label-Free Detection of Exon-19 Mutations in Lung Cancer: A Preliminary Study. In Proceedings of the 2019 9th International Conference on Biomedical Engineering and Technology, Tokyo, Japan, 28–30 March 2019; pp. 223–228. [Google Scholar] [CrossRef]
- Ji, T.; Liu, Z.; Wang, G.Q.; Guo, X.; Akbar Khan, S.; Lai, C.; Chen, H.; Huang, S.; Xia, S.; Chen, B.; et al. Detection of COVID-19: A review of the current literature and future perspectives. Biosens. Bioelectron. 2020, 166, 112455. [Google Scholar] [CrossRef]
- Li, B.; Tan, H.; Anastasova, S.; Power, M.; Seichepine, F.; Yang, G.Z. A bio-inspired 3D micro-structure for graphene-based bacteria sensing. Biosens. Bioelectron. 2019, 123, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Aspermair, P.; Mishyn, V.; Bintinger, J.; Happy, H.; Bagga, K.; Subramanian, P.; Knoll, W.; Boukherroub, R.; Szunerits, S. Reduced graphene oxide–based field effect transistors for the detection of E7 protein of human papillomavirus in saliva. Anal. Bioanal. Chem. 2021, 413, 779–787. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, J.; Hussain, C.M. Graphene-based field-effect transistor biosensors for the rapid detection and analysis of viruses: A perspective in view of COVID-19. Carbon Trends 2021, 2, 100011. [Google Scholar] [CrossRef]
- Thévenot, D.R.; Toth, K.; Durst, R.A.; Wilson, G.S. Electrochemical biosensors: Recommended definitions and classification. Biosens. Bioelectron. 2001, 16, 121–131. [Google Scholar] [CrossRef]
- Campuzano, S.; Yáñez-Sedeño, P.; Pingarrón, J.M. Electrochemical Affinity Biosensors Based on Selected Nanostructures for Food and Environmental Monitoring. Sensors 2020, 20, 5125. [Google Scholar] [CrossRef]
- Wang, J. Carbon-Nanotube Based Electrochemical Biosensors: A Review. Electroanalysis 2005, 17, 7–14. [Google Scholar] [CrossRef]
- Benavente, J. Electrochemical Impedance Spectroscopy as a Tool for Electrical and Structural Characterizations of Membranes in Contact with Electrolyte Solutions. In Recent Advances in Multidisciplinary Applied Physics; Elsevier Ltd.: Amsterdam, The Netherlands, 2005; pp. 463–471. [Google Scholar] [CrossRef]
- Joshi, S.R.; Sharma, A.; Kim, G.H.; Jang, J. Low cost synthesis of reduced graphene oxide using biopolymer for influenz a virus sensor. Mater. Sci. Eng. C 2020, 108, 110465. [Google Scholar] [CrossRef]
- Imran, S.; Ahmadi, S.; Kerman, K. Electrochemical Biosensors for the Detection of SARS-CoV-2 and Other Viruses. Micromachines 2021, 12, 174. [Google Scholar] [CrossRef]
- Tripathy, S.; Singh, S.G. Label-Free Electrochemical Detection of DNA Hybridization: A Method for COVID-19 Diagnosis. Trans. Indian Natl. Acad. Eng. 2020, 5, 205–209. [Google Scholar] [CrossRef]
- Alafeef, M.; Dighe, K.; Moitra, P.; Pan, D. Rapid, ultrasensitive, and quantitative detection of SARS-CoV-2 using antisense oligonucleotides directed electrochemical biosensor chip. ACS Nano 2020, 14, 17028–17045. [Google Scholar] [CrossRef]
- Mahari, S.; Roberts, A.; Shahdeo, D.; Gandhi, S. eCovSens-Ultrasensitive Novel In-House Built Printed Circuit Board Based Electrochemical Device for Rapid Detection of nCovid-19 antigen, a spike protein domain 1 of SARS-CoV-2. bioRxiv 2020. [Google Scholar] [CrossRef]
- Cho, I.H.; Kim, D.H.; Park, S. Electrochemical Biosensors: Perspective on Functional Nanomaterials for on-Site Analysis. Biomater. Res. 2020, 24, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Kaushik, A.K.; Dhau, J.S.; Gohel, H.; Mishra, Y.K.; Kateb, B.; Kim, N.Y.; Goswami, D.Y. Electrochemical SARS-CoV-2 Sensing at Point-of-Care and Artificial Intelligence for Intelligent COVID-19 Management. ACS Appl. Bio Mater. 2020, 3, 7306–7325. [Google Scholar] [CrossRef] [PubMed]
- Ahmadivand, A.; Gerislioglu, B.; Ramezani, Z.; Kaushik, A.; Manickam, P.; Ghoreishi, S.A. Functionalized terahertz plasmonic metasensors: Femtomolar-level detection of SARS-CoV-2 spike proteins. Biosens. Bioelectron. 2021, 177, 112971. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Hitzky, E.; Darder, M.; Wicklein, B.; Ruiz-Garcia, C.; Martín-Sampedro, R.; del Real, G.; Aranda, P. Nanotechnology Responses to COVID-19. Adv. Healthc. Mater. 2020, 9, 2000979. [Google Scholar] [CrossRef]
- Medhi, R.; Srinoi, P.; Ngo, N.; Tran, H.V.; Lee, T.R. Nanoparticle-Based Strategies to Combat COVID-19. ACS Appl. Nano Mater. 2020, 3, 8557–8580. [Google Scholar] [CrossRef]
- Al-Hatamleh, M.A.; Ma’mon, M.H.; Alshaer, W.; Rahman, E.N.; Mohd-Zahid, M.H.; Alhaj-Qasem, D.M.; Yean, C.Y.; Alias, I.Z.; Jaafar, J.; Ferji, K.; et al. COVID-19 infection and nanomedicine applications for development of vaccines and therapeutics: An overview and future perspectives based on polymersomes. Eur. J. Pharmacol. 2021, 896, 173930. [Google Scholar] [CrossRef]
- Du, T.; Liang, J.; Dong, N.; Fang, L.; Xiao, S.; Han, H. Antiviral activity of graphene oxide–silver nanocomposites by preventing viral entry and activation of the antiviral innate immune response. ACS Appl. Bio Mater. 2018, 1, 1286–1293. [Google Scholar] [CrossRef]
- Rashidzadeh, H.; Danafar, H.; Rahimi, H.; Mozafari, F.; Salehiabar, M.; Rahmati, M.A.; Rahamooz-Haghighi, S.; Mousazadeh, N.; Mohammadi, A.; Ertas, Y.N.; et al. Nanotechnology against the novel coronavirus (severe acute respiratory syndrome coronavirus 2): Diagnosis, treatment, therapy and future perspectives. Nanomedicine 2021, 16, 497–516. [Google Scholar] [CrossRef]
- Shehu, I.A.; Auwal, N.M.; Musa, M.K.; Mukhtar, A.; Yusuf, M.S.; Yau, A.A.; Muhammad, M.; Dala, Y.B.; Sani, S.A.; Ahmad, M.S.; et al. Innovative Nanotechnology a Boon for Fight Against Pandemic COVID–19. Front. Nanotechnol. 2021, 3, 1308. [Google Scholar] [CrossRef]
- Rabiee, N.; Ahmadi, S.; Soufi, G.J.; Hekmatnia, A.; Khatami, M.; Fatahi, Y.; Iravani, S.; Varma, R.S. Quantum dots against SARS-CoV-2: Diagnostic and therapeutic potentials. J. Chem. Technol. Biotechnol. 2022, 97, 1640. [Google Scholar] [CrossRef] [PubMed]
- Yao, X.; Ye, F.; Zhang, M.; Cui, C.; Huang, B.; Niu, P.; Liu, X.; Zhao, L.; Dong, E.; Song, C.; et al. In Vitro Antiviral Activity and Projection of Optimized Dosing Design of Hydroxychloroquine for the Treatment of Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Clin. Infect. Dis. 2020, 71, 732–739. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, S.F.; Quadeer, A.A.; McKay, M.R. Preliminary identification of potential vaccine targets for the COVID-19 coronavirus (SARS-CoV-2) based on SARS-CoV immunological studies. Viruses 2020, 12, 254. [Google Scholar] [CrossRef]
- Manivannan, S.; Ponnuchamy, K. Quantum dots as a promising agent to combat COVID-19. Appl. Organomet. Chem. 2020, 34, e5887. [Google Scholar] [CrossRef]
- Bidram, E.; Esmaeili, Y.; Amini, A.; Sartorius, R.; Tay, F.R.; Shariati, L.; Makvandi, P. Nanobased Platforms for Diagnosis and Treatment of COVID-19: From Benchtop to Bedside. ACS Biomater. Sci. Eng. 2021, 14, 2150–2176. [Google Scholar] [CrossRef]
- Sportelli, M.C.; Izzi, M.; Kukushkina, E.A.; Hossain, S.I.; Picca, R.A.; Ditaranto, N.; Cioff, N. Can Nanotechnology and Materials Science Help the Fight against SARS-CoV-2? Nanomaterials 2020, 10, 802. [Google Scholar] [CrossRef]
- Astuti, I.; Ysrafil, Y. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab. Syndr. 2020, 14, 407–412. [Google Scholar] [CrossRef]
- Raha, S.; Mallick, R.; Basak, S.; Duttaroy, A.K. Is copper beneficial for COVID-19 patients? Med. Hypotheses 2020, 142, 109814. [Google Scholar] [CrossRef]
- Kim, H.; Park, M.; Hwang, J.; Kim, J.H.; Chung, D.-R.; Lee, K.-S.; Kang, M. Development of label-free colorimetric assay for MERS-CoV using gold nanoparticles. ACS Sens. 2019, 4, 1306–1312. [Google Scholar] [CrossRef]
- Moitra, P.; Alafeef, M.; Alafeef, M.; Alafeef, M.; Dighe, K.; Frieman, M.B.; Pan, D. Selective Naked-Eye Detection of SARS-CoV-2 Mediated by N Gene Targeted Antisense Oligonucleotide Capped Plasmonic Nanoparticles. ACS Nano 2020, 14, 7617–7627. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Sagar, P.; Priyadarshi, N.; Kaul, S.; Sandhir, R.; Rishi, V.; Singhal, N.K. Nanotechnology-based approaches for the detection of SARS-CoV-2. Front. Nanotechnol. 2020, 2, 589832. [Google Scholar] [CrossRef]
- Untracht, Z.T.; Ozcan, A.; Santra, S.; Kang, E.H. SDS-PAGE for Monitoring the Dissolution of Zinc Oxide Bactericidal Nanoparticles (Zinkicide) in Aqueous Solutions. ACS Omega 2020, 5, 1402–1407. [Google Scholar] [CrossRef]
- Pedone, D.; Moglianetti, M.; De Luca, E.; Bardi, G.; Pompa, P.P. Platinum Nanoparticles in Nanobiomedicine. Chem. Soc. Rev. 2017, 46, 4951–4975. [Google Scholar] [CrossRef] [PubMed]
- Yonezawa, T. Preparation of Metal Nanoparticles and Their Application for Materials. In Nanoparticle Technology Handbook; Elsevier: Amsterdam, The Netherlands, 2018; pp. 829–837. [Google Scholar] [CrossRef]
- Yi, D.K.; Selvan, S.T.; Lee, S.S.; Papaefthymiou, G.C.; Kundaliya, D.; Ying, J.Y.J. Silica-Coated Nanocomposites of Magnetic Nanoparticles and Quantum Dots. J. Am. Chem. Soc. 2005, 127, 4990–4991. [Google Scholar] [CrossRef] [PubMed]
- Iwahori, K.; Enomoto, T.; Furusho, H.; Miura, A.; Nishio, K.; Mishima, Y.; Yamashita, I. Cadmium Sulfide Nanoparticle Synthesis in Dps Protein from Listeria innocua. Chem. Mater. 2007, 19, 3105–3111. [Google Scholar] [CrossRef]
- Mukherjee, S.; Ray, S.; Thakur, R.S. Solid Lipid Nanoparticles: A Modern Formulation Approach in Drug Delivery System. Indian J. Pharm. Sci. 2009, 71, 349–358. [Google Scholar] [CrossRef]
- Edagwa, B.; Zhou, T.; McMillan, J.; Liu, X.-M.; Gendelman, H. Development of HIV Reservoir Targeted Long Acting Nanoformulated Antiretroviral Therapies. Curr. Med. Chem. 2014, 21, 4186–4198. [Google Scholar] [CrossRef]
- Telwatte, S.; Moore, K.; Johnson, A.; Tyssen, D.; Sterjovski, J.; Aldunate, M.; Gorry, P.R.; Ramsland, P.A.; Lewis, G.R.; Paull, J.R.A.; et al. Virucidal activity of the dendrimer microbicide SPL7013 against HIV-1. Antivir. Res. 2011, 90, 195–199. [Google Scholar] [CrossRef]
- Kunz, Y.; Horninger, W.; Pinggera, G.M. Are urologists in trouble with SARS-CoV-2? Reflections and recommendations for specific interventions. Br. J. Urol. 2020, 126, 670–678. [Google Scholar] [CrossRef]
- Xue, X.; Ball, J.K.; Alexander, C.; Alexander, M.R. All Surfaces Are Not Equal in Contact Transmission of SARS-CoV-2. Matter 2020, 3, 1433–1441. [Google Scholar] [CrossRef] [PubMed]
- Feng, S.; Shen, C.; Xia, N.; Song, W.; Fan, M.; Cowling, B.J. Rational Use of Face Masks in the COVID-19 Pandemic. Lancet Respir. Med. 2020, 8, 434–436. [Google Scholar] [CrossRef]
- Bhimanapati, G.R.; Lin, Z.; Meunier, V.; Jung, Y.; Cha, J.; Das, S.; Xiao, D.; Son, Y.; Strano, M.S.; Cooper, V.R.; et al. Recent Advances in Two-Dimensional Materials beyond Graphene. ACS Nano 2015, 9, 11509–11539. [Google Scholar] [CrossRef] [PubMed]
- Sportelli, M.C.; Picca, R.A.; Cioffi, N. Recent Advances in the Synthesis and Characterization of Nano-Antimicrobials. TrAC—Trends Anal. Chem. 2016, 84, 131–138. [Google Scholar] [CrossRef]
- Muthiah, G.; Sarkar, A.; Roy, S.; Singh, P.; Kumar, P.; Bhardwaj, K.; Jaiswal, A. Nanotechnology Toolkit for Combating COVID-19 and Beyond. ChemNanoMat 2022, 8, e202100505. [Google Scholar] [CrossRef]
- O’Dowd, K.; Nair, K.M.; Forouzandeh, P.; Mathew, S.; Grant, J.; Moran, R.; Bartlett, J.; Bird, J.; Pillai, S.C. Face Masks and Respirators in the Fight against the COVID-19 Pandemic: A Review of Current Materials, Advances and Future Perspectives. Materials 2020, 13, 3363. [Google Scholar] [CrossRef]
- Matuschek, C.; Moll, F.; Fangerau, H.; Fischer, J.C.; Zänker, K.; van Griensven, M.; Schneider, M.; Kindgen-Milles, D.; Knoefel, W.T.; Lichtenberg, A.; et al. Face masks: Benefits and risks during the COVID-19 crisis. Eur. J. Med. Res. 2020, 25, 32. [Google Scholar] [CrossRef]
- MacIntyre, C.R.; Seale, H.; Dung, T.C.; Hien, N.T.; Nga, P.T.; Chughtai, A.A.; Rahman, B.; Dwyer, D.E.; Wang, Q. A cluster randomised trial of cloth masks compared with medical masks in healthcare workers. BMJ Open 2015, 5, e006577. [Google Scholar] [CrossRef]
- Abbasinia, M.; Karimie, S.; Haghighat, M.; Mohammadfam, I. Application of Nanomaterials in Personal Respiratory Protection Equipment: A Literature Review. Safety 2018, 4, 47. [Google Scholar] [CrossRef]
- Elechiguerra, J.L.; Burt, J.L.; Morones, J.R.; Camacho-Bragado, A.; Gao, X.; Lara, H.H.; Yacaman, M.J.J. Interaction of silver nanoparticles with HIV-1. J. Nanobiotechnol. 2005, 3. [Google Scholar] [CrossRef]
- Li, Y.; Leung, P.; Yao, L.; Song, Q.W.; Newton, E.J. Antimicrobial effect of surgical masks coated with nanoparticles. J. Hosp. Infect. 2006, 62, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Wahab, M.A.; Li, L.; Li, H.; Abdala, A. Silver Nanoparticle-Based Nanocomposites for Combating Infectious Pathogens: Recent Advances and Future Prospects. Nanomaterials 2021, 11, 581. [Google Scholar] [CrossRef] [PubMed]
- Ramaiah, G.B.; Tegegne, A.; Melese, B. Developments in Nano-materials and Analysing its role in Fighting COVID-19. Mater. Today Proc. 2021, 47, 4357–4363. [Google Scholar] [CrossRef] [PubMed]
- Wainwright, M. Photoinactivation of viruses. Photochem. Photobiol. Sci. 2004, 3, 406–411. [Google Scholar] [CrossRef]
- Wainwright, M. Local treatment of viral disease using photodynamic therapy. Int. J. Antimicrob. Agents 2003, 21, 510–520. [Google Scholar] [CrossRef]
- Naguib, M.; Kurtoglu, M.; Presser, V.; Lu, J.; Niu, J.; Heon, M.; Hultman, L.; Gogotsi, Y.; Barsoum, M.W. Two-Dimensional Nanocrystals Produced by Exfoliation of Ti3AlC2. Adv. Mater. 2011, 23, 4248–4253. [Google Scholar] [CrossRef]
- Lim, M.E.; Lee, Y.L.; Zhang, Y.; Chu, J.J. Photodynamic inactivation of viruses using upconversion nanoparticles. Biomaterials 2012, 33, 1912–1920. [Google Scholar] [CrossRef]
- Wiehe, A.; O’Brien, J.M.; Senge, M.O. Trends and targets in antiviral phototherapy. Photochem. Photobiol. Sci. 2019, 18, 2565–2612. [Google Scholar] [CrossRef]
- Moshfegh, F.; Khosraviani, F.; Moghaddasi, N.; Limoodi, S.F.S.J.; Boluki, E. Antiviral optical techniques as a possible novel approach to COVID-19 treatment. J. Innov. Opt. Health Sci. 2021, 14, 1–18. [Google Scholar] [CrossRef]
- Koirala, A.; Joo, Y.J.; Khatami, A.; Chiu, C.; Britton, P.N. Vaccines for COVID-19: The current state of play. Paediatr. Respir. Rev. 2020, 35, 43–49. [Google Scholar] [CrossRef]
- Roper, R.L.; Rehm, K.E. SARS Vaccines: Where Are We? Expert Rev. Vaccines 2009, 8, 887–898. [Google Scholar] [CrossRef] [PubMed]
- Jackson-Thompson, B.M.; Goguet, E.; Laing, E.D.; Mitre, E. Prospective Assessment of SARS-CoV-2 Seroconversion (PASS) study: An observational cohort study of SARS-CoV-2 infection and vaccination in healthcare workers. BMC Infect Dis. 2021, 21, 544. [Google Scholar] [CrossRef] [PubMed]
- Armengol, G.; Ruiz, L.M.; Orduz, S. Applied Biochemistry and Biotechnology—Part B. Mol. Biotechnol. 2004, 27, 109–118. [Google Scholar] [CrossRef]
- Donnelly, J.J.; Wahren, B.; Liu, M.A.J. DNA Vaccines: Progress and Challenges. J. Immunol. 2005, 175, 633–639. [Google Scholar] [CrossRef]
- Lutz, J.; Lazzaro, S.; Habbeddine, M.; Schmidt, K.E.; Baumhof, P.; Mui, B.L.; Tam, Y.K.; Madden, T.D.; Hope, M.J.; Heidenreich, R.; et al. Unmodified mRNA in LNPs constitutes a competitive technology for prophylactic vaccines. NPJ Vaccines 2017, 2. [Google Scholar] [CrossRef]
- Ma, C.; Wang, L.; Tao, X.; Zhang, N.; Yang, Y.; Tseng, C.T.K.; Li, F.; Zhou, Y.; Jiang, S.; Du, L. Searching for an ideal vaccine candidate among different MERS coronavirus receptor-binding fragments—The importance of immunofocusing in subunit vaccine design. Vaccine 2014, 32, 6170–6176. [Google Scholar] [CrossRef] [PubMed]
- World Health Organization. World Health Organization. Coronavirus Disease 2019 (COVID-19); Situation Report; World Health Organization: New York, NY, USA, 2020; p. 32. [Google Scholar]
- Airtable—Milken Institute. Available online: https://airtable.com/shrSAi6t5WFwqo3GM/tblEzPQS5fnc0FHYR/viwDBH7b6FjmIBX5x?blocks=hide (accessed on 14 November 2020).
- Ghosh, A.K.; Brindisi, M.; Shahabi, D.; Chapman, M.E.; Mesecar, A.D. Drug Development and Medicinal Chemistry Efforts toward SARS-Coronavirus and Covid-19 Therapeutics. ChemMedChem 2020, 15, 907–932. [Google Scholar] [CrossRef]
- Kim, Y.S.; Son, A.; Kim, J.; Kwon, S.B.; Kim, M.H.; Kim, P.; Kim, J.; Byun, Y.H.; Sung, J.; Lee, J.; et al. Chaperna-Mediated Assembly of Ferritin-Based Middle East Respiratory Syndrome-Coronavirus Nanoparticles. Front. Immunol. 2018, 9, 1093. [Google Scholar] [CrossRef]
- Shah, V.K.; Firmal, P.; Alam, A.; Ganguly, D.; Chattopadhyay, S. Overview of immune response during SARS-CoV-2 infection: Lessons from the past. Front. Immunol. 2020, 11, 1949. [Google Scholar] [CrossRef]
- Jung, S.Y.; Kang, K.W.; Lee, E.Y.; Seo, D.W.; Kim, H.L.; Kim, H.; Kwon, T.; Park, H.L.; Kim, H.; Lee, S.M.; et al. Heterologous prime–boost vaccination with adenoviral vector and protein nanoparticles induces both Th1 and Th2 responses against Middle East respiratory syndrome coronavirus. Vaccine 2018, 36, 3468–3476. [Google Scholar] [CrossRef]
- Patra, J.K.; Das, G.; Fraceto, L.F.; Campos, E.V.; Rodriguez-Torres, M.D.; Acosta-Torres, L.S.; Diaz-Torres, L.A.; Grillo, R.; Swamy, M.K.; Sharma, S.; et al. Nano based drug delivery systems: Recent developments and future prospects. J. Nanobiotechnol. 2018, 16, 71. [Google Scholar] [CrossRef] [PubMed]
- Sekimukai, H.; Iwata-Yoshikawa, N.; Fukushi, S.; Tani, H.; Kataoka, M.; Suzuki, T.; Hasegawa, H.; Niikura, K.; Arai, K.; Nagata, N. Gold nanoparticle-adjuvanted S protein induces a strong antigen-specific IgG response against severe acute respiratory syndrome-related coronavirus infection, but fails to induce protective antibodies and limit eosinophilic infiltration in lungs. Microbiol. Immunol. 2020, 64, 33–51. [Google Scholar] [CrossRef] [PubMed]
- Madar-Balakirski, N.; Rosner, A.; Melamed, S.; Politi, B.; Steiner, M.; Tamir, H.; Yahalom-Ronen, Y.; Bar-David, E.; Ben-Shmuel, A.; Sittner, A.; et al. Preliminary nonclinical safety and immunogenicity of an rVSV-ΔG-SARS-CoV-2-S vaccine in mice, hamsters, rabbits and pigs. Arch. Toxicol. 2022, 96, 859–875. [Google Scholar] [CrossRef] [PubMed]
- Poon, C.; Patel, A.A. Organic and inorganic nanoparticle vaccines for prevention of infectious diseases. Nano Express 2020, 1, 012001. [Google Scholar] [CrossRef]
- Pardi, N.; Hogan, M.J.; Porter, F.W.; Weissman, D. MRNA Vaccines-a New Era in Vaccinology. Nat. Rev. Drug Discov. 2018, 17, 261–279. [Google Scholar] [CrossRef]
- Hess, K.L.; Medintz, I.L.; Jewell, C.M. Designing inorganic nanomaterials for vaccines and immunotherapies. Nano Today 2019, 27, 73–98. [Google Scholar] [CrossRef]
- Business Insider India. Moderna’s Groundbreaking Coronavirus Vaccine Was Designed in Just 2 Days. Available online: https://www.businessinsider.in/science/news/modernas-groundbreaking-coronavirus-vaccine-was-designed-in-just-2-days/articleshow/79433184.cms (accessed on 29 November 2020).
- Holm, L.; Reitelseder, S.; Pedersen, T.G.; Doessing, S.; Petersen, S.G.; Flyvbjerg, A.; Andersen, J.L.; Aagaard, P.; Kjaer, M.J. Changes in muscle size and MHC composition in response to resistance exercise with heavy and light loading intensity. Appl. Physiol. 2008, 105, 1454–1461. [Google Scholar] [CrossRef]
- Talebian, S.; Conde, J. Why Go NANO on COVID-19 Pandemic? Matter 2020, 3, 598–601. [Google Scholar] [CrossRef]
- RAPS. COVID-19 Vaccine Tracker. Available online: https://www.raps.org/news-and-articles/news-articles/2020/3/covid-19-vaccine-tracker (accessed on 1 December 2020).
- Bloomberg. More Than 37.9 Million Shots Given: Covid-19 Vaccine Tracker. Available online: https://www.bloomberg.com/graphics/covid-vaccine-tracker-global-distribution/ (accessed on 16 January 2021).
- The Indian Express. World’s Largest Covid-19 Inoculation Drive Begins in India. Available online: https://indianexpress.com/photos/india-news/covid-19-vaccine-india-inoculation-drive-7148817/2/ (accessed on 16 January 2021).
- Fang, E.; Liu, X.; Li, M.; Zhang, Z.; Song, L.; Zhu, B.; Wu, X.; Liu, J.; Zhao, D.; Li, Y. Advances in COVID-19 mRNA vaccine development. Signal Transduct. Target Ther. 2022, 7, 94. [Google Scholar] [CrossRef]
- Duan, Y.; Wang, S.; Zhang, Q.; Gao, W.; Zhang, L. Nanoparticle approaches against SARS-CoV-2 infection. Curr. Opin. Solid State Mater. Sci. 2021, 25, 100964. [Google Scholar] [CrossRef]
- Rauf, M.A.; Tasleem, M.; Bhise, K.; Tatiparti, K.; Sau, S.; Iyer, A.K. Nano-therapeutic strategies to target coronavirus. View 2021, 2, 20200155. [Google Scholar] [CrossRef]
- Koppers-Lalic, D.; Hogenboom, M.M.; Middeldorp, J.M.; Pegtel, D.M. Virus-Modified Exosomes for Targeted RNA Delivery; A New Approach in Nanomedicine. Adv. Drug Deliv. Rev. 2013, 65, 348–356. [Google Scholar] [CrossRef] [PubMed]
- Li, E.; Chi, H.; Huang, P.; Yan, F.; Zhang, Y.; Liu, C.; Wang, Z.; Li, G.; Zhang, S.; Mo, R.; et al. A Novel Bacterium-Like Particle Vaccine Displaying the MERS-CoV Receptor-Binding Domain Induces Specific Mucosal and Systemic Immune Responses in Mice. Viruses 2019, 11, 799. [Google Scholar] [CrossRef]
- Park, H.J.; Lee, E.Y.; Jung, S.; Ko, H.L.; Lee, S.M.; Nam, J.H. Spike nanoparticle and recombinant adenovirus 5 vaccines induce specific antibodies against the Middle East respiratory syndrome coronavirus (MERS-CoV). J. Immunol. 2017, 198, 255. [Google Scholar]
- Attia, M.A.; Essa, E.A.; Elebyary, T.T.; Faheem, A.M.; Elkordy, A.A. Brief on Recent Application of Liposomal Vaccines for Lower Respiratory Tract Viral Infections: From Influenza to COVID-19 Vaccines. Pharmaceuticals 2021, 14, 1173. [Google Scholar] [CrossRef] [PubMed]
- Rezabakhsh, A.; Mahdipour, M.; Nourazarian, A.; Habibollahi, P.; Sokullu, E.; Avci, Ç.B.; Rahbarghazi, R. Application of exosomes for the alleviation of COVID-19-related pathologies. Cell Biochem. Funct. 2022, 40, 430–438. [Google Scholar] [CrossRef]
- Mao, L.; Chen, Z.; Wang, Y.; Chen, C. Design and application of nanoparticles as vaccine adjuvants against human corona virus infection. J. Inorg. Biochem. 2021, 219, 111454. [Google Scholar] [CrossRef]
- Schoenmaker, L.; Witzigmann, D.; Kulkarni, J.A.; Verbeke, R.; Kersten, G.; Jiskoot, W.; Crommelin, D.J.A. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int. J. Pharm. 2021, 601, 120586. [Google Scholar] [CrossRef]
- Mazini, L.; Rochette, L.; Malka, G. Exosomes contribution in COVID-19 patients’ treatment. J. Transl. Med. 2021, 19, 1–8. [Google Scholar] [CrossRef]
- Hou, X.; Zaks, T.; Langer, R.; Dong, Y. Lipid nanoparticles for mRNA delivery. Nat. Rev. Mater. 2021, 6, 1078–1094. [Google Scholar] [CrossRef]
| Nano-Assisted Drugs | Mechanism |
|---|---|
| 1. ACE 2 receptors | It has been confirmed that SARS-CoV-2 enters cells via the ACE2 receptor. This implies that the blockage or modulation of ACE2 could be a potential means to halt the cellular entry of SARS-CoV-2 and prevent subsequent infection. The mechanism of blockage involves either inhibiting the virus ACE2-RBD interaction, directly inhibiting ACE2-receptors, or using recombinant human ACE2 (rhACE2); APN01 to disrupt virus endocytosis [21]. |
| 2. TMPRSS2 | Human TMPRSS2 more powerfully activates the COVID-19 spike protein and facilitates its fusion across the host cell membrane via ACE2 than endosomal cathepsin and other protease enzymes. As a result, targeting TMPRSS2 may be critical for developing anti-SARS-CoV-2 molecules. Some potent TMPRSS2 inhibitors, such as camostate and bromhexine, have been shown to be repurposed against COVID-19. Stopsack K investigated the anti-TMPRSS2 activity of androgens and other corticosteroids and demonstrated their potential for repurposing in the treatment of COVID-19. Furthermore, determining the crystal structure of the TMPRSS2 protein could aid in the development of COVID-19 [23]. |
| 3. Furin | The SARS-CoV-2 S-protein possessed a Furin-like cleavage site (FCS), which is responsible for the S-structural protein’s form, electrostatic interaction, and furin binding affinity. The study in Zhejiang province examined the effect of FCS mutation (F1-2) Hasan A and demonstrated the impact of furin-ACE2 enzyme interaction and its pathogenic role in COVID-19 infection, determining its relative proclivity to influence the structure of S-protein and the interaction between FCS and Furin [24]. |
| 4. Capthepsin L | The activation of S-protein by lysosomal cathepsin facilitates COVID-19 cellular fusion and endocytosis. Targeting the cathepsin L enzyme could be a therapeutic option for developing novel anti-coronavirus moiety. The Hoffmann M study demonstrated the effectiveness of cathepsin L inhibitor (SID26681509) in vitro and concluded that the compound could selectively prevent pseudovirus cellular entry by 76 % [25]. It may help to prevent the progression of pulmonary fibrosis. Furthermore, combining cathepsin L with TMPRSS2 inhibition to target COVID-19 could provide an effective cure for coronavirus infection. However, the lack of a crystal structure for cathepsin L could aid in the development of specific antibodies against the cathepsin L enzyme and thus the COVID-19 infection [26]. |
| 5. Two-Pore Channel (TPC2) | This is a voltage-gated channel essential for sodium and calcium exchange across the cell membrane, and it was reported to play a vital role in cellular trafficking of the Ebola virus [27]. The combination of antidepressants (such as pimozide and fluphenazine) and selective estrogen receptor modulators SERM (such as raloxifene, tamoxifen and clomiphene) is reported to have shown a potential blockage effect on TPC2 [28], and hence, it could be regarded as a possible target to curb COVID-19 infection. |
| 6. AAK1 and GKA | AAK1 and GKA are serine-threonine kinase enzymes responsible for virus intracellular endocytosis and genomic release in hepatitis c, Ebola, Dengue and prospectively COVID-19 viruses [29]. Baricitanib, an AAK1 and GKA inhibitor, was suggested for the treatment of COVID-19 by interfering with the virus’s cellular entrance mechanism, although the proposal lacked clinical validation. However, the clinical trial conducted in Europe for the use of Baricitanib in COVID-19 patients has shown a negative prognosis [30]. Structural-based studies of AAK1 and GKA revealed the possibility of developing or modifying drug molecules effective for the treatment of COVID-19 infection [31]. |
| 7. Nonstructural protein: | Coronavirus possessed 16 various nsp rendering functions, although the specific functions elicited by nsp remain unclear. The crystal structure of nsp has been established, aiding the development of novel molecules in blocking its action [32,33]. |
| 8. Proteases 3CLpro and PLpro | The COVID-19 main proteases 3CLpro and PLpro have a vital role in its replication processes. Inhibition of 3CLpro and PLpro with peptides and peptidomimetics would be a hot target for COVID-19 prevention [34]. Previously, the FDA approved two potent inhibitors of 3CLpro (lopinavir and ritonavir) as repurposed drugs for use against COVID-19 [35]. |
| 9. RNA-Dependent RNA Polymerase (RdRp) | RdRp plays a crucial role in facilitating the process of genome replication and transcription in the COVID-19 life cycle [36]. The development of the RdRp structure in complex with its cofactors (nsp7 and nsp8) is encouraging. Therefore, interfering with the function of these enzymes would help the drug design for the treatment of COVID-19 infection [37]. |
| Biosensor | Principle | Targeted Component | Detection Time | Ref. |
|---|---|---|---|---|
| Optical fiber-based biosensor | Act by the measurement of absorbance, refractive Index, fluorescence, and chemiluminescence | Bio-receptor | Immediate | [84,85] |
| Indirect immunofluorescence biosensors | Determination of antibody and antigen substrate against infectious disease | Epithelial sodium channel | Several hours | [86] |
| Nanowire biosensors | Field-effect transistor application | DNA sequence and specific protein | 44–46 min | [87] |
| RT-LAMP-VF | Isothermal amplification labeled using fluorescence iso-thiocyanate | N-gene of the MERS-CoV | 30 min | [88] |
| Amperometric biosensor | Production of a potential or current energy, which is proportional to the concentration of the detected substance | Glucose concentration | 15 min | [89] |
| Piezoelectric biosensor | Acoustics (sound vibration) | Alters crystal surface frequency | 10 min | [90] |
| Conductormetric biosensor | Cheap thin-film standard technology | Enzyme, whole-cell and DNA | Short time | [91] |
| Localized surface plasmon resonance | Electromagnetic incident light excited the electrons of metal nanoparticles to oscillate | Nucleic Acid | NA | [92] |
| Lanthanide-Doped Nanoparticle-based Lateral Flow Immunoassay | Detection Antibody Conjugated with AuNP | Phosphoprotein, Specific IgM anti-P. jirovecii antibody | 10 min | [93] |
| DETECTOR (Lateral Flow detection) | Amplification of viral genetic material | Nucleic Acid | 45 min | [76] |
| CRISPER/Cas-12 a based detection with the naked eye | Generation of green fluorescence in blue light observable with the naked eye | Nucleic Acid | 45 min | [94] |
| Field-Effect Transistor-based Biosensor | The semiconductor devices work by the change in potential differences that result in the sensitivity of the indicator | Spike protein | Few minutes | [95] |
| Reverse Transcription Loop-Mediated Isothermal Amplification (RT-LAMP) | Synthesize and amplify cDNA from template RNA to detect them using LAMP technology | Nucleic Acid | 30–45 min | [96,97] |
| RT-LAMP + lateral flow | The assays containing positive COVID-19 produce a green light using Visual detection reagent | ORF 1a/b | 60 min | [98] |
| Nanomaterials | The Mechanism of Action | Remarks | Reference |
|---|---|---|---|
| Silver nanoparticles (Ag+) | AgNPs act by promoting and forming a process for Ag+ coordinating to the iodine probe base on irreversible tandem ring-opening in which the color and fluorescence changed | It has a veridical effect when used in a facial mask, and it reduces the titter of COVID-19 (SARS-CoV-2) to zero | [170] |
| Gold nanoparticles (AU) | Detect MASS-COV using double-stranded DNA by calorimeter assay | Eosinophilic infiltration in lungs | [171] |
| Zinc oxide (ZnO) nanoparticles | Lowe bacterial and viral detection capacity in patients with minimal viral load | Limit the cytokine storm in COVID-19 and limit the damage to the tissue | [172] |
| Zinc sulfate (ZnSNps), nanoparticles | Substrate specific preferences of SARS-CoV and SARS-CoV-2, but mechanism remains unclear | Increases the alcohol content beyond the comfort zone in yeast pushing | [173] |
| Platinum nanoparticles | The nanocomposite of platinum characterizes by a transmission electron microscope in which the average particle size of 2.2 nm ± 0.6 | Very low tolerable drug payload in antibody drugs conjugated | [174] |
| Magnetics nanoparticles | Detection of virus and virus genome in immunoassay due to the alternation of magnetization with changing electrical resistance from the high volts into low volt | Its application limited to in vivo tests that demonstrate the efficacy of graphene oxide | [175] |
| Silica-coated magnetic nanoparticles | Work in sandwich hybridization assay for cDNA target detection with a limit value of 2.0 × 103 copies within 6 h | Do not have an antifungal effect; it is limited only to the microorganism’s circulation in the air | [176] |
| Cadmium sulfide (CdS)-NPs | In the ambient temperature, the core-shell nanowire of CdS possesses superior intense green luminescence emission, which demonstrates charge performing carrier’s recombination inside the CdS core | The detection limit in the fluorescence intensity range is 1.2 × 10−11 | [177] |
| Lipid-based NPs | It enhances drug delivery in a controlled manner with high drug loading capacity | Because of its crystallization, it has the possibility of drug expulsion upon storage and low drug loading efficiency | [178] |
| Solid lipid NPs | It decreases the effect of first-pass metabolism and hence enhanced tissue distribution of drugs | ||
| Nanoemulsions (NEs), | It works by increasing bioavailability and lymphatic uptake of some drugs such as saquinavir or indinavir | Instability, low solubilizing capacity for high-melting substance | [65] |
| Polymer-based NPs | It improves the pharmacokinetic parameters of the drug preventing its early degradation and side effects | Biphasic drug release | [179] |
| Dendrimers | They have excellent cellular uptake with longer circulation times, and inhibiting viral entry fusion has been successful in treating HIV and HSV2 infections | Low aqueous solubility and high nonspecific toxicity | [180] |
| Sponsor | Vaccine | Description | Effectiveness | Status | Ref. |
|---|---|---|---|---|---|
| Moderna | mRNA-1273 | mRNA-based vaccine capped with LNPs | 94.1% | Approved | [226] |
| Oxford University AstraZeneca; IQVIA; and Serum Institute of India | AZD1222 | Replication-deficient viral vector-based vaccine (adenovirus from chimpanzees) | 70.4% | ||
| Pfizer, BioNTech | BNT162 | mRNA-based vaccine | 90% | ||
| CanSino and Biologics | Ad5-nCoV | Recombinant vaccine (adenovirus type 5 vector) | Encouraging | ||
| Sinovac | CoronaVac | Inactivated vaccine (formalin with alum adjuvant) | 92.4% | ||
| Bharat Biotech; National Institute of Virology | Covaxin | Inactivated vaccine | Encouraging | ||
| Johnson & Johnson | JNJ-78436735 (formerly Ad26.COV2.S) | Non-replicating viral vector | Hold on pending the outcome of ADR | ||
| Novavax | NVX-CoV2373 | Nanovaccine | Encouraging | ||
| Gamaleya Research Institute, Acellena Contract Drug Research and Development | Sputnik V | Non-replicating viral vector | 92% | ||
| Federal Budgetary Research Institution State Research Center of Virology and Biotechnology | EpiVacCorona | Peptide vaccine | Encouraging | ||
| Beijing Institute of Biological Products; China National Pharmaceutical Group (Sinopharm) | BBIBP-CorV | Inactivated vaccine | 86% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shehu, I.A.; Musa, M.K.; Datta, A.; Verma, A. Application of Nanotechnology in COVID-19 Infection: Findings and Limitations. J. Nanotheranostics 2022, 3, 203-232. https://doi.org/10.3390/jnt3040014
Shehu IA, Musa MK, Datta A, Verma A. Application of Nanotechnology in COVID-19 Infection: Findings and Limitations. Journal of Nanotheranostics. 2022; 3(4):203-232. https://doi.org/10.3390/jnt3040014
Chicago/Turabian StyleShehu, Ibrahim A., Muhammad K. Musa, Aparna Datta, and Amita Verma. 2022. "Application of Nanotechnology in COVID-19 Infection: Findings and Limitations" Journal of Nanotheranostics 3, no. 4: 203-232. https://doi.org/10.3390/jnt3040014
APA StyleShehu, I. A., Musa, M. K., Datta, A., & Verma, A. (2022). Application of Nanotechnology in COVID-19 Infection: Findings and Limitations. Journal of Nanotheranostics, 3(4), 203-232. https://doi.org/10.3390/jnt3040014