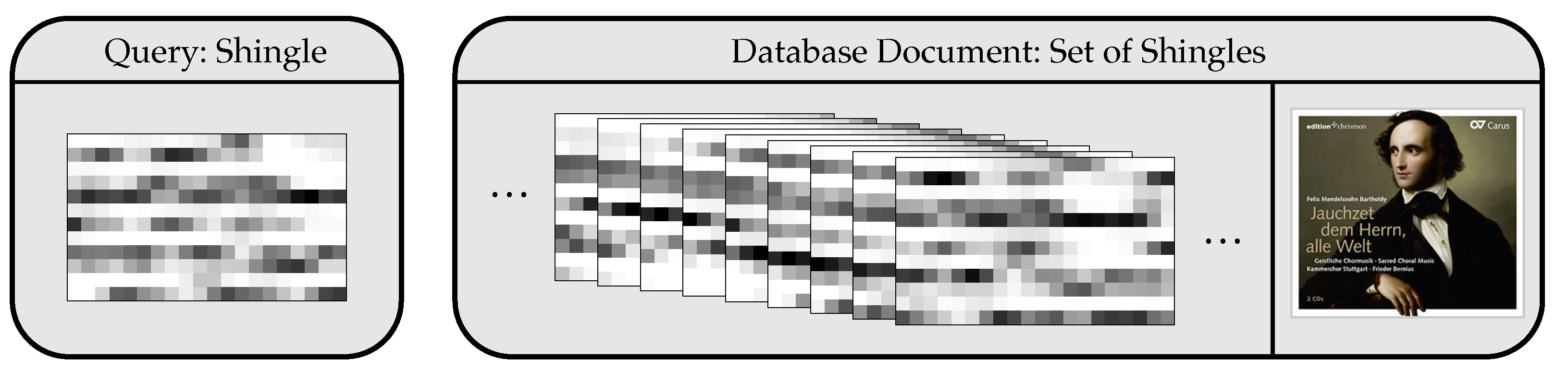
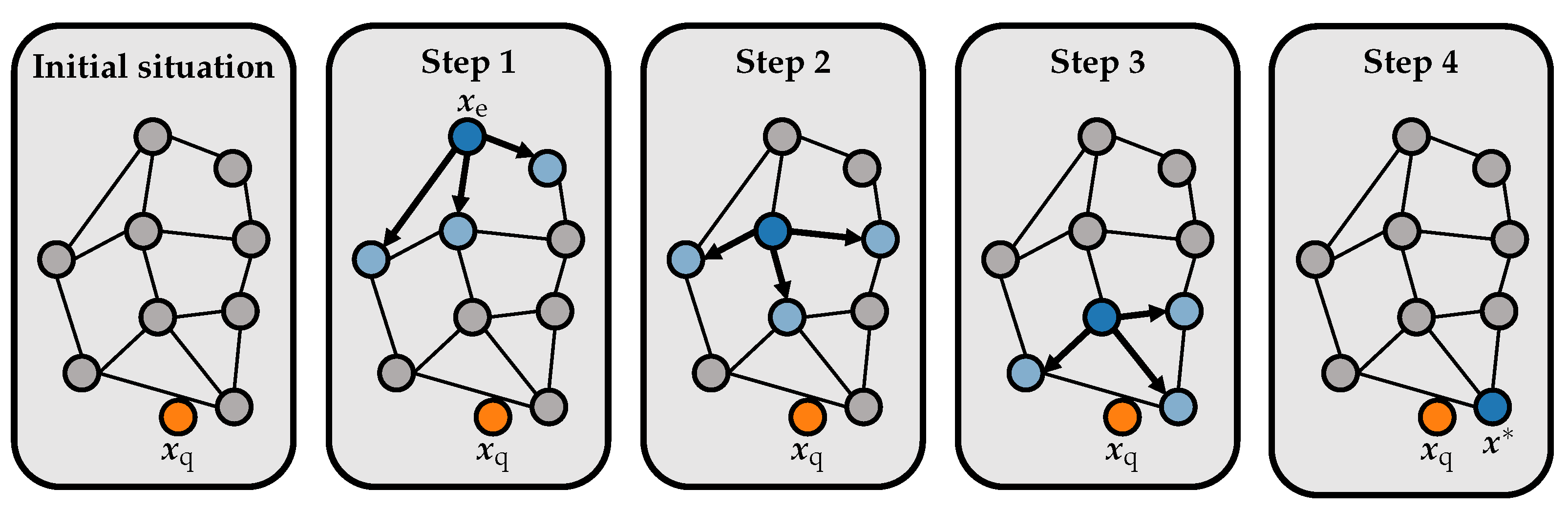
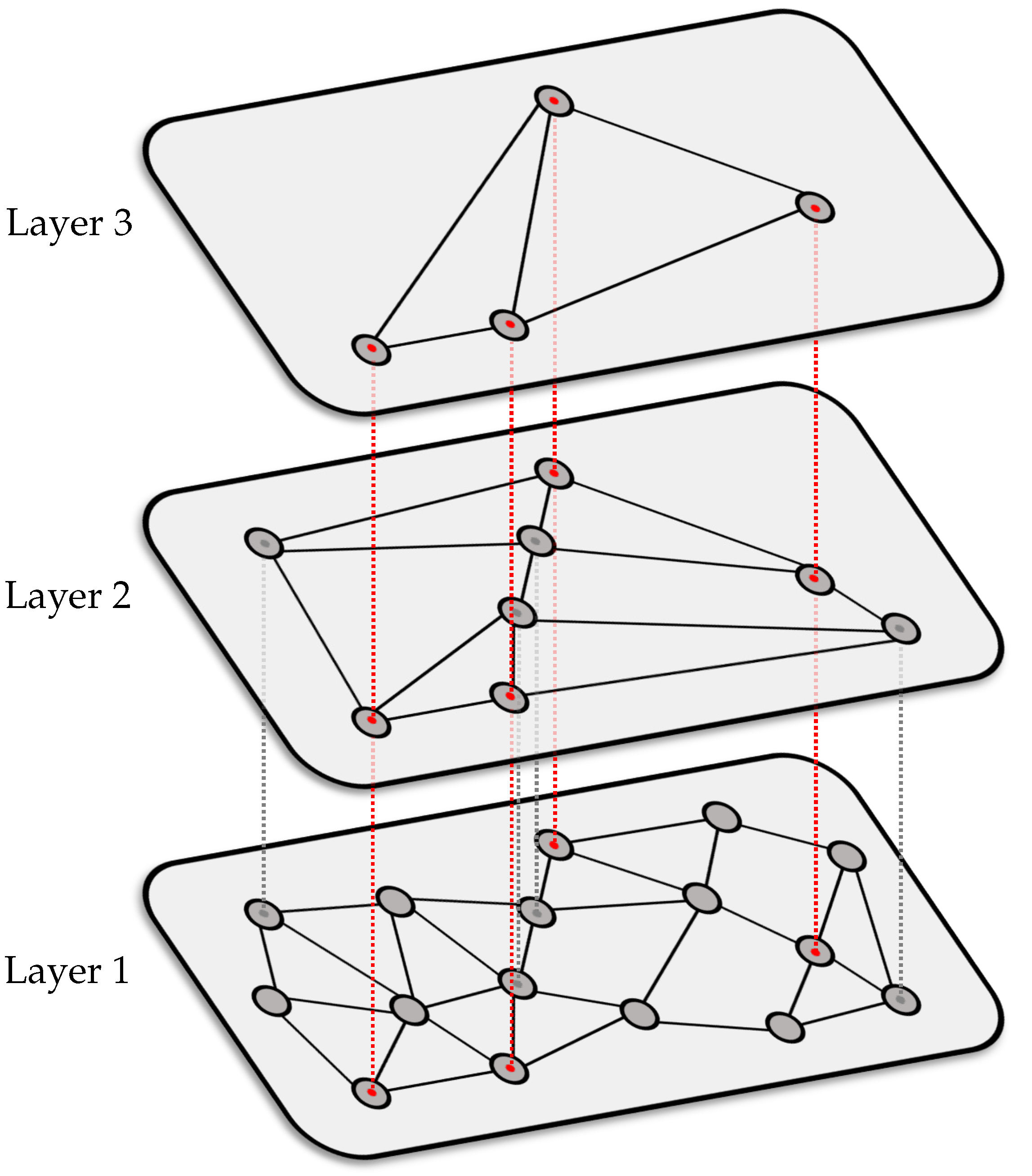
We now use the HNSW graph as an index structure in our music retrieval application. Here, a node of the graph corresponds to a database shingle with or without dimensionality reduction.
4.1. Experimental Setup
In the following, we analyze the possible decrease in retrieval quality and the improvements of the retrieval runtime introduced by the HNSW graph. To this end, we consider quantitative performance measures, which we list in
Table 2. To evaluate the retrieval quality, we use standard precision-based measures (more details in
Section 4.2). To analyze the impact of the index structures on the retrieval speed, we consider several steps that are involved in our retrieval scenario. Some steps need to be computed offline when processing the database documents, and other steps need to be computed online when processing a query. In the offline phase, we need to compute features for all audio files of the database, construct an index and save the index file to disk. These steps can be carried out at any time and on any system (offline). When applying our index, we first need to load the index into the computer’s main memory (RAM). This loading step can be considered as being in-between the online and offline stages. The index loading needs to be performed on the actual system where the retrieval service is provided. When the index structure can be kept in the main memory, it does not have to be reloaded for each query. Therefore, we still consider it as part of the offline stage. In the actual online phase, we need to compute the query features and perform the nearest neighbor search procedure using our index structure. In the following sections, we analyze these steps in the order given in
Table 2.
If not mentioned otherwise, we always report on average time measurements (
) for 100 iterations of the experiment. Note that runtime evaluation is a delicate topic on its own [
39]. For example, one may argue that it is more meaningful to report minimum instead of average runtime measurements because other processes running in parallel affect the mean more than the minimum. In our case, this is not a major issue because the standard deviation
is always relatively low. We want to highlight that we take a practical perspective by measuring runtimes using distinct implementations of the respective index structures, implemented in different programming languages. The absolute runtimes obtained may vary when using different implementations or hardware systems. Our study gives practical insights into the runtimes obtained by specific implementations on specific platforms for our specific application. In general, we are interested in the orders of magnitude, the relative differences between the time measurements, and the relationships between index size and runtime.
We compare three different search approaches: an exhaustive search approach (full search, exact search solution), an indexing strategy using
K-d trees (KD, exact search solution), and the graph-based index structure (HNSW, approximative search solution). We perform our experiments using Python 3.6.5 on a computer with an Intel Xeon CPU E5-2620 v4 (2.10 GHz) and 31 GiB RAM. We use the efficient pairwise-distance calculation of scipy 1.0.1 [
40] for the full search, which is calling a highly optimized implementation in C. For the
K-d trees (using a default leaf size of 30), we use the implementation of scikit-learn 0.20.1 [
41], which is written in Cython. For the HNSW graph, we use the efficient hnswlib implementation in C++ by the authors of the original paper [
25] (
https://github.com/nmslib/hnswlib, accessed on 12 May 2021), using the Python wrapper version 0.4.0. We use librosa 0.7.1 [
42] for the audio processing pipelines and TensorFlow 1.7.0 [
43] for the deep neural network implementation.
4.2. Retrieval Quality
In contrast to the
K-d tree approach, the HNSW graph only provides an approximate search solution. To understand the impact of this approximation within our retrieval scenario, we measure our retrieval system’s quality using the dataset
, closely following [
9].
We consider a document-level rather than a shingle-level retrieval. Here, the distance between a query shingle and a document is given by the minimizing distance between the query and all document shingles. We construct a single index structure for the entire dataset
(using either a
K-d tree or an HNSW graph) and search for the 10,000 nearest items in the database to a given query. Using the distances of the returned items, we create a ranked list of documents, ordered by ascending distances. Note that we were not able to rank all database documents as some documents may not have a corresponding item among the items returned (this did not affect the evaluation measures in our experiments). For evaluating the ranked list, we consider three standard retrieval evaluation measures [
44]. First, we use precision at one (P@1), which is 1 if the top-ranked document is relevant (i.e., being a version of the same musical work as that of the query), and 0 otherwise. Note that, for exact nearest neighbor searches, the top match is always identical to the query because, in our experiments, the query is part of the database (which leads to a P@1-value of 1). We still use this measure to check whether the approximate search approach is able to find the “trivial” match. Second, we use precision at three (P@3), which is the proportion of relevant documents among the top 3 documents of the ranked list. Third, we use
R-precision (P
), which is the proportion of relevant documents among the first
R ranks, where
denotes the number of relevant documents for the given query (which may differ for each query, between 4 and 67).
We generate 3300 query shingles from
by an equidistant sampling of ten queries from each recording of
, resulting in 3300 queries. Each evaluation measure is finally averaged over the 3300 query shingles used.
Table 3 shows the resulting evaluation measures. A row in this table specifies the dimensionality reduction approach (no reduction, PCA-, or DNN-based embedding), the dimensionality
K, and the search strategy (Full Search, or HNSW). The retrieval results for the exhaustive search (Full Search) and the
K-d tree strategy (KD) are identical because both approaches are exact nearest neighbor searches (with different properties in terms of runtime, as decribed in
Section 4.5). For example, without dimensionality reduction, we obtain a P@1-value of 1.0, a P@3-value of 0.9965, and a P
-value of 0.9434. This result shows that the shingle-based retrieval approach is able to identify most of the versions correctly, but there are a few false positives. Reducing the shingle dimensions (which is important for some indexing approaches, such as
K-d trees) leads to further degradations of the retrieval quality, as already shown in previous work [
9]. For example, reducing the dimensionality from 240 to 30 with the PCA-based embedding, we obtain a P@3-value of 0.9910. For smaller dimensionalities, the DNN is beneficial over PCA for embedding the shingles, e.g., resulting in P
-values of 0.7350 (PCA) and 0.8333 (DNN) for
. Using the HNSW graph as an index structure, we obtain more or less the same evaluation metrics for all settings. This finding demonstrates that the approximate search approach of the HNSW graph has almost no negative impact on the retrieval results within our application scenario. When we analyze the runtime improvements in the following sections, we can bear in mind that they come without substantial loss in retrieval quality.
4.3. Feature Computation
In this section, we report on the runtimes for the various steps involved in the feature computation. This computation procedure has to be performed in the offline stage (for the whole database) and online stage (for the query). In contrast to the document-based analysis of the retrieval quality (evaluating a ranked list of documents), we now use an item-based evaluation (runtime to process a database item). To compute our measurements, we first load 20 s of an audio file (using
librosa.load). In general, our audio files are longer, but for our runtime experiments, we only use a 20-second segment, corresponding to the length of a shingle. Then, we compute the spectral features [
33] (with
librosa.iirt). Next, we compute the CENS features [
34] (using
librosa.feature.chroma_cens). This step concludes the feature computation if no dimensionality reduction is applied. An additional step is performed in the embedding-based retrieval approaches (PCA-based embedding using
sklearn.decomposition.PCA or DNN-based embedding as described in [
9]).
Table 4 shows the time measurements. The loading of the audio segment only takes 44.8 ms. The next step is the spectral feature computation, which needs more than a second (1171.5 ms). The time required for the CENS computation is not significant (0.9 ms). In the table, we list the times to embed the shingle for some selected dimensionalities. In general, the PCA-based embedding is faster than 0.1 ms, and the DNN-based embedding does not take more than 2 ms.
The numbers of the table show that the major bottleneck of the feature computation is the spectral transform. Compared to this, the times of the other steps are not significant. The runtime for the query feature transform is not our focus in this paper. Obviously, it only scales linearly with the query length, which is usually short (i.e., no scalability issue). The runtime for computing the database documents’ features is also not critical because it can be computed offline. A possible future research direction could be to compute embeddings from spectral representations that are less expensive to compute, e.g., using the short-time Fourier transform (STFT) with the FFT. Having the same window and hop length settings as the spectral transform used, computing the magnitude STFT for the 20-s audio snippet only takes 19.2 ms on average. However, using the STFT-based features may go along with a decrease of feature quality, which may affect the retrieval results.
4.4. Constructing, Saving, and Loading the Index
We now address various performance measures for the offline stage, i.e., for constructing, saving, and loading the index structures. For these steps, we restrict our analysis to one embedding technique (PCA) because the specific embedding strategy used has only a minor influence on these measures. The first step is to construct the index structure (either a K-d tree or an HNSW graph). We report on the times required to construct the index, given that the data to be indexed is already in the computer’s main memory (i.e., pre-computed shingle embeddings, without having any distances pre-computed). When this data needs to be read from disk, it will cause some additional overhead. For example, loading all pre-computed shingle embeddings () of takes 0.9 ms on average. Loading the full shingles () of requires one second on average.
Columns 4 and 5 of
Table 5 show the time needed to construct the index structures for various dimensionalities. We include time measurements for the smaller dataset
and the larger dataset
. The first row in the table refers to the
K-d tree index for shingles without dimensionality reduction (
). This setting leads to construction times of 0.54 s for
and 43.03 s for
. For lower dimensions, this time decreases. For example, constructing the
K-d tree index for
takes 3.89 s, 1.66 s, and 0.99 s for the dimensionalities of 30, 12, and 6, respectively. Constructing an HNSW graph for
requires 0.65 s for the smaller dataset
and 26.66 s for the larger dataset
. For this large dataset and a high dimensionality of
, constructing an HNSW graph is faster (26.66 s) than constructing a
K-d tree (43.03 s) in the implementations used. However, this is not the case for lower dimensions. For example, constructing the index structures for the larger dataset
using
requires 3.89 s for the KD and 16.38 s for the HNSW approach. In general, the construction time grows approximately in a linear fashion with the dimensionality
K for lower dimensions. Only for large dimensionalities, the time for constructing a
K-d tree explodes, which agrees with the fact that
K-d trees are not suited for high-dimensional data [
21,
37]. In contrast, the HNSW approach behaves stable for all considered dimensionalities. In all our settings, constructing an index takes less than a minute. We do not consider this duration critical in our application because the step is performed offline.
The next step is to save the index structure to the hard disk, where it requires disk space. In the case of the
K-d tree, we use scikit-learn’s [
41] recommended default persistence format based on the Python package joblib without compression. In the case of the HNSW graph, we use the default storage format of hnswlib, which is a custom binary format (also without compression). Columns 6 and 7 of
Table 5 show the required disk space used for storing the index structures. Without dimensionality reduction (
), storing the
K-d tree requires 209.7 MB and 5161.2 MB of disk space for
and
, respectively. The HNSW graph takes 53.9 MB and 1310.9 MB for the same data. In general, the required disk space scales roughly linearly with the dataset size as well as with the dimensionality in both indexing approaches. Furthermore, the graph-based index is generally more space-efficient than the tree-based structure in the given formats.
To apply a pre-computed index for retrieval, we need to load it into the computer’s main memory. We perform this step of loading the index with the functions required for the respective file formats used in the previous step (using functions from joblib and hnswlib, respectively). Columns 8 and 9 of
Table 5 show the required time to load the index files. Loading a
K-d tree without dimensionality reduction requires 131.5 ms for
and 3209.1 ms for
. Using the same data, loading an HNSW graph takes 108.1 ms and 2680.0 ms, respectively. For smaller dimensions, loading a
K-d tree is faster than loading an HNSW graph (e.g.,
and
: 167.0 ms for KD and 2071.5 ms for HNSW). The time to load an index scales linearly with the dimensionality in both KD and HNSW approaches (with a much flatter slope for HNSW).
With our system (see
Section 4.1 for specifications), we did not have any issues with loading the index structures into the main memory (31 GiB RAM) in all settings. On other systems with less memory (8 GiB RAM), we could not fully load the index structures into the main memory for
. In this case, dimensionality reduction becomes crucial for practical reasons.
4.5. Retrieval Time
In this section, we report on our runtime experiments for searching the nearest neighbors in our datasets. Regarding efficiency, these experiments refer to the most critical part of the retrieval pipeline because this step is part of the online stage (where the runtime affects the user experience). An efficient search is of major importance for scalability because, in general, the search runtime scales with the dataset size. The aim of our index structures is to improve the efficiency of this step.
We can also consider the complexity of the search approaches from a theoretical perspective (which is not the focus of this paper, being a practice report). The runtime of the full search increases linearly with the dataset size. In other words, its search complexity is in the order of
. The expected search complexity for
K-d trees is in the order of
[
13]. However, it is well known [
45] that the actual performance may be equivalent or worse than an exhaustive search, depending on the data distribution (which influences the tree structure). Especially for high-dimensional data, the search performance degenerates. According to [
25], the overall complexity scaling of the search for the HNSW graph is
, which does not degenerate for high-dimensional data.
For the experiments of this section, we search for the
nearest items in our dataset (either
or
) to a given query, where we consider
. We again use 3300 queries (as in
Section 4.2) and perform several repetitions of this experiment. Then, we normalize the measured runtimes with respect to the number of queries and repetitions, such that the reported measures refer to the time needed for a single query.
Table 6 shows the results of our experiments. Let us first consider the PCA-based dimensionality reduction using
for the smaller dataset
. The exhaustive search requires 4.803 ms. The indexing approaches are much faster, taking 0.009 ms using a
K-d tree and 0.007 ms using an HNSW graph. In this setting, there is no large difference between the indexing approaches. When we search for more neighbors (increasing
), the KD approach slows down substantially (0.621 ms, 1.001 ms, and 2.025 ms for
values of 10, 100, and 1000, respectively). The runtime does not increase to the same extent for the HNSW approach (0.007 ms, 0.007 ms, and 0.056 ms). Searching in the larger dataset
shows the potential of the HNSW index even more clearly. For example, using the PCA-based embedding (
), for
, the
K-d tree requires 66.499 ms and the HNSW graph needs only 0.019 ms. While the increased dataset size only has a minor effect on the runtime for the HNSW approach, it dramatically increases the KD strategy’s runtime. For lower dimensionalities, the runtime differences between the
K-d tree and the HNSW graph are less extreme. For example, the KD approach requires 0.201 ms for
(
,
). Still, the HNSW graph is much faster (0.011 ms). Without dimensionality reduction, the KD approach breaks down (more than 700 ms for
), which is a known fact [
21,
37]. However, the HNSW graph still facilitates fast retrieval (e.g., 0.021 ms for
). This substantial decrease in retrieval time shows the power of the graph-based search approach. In general, the tendencies discussed for the PCA reduction are similar when using the DNN-based embedding.
Note that the exhaustive search involves computing all pairwise distances between the query and the database items. As a consequence, the parameter does not influence the search time. Furthermore, we did not perform the full search for because of excessive memory requirements.
Figure 5 shows the search runtimes for various dimensionalities
K. We observe that the runtime increases more than linearly with dimensionality
K for the
K-d tree. In contrast, the runtime for the HNSW graph increases only slightly with increasing dimensionality
K. Note the different scales of the vertical axes, which again underline the substantial search time improvements caused by the HNSW graph. We see that the HNSW approach requires nearly the same time for searching 1, 10, or 100 database items (resulting in overlapping curves). The reason for this is the parameter setting
(number of intermediate neighbor candidates, described in
Section 3.2), which leads us to search internally for 100 neighbors anyway.
To summarize, we can conclude that the dimensionality reduction approach (PCA or DNN) has only a minor influence on the runtime, which is expected. The dimensionality
K of the index items has a substantial impact on the runtime. Still, the indexing approach (KD or HNSW) has the most important effect on the runtime. Our experiments show that, compared to
K-d trees, the HNSW index is much faster and more stable concerning the dimensionality and the number of items to be indexed. This increase in retrieval efficiency comes without substantial loss in retrieval quality (as shown in
Section 4.2), which makes the HNSW graph a powerful tool for music retrieval.