SkinVisualNet: A Hybrid Deep Learning Approach Leveraging Explainable Models for Identifying Lyme Disease from Skin Rash Images
Abstract
1. Introduction
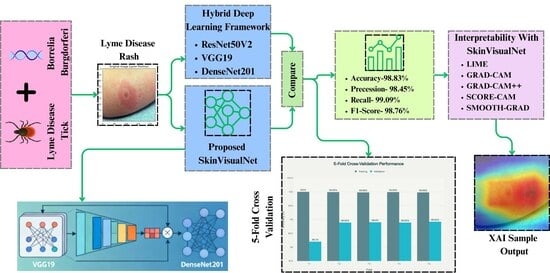
- Developed the SkinVisualNet model by combining VGG19 and DenseNet201, achieving 98.83% accuracy and 98.31% F1 score in Lyme disease rash classification which outperformed other related works that used same dataset.
- Applied gamma correction, Contrast stretching and data augmentation, boosting all model’s accuracy, Precision, recall and f1 score up to 10–13%.
- Applied LIME to provide localized, interpretable explanations for individual predictions by highlighting the specific visual features that influenced the model’s decision, thereby enhancing user understanding and trust.
- Applied Grad-CAM, Grad-CAM++, Score CAM and Smooth Grad for visual explanations by generating heatmaps that highlight important regions in input images, enhancing interpretability.
2. Related Work
Background
3. Materials and Method
3.1. Dataset Description
3.2. Data Preprocessing
3.2.1. Gamma Correction
3.2.2. Contrast Stretching
3.2.3. Data Augmentation
3.3. Dataset Splitting
3.4. Data Verification
3.5. Model Description
3.5.1. VGG19
3.5.2. DenseNet201
3.5.3. ResNet50V2
3.5.4. SkinVisualNet (Proposed Hybrid Model)
3.5.5. Parameters of Hybrid Model
4. Results
4.1. Result Analysis Before Preprocessing
4.2. Result Analysis After Preprocessing
4.2.1. Performance of the VGG19
4.2.2. Performance of the DenseNet201
4.2.3. Performance of the ResNet50V2
4.2.4. Performance of the SkinVisualNet
4.2.5. Fine-Tuning and Parameter Optimization of All Model
4.3. Explainable AI
4.3.1. LIME
4.3.2. Grad-CAM, Grad-CAM++, Score CAM and Smooth Grad Visualizations
4.3.3. Implications of Applying Explainable AI
4.4. Cross Validation
4.5. Comparison of Performance Before and After Preprocessing
4.6. Comparison of SkinVisualNet with Related Lyme Disease Detection Studies
5. Discussion
5.1. Result Analysis of the Models
5.2. Potentiality of Comparison with Transformer Models
5.3. Constraints of Using Single Source Dataset
5.4. Effectiveness of Pre-Processing & Post-Processing Steps
5.5. Computational Complexity and Deployment Considerations
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Dong, Y.; Zhou, G.; Cao, W.; Xu, X.; Zhang, Y.; Ji, Z.; Yang, J.; Chen, J.; Liu, M.; Fan, Y.; et al. Global seroprevalence and sociodemographic characteristics of Borrelia burgdorferi sensu lato in human populations: A systematic review and meta-analysis. BMJ Glob. Health 2022, 7, e007744. [Google Scholar] [CrossRef]
- Lantos, P.M.; Rumbaugh, J.; Bockenstedt, L.K.; Falck-Ytter, Y.T.; Aguero-Rosenfeld, M.E.; Auwaerter, P.G.; Baldwin, K.; Bannuru, R.R.; Belani, K.K.; Bowie, W.R.; et al. Clinical practice guidelines by the infectious diseases society of America (IDSA), American Academy of Neurology (AAN), and American College of Rheumatology (ACR): 2020 Guidelines for the prevention, diagnosis and treatment of lyme disease. Clin. Infect. Dis. 2020, 72, e1–e48. [Google Scholar] [CrossRef]
- Bobe, J.R.; Jutras, B.L.; Horn, E.J.; Embers, M.E.; Bailey, A.; Moritz, R.L.; Zhang, Y.; Soloski, M.J.; Ostfeld, R.S.; Marconi, R.T.; et al. Recent Progress in Lyme Disease and Remaining Challenges. Front. Med. 2021, 8, 666554. [Google Scholar] [CrossRef]
- Vinayaraj, E.V.; Gupta, N.; Sreenath, K.; Thakur, C.K.; Gulati, S.; Anand, V.; Tripathi, M.; Bhatia, R.; Vibha, D.; Dash, D.; et al. Clinical and laboratory evidence of Lyme disease in North India, 2016–2019. Travel Med. Infect. Dis. 2021, 43, 102134. [Google Scholar] [CrossRef]
- American Academy of Dermatology Association. Signs of Lyme Disease. Available online: https://www.aad.org/public/diseases/a-z/lyme-disease-signs (accessed on 5 March 2024).
- Shandilya, G.; Anand, V. Optimizing Lyme Bacterial Disease Identification with Fine-Tuned EfficientnetB0 Model. In Proceedings of the 2024 4th International Conference on Technological Advancements in Computational Sciences (ICTACS), Tashkent, Uzbekistan, 13–15 November 2024; pp. 682–686. [Google Scholar] [CrossRef]
- Burlina, P.M.; Joshi, N.J.; Ng, E.; Billings, S.D.; Rebman, A.W.; Aucott, J.N. Automated detection of erythema migrans and other confounding skin lesions via deep learning. Comput. Biol. Med. 2019, 105, 151–156. [Google Scholar] [CrossRef]
- Mohan, J.; Sivasubramanian, A.; V., S.; Ravi, V. Enhancing skin disease classification leveraging transformer-based deep learning architectures and explainable AI. Comput. Biol. Med. 2025, 190, 110007. [Google Scholar] [CrossRef]
- A, R.S.; Chamola, V.; Hussain, Z.; Albalwy, F.; Hussain, A. A novel end-to-end deep convolutional neural network based skin lesion classification framework. Expert Syst. Appl. 2024, 246, 123056. [Google Scholar] [CrossRef]
- Jerrish, D.J.; Nankar, O.; Gite, S.; Patil, S.; Kotecha, K.; Selvachandran, G.; Abraham, A. Deep learning approaches for lyme disease detection: Leveraging progressive resizing and self-supervised learning models. Multimedia Tools Appl. 2023, 83, 21281–21318. [Google Scholar] [CrossRef]
- Priyan, S.V.; Dhanasekaran, S.; Karthick, P.V.; Silambarasan, D. A new deep neuro-fuzzy system for Lyme disease detection and classification using UNet, Inception, and XGBoost model from medical images. Neural Comput. Appl. 2024, 36, 9361–9374. [Google Scholar] [CrossRef]
- Hossain, S.I.; de Herve, J.D.G.; Hassan, M.S.; Martineau, D.; Petrosyan, E.; Corbin, V.; Beytout, J.; Lebert, I.; Durand, J.; Carravieri, I.; et al. Exploring convolutional neural networks with transfer learning for diagnosing Lyme disease from skin lesion images. Comput. Methods Programs Biomed. 2022, 215, 106624. [Google Scholar] [CrossRef] [PubMed]
- Radtke, F.A.; Ramadoss, N.; Garro, A.; Bennett, J.E.; Levas, M.N.; Robinson, W.H.; Nigrovic, P.A.; Nigrovic, L.E. Serologic response to Borrelia antigens varies with clinical phenotype in children and young adults with Lyme disease. J. Clin. Microbiol. 2021, 59, e0134421. [Google Scholar] [CrossRef] [PubMed]
- Dipakkumar, S.V.; Shah, V.D.; Shah, D.; Deshmukh, S. Automated Detection of Lyme Disease using Transfer Learning Techniques. In Proceedings of the 2024 3rd International Conference on Applied Artificial Intelligence and Computing (ICAAIC), Salem, India, 5–7 June 2024; pp. 329–333. [Google Scholar] [CrossRef]
- Saravanan, M.; Shakti, S. Performance Enhancement to Improve Accuracy for the Identification of Lyme Disease by using Novel ANN Algorithm by Comparing with K-Mean Algorithm. In Proceedings of the 2023 6th International Conference on Recent Trends in Advance Computing (ICRTAC), Chennai, India, 14–15 December 2023; pp. 690–695. [Google Scholar] [CrossRef]
- Shikamaru, S. “Lyme Disease Rashes Dataset,” Kaggle, 2023. Available online: https://www.kaggle.com/datasets/sshikamaru/lyme-disease-rashes (accessed on 5 January 2024).
- Rahman, S.; Rahman, M.M.; Abdullah-Al-Wadud, M.; Al-Quaderi, G.D.; Shoyaib, M. An adaptive gamma correction for image enhancement. Eurasip J. Image Video Process. 2016, 2016, 35. [Google Scholar] [CrossRef]
- Maragatham, G.; Roomi, S.M.M. A Review of Image Contrast Enhancement Methods and Techniques. Res. J. Appl. Sci. Eng. Technol. 2015, 9, 309–326. [Google Scholar] [CrossRef]
- Shorten, C.; Khoshgoftaar, T.M. A survey on Image Data Augmentation for Deep Learning. J. Big Data 2019, 6, 60. [Google Scholar] [CrossRef]
- Świętokrzyska, P.; Wojcieszak, Ł. Porozumienie regulujące status Morza Kaspijskiego-geneza i znaczenie. [The agreement regulating the Caspian Sea status-genesis and meaning]. Nowa Polityka Wschod. 2019, 20, 54–70. [Google Scholar] [CrossRef]
- Wang, Z.; Bovik, A.C.; Sheikh, H.R.; Simoncelli, E.P. Image quality assessment: From error visibility to structural similarity. IEEE Trans. Image Process. 2004, 13, 600–612. [Google Scholar] [CrossRef]
- Sara, U.; Akter, M.; Uddin, M.S. Image Quality Assessment through FSIM, SSIM, MSE and PSNR—A Comparative Study. J. Comput. Commun. 2019, 7, 8–18. [Google Scholar] [CrossRef]
- Rahman, M.M.; Uzzaman, A.; Sami, S.I.; Khatun, F. Developing a Deep Neural Network-Based Encoder-Decoder Framework in Automatic Image Captioning Systems. 2022. Available online: https://www.researchsquare.com/article/rs-2046359/v1 (accessed on 26 November 2025).
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar] [CrossRef]
- Vijay, C.; Aayush, J.; Neha, G.; Dilbag, S.; Manjit, K. Classification of the COVID-19 infected patients using DenseNet201 based deep transfer learning. J. Biomol. Struct. Dyn. 2020, 39, 5682–5689. [Google Scholar] [CrossRef]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 22–25 July 2017. [Google Scholar]
- Kvak, D. Leveraging Computer Vision Application in Visual Arts: A Case Study on the Use of Residual Neural Network to Classify and Analyze Baroque Paintings. arXiv 2022, arXiv:2210.15300. [Google Scholar] [CrossRef]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. arXiv 2015, arXiv:1512.03385. [Google Scholar] [CrossRef]
- Ioffe, S.; Szegedy, C. Batch normalization: Accelerating deep network training by reducing internal covariate shift. In Proceedings of the 32nd International Conference on International Conference on Machine Learning (ICML), Lille, France, 6–11 July 2015. [Google Scholar]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A simple way to prevent neural networks from overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. In Proceedings of the International Conference on Learning Representations (ICLR), Banff, AB, Canada, 14–16 April 2014. [Google Scholar]
- Banerjee, K.; C, V.P.; Gupta, R.R.; Vyas, K.; H, A.; Mishra, B. Exploring Alternatives to Softmax Function. arXiv 2021, arXiv:2011.11538. [Google Scholar] [CrossRef]
- Tejaswi, K.; Zhang, E. Using Deep Learning in Lyme Disease Diagnosis. J. Stud. Res. 2021, 10. [Google Scholar] [CrossRef]
- Micikevicius, P.; Narang, S.; Alben, J.; Diamos, G.F.; Elsen, E.; García, D.; Ginsburg, B.; Houston, M.; Kuchaiev, O.; Venkatesh, G.; et al. Mixed Precision training. arXiv 2017, arXiv:1710.03740. [Google Scholar] [CrossRef]
- Barron, J.T. A General and Adaptive Robust Loss Function. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Long Beach, CA, USA, 15–20 June 2019. [Google Scholar] [CrossRef]
- Laison, E.K.E.; Ibrahim, M.H.; Boligarla, S.; Li, J.; Mahadevan, R.; Ng, A.; Muthuramalingam, V.; Lee, W.Y.; Yin, Y.; Nasri, B.R. Identifying potential Lyme disease cases using self-reported worldwide Tweets: Deep learning modeling approach enhanced with sentimental words through emojis. J. Med. Internet Res. 2023, 25, e47014. [Google Scholar] [CrossRef]
- Dosovitskiy, A.; Beyer, L.; Kolesnikov, A.; Weissenborn, D.; Zhai, X.; Unterthiner, T.; Dehghani, M.; Minderer, M.; Heigold, G.; Gelly, S.; et al. An Image is Worth 16 × 16 Words: Transformers for Image Recognition at Scale. arXiv 2020, arXiv:2010.11929. [Google Scholar]
- Han, K.; Wang, Y.; Chen, H.; Chen, X.; Guo, J.; Liu, Z.; Tang, Y.; Xiao, A.; Xu, C.; Xu, Y.; et al. A survey on vision transformer. IEEE Trans. Pattern Anal. Mach. Intell. 2022, 45, 87–110. [Google Scholar] [CrossRef]
- Marafioti, A.; Zohar, O.; Farré, M.; Noyan, M.; Bakouch, E.; Cuenca, P.; Zakka, C.; Allal, L.B.; Lozhkov, A.; Tazi, N.; et al. Smolvlm: Redefining small and efficient multimodal models. arXiv 2025, arXiv:2504.05299. [Google Scholar] [CrossRef]











| Author & Year | Dataset | Model Used | Limitations |
|---|---|---|---|
| G. Shandilya and V. Anand (2024) [6] | Skin Disease Dataset 2525 images publicly available to google | EfficientNetB0 |
|
| Philippe et al. (2019) [7] | Lyme Disease 116 images taken of 63 research participants from the Mid-Atlantic region | Deep learning |
|
| Mohan et al. (2025) [8] | Different Skin Disease Dataset | Dino V2 |
|
| Razia et al. (2024) [9] | HAM10000 dataset, which consists of 10,000 dermatoscopic images from diverse populations | S-MobileNet |
|
| Jerrish et al. (2023) [10] | Lyme Disease Rashes Dataset, containing: Original: 359 images (151 Lyme-positive, 208 Lyme-negative) | DenseNet121 with Progressive Resizing |
|
| Priyan et al. (2024) [11] | The study relies on a Kaggle dataset with significant class imbalance (4118 non-Lyme vs. 941 Lyme images in training) | Deep Neuro-Fuzzy System (DNFS) with UNet, InceptionV3, XGBoost, and Mayfly Optimization (MO) |
|
| Hossain et al. (2022) [12] | Lyme Disease | ResNet50V2 |
|
| Radtke et al. (2021) [13] | Lyme Disease (527 Children) | LASSO |
|
| S. V. Dipakkumar et al. (2024) [14] | Erythema Migrans (EM) rashes (the “bull’s-eye rash” characteristic of Lyme disease) | Convolutional Neural Network (CNN) |
|
| Saravanan et al. (2023) [15] | Lyme Disease Image Dataset (889 images) sourced from Kaggle | Artificial Neural Network (ANN) |
|
| Image | SSIM | PSNR | MSE | RMSE |
|---|---|---|---|---|
 | 0.846624 | 21.174823 | 0.007630 | 0.087349 |
 | 0.931588 | 17.612324 | 0.017329 | 0.131639 |
 | 0.950745 | 21.689516 | 0.006777 | 0.082324 |
 | 0.823486 | 20.381293 | 0.009159 | 0.095705 |
 | 0.959904 | 23.746098 | 0.004221 | 0.064967 |
 | 0.938167 | 21.302143 | 0.007409 | 0.086078 |
 | 0.828316 | 20.555193 | 0.008800 | 0.093808 |
 | 0.899446 | 19.873140 | 0.010296 | 0.101471 |
 | 0.759238 | 20.984519 | 0.007972 | 0.089284 |
 | 0.909006 | 19.714105 | 0.010680 | 0.103346 |
| Layer | Type | Trainable Parameters | Input Shape | Output Shape | Description |
|---|---|---|---|---|---|
| Input | Image Tensor | 0 | (224, 224, 3) | (224, 224, 3) | Input image of size 224 × 224 with 3 color channels (RGB). |
| DenseNet201 | ConvNet | 18,328,256 | (224, 224, 3) | (None, 7, 7, 1920) | Pre-trained DenseNet201 model without the top layer. |
| DenseNet201 GAP | GlobalAveragePooling2D | 0 | (None, 7, 7, 1920) | (None, 1920) | Global Average Pooling layer applied to DenseNet201 model’s convolutional base. |
| VGG19 | ConvNet | 20,024,384 | (224, 224, 3) | (None, 7, 7, 512) | Pre-trained VGG19 model without the top layer. |
| VGG19 GAP | GlobalAveragePooling2D | 0 | (None, 7, 7, 512) | (None, 512) | Global Average Pooling layer applied to VGG19 model’s convolutional base. |
| Concatenate | Concatenation | 0 | (None, 1920), (None, 512) | (None, 2432) | Concatenation of DenseNet201 and VGG19 outputs. |
| Batch Normalization 1 | BatchNormalization | 4864 | (None, 2432) | (None, 2432) | Batch Normalization applied after concatenation. |
| Dense 1 | Dense | 1,245,184 | (None, 2432) | (None, 512) | Dense layer with 512 units, applying L2 regularization. |
| Batch Normalization 2 | BatchNormalization | 2048 | (None, 512) | (None, 512) | Batch Normalization applied after the first dense layer. |
| Activation 1 | Activation | 0 | (None, 512) | (None, 512) | ReLU activation applied to the normalized dense layer output. |
| Dropout 1 | Dropout | 0 | (None, 512) | (None, 512) | Dropout layer with a rate of 0.5 applied to the activated dense layer output. |
| Dense 2 | Dense | 131,328 | (None, 512) | (None, 256) | Dense layer with 256 units, applying L2 regularization. |
| Batch Normalization 3 | BatchNormalization | 1024 | (None, 256) | (None, 256) | Batch Normalization applied after the second dense layer. |
| Activation 2 | Activation | 0 | (None, 256) | (None, 256) | ReLU activation applied to the normalized dense layer output. |
| Dropout 2 | Dropout | 0 | (None, 256) | (None, 256) | Dropout layer with a rate of 0.5 applied to the activated second dense layer output. |
| Output | Dense (Softmax) | 514 | (None, 256) | (None, 2) | Softmax output layer providing probability distribution over 2 classes. |
| Model | Accuracy | Precision | Recall | F1 Score | AUC |
|---|---|---|---|---|---|
| ResNet50V2 | 84.47% | 38.30% | 27.91% | 32.29% | 0.86 |
| DenseNet201 | 85.29% | 43.14% | 34.11% | 38.10% | 0.89 |
| SkinVisualNet | 85.49% | 58.14% | 32.26% | 41.49% | 0.90 |
| VGG19 | 86.93% | 60.00% | 4.65% | 8.33% | 0.91 |
| Model | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|
| VGG19 | 89.43% | 95.77% | 92.49% | 92.31% |
| DenseNet201 | 93.34% | 92.98% | 93.16% | 93.54% |
| RestNet50 V2 | 94.92% | 95.27% | 95.10% | 95.14% |
| SkinVisualNet | 98.45% | 99.09% | 98.77% | 98.83% |
| Model Name | Layer | Kernel Size | Activation Function | Pooling Layer | Optimizer | Regularization | Learning Rate | Epoch Time (s) |
|---|---|---|---|---|---|---|---|---|
| SkinVisualNet | DenseNet201, VGG19 | 7 × 7, 3 × 3, 1 × 1 | ReLU (Dense layers), Softmax | Global Average Pooling (GAP), None (Dense layers) | Adam | L2 (0.001), Dropout (0.5) | 0.00001 | 5704.52 |
| Concatenation Layer | - | - | - | - | ||||
| BatchNormalization | - | - | - | BatchNorm | ||||
| Dense Layer 1 | - | ReLU | - | L2 (0.001) | ||||
| BatchNormalization | - | - | - | BatchNorm | ||||
| Dropout (0.5) | - | - | - | Dropout (0.5) | ||||
| Dense Layer 2 | - | ReLU | - | L2 (0.001) | ||||
| BatchNormalization | - | - | - | BatchNorm | ||||
| Dropout (0.5) | - | - | - | Dropout (0.5) | ||||
| Output Layer | - | Softmax | - | |||||
| DenseNet201 | Conv1_x | 7 × 7 | ReLU | MaxPooling (3 × 3, stride 2) | Adam | None | 0.00001 | 1463.42 |
| Dense Block 1 | 1 × 1, 3 × 3 | ReLU | - | BatchNorm | ||||
| Dense Block 2 | 1 × 1, 3 × 3 | ReLU | - | BatchNorm | ||||
| Dense Block 3 | 1 × 1, 3 × 3 | ReLU | - | BatchNorm | ||||
| Dense Block 4 | 1 × 1, 3 × 3 | ReLU | - | BatchNorm | ||||
| Transition Layers | 1 × 1 | ReLU | AvgPooling (2 × 2, stride 2) | BatchNorm | ||||
| Dense Layers | - | ReLU, Softmax | GlobalAveragePooling2D | Dropout (if added) | ||||
| ResNet50V2 | Conv1 | 7 × 7 | ReLU | MaxPooling (3 × 3, stride 2) | Adam | None | 0.00001 | 1430.79 |
| Conv2_x Block1 | 1 × 1, 3 × 3, 1 × 1 | ReLU | - | BatchNorm | ||||
| Conv3_x Block2–4 | 1 × 1, 3 × 3, 1 × 1 | ReLU | - | BatchNorm | ||||
| Conv4_x Block1 | 1 × 1, 3 × 3, 1 × 1 | ReLU | - | BatchNorm | ||||
| Conv5_x Block2–6 | 1 × 1, 3 × 3, 1 × 1 | ReLU | - | BatchNorm | ||||
| Dense Layers | - | ReLU, Softmax | GlobalAveragePooling2D | - | ||||
| VGG19 | Conv1_x | 3 × 3 | ReLU | MaxPooling (2 × 2) | Adam | 0.00001 | 1412.92 | |
| Conv2_x | 3 × 3 | ReLU | MaxPooling (2 × 2) | |||||
| Conv3_x | 3 × 3 | ReLU | MaxPooling (2 × 2) | |||||
| Conv4_x | 3 × 3 | ReLU | MaxPooling (2 × 2) | |||||
| Conv5_x | 3 × 3 | ReLU | MaxPooling (2 × 2) | |||||
| Dense Layers | - | ReLU, Softmax | GlobalAveragePooling2D |
| Accuracy (%) | F1 | F2 | F3 | F4 | F5 |
|---|---|---|---|---|---|
| Training | 100 | 99.99 | 99.98 | 99.99 | 99.98 |
| Validation | 98.20 | 98.89 | 98.90 | 98.89 | 98.92 |
| Model | Precision | Recall | F1 Score | Accuracy | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|---|---|---|---|
| After Preprocessing | Before Preprocessing | |||||||
| VGG19 | 89.43% | 95.77% | 92.49% | 92.31% | 86.93% | 60.00% | 4.65% | 8.33% |
| DenseNet201 | 93.34% | 92.98% | 93.16% | 93.54% | 85.29% | 43.14% | 34.11% | 38.10% |
| ResNet50V2 | 94.92% | 95.27% | 95.10% | 95.14% | 84.47% | 38.30% | 27.91% | 32.29% |
| SkinVisualNet | 98.45% | 99.09% | 98.77% | 98.83% | 85.49% | 58.14% | 32.26% | 41.49% |
| Author (Year) | Dataset Population Focus | Dataset Explanation | Explainability Method | Accuracy (%) |
|---|---|---|---|---|
| Laison et al. [36] (2023) | Worldwide social media users | Self-reported Lyme-related symptom Tweets collected globally; includes textual and emoji-based sentiment features | Not applicable. (Text-based sentiment and feature-importance interpretation only) | ~90% (varies by model) |
| Jerrish et al. [10] (2023) | Clinical skin rash images | Curated Lyme rash datasets using progressive resizing and self-supervised learning | None reported (focus primarily on SSL) | 94–96% |
| Priyan et al. [11] (2024) | Clinical medical imaging patients | Medical image dataset processed using UNet, Inception, and a neuro-fuzzy classifier | Grad-CAM (typical for UNet-based pipelines) | 97.45% |
| SkinVisualNet | Public Kaggle Lyme rash dataset | Kaggle-derived Lyme disease rash images with augmentation and preprocessing | LIME, Grad-CAM, CAM++, Score-CAM, SmoothGrad | 98.83% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sohel, A.; Turjy, R.C.D.; Bappy, S.P.; Assaduzzaman, M.; Marouf, A.A.; Rokne, J.G.; Alhajj, R. SkinVisualNet: A Hybrid Deep Learning Approach Leveraging Explainable Models for Identifying Lyme Disease from Skin Rash Images. Mach. Learn. Knowl. Extr. 2025, 7, 157. https://doi.org/10.3390/make7040157
Sohel A, Turjy RCD, Bappy SP, Assaduzzaman M, Marouf AA, Rokne JG, Alhajj R. SkinVisualNet: A Hybrid Deep Learning Approach Leveraging Explainable Models for Identifying Lyme Disease from Skin Rash Images. Machine Learning and Knowledge Extraction. 2025; 7(4):157. https://doi.org/10.3390/make7040157
Chicago/Turabian StyleSohel, Amir, Rittik Chandra Das Turjy, Sarbajit Paul Bappy, Md Assaduzzaman, Ahmed Al Marouf, Jon George Rokne, and Reda Alhajj. 2025. "SkinVisualNet: A Hybrid Deep Learning Approach Leveraging Explainable Models for Identifying Lyme Disease from Skin Rash Images" Machine Learning and Knowledge Extraction 7, no. 4: 157. https://doi.org/10.3390/make7040157
APA StyleSohel, A., Turjy, R. C. D., Bappy, S. P., Assaduzzaman, M., Marouf, A. A., Rokne, J. G., & Alhajj, R. (2025). SkinVisualNet: A Hybrid Deep Learning Approach Leveraging Explainable Models for Identifying Lyme Disease from Skin Rash Images. Machine Learning and Knowledge Extraction, 7(4), 157. https://doi.org/10.3390/make7040157