West Nile Virus Lineage 2 in Free-Living Corvus cornix Birds in Poland
Abstract
:1. Introduction
2. Materials and Methods
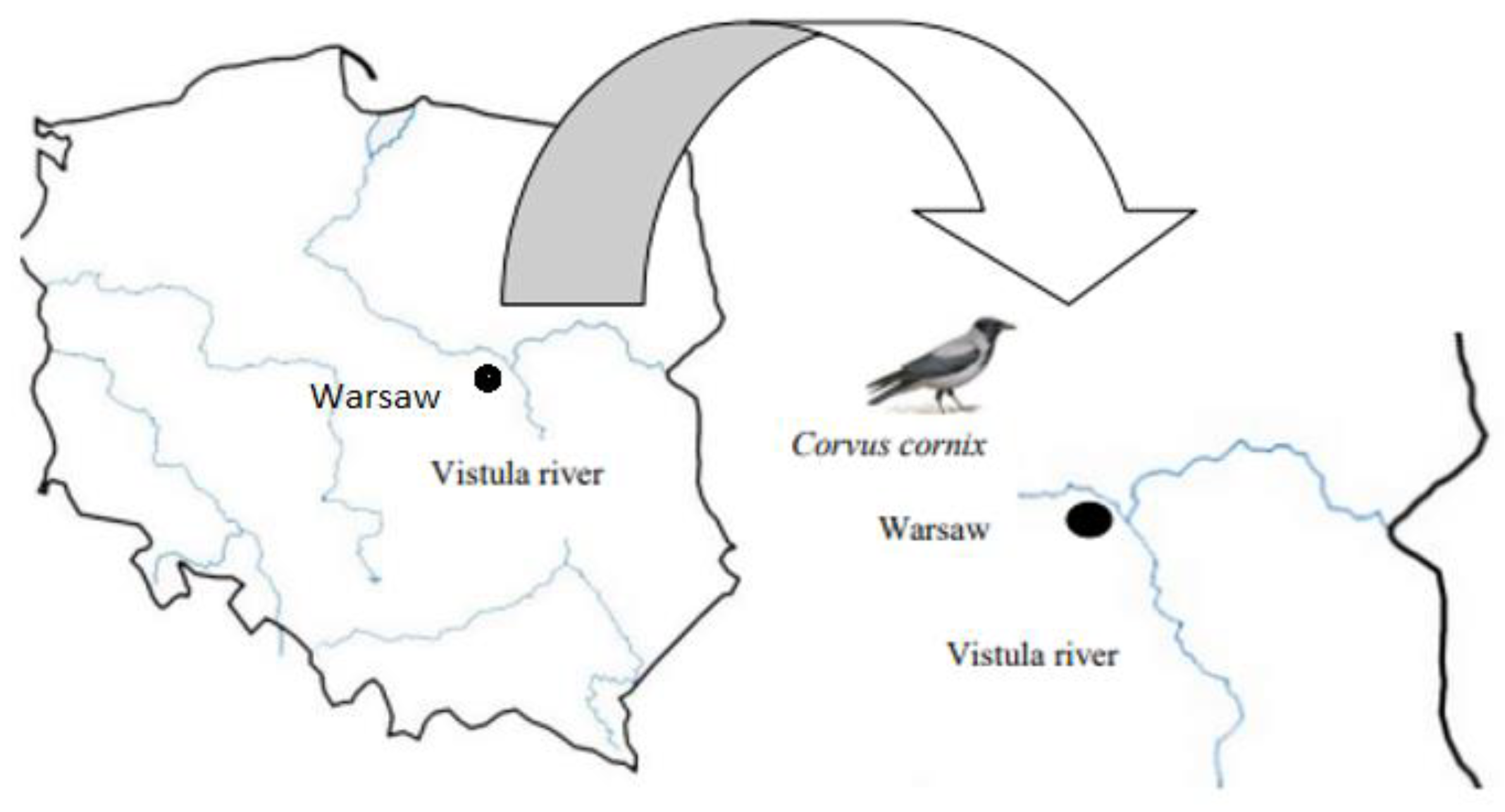
2.1. Study Region
2.2. Ethical Statement
2.3. Sampling Collection
2.4. RNA Extraction
2.5. Control Samples
2.6. Primers and RT-PCR Amplification of WNV
2.7. Molecular Studies
2.8. Full-Length WNV Genome Sequencing—Sequencing Analysis
2.9. Phylogenetic Analysis
3. Results
3.1. Studies
3.2. RT-PCR and Real-Time RT-PCR
3.3. Sequencing Analysis
3.4. Phylogenetic Tree
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Hubalek, Z.; Wegner, E.; Halouzka, J.; Tryjanowski, P.; Jerzak, L.; Sikutova, S.; Rudolf, I.; Kruszewicz, A.G.; Jaworski, Z.; Włodarczyk, R. Serologic Survey of potential vertebrate hosts for West Nile virus in Poland. Viral Immunol. 2008, 21, 247–253. [Google Scholar] [CrossRef] [PubMed]
- Komar, N.; Langevin, S.; Hinten, S.; Nemeth, N.; Edwards, E.; Hettler, D.; Davis, B.; Bowen, R.; Bunning, M. Experimental infection of North American birds with the New York 1999 strain of West Nile virus. Emerg. Infect. Dis. 2003, 9, 311–322. [Google Scholar] [CrossRef] [PubMed]
- Perez-Ramirez, E.; Llorente, F.; Jimenez-Clavero, M.A. Experimantal infections of wild birds with West Nile virus. Viruses 2014, 6, 752–781. [Google Scholar] [CrossRef]
- Ciota, A.T.; Chin, P.A.; Kramer, L.D. The effect of hybridization of Culex pipiens complex mosquitoes on transmission of West Nile virus. Parasitol. Vectors 2013, 6, 305. [Google Scholar] [CrossRef] [PubMed]
- Chancey, C.; Grinev, A.; Volkova, E.; Rios, M. The Global Ecology and Epidemiology of West Nile Virus. BioMed Res. Int. 2015, 2015, 376230. [Google Scholar] [CrossRef] [PubMed]
- Smithburn, K.C.; Hughes, T.P.; Burke, A.W.; Paul, J.H. A neurotropic virus isolated from the blood of a native of Uganda. Am. J. Trop. Med. Hyg. 1940, 20, 471–492. [Google Scholar] [CrossRef]
- Kramer, L.D.; Styer, L.M.; Ebel, G.D. A global perspective on the epidemiology of West Nile virus. Annu. Rev. Entomol. 2008, 53, 61–81. [Google Scholar] [CrossRef]
- Vilibic-Cavlek, T.; Savic, V.; Petrovic, T.; Toplak, I.; Barbic, L.; Petric, D.; Tabain, I.; Hrnjakovic-Cvjetkovic, I.; Bogdanic, M.; Klobucar, A.; et al. Emerging trends in the epidemiology of West Nile and Usutu virus infections in southern Europe. Front. Vet. Sci. 2019, 6, 437. [Google Scholar] [CrossRef]
- Benzarti, E.; Linden, A.; Desmecht, D.; Garigliany, M. Mosquito-borne epornitic flaviviruses: An update and review. J. Gen. Virol. 2019, 100, 119–132. [Google Scholar] [CrossRef]
- Cox, S.L.G.; Campbell, D.; Nemeth, N.N. Outbreaks of West Nile virus in captive waterfowl in Ontario, Canada. Avian Pathol. 2015, 44, 135–141. [Google Scholar] [CrossRef]
- Petrović, T.; Blázquez, A.B.; Lupulović, D.; Lazić, G.; Escribano-Romero, E.; Fabijan, D.; Kapetanov, M.; Lazić, S.; Saiz, J.C. Monitoring West Nile virus (WNV) infection in wild birds in Serbia during 2012: First isolation and characterisation of WNV strains from Serbia. Eurosurvelllance 2013, 18, 20622. [Google Scholar] [CrossRef] [PubMed]
- Jourdain, E.; Gauthier-Clerc, M.; Bicout, D.J.; Sabatier, P. Bird migration routes and risk for pathogen dispersion into Western Mediterranean wetlands. Emerg. Infect. Dis. 2007, 13, 365–372. [Google Scholar] [CrossRef]
- Golding, N.; Nunn, M.; Medlock, J.M.; Purse, B.V.; Vaux, G.C.; Schäffer, S.M. West Nile virus vector Culex modestus established in southern England. Parasitol. Vectors 2012, 5, 32. [Google Scholar] [CrossRef] [PubMed]
- Ciota, A.T.; Kramer, L.D. Vector-Virus Interactions and Transmission Dynamics of West Nile. Viruses 2013, 5, 3021–3047. [Google Scholar] [CrossRef]
- Medlock, J.M.; Hansford, K.M.; Schaffner, F.; Versteirt, V.; Hendrickx, G.; Zeller, H.; Bortel, W.V. A review of the invasive mosquitoes in Europe: Ecology, public health risks, and control options. Vector Borne Zoonotic Dis. 2012, 2, 435–447. [Google Scholar] [CrossRef]
- Rappole, J.H.; Derrickson, S.R.; Hubálek, Z. Migratory birds and spread of West Nile virus in the Western Hemisphere. Emerg. Infect. Dis. 2000, 6, 319–328. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Triana, L.M.; Jeffries, C.L.; Mansfield, K.L.; Carnell, G.; Fooks, A.R.; Johnson, N. Emergence of West Nile Virus Lineage 2 in Europe: A Review on the Introduction and Spread of a Mosquito-Borne Disease. Front. Public Health 2014, 2, 271. [Google Scholar] [CrossRef]
- Niczyporuk, J.S.; Samorek-Salamonowicz, E.; Kozdruń, W.; Mizak, Z. Attempts to detect West Nile Virus in wild birds in Poland. Acta Vet. Hung. 2011, 59, 405–408. [Google Scholar] [CrossRef] [PubMed]
- Niczyporuk, J.S.; Samorek-Salamonowicz, E.; Lecollinet, S.; Pancewicz, A.S.; Kozdruń, W.; Czekaj, H. Occurence of West Nile Virus antibodies in wild birds, horses, and human in Poland. BioMed Res. Int. 2015, 2015, 234181. [Google Scholar] [CrossRef]
- Niczyporuk, J.S. Wirus Zachodniego Nilu w Polsce—Realne zagrożenie w świetle doniesień prezentowanych na konferencji. Aktualne problemy dotyczące czynników zakaźnych przenoszonych przez krew. J. Transfus. Med. 2017, 10, 54–62. [Google Scholar]
- Ziegler, U.; Lühken, R.; Keller, M.; Cadar, D.; Van der Grinten, E.; Michel, F.; Albrecht, K.; Eiden, M.; Rinder, M.; Lachmann, L.; et al. West Nile virus epizootic in Germany 2018. Antivir. Res. 2019, 162, 39–43. [Google Scholar] [CrossRef]
- Sikkema, R.S.; Schrama, M.; van den Berg, T.; Morren, J.; Munger, E.; Krol, L.; van der Beek, J.G.; Blom, R.; Chestakova, I.; van der Linden, A.; et al. Detection of West Nile virus in a common whitethroat (Curruca communis) and Culex mosquitoes in the Netherlands, 2020. Eurosurveillance 2020, 25, 2001704. [Google Scholar] [CrossRef] [PubMed]
- Linke, S.; Ellerbrok, H.; Niedrig, M.; Nitsche, A.; Pauli, G. Detection of West Nile virus lineages 1 and 2 by real-time PCR. J. Virol. Methods 2007, 146, 355–358. [Google Scholar] [CrossRef] [PubMed]
- Del Amo, J.; Sotelo, E.; Fernández-Pinero, J.; Gallardo, C.; Llorente, F.; Agüero, M.; Jiménez-Clavero, M.A. A novel quantitative multiplex real-time RT-PCR for the simultaneous detection and differentiation of West Nile virus lineages 1 and 2, and of Usutu virus. J. Virol. Methods 2013, 189, 321–327. [Google Scholar] [CrossRef] [PubMed]
- Piorkowski, G.; Richard, P.; Baronti, C.; Gallian, P.; Charrel, R.; Leparc-Goffart, I.; de Lamballerie, X. Complete coding sequence of Zika virus from Martinique outbreak in 2015. New Microbes New Infect. 2016, 11, 52–53. [Google Scholar] [CrossRef]
- Nei, M.; Kumar, S. Molecular Evolution and Phylogenetics; Oxford University Press: New York, NY, USA, 2000. [Google Scholar]
- Tamura, K.; Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in human and chimpanzez. Mol. Biol. Evol. 1993, 10, 512–526. [Google Scholar] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA 11: Molecular Evolutionary Genetics Analysis Version 11. Mol. Biol. Evol. 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Schvartz, G.; Farnoushi, Y.; Berkowitz, A.; Edery, N.; Hahn, S.; Steinman, A.; Lublin, A.; Erster, O. Molecular characterization of the re-emerging West Nile virus in avian species and equids in Israel, 2018, and pathological description of the disease. Parasites Vectors 2020, 13, 528. [Google Scholar] [CrossRef] [PubMed]
- Schvartz, G.; Tirosh-Levy, S.; Bider, S.; Lublin, A.; Farnoushi, Y.; Erster, O.; Steinman, A. West Nile Virus in Common Wild Avian Species in Israel. Pathogens 2022, 11, 107. [Google Scholar] [CrossRef]
- Paz, S. Climate change impacts on West Nile virus transmission in a global context. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2015, 370, 20130561. [Google Scholar] [CrossRef]
- Barzon, L.; Montarsi, F.; Quaranta, E.; Monne, I.; Pacenti, M.; Michelutti, A.; Toniolo, F.; Danesi, P.; Marchetti, G.; Gobbo, F.; et al. Early start of seasonal transmission and co-circulation of West Nile virus lineage 2 and newly introduced lineage 1 strain, northern Italy. Eurosurvellance 2022, 27, 2200548. [Google Scholar]
- Ziegler, U.; Angenvoort, J.; Klaus, C.; Nagel-Kohl, U.; Sauerwald, C.; Thalheim, S.; Horner, S.; Braun, B.; Kenklies, S.; Tyczka, J.; et al. Use of Competition ELISA for Monitoring of West Nile Virus infections in Horses in Germany. Int. J. Environ. Res. Public Health 2013, 10, 3112–3120. [Google Scholar] [CrossRef] [PubMed]
- European Centre for Disease Prevention and Control. Epidemiological Update: West Nile Virus Transmission Season in Europe. 2022. Available online: https://www.ecdc.europa.eu/en/news-events/epidemiologicalupdate-west-nile-virus-transmission-seasoneurope-2022 (accessed on 26 November 2022).
- Glavitis, R.; Ferenci, E.; Ivanics, E.; Bakony, T.; Mato, T.; Zarka, P.V. Co-occurrence of West Nile Fever and circovirus infection in a goose flock in Hungary. Avian Pathol. 2005, 34, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Balança, G.; Gaidet, N.; Savini, G.; Vollot, B.; Foucart, A.; Reiter, P.; Boutonnier, A.; Lelli, R.; Monicat, F. Low West Nile virus circulation in wild birds in an area of recurring outbreaks in Southern France. Vector Borne Zoonotic Dis. 2009, 9, 737–741. [Google Scholar] [CrossRef] [PubMed]
- Erdélyi, K.; Ursu, K.; Ferenczi, E.; Szeredi, L.; Rátz, F.; Skáre, J.; Bakonyi, T. Clinical and pathologic features of lineage 2 West Nile virus infections in birds of prey in Hungary. Vector Borne Zoonotic Dis. 2007, 7, 181–188. [Google Scholar] [CrossRef]
- Wodak, E.; Richter, S.; Bagó, Z.; Revilla-Fernández, S.; Weissenböck, H.; Nowotny, N.; Winter, P. Detection and molecular analysis of West Nile virus infections in birds of prey in the eastern part of Austria in 2008–2009. Vet. Microbiol. 2011, 149, 358–366. [Google Scholar] [CrossRef]
- Valiakos, G.; Plavos, K.; Vontas, A.; Sofia, M.; Giannakopoulos, A.; Giannoulis, G.; Spyrou, V.; Tsokana, N.; Chatzopoulos, D.; Kantere, M.; et al. Phylogenetic Analysis of Bird-Virulent West Nile Virus Strain, Greece. Emerg. Infect. Dis. 2019, 25, 2323–2325. [Google Scholar] [CrossRef]
- Bravo-Barriga, D.; Aguilera-Sepúlveda, P.; Carvajal, F.G.; Llorente, F.; Reina, D.; Jiménez-Clavero, M.A.; Frontera, E. West Nile and Usutu virus infections in wild birds admitted to rehabilitation centres in Extremadura, western Spain, 2017–2019. Vet. Microbiol. 2021, 255, 109020. [Google Scholar] [CrossRef]

| Avian Order | Avian Species | Analysed (n) | Result |
|---|---|---|---|
| Anseriformes (Waterfowl) | Cygnus olor Anas platyrhynchos | 12 5 | Negative Negative |
| Passeriformes (Corvidae) | Corvus cornix Corvus frugilegus Corvus monedula | 49 4 2 | Positive 17 (34.69%) Negative Negative |
| Strigiformes (Owls) | Strix aluco Bubo scandiacus | 1 1 | Negative Negative |
| Falconiformes (Falconidae) | Falco peregrinus | 3 | Negative |
| Ciconiiformes (Storks) | Ciconia ciconia | 18 | Negative |
| 99 | Positive 17 (17.17%) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Niczyporuk, J.S.; Kozdrun, W.; Czujkowska, A.; Blanchard, Y.; Helle, M.; Dheilly, N.M.; Gonzalez, G. West Nile Virus Lineage 2 in Free-Living Corvus cornix Birds in Poland. Trop. Med. Infect. Dis. 2023, 8, 417. https://doi.org/10.3390/tropicalmed8080417
Niczyporuk JS, Kozdrun W, Czujkowska A, Blanchard Y, Helle M, Dheilly NM, Gonzalez G. West Nile Virus Lineage 2 in Free-Living Corvus cornix Birds in Poland. Tropical Medicine and Infectious Disease. 2023; 8(8):417. https://doi.org/10.3390/tropicalmed8080417
Chicago/Turabian StyleNiczyporuk, Jowita S., Wojciech Kozdrun, Agnieszka Czujkowska, Yannick Blanchard, Mariteragi Helle, Nolwenn M. Dheilly, and Gaelle Gonzalez. 2023. "West Nile Virus Lineage 2 in Free-Living Corvus cornix Birds in Poland" Tropical Medicine and Infectious Disease 8, no. 8: 417. https://doi.org/10.3390/tropicalmed8080417
APA StyleNiczyporuk, J. S., Kozdrun, W., Czujkowska, A., Blanchard, Y., Helle, M., Dheilly, N. M., & Gonzalez, G. (2023). West Nile Virus Lineage 2 in Free-Living Corvus cornix Birds in Poland. Tropical Medicine and Infectious Disease, 8(8), 417. https://doi.org/10.3390/tropicalmed8080417







