Machine Learning Prediction Models of Beneficial and Toxicological Effects of Zinc Oxide Nanoparticles in Rat Feed
Abstract
1. Introduction
2. Literature Review
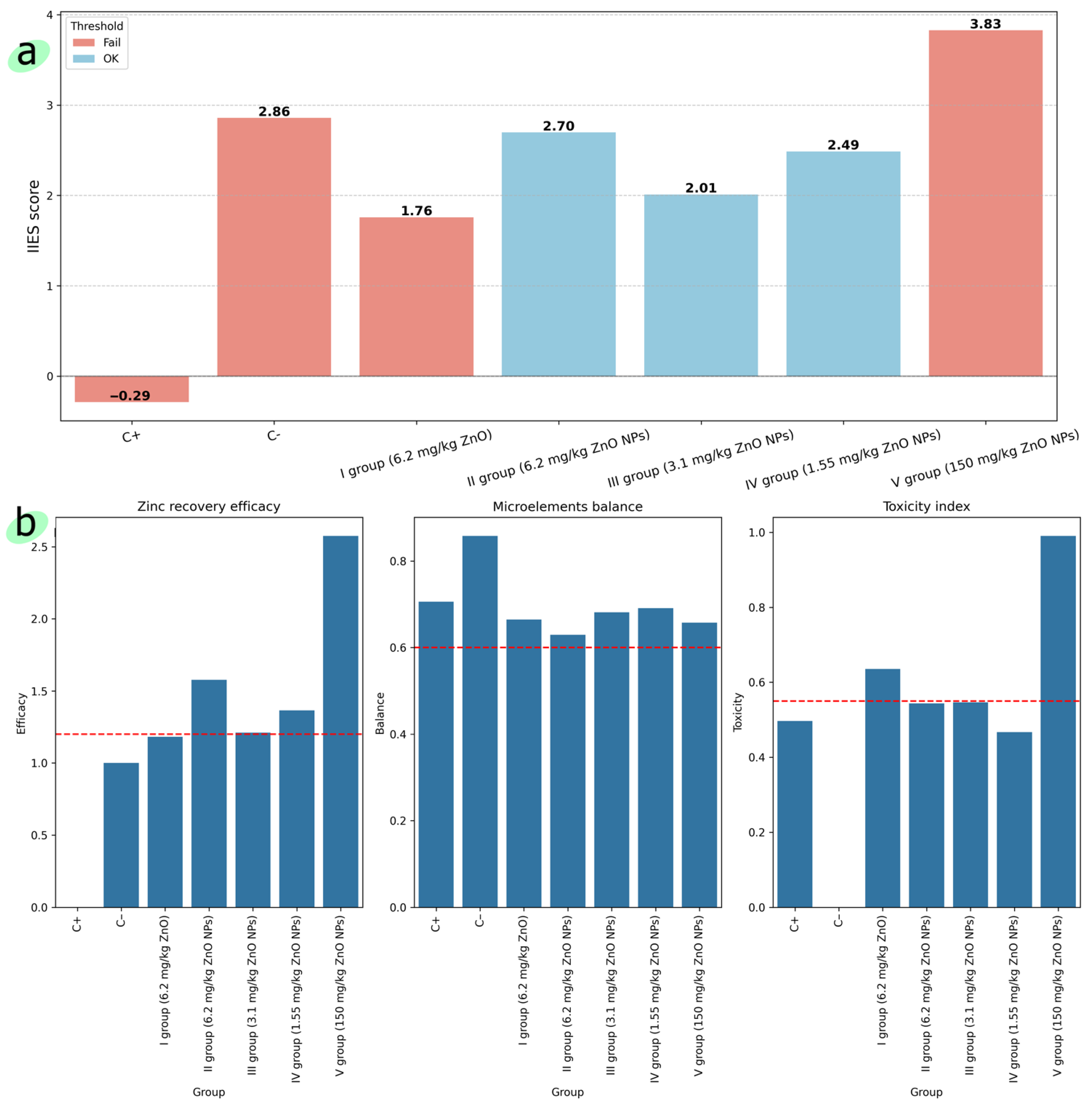
- We developed a weighted importance score based on feature importance and element groups, and an integral indicator of efficiency and safety to assess the optimal dosage of zinc oxide nanoparticles in rat feed;
- We implemented a fully connected feedforward artificial neural network and kernel ridge regression model with a new custom loss function;
- We studied the possibility of generating reliable synthetic data for the problem of low-dimensional experimental data.
- We studied the effect of zinc oxide nanoparticles in feed on elemental homeostasis in rats;
- We determined the optimal dosage of zinc oxide nanoparticles in terms of beneficial and toxicological effects;
- We built predictions of the levels of essential and toxic elements, proteins, and enzymes, depending on the concentration of zinc oxide nanoparticles in the range from 1 mg/kg to 150 mg/kg.
3. Materials and Methods
3.1. Data Preprocessing
- Essential microelements: Fe, Zn, Cu, Mn, Co, Se, I, Cr;
- Toxic microelements: Sn, Al, As, Hg, Pb, Be, Cd;
- Macroelements: Na, Ca, P, K, Mg;
- Partially essential elements: B, Si, V, Ni, Li.
- EH_1—original data on element content;
- EH_2—original data on proteins and enzymes content;
- EH_Synt_1—synthetic data on element content;
- EH_Synt_2—synthetic data on proteins and enzymes content.
- 1. Zinc recovery efficacy (relative to the norm), defined as follows:
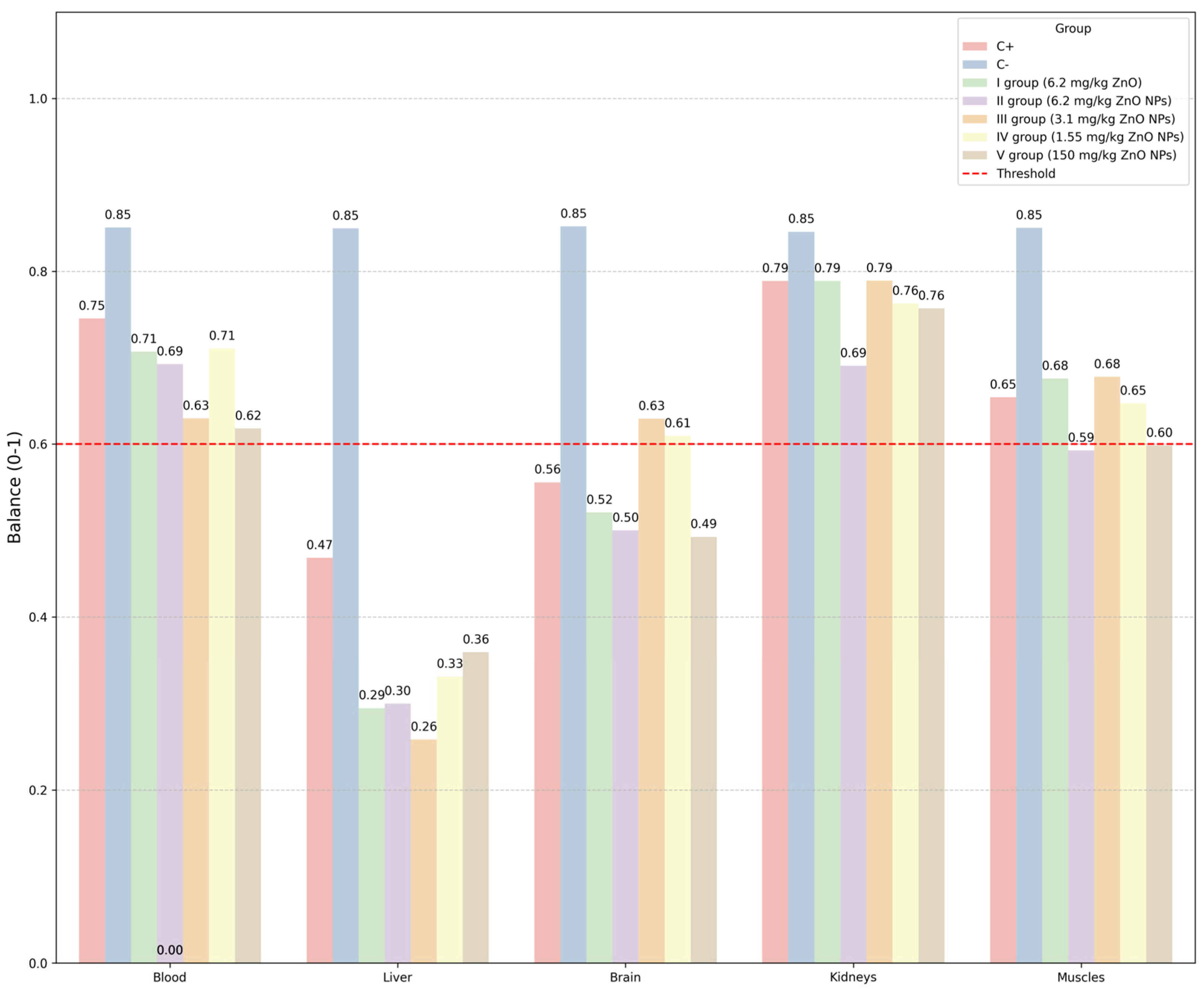
- 2. Balance of elemental composition (average deviation from the norm), defined as follows:where L is the number of elements, is the average level of the l-th element in group d, and denote the average level and standard deviation of the l-th element in the negative control group C−.
- 3. Toxicity (deviation of toxic elements) is defined as follows:where P is the number of toxic microelements, is the average level of the p-th toxic element in group d, and and are the average level and standard deviation of the p-th toxic element in the negative control group C−.
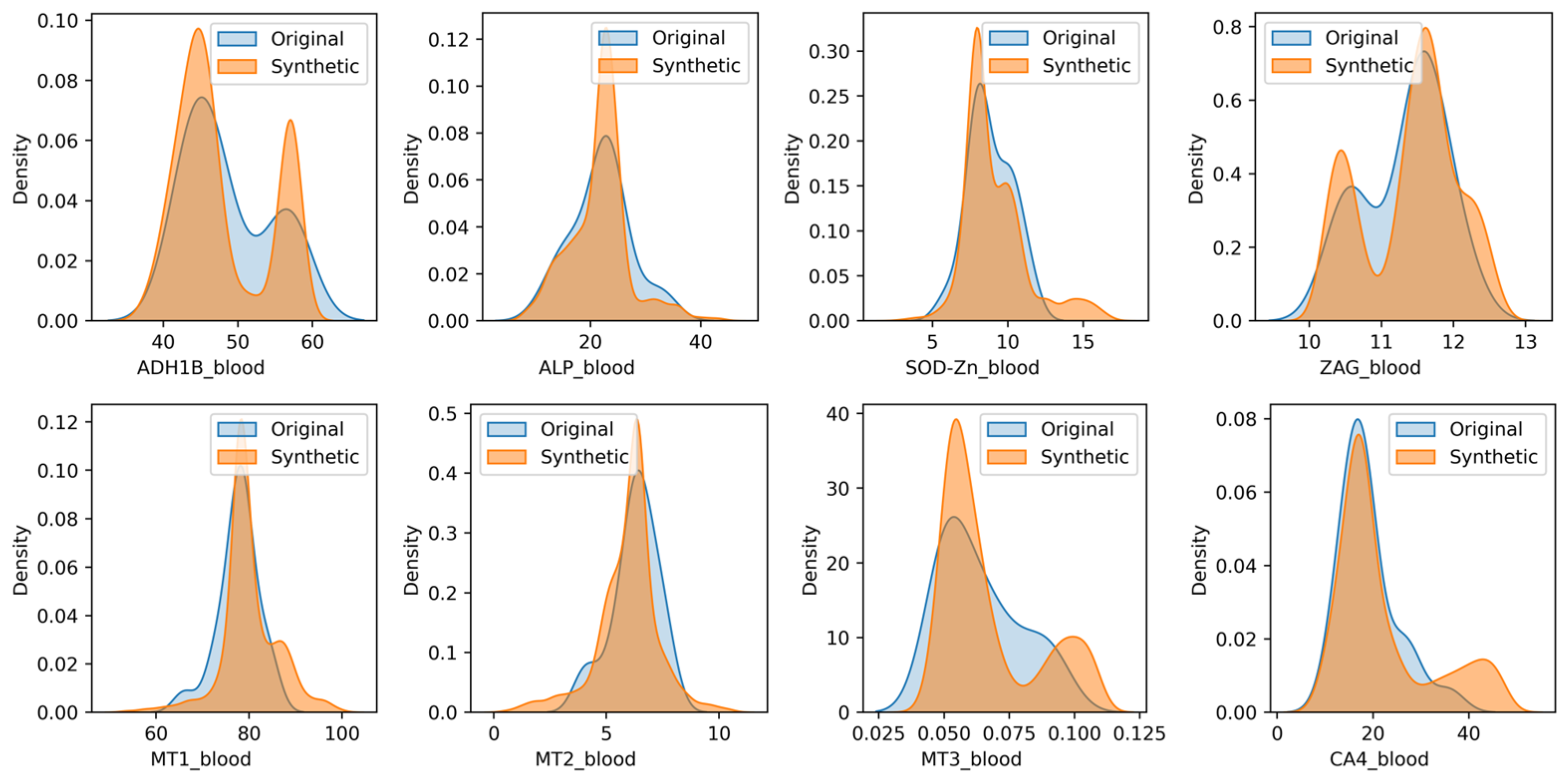
3.2. Synthetic Data Generation and Evaluation
3.3. Non-Linear Regression Models
4. Results
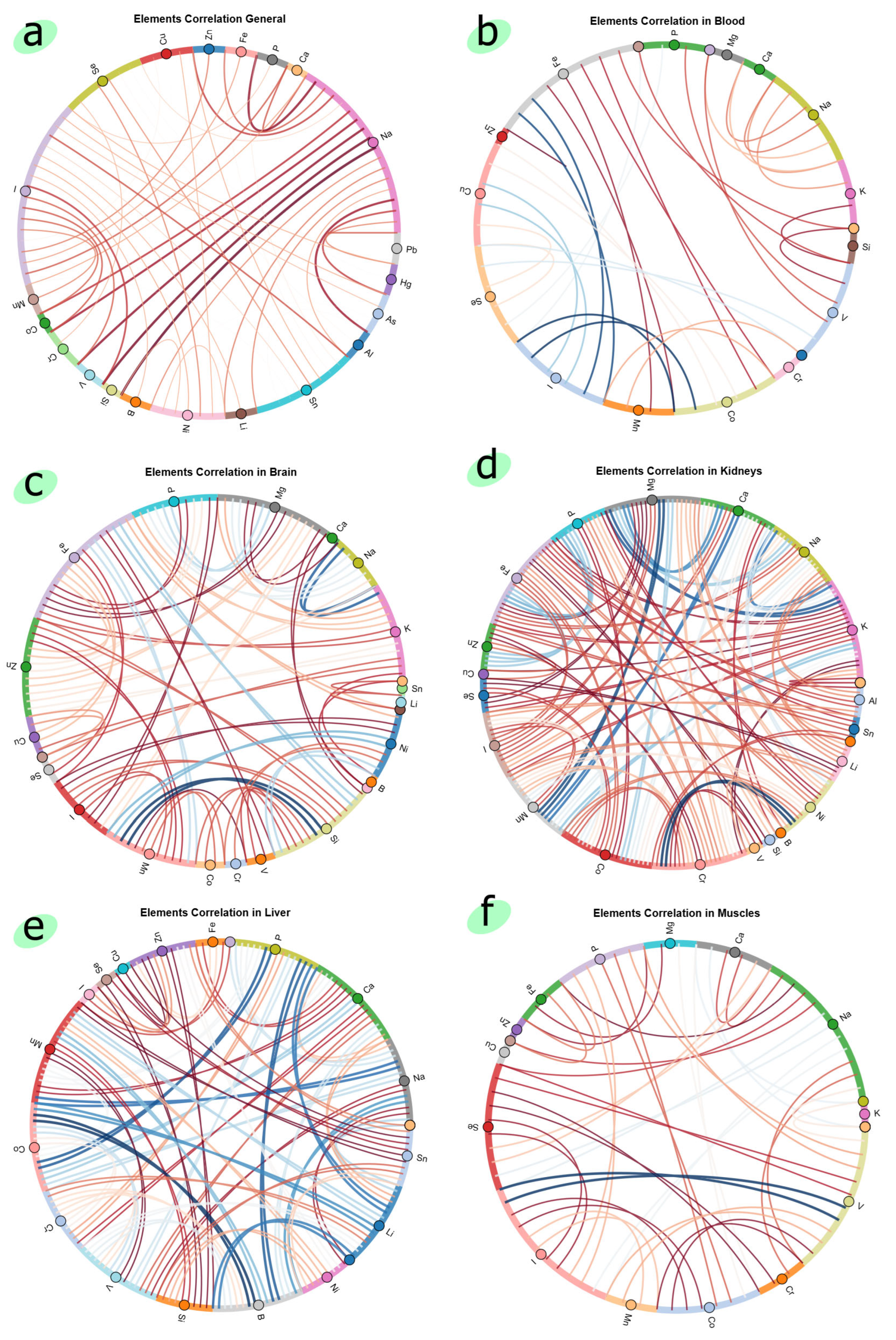
4.1. Data Analysis of Elemental Homeostasis of Wistar Rats
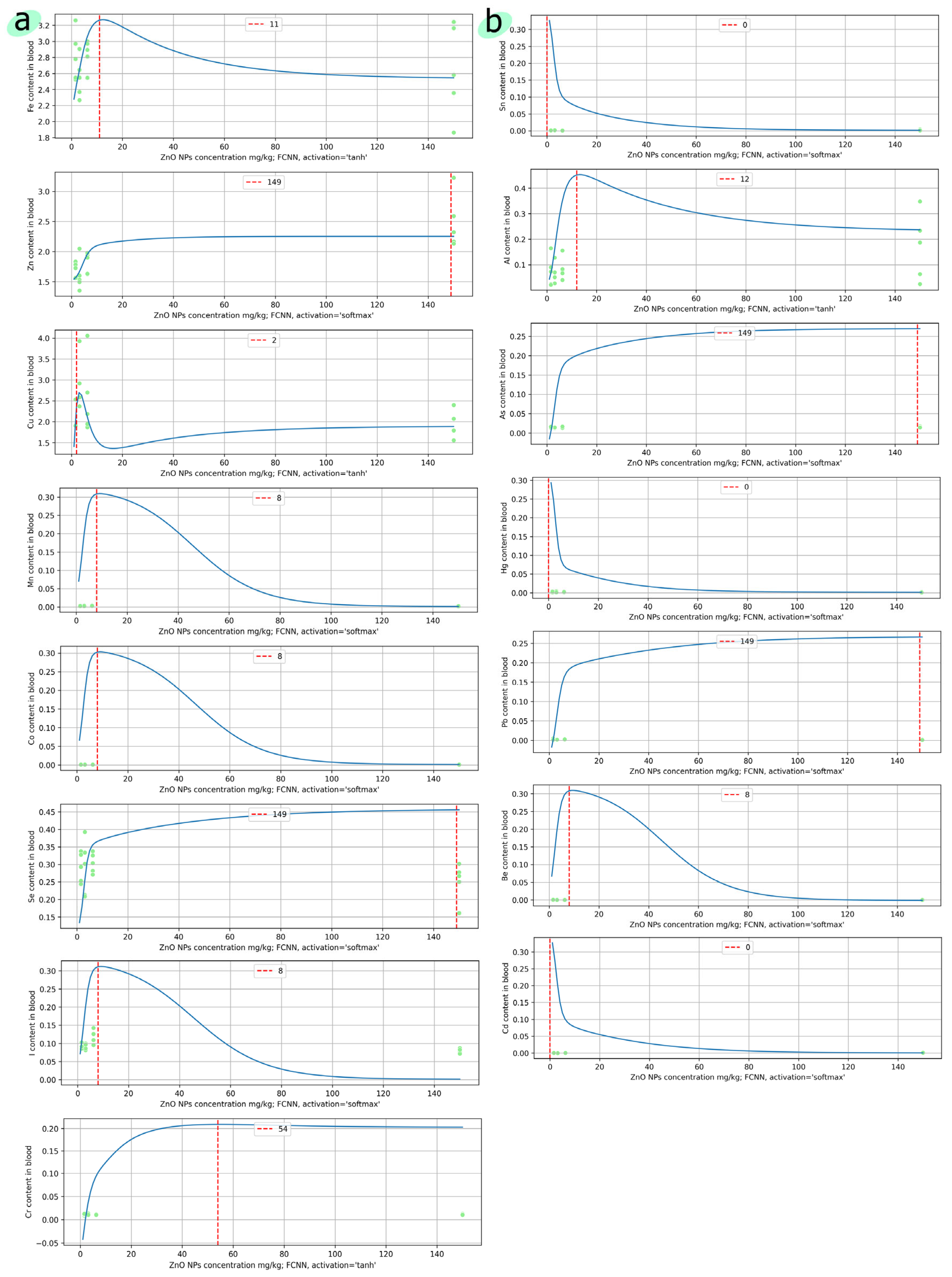
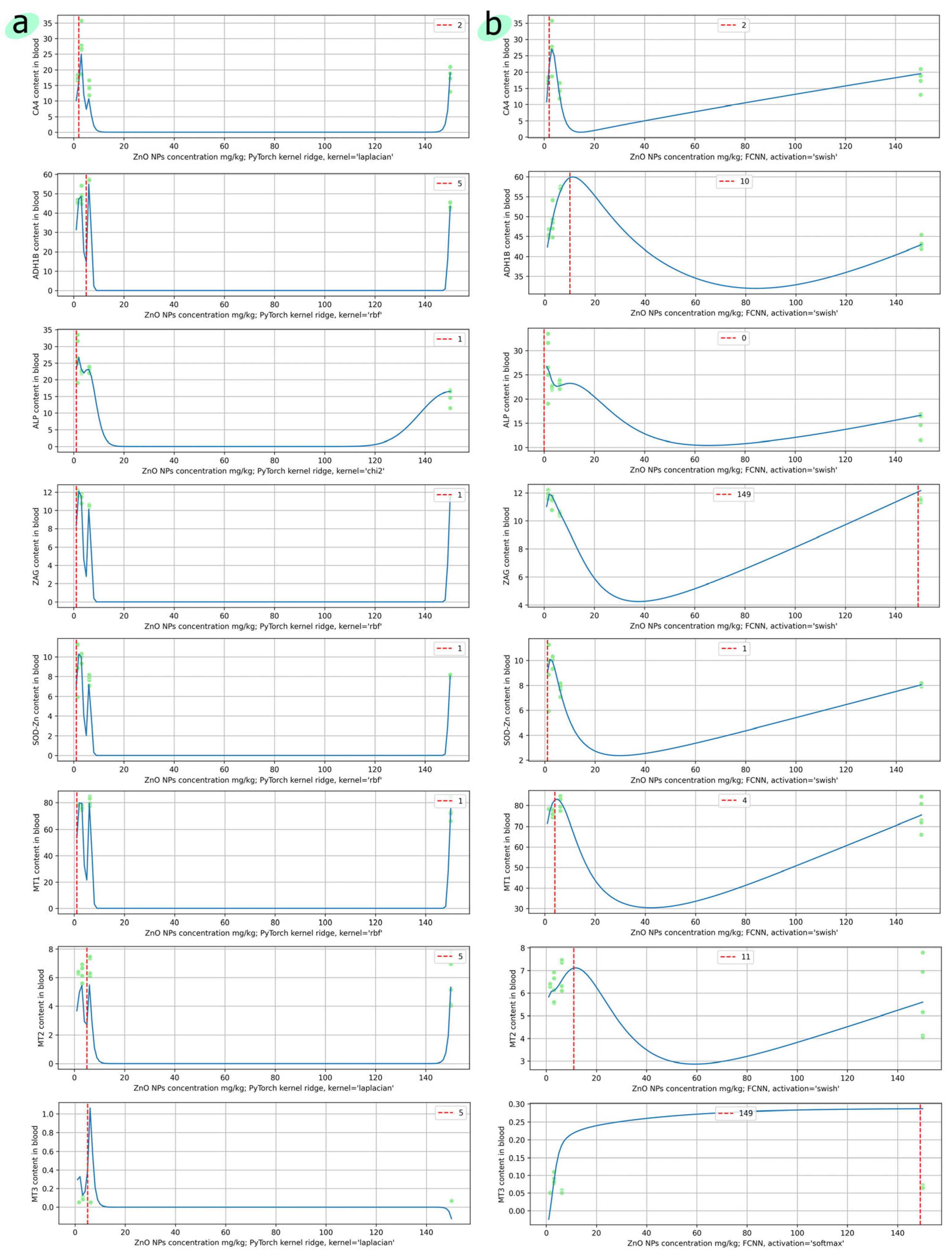
4.2. Predictive Models of Toxic and Essential Elements in the Blood of Laboratory Animals Based on the Dosage of Zinc Oxide Nanoparticles
5. Discussion
6. Conclusions
- We rigorously evaluated the similarity between the original and synthetic datasets using the SDMetrics tool and KDE plots;
- During training, we incorporated domain-informed constraints to prevent the generation of physiologically implausible values with a custom loss function;
- The utility of the augmented dataset was assessed through the performance of two predictive models (FCNN and Kernel Ridge).
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ADASYN | Adaptive Synthetic |
| ALP | Alkaline Phosphatase |
| ALT | Alanine Transaminase |
| ANN | Artificial Neural Network |
| AST | Aspartate Aminotransferase |
| CGW | Correlation Graph Weight |
| CTGAN | Conditional Tabular Generative Adversarial Network |
| EH | Elemental Homeostasis |
| FCNN | Fully Connected Neural Network |
| IIES | Integral Indicator of Efficiency and Safety |
| IQR | Interquartile Range |
| KDE | Kernel Density Estimate |
| LDH | Lactate Dehydrogenase |
| MAE | Mean Absolute Error |
| ML | Machine Learning |
| MLP | Multi-layer Perceptron |
| MSE | Mean Squared Error |
| NPs | Nanoparticle(s) |
| RBF | Radial Basis Function |
| RMSE | Root Mean Squared Error |
| SMOTE | Synthetic Minority Oversampling Technique |
References
- Huang, S.; Wang, L.; Liu, L.; Hou, Y.; Li, L. Nanotechnology in agriculture, livestock, and aquaculture in China. A review. Agron. Sustain. Dev. 2015, 35, 369–400. [Google Scholar] [CrossRef]
- Bai, D.P.; Lin, X.Y.; Huang, Y.F.; Zhang, X.F. Theranostics aspects of various nanoparticles in veterinary medicine. Int. J. Mol. Sci. 2018, 19, 3299. [Google Scholar] [CrossRef]
- Gelaye, Y. Application of nanotechnology in animal nutrition: Bibliographic review. Cogent Food Agric. 2024, 10, 2290308. [Google Scholar] [CrossRef]
- Kolesnikov, S.; Timoshenko, A.; Minnikova, T.; Tsepina, N.; Kazeev, K.; Akimenko, Y.; Zhadobin, A.; Shuvaeva, V.; Rajput, V.D.; Mandzhieva, S.; et al. Impact of Metal-Based Nanoparticles on Cambisol Microbial Functionality, Enzyme Activity, and Plant Growth. Plants 2021, 10, 2080. [Google Scholar] [CrossRef]
- Paramo, L.A.; Feregrino-Pérez, A.A.; Guevara, R.; Mendoza, S.; Esquivel, K. Nanoparticles in agroindustry: Applications, toxicity, challenges, and trends. Nanomaterials 2020, 10, 1654. [Google Scholar] [CrossRef]
- Singh, A.V.; Ansari, M.H.D.; Rosenkranz, D.; Maharjan, R.S.; Kriegel, F.L.; Gandhi, K.; Kanase, A.; Singh, R.; Laux, P.; Luch, A. Artificial intelligence and machine learning in computational nanotoxicology: Unlocking and empowering nanomedicine. Adv. Healthc. Mater. 2020, 9, 1901862. [Google Scholar] [CrossRef] [PubMed]
- Saeed, S.; Afzal, G.; Kiran, S.; Ahmad, H.I.; Haider, M.Z.; Naz, S. Role of Nanoparticles in Veterinary Medicine and as Feed Additive in Livestock. In Recent Advances in Biotechnology; Bentham Science Publishers: Sharjah, United Arab Emirates, 2023; pp. 229–242. [Google Scholar] [CrossRef]
- Naumenko, S.; Koshevoy, V.; Matsenko, O.; Miroshnikova, O.; Zhukova, I.; Bespalova, I. Antioxidant properties and toxic risks of using metal nanoparticles on health and productivity in poultry. J. World’s Poult. Res. 2023, 13, 292–306. [Google Scholar] [CrossRef]
- Rahman, H.S.; Othman, H.H.; Abdullah, R.; Edin, H.Y.A.S.; AL-Haj, N.A. Beneficial and toxicological aspects of zinc oxide nanoparticles in animals. Vet. Med. Sci. 2022, 8, 1769–1779. [Google Scholar] [CrossRef]
- Bąkowski, M.; Kiczorowska, B.; Samolińska, W.; Klebaniuk, R.; Lipiec, A. Silver and zinc nanoparticles in animal nutrition—A review. Ann. Anim. Sci. 2018, 18, 879–898. [Google Scholar] [CrossRef]
- Tsekhmistrenko, O.; Bityutskyy, V.; Tsekhmistrenko, S.; Melnychenko, O.; Tymoshok, N.; Spivak, M. Use of nanoparticles of metals and nonmetals in poultry farming. Technol. Prod. Process. Livest. Prod. 2019, 2, 113–130. [Google Scholar] [CrossRef]
- Hardy, A.; Benford, D.; Halldorsson, T.; Jeger, M.J.; Knutsen, H.K.; More, S.; Naegeli, H.; Noteborn, H.; Ockleford, C.; EFSA Scientific Committee; et al. Guidance on risk assessment of the application of nanoscience and nanotechnologies in the food and feed chain: Part 1, human and animal health. EFSA J. 2018, 16, 05327. [Google Scholar] [CrossRef] [PubMed]
- Elalfy, M.M.; El-hadidy, M.G.; Abouelmagd, M.M. Update of usefulness and adverse effects of nanoparticles on animals and human health. J. Vet. Med. Health 2018, 2, 106. [Google Scholar]
- Lutkovskaya, Y.; Sizova, E.; Kamirova, A. Ultrafine forms of trace elements in the diet of ruminants: Impact on productivity and health. Agrar. Sci. J. 2024, 5, 96–104. [Google Scholar] [CrossRef]
- Vanivska, K.; Dianová, L.; Halo, M., Jr.; Štefunková, N.; Lenický, M.; Slanina, T.; Tirpák, F.; Ivanič, P.; Stawarz, R.; Massányi, P. Toxicity of nanoparticles on animal and human organism: Cell response. J. Microbiol. Biotechnol. Food Sci. 2024, 14, e10844. [Google Scholar] [CrossRef]
- Shivaswamy, M.S.; Yashkamal, K.; Shivakumar, M.S. In vivo and in vitro toxicity of nanomaterials in animal systems. In Nanotoxicology for Agricultural and Environmental Applications; Academic Press: Cambridge, MA, USA, 2024; pp. 159–169. [Google Scholar] [CrossRef]
- Sharma, N.; Kurmi, B.D.; Singh, D.; Mehan, S.; Khanna, K.; Karwasra, R.; Kumar, S.; Chaudhary, A.; Jakhmola, V.; Sharma, A.; et al. Nanoparticles toxicity: An overview of its mechanism and plausible mitigation strategies. J. Drug Target. 2024, 32, 457–469. [Google Scholar] [CrossRef]
- Pradhan, K.; Mishra, L.; Mishra, M. Nanotoxicology and Its Remediation. In Smart Nanomaterials for Infectious Diseases; Book Sales Department, Royal Society of Chemistry, Thomas Graham House: Cambridge, UK, 2024. [Google Scholar] [CrossRef]
- Parashar, S.; Raj, S.; Srivastava, P.; Singh, A.K. Comparative toxicity assessment of selected nanoparticles using different experimental model organisms. J. Pharmacol. Toxicol. Methods 2024, 130, 107563. [Google Scholar] [CrossRef]
- Bisla, A.; Honparkhe, M.; Srivastava, N. A review on applications and toxicities of metallic nanoparticles in mammalian semen biology. Andrologia 2022, 54, e14589. [Google Scholar] [CrossRef]
- Dianová, L.; Tirpák, F.; Halo, M.; Slanina, T.; Massányi, M.; Stawarz, R.; Formicki, G.; Madeddu, R.; Massányi, P. Effects of selected metal nanoparticles (Ag, ZnO, TiO2) on the structure and function of reproductive organs. Toxics 2022, 10, 459. [Google Scholar] [CrossRef]
- Sial, B.E.; Ali, S.A.; Aslam, N.; Maqsood, R.; Iqbal, S.; Mehmood, Y.; Mustafa, G. ZnO nanoparticles impact on organ systems in rats: A comprehensive exploration of diverse exposure pathways. J. Zool. Syst. 2023, 2023, 37–51. [Google Scholar] [CrossRef]
- Sharif, S.; Fatima, R.; Naz, S. Toxic Effects of Metallic Nanoparticles on Rat’s Spleen; a Review. Pol. J. Environ. Stud. 2023, 32, 991–997. [Google Scholar] [CrossRef] [PubMed]
- Shafiq, A.A.; Luaib, N.M.; Mahdi, R.H.; Alsaimary, I.E. Oral Toxicity of Magnesium Oxide Nanoparticles, MgO NPs on Liver in Male Rats. Iraqi J. Sci. 2024, 65, 6323–6335. [Google Scholar] [CrossRef]
- Verma, G.S.; Nirmal, N.K.; Gunpal, D.; Gupta, H.; Yadav, M.; Kumar, N.; John, P.J. Intraperitoneal exposure of iron oxide nanoparticles causes dose-dependent toxicity in Wistar rats. Toxicol. Ind. Health 2021, 37, 763–775. [Google Scholar] [CrossRef]
- Ali, A.; Saeed, S.; Hussain, R.; Saif, M.S.; Waqas, M.; Asghar, I.; Xue, X.; Hasan, M. Exploring the impact of silica and silica-based nanoparticles on serological parameters, histopathology, organ toxicity, and genotoxicity in Rattus norvegicus. Appl. Surf. Sci. Adv. 2024, 19, 100551. [Google Scholar] [CrossRef]
- Yousef, M.I.; Roychoudhury, S.; Jafaar, K.S.; Slama, P.; Kesari, K.K.; Kamel, M.A.E. N Aluminum oxide and zinc oxide induced nanotoxicity in rat brain, heart, and lung. Physiol. Res. 2022, 71, 677. [Google Scholar] [CrossRef]
- Moshrefi, A.H.; Hosseini, S.M.; Parvizi, M.R. Toxicological effect of exposure to different doses of zinc oxide nanoparticles on brain and heart structures of male Wistar rats. Bulg. J. Vet. Med. 2023, 26, 510–523. [Google Scholar] [CrossRef]
- Rehman, R.; Khanam, S.; Khan, Y.; Hayat, F. Toxicological Effects of Colloidal Silver Nanoparticles on Rat Health: Assessing Physiological, Hematological, Biochemical, and Behavioral Parameters. Innov. Res. Appl. Biol. Chem. Sci. 2024, 2, 101–107. [Google Scholar] [CrossRef]
- Jarrar, B.; Almansour, M.; Al-Doaiss, A.; Lee, S.Y.; Melhem, W.; Jarrar, Q.; Sewelam, A. Metallic and metallic oxide nanoparticles toxicity primarily targets the mitochondria of hepatocytes and renal cells. Toxicol. Ind. Health 2024, 40, 667–678. [Google Scholar] [CrossRef] [PubMed]
- Sutunkova, M.; Klinova, S.; Ryabova, Y.; Tazhigulova, A.; Shabardina, L.; Bateneva, V.; Shelomentsev, I.; Privalova, L. Acute Toxicity Induced by Inhalation Exposure to Lead Oxide Nanoparticles in Rats. Public Health Life Environ. PHLE 2023, 31, 24–30. (In Russian) [Google Scholar] [CrossRef]
- Rangel DaCosta, L.; Sytwu, K.; Groschner, C.K.; Scott, M.C. A robust synthetic data generation framework for machine learning in high-resolution transmission electron microscopy (HRTEM). Npj Comput. Mater. 2024, 10, 165. [Google Scholar] [CrossRef]
- Varsou, D.D.; Kolokathis, P.D.; Antoniou, M.; Sidiropoulos, N.K.; Tsoumanis, A.; Papadiamantis, A.G.; Melagraki, G.; Lynch, I.; Afantitis, A. In silico assessment of nanoparticle toxicity powered by the Enalos Cloud Platform: Integrating automated machine learning and synthetic data for enhanced nanosafety evaluation. Comput. Struct. Biotechnol. J. 2024, 25, 47–60. [Google Scholar] [CrossRef]
- Khadka, K.; Chandrasekaran, J.; Lei, Y.; Kacker, R.N.; Kuhn, D.R. Synthetic data generation using combinatorial testing and variational autoencoder. In Proceedings of the 2023 IEEE International Conference on Software Testing, Verification and Validation Workshops (ICSTW), Dublin, Ireland, 16–20 April 2023; IEEE: New York, NY, USA; pp. 228–236. [Google Scholar] [CrossRef]
- Dou, B.; Zhu, Z.; Merkurjev, E.; Ke, L.; Chen, L.; Jiang, J.; Zhu, Y.; Liu, J.; Zhang, B.; Wei, G.-W. Machine learning methods for small data challenges in molecular science. Chem. Rev. 2023, 123, 8736–8780. [Google Scholar] [CrossRef] [PubMed]
- Goyal, M.; Mahmoud, Q.H. A systematic review of synthetic data generation techniques using generative AI. Electronics 2024, 13, 3509. [Google Scholar] [CrossRef]
- Reeves, P.G.; Nielsen, F.H.; Fahey, G.C., Jr. AIN-93 purified diets for laboratory rodents: Final report of the American institute of nutrition ad hoc writing committee on the reformulation of the AIN-76A rodent diet. J. Nutr. 1993, 123, 1939–1951. [Google Scholar] [CrossRef] [PubMed]
- Skalny, A.V. Bioelementology as an interdisciplinary integrative approach in life sciences: Terminology, classification, perspectives. J. Trace Elem. Med. Biol. 2011, 25, S3–S10. [Google Scholar] [CrossRef]
- Da Silva, J.F.; Williams, R.J.P. The Biological Chemistry of the Elements: The Inorganic Chemistry of Life; Oxford University Press: Oxford, UK, 2001. [Google Scholar] [CrossRef]
- Xu, L.; Skoularidou, M.; Cuesta-Infante, A.; Veeramachaneni, K. Modeling Tabular Data Using Conditional GAN. Adv. Neural Inf. Process. Syst. 2019, 32, 1–11. [Google Scholar]
- Smirnova, E. Mathematical Modeling of Adaptation to Extreme Conditions, the Effect of Group Stress and Correlation Adaptometry [Dissertation Abstract]. Krasnoyarsk; 2000. Available online: http://elibrary.ru/item.asp?id=15980174 (accessed on 9 June 2025).

| Nanoparticle Type | Effects on Animal Husbandry | Ref. |
|---|---|---|
| Zinc oxide (ZnO) | Improved growth, antioxidant properties, and enhanced immune function; potential liver and kidney damage | [8,9] |
| Silver (Ag), zinc (Zn) | Antimicrobial activity, improved productivity; potential neurotoxicity and reproductive toxicity | [10] |
| Zinc (Zn), silver (Ag), selenium (Se), cerium (Ce), iron (Fe) | Improved hemoglobin levels, enhanced productivity, potential oxidative stress, and tissue damage | [11] |
| Selenium (Se) | Enhanced antioxidant system, improved immune function; potential oxidative stress and inflammation | [14] |
| Silver (Ag), zinc oxide (ZnO), titanium dioxide (TiO2) | Improved productivity, antimicrobial activity, potential oxidative stress, and inflammation | [21] |
| Nanoparticle Type | Rats Type | Main Outcomes | Ref. |
|---|---|---|---|
| Magnesium oxide nanoparticles (MgO) | Sprague–Dawley albino rats (Rattus norvegicus) | 250 and 1000 mg/kg doses; a highly significant increase in AST, ALT, ALP, and total bilirubin | [24] |
| Iron oxide (IO) | Wistar rats | 20, 40, and 80 mg/kg doses; significant changes in lungs, liver, and kidney; LDH increased significantly | [25] |
| Crystalline silica (C-SiO2) NPs, silver-silica (Ag-SiO2), zinc-oxide silica (ZnO-SiO2) | Rattus norvegicus | 500 µg/kg doses; significantly altered 95% of serological, DNA damage, histopathological, and oxidative stress parameters | [26] |
| Aluminum oxide (Al2O3), zinc oxide (ZnO) | Male Wistar rats | 70 mg/kg of Al2O3 and 100 mg/kg of ZnO NPs doses; increase in cytokines, p53, oxidative stress, creatine kinase, norepinephrine, acetylcholine, and lipid profile; significant decrease in the levels of antioxidant enzymes, total antioxidant capacity, and the activity of acetylcholine esterase in the brain, heart, and lung | [27] |
| Zinc oxide (ZnO) | Male Wistar rats | 4, 8, 25, 50, 100, and 200 mg/kg doses; histopathologic lesions in the heart structures of 200 mg/kg; doses higher than 25 mg/kg are not recommended in terms of toxicity. | [28] |
| Colloidal silver nanoparticles (AgNPs) | Male Wistar rats | 10 μg/kg/day and 100 μg/kg/day doses; a notable reduction in body weight and exhibited potential elevation in liver enzymes for the 100 μg/kg/day dose | [29] |
| Zinc oxide (ZnO) | Male Wistar rats | 3.1 mg/kg dose is optimal in terms of balanced elemental homeostasis; good predictive results for essential elements, proteins, and enzymes in the range from 1 mg/kg to 150 mg/kg | Our research |
| Number of Records | Total Score | Column Shapes Score | Column Pair Trends Score |
|---|---|---|---|
| EH_Synt_1 data | |||
| 100 | 0.8069 | 0.7217 | 0.8922 |
| 200 | 0.8059 | 0.7213 | 0.8904 |
| 300 | 0.8132 | 0.7302 | 0.8961 |
| 500 | 0.8120 | 0.7288 | 0.8953 |
| EH_Synt_2 data | |||
| 100 | 0.8270 | 0.7729 | 0.8811 |
| 200 | 0.8285 | 0.7724 | 0.8845 |
| 300 | 0.8254 | 0.7715 | 0.8793 |
| 500 | 0.8269 | 0.7730 | 0.8807 |
| № | Experimental Group | CGW | CGW, α = 0.5 |
|---|---|---|---|
| Both EH_1 and EH_2 datasets | |||
| 1 | C+ | 457.35 | 293.23 |
| 2 | C− | 470.82 | 315.21 |
| 3 | I group (6.2 mg/kg ZnO) | 461.12 | 300.87 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 440.52 | 279.65 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 474.28 | 302.99 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 455.50 | 294.54 |
| 7 | V group (150 mg/kg ZnO NPs) | 466.35 | 299.43 |
| Blood data | |||
| 1 | C+ | 23.93 | 15.82 |
| 2 | C− | 25.18 | 17.52 |
| 3 | I group (6.2 mg/kg ZnO) | 24.18 | 15.56 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 25.04 | 16.74 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 23.40 | 15.44 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 25.44 | 17.26 |
| 7 | V group (150 mg/kg ZnO NPs) | 27.27 | 19.70 |
| Brain data | |||
| 1 | C+ | 48.77 | 34.56 |
| 2 | C− | 52.12 | 41.30 |
| 3 | I group (6.2 mg/kg ZnO) | 45.41 | 33.43 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 47.69 | 34.02 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 44.50 | 30.73 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 47.75 | 36.21 |
| 7 | V group (150 mg/kg ZnO NPs) | 51.54 | 37.71 |
| Kidney data | |||
| 1 | C+ | 14.79 | 10.47 |
| 2 | C− | 19.62 | 17.20 |
| 3 | I group (6.2 mg/kg ZnO) | 13.90 | 9.83 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 15.91 | 11.48 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 12.69 | 8.56 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 16.43 | 12.64 |
| 7 | V group (150 mg/kg ZnO NPs) | 15.53 | 11.40 |
| Liver data | |||
| 1 | C+ | 24.29 | 19.18 |
| 2 | C− | 23.76 | 17.65 |
| 3 | I group (6.2 mg/kg ZnO) | 25.25 | 18.48 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 21.33 | 14.16 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 21.19 | 15.25 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 24.55 | 18.74 |
| 7 | V group (150 mg/kg ZnO NPs) | 21.54 | 14.95 |
| Muscle data | |||
| 1 | C+ | 26.70 | 20.00 |
| 2 | C− | 13.78 | 8.70 |
| 3 | I group (6.2 mg/kg ZnO) | 14.38 | 10.93 |
| 4 | II group (6.2 mg/kg ZnO NPs) | 13.41 | 8.21 |
| 5 | III group (3.1 mg/kg ZnO NPs) | 15.37 | 10.77 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 13.43 | 8.58 |
| 7 | V group (150 mg/kg ZnO NPs) | 15.21 | 10.78 |
| № | Experimental Group | Weighted Importance Score |
|---|---|---|
| 1 | III group (3.1 mg/kg ZnO NPs) | 1.319 |
| 2 | C− | 1.224 |
| 3 | II group (6.2 mg/kg ZnO NPs) | 1.095 |
| 4 | I group (6.2 mg/kg ZnO) | 1.063 |
| 5 | V group (150 mg/kg ZnO NPs) | 1.061 |
| 6 | IV group (1.55 mg/kg ZnO NPs) | 1.011 |
| 7 | C+ | 0.58 |
| № | Element | Model | Parameters | RMSE | MAE | Residuals Distribution | Predicted Max Content |
|---|---|---|---|---|---|---|---|
| 1 | Fe | Kernel Ridge | kernel = ‘rbf’ | 0.2787 | 0.3995 | Asymmetrical | 2.9512 |
| kernel = ‘chi2’ | 0.3195 | 0.4205 | Asymmetrical | 2.7698 | |||
| kernel = ‘laplacian’ | 0.3515 | 0.4443 | Asymmetrical | 2.6436 | |||
| FCNN | activation = ‘tanh’ | 0.2619 | 0.3905 | Asymmetrical | 3.2968 | ||
| activation = ‘sigmoid’ | 0.3102 | 0.4442 | Asymmetrical | 2.9483 | |||
| activation = ‘softmax’ | 0.2841 | 0.4203 | Asymmetrical | 2.7619 | |||
| activation = ‘swish’ | 0.3498 | 0.4237 | Asymmetrical | 3.5429 | |||
| 2 | Zn | Kernel Ridge | kernel = ‘rbf’ | 0.2621 | 0.3988 | Asymmetrical | 2.0894 |
| kernel = ‘chi2’ | 0.2246 | 0.3739 | Multimodal | 2.4197 | |||
| kernel = ‘laplacian’ | 0.2591 | 0.4104 | Asymmetrical | 2.5861 | |||
| FCNN | activation = ‘tanh’ | 0.1029 | 0.2556 | Asymmetrical | 2.3542 | ||
| activation = ‘sigmoid’ | 0.1644 | 0.3198 | Asymmetrical | 2.0265 | |||
| activation = ‘softmax’ | 0.1016 | 0.2531 | Asymmetrical | 2.2347 | |||
| activation = ‘swish’ | 0.1797 | 0.3071 | Asymmetrical | 2.1959 | |||
| 3 | Cu | Kernel Ridge | kernel = ‘rbf’ | 0.6491 | 0.6345 | Asymmetrical | 3.0081 |
| kernel = ‘chi2’ | 0.6063 | 0.6091 | Asymmetrical | 2.7348 | |||
| kernel= ‘laplacian’ | 0.6079 | 0.6043 | Asymmetrical | 2.7927 | |||
| FCNN | activation = ‘tanh’ | 0.5197 | 0.5419 | Asymmetrical | 2.7821 | ||
| activation = ‘sigmoid’ | 0.5959 | 0.5853 | Asymmetrical | 2.2453 | |||
| activation = ‘softmax’ | 0.6274 | 0.5906 | Asymmetrical | 2.3305 | |||
| activation = ’swish’ | 0.6028 | 0.5955 | Asymmetrical | 2.5059 | |||
| 4 | Mn | Kernel Ridge | kernel = ‘rbf’ | 0.3409 | 0.4838 | Multimodal | 1.2808 |
| kernel = ‘chi2’ | 0.2695 | 0.4232 | Multimodal | 0.6692 | |||
| kernel = ‘laplacian’ | 0.3207 | 0.4336 | Multimodal | 0.5349 | |||
| FCNN | activation = ‘tanh’ | 0.0309 | 0.1412 | Multimodal | 0.2166 | ||
| activation = ‘sigmoid’ | 0.0323 | 0.1461 | Multimodal | 0.2598 | |||
| activation = ‘softmax’ | 0.0301 | 0.1368 | Multimodal | 0.3187 | |||
| activation = ’swish’ | 0.0413 | 0.1470 | Multimodal | 0.2062 | |||
| 5 | Co | Kernel Ridge | kernel = ‘rbf’ | 0.3930 | 0.4909 | Multimodal | 0.3964 |
| kernel = ‘chi2’ | 0.2161 | 0.3426 | Multimodal | 0.6996 | |||
| kernel = ‘laplacian’ | 0.3082 | 0.4400 | Multimodal | 0.8097 | |||
| FCNN | activation = ‘tanh’ | 0.0336 | 0.1484 | Multimodal | 0.2830 | ||
| activation = ‘sigmoid’ | 0.0363 | 0.1584 | Multimodal | 0.3190 | |||
| activation = ‘softmax’ | 0.0305 | 0.1298 | Multimodal | 0.3104 | |||
| activation = ’swish’ | 0.1126 | 0.2129 | Multimodal | 0.5359 | |||
| 6 | Se | Kernel Ridge | kernel = ‘rbf’ | 0.1816 | 0.3158 | Multimodal | 1.6265 |
| kernel = ‘chi2’ | 0.3138 | 0.4278 | Multimodal | 2.0551 | |||
| kernel = ‘laplacian’ | 0.3660 | 0.4779 | Multimodal | 1.1546 | |||
| FCNN | activation = ‘tanh’ | 0.0224 | 0.1242 | Asymmetrical | 0.5166 | ||
| activation = ‘sigmoid’ | 0.0243 | 0.1283 | Asymmetrical | 0.4496 | |||
| activation = ‘softmax’ | 0.0193 | 0.1164 | Asymmetrical | 0.4557 | |||
| activation = ’swish’ | 0.0636 | 0.1474 | Asymmetrical | 0.5595 | |||
| 7 | I | Kernel Ridge | kernel = ‘rbf’ | 0.3719 | 0.4487 | Multimodal | 1.1298 |
| kernel = ‘chi2’ | 0.3036 | 0.4342 | Multimodal | 0.7566 | |||
| kernel = ‘laplacian’ | 0.2773 | 0.4258 | Multimodal | 1.2144 | |||
| FCNN | activation = ‘tanh’ | 0.0210 | 0.1134 | Multimodal | 0.3210 | ||
| activation = ‘sigmoid’ | 0.0209 | 0.11511 | Multimodal | 0.4038 | |||
| activation = ‘softmax’ | 0.0161 | 0.1030 | Multimodal | 0.3103 | |||
| activation = ’swish’ | 0.0582 | 0.1437 | Multimodal | 0.3563 | |||
| 8 | Cr | Kernel Ridge | kernel = ‘rbf’ | 0.4244 | 0.5110 | Multimodal | 0.3600 |
| kernel = ‘chi2’ | 0.1800 | 0.3301 | Multimodal | 1.0060 | |||
| kernel = ‘laplacian’ | 0.2813 | 0.4280 | Multimodal | 0.8072 | |||
| FCNN | activation = ‘tanh’ | 0.0291 | 0.1362 | Multimodal | 0.2161 | ||
| activation = ‘sigmoid’ | 0.0340 | 0.1504 | Multimodal | 0.3866 | |||
| activation = ‘softmax’ | 0.0297 | 0.1378 | Multimodal | 0.2245 | |||
| activation = ’swish’ | 0.0648 | 0.1715 | Multimodal | 0.4179 |
| № | Element | Model | Parameters | RMSE | MAE | Residuals Distribution | Predicted Max content |
|---|---|---|---|---|---|---|---|
| 1 | Sn | Kernel Ridge | kernel = ‘rbf’ | 0.3273 | 0.4493 | Multimodal | 0.6358 |
| kernel = ‘chi2’ | 0.3228 | 0.4881 | Multimodal | 0.5155 | |||
| kernel = ‘laplacian’ | 0.2566 | 0.4080 | Multimodal | 0.8477 | |||
| FCNN | activation = ‘tanh’ | 0.0325 | 0.1469 | Multimodal | 0.2775 | ||
| activation = ‘sigmoid’ | 0.0315 | 0.1438 | Multimodal | 0.2979 | |||
| activation = ‘softmax’ | 0.0299 | 0.1368 | Multimodal | 0.3272 | |||
| activation = ‘swish’ | 0.0761 | 0.1931 | Multimodal | 0.5563 | |||
| 2 | Al | Kernel Ridge | kernel = ‘rbf’ | 0.2435 | 0.4112 | Multimodal | 0.8085 |
| kernel = ‘chi2’ | 0.4254 | 0.4732 | Asymmetrical | 0.3071 | |||
| kernel = ‘laplacian’ | 0.3558 | 0.4688 | Multimodal | 0.3434 | |||
| FCNN | activation = ‘tanh’ | 0.0248 | 0.1264 | Asymmetrical | 0.3146 | ||
| activation = ‘sigmoid’ | 0.0308 | 0.1452 | Asymmetrical | 0.3201 | |||
| activation = ‘softmax’ | 0.0325 | 0.1494 | Asymmetrical | 0.3039 | |||
| activation = ‘swish’ | 0.1629 | 0.2027 | Multimodal | 0.6211 | |||
| 3 | As | Kernel Ridge | kernel = ‘rbf’ | 0.3767 | 0.4672 | Multimodal | 1.8047 |
| kernel = ‘chi2’ | 0.3368 | 0.4689 | Multimodal | 0.5911 | |||
| kernel = ‘laplacian’ | 0.2887 | 0.4435 | Multimodal | 0.7845 | |||
| FCNN | activation = ‘tanh’ | 0.0288 | 0.1367 | Multimodal | 0.2441 | ||
| activation = ‘sigmoid’ | 0.0349 | 0.1538 | Multimodal | 0.2718 | |||
| activation = ‘softmax’ | 0.0272 | 0.1314 | Multimodal | 0.2665 | |||
| activation = ’swish’ | 0.1043 | 0.1979 | Multimodal | 0.5054 | |||
| 4 | Hg | Kernel Ridge | kernel = ‘rbf’ | 0.3155 | 0.4647 | Multimodal | 0.6869 |
| kernel = ‘chi2’ | 0.3728 | 0.4747 | Multimodal | 1.0555 | |||
| kernel = ‘laplacian’ | 0.3018 | 0.4151 | Multimodal | 0.4964 | |||
| FCNN | activation = ‘tanh’ | 0.0342 | 0.1497 | Multimodal | 0.3177 | ||
| activation = ‘sigmoid’ | 0.0353 | 0.1552 | Multimodal | 0.3263 | |||
| activation = ‘softmax’ | 0.0292 | 0.1337 | Multimodal | 0.3183 | |||
| activation = ’swish’ | 0.0679 | 0.1671 | Multimodal | 0.1973 | |||
| 5 | Pb | Kernel Ridge | kernel = ‘rbf’ | 0.2680 | 0.4015 | Multimodal | 1.3640 |
| kernel = ‘chi2’ | 0.2704 | 0.4107 | Multimodal | 0.6657 | |||
| kernel = ‘laplacian’ | 0.3206 | 0.4354 | Multimodal | 0.9522 | |||
| FCNN | activation = ‘tanh’ | 0.0322 | 0.1455 | Multimodal | 0.3568 | ||
| activation = ‘sigmoid’ | 0.0330 | 0.1484 | Multimodal | 0.3164 | |||
| activation = ‘softmax’ | 0.0294 | 0.1350 | Multimodal | 0.3146 | |||
| activation = ’swish’ | 0.1638 | 0.2251 | Multimodal | 0.3828 | |||
| 6 | Be | Kernel Ridge | kernel = ‘rbf’ | 0.3186 | 0.4307 | Multimodal | 0.8344 |
| kernel = ‘chi2’ | 0.4514 | 0.5470 | Multimodal | 0.9656 | |||
| kernel = ‘laplacian’ | 0.3062 | 0.4377 | Multimodal | 0.1190 | |||
| FCNN | activation = ‘tanh’ | 0.0382 | 0.1597 | Multimodal | 0.2436 | ||
| activation = ‘sigmoid’ | 0.0328 | 0.1487 | Multimodal | 0.3366 | |||
| activation = ‘softmax’ | 0.0313 | 0.1340 | Multimodal | 0.2603 | |||
| activation = ’swish’ | 0.1704 | 0.2423 | Multimodal | 0.3919 | |||
| 7 | Cd | Kernel Ridge | kernel = ‘rbf’ | 0.2955 | 0.4306 | Multimodal | 0.4646 |
| kernel = ‘chi2’ | 0.2932 | 0.4290 | Multimodal | 0.8920 | |||
| kernel = ‘laplacian’ | 0.3988 | 0.4934 | Multimodal | 0.8130 | |||
| FCNN | activation = ‘tanh’ | 0.0322 | 0.1434 | Multimodal | 0.4029 | ||
| activation = ‘sigmoid’ | 0.0335 | 0.1493 | Multimodal | 0.2838 | |||
| activation = ‘softmax’ | 0.0268 | 0.1269 | Multimodal | 0.3301 | |||
| activation = ’swish’ | 0.1734 | 0.2314 | Multimodal | 0.4846 |
| № | Element | Model | Parameters | RMSE | MAE | Residuals Distribution | Predicted Max Content |
|---|---|---|---|---|---|---|---|
| 1 | CA4 | Kernel Ridge | kernel = ‘rbf’ | 14.6207 | 2.5678 | Multimodal | 26.9611 |
| kernel = ‘chi2’ | 15.6488 | 2.6430 | Asymmetrical | 26.0791 | |||
| kernel = ‘laplacian’ | 13.8650 | 2.4912 | Multimodal | 24.5711 | |||
| FCNN | activation = ‘tanh’ | 40.6357 | 4.6784 | Asymmetrical | 19.3162 | ||
| activation = ‘sigmoid’ | 40.8586 | 4.7742 | Asymmetrical | 19.1345 | |||
| activation = ‘softmax’ | 263.2020 | 14.917 | Asymmetrical | 4.0573 | |||
| activation = ‘swish’ | 15.0017 | 2.5964 | Asymmetrical | 26.8949 | |||
| 2 | ADH1B | Kernel Ridge | kernel = ‘rbf’ | 3.5569 | 1.2353 | Multimodal | 54.9218 |
| kernel = ‘chi2’ | 4.1477 | 1.3417 | Multimodal | 57.6527 | |||
| kernel = ‘laplacian’ | 3.8703 | 1.2907 | Multimodal | 47.6248 | |||
| FCNN | activation = ‘tanh’ | 30.7518 | 4.7432 | Multimodal | 48.5105 | ||
| activation = ‘sigmoid’ | 37.8307 | 4.5278 | Multimodal | 45.5706 | |||
| activation = ‘softmax’ | 2001.93 | 44.3947 | Multimodal | 4.1230 | |||
| activation = ‘swish’ | 6.0534 | 1.8218 | Asymmetrical | 59.6373 | |||
| 3 | ALP | Kernel Ridge | kernel = ‘rbf’ | 9.9965 | 1.9220 | Multimodal | 26.6896 |
| kernel = ‘chi2’ | 9.4867 | 1.8509 | Multimodal | 26.8992 | |||
| kernel = ‘laplacian’ | 10.3592 | 1.9001 | Multimodal | 21.3801 | |||
| FCNN | activation = ‘tanh’ | 22.8733 | 3.2491 | Asymmetrical | 22.3343 | ||
| activation = ‘sigmoid’ | 22.2092 | 3.1820 | Multimodal | 22.7722 | |||
| activation = ‘softmax’ | 357.2593 | 18.2905 | Asymmetrical | 3.9571 | |||
| activation = ’swish’ | 10.1903 | 2.1047 | Asymmetrical | 26.9728 | |||
| 4 | ZAG | Kernel Ridge | kernel = ‘rbf’ | 0.0772 | 0.1955 | Multimodal | 12.0935 |
| kernel = ‘chi2’ | 0.0917 | 0.2112 | Multimodal | 12.2758 | |||
| kernel = ‘laplacian’ | 0.0824 | 0.1989 | Multimodal | 11.5086 | |||
| FCNN | activation = ‘tanh’ | 0.2983 | 0.4494 | Multimodal | 11.6732 | ||
| activation = ‘sigmoid’ | 0.3957 | 0.5246 | Multimodal | 11.4666 | |||
| activation = ‘softmax’ | 55.9166 | 7.4532 | Multimodal | 3.9647 | |||
| activation = ’swish’ | 0.2390 | 0.3785 | Multimodal | 12.1311 | |||
| 5 | SOD-Zn | Kernel Ridge | kernel = ‘rbf’ | 1.4288 | 0.6696 | Multimodal | 10.2444 |
| kernel= ‘chi2’ | 1.5202 | 0.6944 | Multimodal | 10.5237 | |||
| kernel = ‘laplacian’ | 1.4431 | 0.6727 | Multimodal | 9.1893 | |||
| FCNN | activation = ‘tanh’ | 1.5072 | 0.7509 | Asymmetrical | 10.1673 | ||
| activation = ‘sigmoid’ | 1.6879 | 0.8332 | Asymmetrical | 10.0199 | |||
| activation = ‘softmax’ | 28.4987 | 5.0826 | Asymmetrical | 3.7381 | |||
| activation = ’swish’ | 1.4718 | 0.6924 | Asymmetrical | 10.1762 | |||
| 6 | MT1 | Kernel Ridge | kernel = ‘rbf’ | 18.6226 | 2.6608 | Asymmetrical | 79.6881 |
| kernel = ‘chi2’ | 20.9568 | 2.8212 | Asymmetrical | 83.6053 | |||
| kernel = ‘laplacian’ | 20.3503 | 2.8068 | Asymmetrical | 75.3316 | |||
| FCNN | activation = ‘tanh’ | 67.1140 | 7.3396 | Multimodal | 72.1062 | ||
| activation = ‘sigmoid’ | 395.9441 | 19.1312 | Asymmetrical | 64.6869 | |||
| activation = ‘softmax’ | 5555.85 | 74.3725 | Asymmetrical | 4.0292 | |||
| activation = ’swish’ | 25.1968 | 3.9657 | Asymmetrical | 83.0218 | |||
| 7 | MT2 | Kernel Ridge | kernel = ‘rbf’ | 0.9364 | 0.6131 | Asymmetrical | 6.3687 |
| kernel = ‘chi2’ | 0.9325 | 0.6285 | Asymmetrical | 7.1156 | |||
| kernel = ‘laplacian’ | 0.8081 | 0.5622 | Asymmetrical | 5.5452 | |||
| FCNN | activation = ‘tanh’ | 1.0611 | 0.7543 | Asymmetrical | 6.7182 | ||
| activation = ‘sigmoid’ | 1.0325 | 0.7113 | Asymmetrical | 6.4843 | |||
| activation = ‘softmax’ | 6.9113 | 2.4723 | Asymmetrical | 3.6365 | |||
| activation = ‘swish’ | 1.0315 | 0.7060 | Asymmetrical | 7.2760 | |||
| 8 | MT3 | Kernel Ridge | kernel = ‘rbf’ | 0.3527 | 0.4698 | Multimodal | 1.0865 |
| kernel = ‘chi2’ | 0.3199 | 0.4196 | Multimodal | 0.9241 | |||
| kernel = ‘laplacian’ | 0.2291 | 0.3630 | Multimodal | 0.9298 | |||
| FCNN | activation = ‘tanh’ | 0.0211 | 0.1125 | Multimodal | 0.4658 | ||
| activation = ‘sigmoid’ | 0.0242 | 0.1233 | Multimodal | 0.3263 | |||
| activation = ‘softmax’ | 0.0182 | 0.1104 | Multimodal | 0.2827 | |||
| activation = ‘swish’ | 0.0895 | 0.1798 | Multimodal | 0.4480 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Legashev, L.; Khokhlov, I.; Bolodurina, I.; Shukhman, A.; Kolesnik, S. Machine Learning Prediction Models of Beneficial and Toxicological Effects of Zinc Oxide Nanoparticles in Rat Feed. Mach. Learn. Knowl. Extr. 2025, 7, 91. https://doi.org/10.3390/make7030091
Legashev L, Khokhlov I, Bolodurina I, Shukhman A, Kolesnik S. Machine Learning Prediction Models of Beneficial and Toxicological Effects of Zinc Oxide Nanoparticles in Rat Feed. Machine Learning and Knowledge Extraction. 2025; 7(3):91. https://doi.org/10.3390/make7030091
Chicago/Turabian StyleLegashev, Leonid, Ivan Khokhlov, Irina Bolodurina, Alexander Shukhman, and Svetlana Kolesnik. 2025. "Machine Learning Prediction Models of Beneficial and Toxicological Effects of Zinc Oxide Nanoparticles in Rat Feed" Machine Learning and Knowledge Extraction 7, no. 3: 91. https://doi.org/10.3390/make7030091
APA StyleLegashev, L., Khokhlov, I., Bolodurina, I., Shukhman, A., & Kolesnik, S. (2025). Machine Learning Prediction Models of Beneficial and Toxicological Effects of Zinc Oxide Nanoparticles in Rat Feed. Machine Learning and Knowledge Extraction, 7(3), 91. https://doi.org/10.3390/make7030091







