Shaping the Cell and the Future: Recent Advancements in Biophysical Aspects Relevant to Regenerative Medicine
Abstract
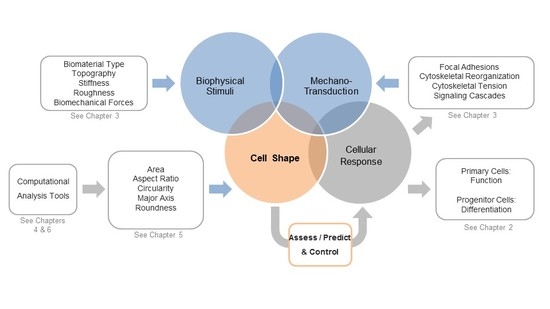
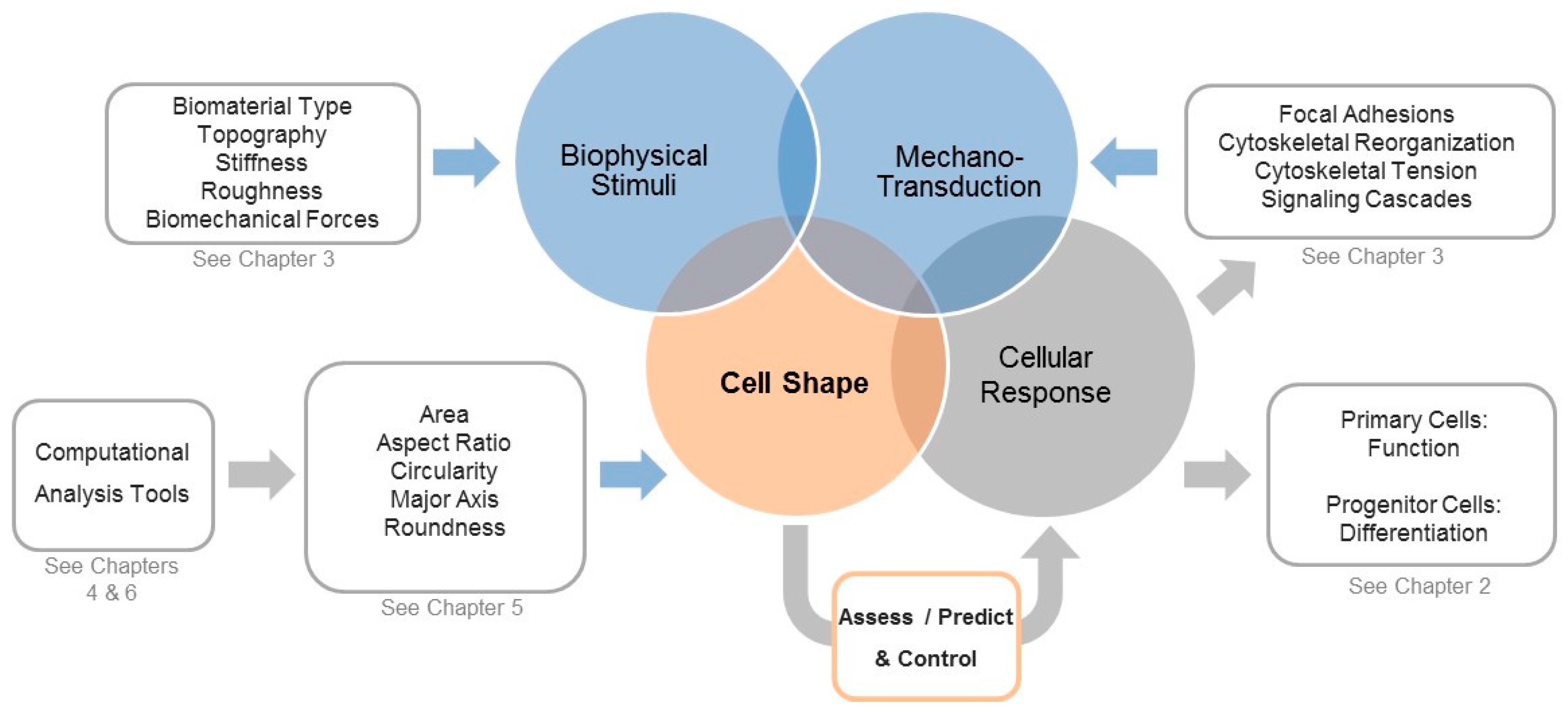
1. Is Shaping the Cell Also Shaping Regenerative Medicine?
2. Cell Morphology as a Novel Tool to Assess Biological Responses in Tissue Engineering
3. The Cellular Components that Sense and Respond to Biophysical Stimuli and Affect Cell Shape
4. Computational Tools to Segment Images for Recognizing Cells
5. Measuring Cellular Morphology with a Panel of Cell Shape Descriptors
6. Computational Approaches to Classify Shape Profiles into Biologically Interpretable Groups
7. A View into the Future: Controlling Shape for Regenerative Medicine Applications
Conflicts of Interest
References
- Bianco, P. “Mesenchymal” stem cells. Annu. Rev. Cell. Dev. Biol. 2014, 30, 677–704. [Google Scholar] [CrossRef] [PubMed]
- Klein, G.; Hart, M.L.; Brinchmann, J.E.; Rolauffs, B.; Stenzl, A.; Sievert, K.-D.; Aicher, W.K. Mesenchymal stromal cells for sphincter regeneration. Adv. Drug Deliv. Rev. 2015, 82–83C, 123–136. [Google Scholar] [CrossRef] [PubMed]
- Caplan, A.I.; Mason, C.; Reeve, B. The 3Rs of Cell Therapy. Stem Cells Transl. Med. 2016. [Google Scholar] [CrossRef] [PubMed]
- Bianco, P.; Cao, X.; Frenette, P.S.; Mao, J.J.; Robey, P.G.; Simmons, P.J.; Wang, C.Y. The meaning, the sense and the significance: Translating the science of mesenchymal stem cells into medicine. Nat. Med. 2013, 19, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Iskratsch, T.; Wolfenson, H.; Sheetz, M.P. Appreciating force and shape-the rise of mechanotransduction in cell biology. Nat. Rev. Mol. Cell Biol. 2014, 15, 825–833. [Google Scholar] [CrossRef] [PubMed]
- Paluch, E.K.; Nelson, C.M.; Biais, N.; Fabry, B.; Moeller, J.; Pruitt, B.L.; Wollnik, C.; Kudryasheva, G.; Rehfeldt, F.; Federle, W. Mechanotransduction: Use the force(s). BMC Biol. 2015, 13, 47. [Google Scholar] [CrossRef] [PubMed]
- Strzyz, P. Mechanotransduction: May the force be with you. Nat. Rev. Mol. Cell Biol. 2016, 17, 533. [Google Scholar] [CrossRef] [PubMed]
- Albert, P.J.; Schwarz, U.S. Modeling cell shape and dynamics on micropatterns. Cell Adhes. Migr. 2016, 10, 516–528. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Relan, N.K.; Przywara, D.A.; Schuger, L. Embryonic mesenchymal cells share the potential for smooth muscle differentiation: Myogenesis is controlled by the cell’s shape. Development 1999, 126, 3027–3033. [Google Scholar] [PubMed]
- Manasek, F.J.; Burnside, M.B.; Waterman, R.E. Myocardial cell shape change as a mechanism of embryonic heart looping. Dev. Biol. 1972, 29, 349–371. [Google Scholar] [CrossRef]
- Gao, L.; McBeath, R.; Chen, C.S. Stem cell shape regulates a chondrogenic versus myogenic fate through Rac1 and N-cadherin. Stem Cells 2010, 28, 564–572. [Google Scholar] [CrossRef] [PubMed]
- Singhvi, R.; Kumar, A.; Lopez, G.P.; Stephanopoulos, G.N.; Wang, D.I.; Whitesides, G.M.; Ingber, D.E. Engineering cell shape and function. Science 1994, 264, 696–698. [Google Scholar] [CrossRef] [PubMed]
- Folkman, J.; Moscona, A. Role of cell shape in growth control. Nature 1978, 273, 345–349. [Google Scholar] [CrossRef] [PubMed]
- Costa-Almeida, R.; Carvalho, D.T.O.; Ferreira, M.J.S.; Pesqueira, T.; Monici, M.; Van Loon, J.J.W.A.; Granja, P.L.; Gomes, M.E. Simulated hypergravity induces changes in human tendon-derived cells: From cell morphology to gene expression. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine, TERMIS-EU 2017 Conference, Davos, Switzerland, 26–30 June 2017; AO Research Institute Davos: Davos, Switzerland, 2017; p. 429. [Google Scholar]
- Zhang, J.; Rubert, M.; Baumgartner, J.; Wehrle, E.; Paul, G.; Müller, R. The influence of mechanical properties of 3D bioprinted hMSCs-laden alginate composite scaffolds on cell viability and morphology. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine, TERMIS-EU 2017 Conference, Davos, Switzerland, 26–30 June 2017; AO Research Institute Davos: Davos, Switzerland, 2017; p. 608. [Google Scholar]
- Rocca, A.; Marino, A.; Rocca, V.; Moscato, S.; De Vito, G.; Piazza, V.; Mazzolai, B.; Mattoli, V.; Ngo-Anh, T.J.; Ciofani, G. Barium titanate nanoparticles and hypergravity stimulation improve differentiation of mesenchymal stem cells into osteoblasts. Int. J. Nanomed. 2015, 10, 433–445. [Google Scholar]
- Jain, N.; Vogel, V. Cell geometry regulates temporal regulation of gene expression during macrophage activation. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine TERMIS-EU 2017 Conference, Davos, Switzerland 26–30 June 2017; AO Research Institute Davos: Davos, Switzerland, 2017. [Google Scholar]
- Hart, M.L.; Schmid, L.; Walters, B.; Rolauffs, B. Implementation and verification of an efficient, precise, reproducible and accurate automatic quantification method for measuring mesenchymal stromal cell morphological parameters. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine TERMIS-EU 2017 Conference, Davos, Switzerland, 26–30 June 2017; AO Research Institute Davos: Davos, Switzerland, 2017; p. 567. [Google Scholar]
- Uynuk-Ool, T.; Rothdiener, M.; Walters, B.; Hegemann, M.; Palm, J.; Nguyen, P.; Seeger, T.; Stöckle, U.; Stegemann, J.P.; Aicher, W.K.; et al. The geometrical shape of mesenchymal stromal cells measured by quantitative shape descriptors is determined by the stiffness of the biomaterial and by cyclic tensile forces. J. Tissue Eng. Regen. Med. 2017. [Google Scholar] [CrossRef] [PubMed]
- Kato, R. Application and understanding of morphological data for non-invasive quality control of stem cell manufacturing. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine TERMIS-EU 2017 Conference, 26–30 June 2017, Davos, Switzerland; AO Research Institute Davos: Davos, Switzerland, 2017; p. 798. [Google Scholar]
- Fujitani, M.; Kawai, S.; Kanie, K.R.K. Morphology-based real-time evaluation of culture condition. In eCells & Materials (eCM) Meeting Abstracts 2017, Collection 2. Personalised Therapies for Regenerative Medicine TERMIS-EU 2017 Conference, Davos, Switzerland, 26–30 June 2017; AO Research Institute Davos: Davos, Switzerland, 2017; p. 797. [Google Scholar]
- Kato, R.; Matsumoto, M.; Sasaki, H.; Joto, R.; Okada, M.; Ikeda, Y.; Kanie, K.; Suga, M.; Kinehara, M.; Yanagihara, K.; et al. Parametric analysis of colony morphology of non-labelled live human pluripotent stem cells for cell quality control. Sci. Rep. 2016, 6, 34009. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, H.; Takeuchi, I.; Okada, M.; Sawada, R.; Kanie, K.; Kiyota, Y.; Honda, H.; Kato, R. Label-free morphology-based prediction of multiple differentiation potentials of human mesenchymal stem cells for early evaluation of intact cells. PLoS ONE 2014, 9, e93952. [Google Scholar] [CrossRef] [PubMed]
- Walters, B.; Uynuk-Ool, T.; Rothdiener, M.; Palm, J.; Hart, M.L.; Stegemann, J.P.; Rolauffs, B. Engineering the geometrical shape of mesenchymal stromal cells through defined cyclic stretch regimens. Sci. Rep. 2017, 7, 6640. [Google Scholar] [CrossRef] [PubMed]
- Marklein, R.A.; Lo Surdo, J.L.; Bellayr, I.H.; Godil, S.A.; Puri, R.K.; Bauer, S.R. High Content Imaging of Early Morphological Signatures Predicts Long Term Mineralization Capacity of Human Mesenchymal Stem Cells upon Osteogenic Induction. Stem Cells 2016, 34, 935–947. [Google Scholar] [CrossRef] [PubMed]
- Matsuoka, F.; Takeuchi, I.; Agata, H.; Kagami, H.; Shiono, H.; Kiyota, Y.; Honda, H.; Kato, R. Morphology-based prediction of osteogenic differentiation potential of human mesenchymal stem cells. PLoS ONE 2013, 8, e55082. [Google Scholar] [CrossRef] [PubMed]
- Seiler, C.; Gazdhar, A.; Reyes, M.; Benneker, L.M.; Geiser, T.; Siebenrock, K.A.; Gantenbein-Ritter, B. Time-lapse microscopy and classification of 2D human mesenchymal stem cells based on cell shape picks up myogenic from osteogenic and adipogenic differentiation. J. Tissue Eng. Regen. Med. 2014, 8, 737–746. [Google Scholar] [CrossRef] [PubMed]
- Pearson, Y.E.; Lund, A.W.; Lin, A.W.; Ng, C.P.; Alsuwaidi, A.; Azzeh, S.; Gater, D.L.; Teo, J.C. Non-invasive single-cell biomechanical analysis using live-imaging datasets. J. Cell Sci. 2016, 129, 3351–3364. [Google Scholar] [CrossRef] [PubMed]
- Zanier, E.R.; Fumagalli, S.; Perego, C.; Pischiutta, F.; De Simoni, M.G. Shape descriptors of the “never resting” microglia in three different acute brain injury models in mice. Intensive Care Med. Exp. 2015, 3, 39. [Google Scholar] [CrossRef] [PubMed]
- Comin, C.H.; Xu, X.; Wang, Y.; Costa Lda, F.; Yang, Z. An image processing approach to analyze morphological features of microscopic images of muscle fibers. Comput. Med. Imaging Graph. 2014, 38, 803–814. [Google Scholar] [CrossRef] [PubMed]
- Dahl, K.N.; Ribeiro, A.J.; Lammerding, J. Nuclear shape, mechanics, and mechanotransduction. Circ. Res. 2008, 102, 1307–1318. [Google Scholar] [CrossRef] [PubMed]
- Lozoya, O.A.; Gilchrist, C.L.; Guilak, F. Universally Conserved Relationships between Nuclear Shape and Cytoplasmic Mechanical Properties in Human Stem Cells. Sci. Rep. 2016, 6, 23047. [Google Scholar] [CrossRef] [PubMed]
- Liu, E.; Gordonov, S.; Treiser, M.D.; Moghe, P.V. Parsing the early cytoskeletal and nuclear organizational cues that demarcate stem cell lineages. Cell Cycle 2010, 9, 2108–2117. [Google Scholar] [CrossRef] [PubMed]
- Vega, S.L.; Liu, E.; Arvind, V.; Bushman, J.; Sung, H.J.; Becker, M.L.; Lelievre, S.; Kohn, J.; Vidi, P.A.; Moghe, P.V. High-content image informatics of the structural nuclear protein NuMA parses trajectories for stem/progenitor cell lineages and oncogenic transformation. Exp. Cell Res. 2017, 351, 11–23. [Google Scholar] [CrossRef] [PubMed]
- Vega, S.L.; Liu, E.; Patel, P.J.; Kulesa, A.B.; Carlson, A.L.; Ma, Y.; Becker, M.L.; Moghe, P.V. High-content imaging-based screening of microenvironment-induced changes to stem cells. J. Biomol. Screen 2012, 17, 1151–1162. [Google Scholar] [CrossRef] [PubMed]
- Nikkhah, M.; Edalat, F.; Manoucheri, S.; Khademhosseini, A. Engineering microscale topographies to control the cell-substrate interface. Biomaterials 2012, 33, 5230–5246. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Chen, C.S.; Fu, J. Forcing stem cells to behave: A biophysical perspective of the cellular microenvironment. Annu. Rev. Biophys. 2012, 41, 519–542. [Google Scholar] [CrossRef] [PubMed]
- Brun, J.; Abruzzese, T.; Rolauffs, B.; Aicher, W.K.; Hart, M.L. Choice of xenogenic-free expansion media significantly influences the myogenic differentiation potential of human bone marrow-derived mesenchymal stromal cells. Cytotherapy 2016, 18, 344–359. [Google Scholar] [CrossRef] [PubMed]
- Brun, J.; Lutz, K.A.; Neumayer, K.M.; Klein, G.; Seeger, T.; Uynuk-Ool, T.; Worgotter, K.; Schmid, S.; Kraushaar, U.; Guenther, E.; et al. Smooth Muscle-Like Cells Generated from Human Mesenchymal Stromal Cells Display Marker Gene Expression and Electrophysiological Competence Comparable to Bladder Smooth Muscle Cells. PLoS ONE 2015, 10, e0145153. [Google Scholar] [CrossRef] [PubMed]
- Geiger, B.; Spatz, J.P.; Bershadsky, A.D. Environmental sensing through focal adhesions. Nat. Rev. Mol. Cell Biol. 2009, 10, 21–33. [Google Scholar] [CrossRef] [PubMed]
- Engler, A.J.; Sen, S.; Sweeney, H.L.; Discher, D.E. Matrix elasticity directs stem cell lineage specification. Cell 2006, 126, 677–689. [Google Scholar] [CrossRef] [PubMed]
- Khalili, A.A.; Ahmad, M.R. A Review of Cell Adhesion Studies for Biomedical and Biological Applications. Int. J. Mol. Sci. 2015, 16, 18149–18184. [Google Scholar] [CrossRef] [PubMed]
- Pincus, Z.; Theriot, J.A. Comparison of quantitative methods for cell-shape analysis. J. Microsc. 2007, 227, 140–156. [Google Scholar] [CrossRef] [PubMed]
- Meijering, E.; Dzyubachyk, O.; Smal, I.; Van Cappellen, W.A. Tracking in cell and developmental biology. Semin. Cell Dev. Biol. 2009, 20, 894–902. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Yu, S.; Yang, Y.; Xie, Y. Feature and contrast enhancement of mammographic image based on multiscale analysis and morphology. Comput. Math. Methods Med. 2013, 2013, 716948. [Google Scholar] [CrossRef] [PubMed]
- Jirapatnakul, A.C.; Fotin, S.V.; Reeves, A.P.; Biancardi, A.M.; Yankelevitz, D.F.; Henschke, C.I. Automated nodule location and size estimation using a multi-scale Laplacian of Gaussian filtering approach. Conf. Proc. IEEE Eng. Med. Biol. Soc. 2009, 2009, 1028–1031. [Google Scholar] [PubMed]
- Kong, H.; Akakin, H.C.; Sarma, S.E. A generalized Laplacian of Gaussian filter for blob detection and its applications. IEEE Trans. Cybern. 2013, 43, 1719–1733. [Google Scholar] [CrossRef] [PubMed]
- Meijering, E. Cell Segmentation: 50 Years Down the Road. IEEE Signal Process. Mag. 2012, 29, 140–145. [Google Scholar] [CrossRef]
- Krause, M.; Gautreau, A. Steering cell migration: Lamellipodium dynamics and the regulation of directional persistence. Nat. Rev. Mol. Cell Biol. 2014, 15, 577–590. [Google Scholar] [CrossRef] [PubMed]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH Image to ImageJ: 25 Years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Schindelin, J.; Rueden, C.T.; Hiner, M.C.; Eliceiri, K.W. The ImageJ ecosystem: An open platform for biomedical image analysis. Mol. Reprod. Dev. 2015, 82, 518–529. [Google Scholar] [CrossRef] [PubMed]
- Shapiro, W.L.; Anglin, L.W. Metamorph: Computer Support for Qualitative Research. Mid-West. Educ. Res. 1993, 6, 30–34. [Google Scholar]
- Schindelin, J.; Arganda-Carreras, I.; Frise, E.; Kaynig, V.; Longair, M.; Pietzsch, T.; Preibisch, S.; Rueden, C.; Saalfeld, S.; Schmid, B.; et al. Fiji: An open-source platform for biological-image analysis. Nat. Methods 2012, 9, 676–682. [Google Scholar] [CrossRef] [PubMed]
- Soliman, K. CellProfiler: Novel Automated Image Segmentation Procedure for Super-Resolution Microscopy. Biol. Proced. Online 2015, 17, 11. [Google Scholar] [CrossRef] [PubMed]
- Sommer, C.; Gerlich, D.W. Machine learning in cell biology—Teaching computers to recognize phenotypes. J. Cell Sci. 2013, 126, 5529–5539. [Google Scholar] [CrossRef] [PubMed]
- Grys, B.T.; Lo, D.S.; Sahin, N.; Kraus, O.Z.; Morris, Q.; Boone, C.; Andrews, B.J. Machine learning and computer vision approaches for phenotypic profiling. J. Cell Biol. 2017, 216, 65–71. [Google Scholar] [CrossRef] [PubMed]
- Logan, D.J.; Shan, J.; Bhatia, S.N.; Carpenter, A.E. Quantifying co-cultured cell phenotypes in high-throughput using pixel-based classification. Methods 2016, 96, 6–11. [Google Scholar] [CrossRef] [PubMed]
- Shan, J.; Schwartz, R.E.; Ross, N.T.; Logan, D.J.; Thomas, D.; Duncan, S.A.; North, T.E.; Goessling, W.; Carpenter, A.E.; Bhatia, S.N. Identification of small molecules for human hepatocyte expansion and iPS differentiation. Nat. Chem. Biol. 2013, 9, 514–520. [Google Scholar] [CrossRef] [PubMed]
- Hilsenbeck, O.; Schwarzfischer, M.; Loeffler, D.; Dimopoulos, S.; Hastreiter, S.; Marr, C.; Theis, F.J.; Schroeder, T. fastER: A user-friendly tool for ultrafast and robust cell segmentation in large-scale microscopy. Bioinformatics 2017, 33, 2020–2028. [Google Scholar] [CrossRef] [PubMed]
- Juneau, P.M.; Garnier, A.; Duchesne, C. Monitoring of adherent live cells morphology using the undecimated wavelet transform multivariate image analysis (UWT-MIA). Biotechnol. Bioeng. 2017, 114, 141–153. [Google Scholar] [CrossRef] [PubMed]
- Chowdhury, S.; Kandhavelu, M.; Yli-Harja, O.; Ribeiro, A.S. Cell segmentation by multi-resolution analysis and maximum likelihood estimation (MAMLE). BMC Bioinform. 2013, 14, S8. [Google Scholar]
- Jaccard, N.; Griffin, L.D.; Keser, A.; Macown, R.J.; Super, A.; Veraitch, F.S.; Szita, N. Automated method for the rapid and precise estimation of adherent cell culture characteristics from phase contrast microscopy images. Biotechnol. Bioeng. 2014, 111, 504–517. [Google Scholar] [CrossRef] [PubMed]
- McBeath, R.; Pirone, D.M.; Nelson, C.M.; Bhadriraju, K.; Chen, C.S. Cell shape, cytoskeletal tension, and RhoA regulate stem cell lineage commitment. Dev. Cell 2004, 6, 483–495. [Google Scholar] [CrossRef]
- Newman, P.; Galenano-Nino, J.L.; Graney, P.; Razal, J.M.; Minett, A.I.; Ribas, J.; Ovalle-Robles, R.; Biro, M.; Zreiqat, H. Relationship between nanotopographical alignment and stem cell fate with live imaging and shape analysis. Sci. Rep. 2016, 6, 37909. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Abdeen, A.A.; Kilian, K.A. Rewiring mesenchymal stem cell lineage specification by switching the biophysical microenvironment. Sci. Rep. 2014, 4, 5188. [Google Scholar] [CrossRef] [PubMed]
- Costa-Almeida, R.; Carvalho, D.T.; Ferreira, M.J.; Aresta, G.; Gomes, M.E.; van Loon, J.J.W.A.; Van der Heiden, K.; Granja, P.L. Effects of hypergravity on the angiogenic potential of endothelial cells. J. R. Soc. Interface 2016, 13. [Google Scholar] [CrossRef] [PubMed]
| Cell Type/Tissue | Sp. | Shape Inducer | Correlation to Biological Function | Shape Visualization | Tool | Shape Descriptors | Ref |
|---|---|---|---|---|---|---|---|
| MSCs | Hu | Biomaterial stiffness and type, sinusoidal cyclic stretch | SMC differentiation | Calcein staining | ImageJ | Aspect ratio, circularity, roundness, solidity | [19] |
| MSCs | Hu | Biomaterial stiffness and type | SMC differentiation | Calcein staining | ImageJ | Aspect ratio, circularity, major axis, roundness, solidity | [18] |
| MSCs | Hu | Contraction stimulant | SMC contraction | Calcein staining | ImageJ | Length | [38] |
| MSCs | Rat | Hyper-gravity, nanoparticles | Osteogenic differentiation | Coomassie brilliant blue staining | ImageJ | Area, circularity, convex area, roundness, solidity | [16] |
| MSCs | Hu | Differentiation media | Osteogenic differentiation | Phase-contrast microscopy using BioStation CT | MetaMorph | Breadth, elliptical form factor, fiber breadth, fiber length, hole area, inner radius, relative hole area, shape factor, total area | [26] |
| MSCs | Hu | Expansion and Differentiation media | Osteogenic differentiation and mineralization | FITC maleimide for cell shape visualization and Hoechst staining for nuclear morphology | CellProfiler | Area, compactness, eccentricity, Euler number, extent, form factor, major axis, minor axis, orientation, perimeter, solidity, 30 Zernike shape features from orders 0 to 9 | [25] |
| MSCs | Hu | Differentiation media, biomaterial type, beads | Osteo-, adipo-, chondro- and myogenic differentiation | Phase-contrast microscopy using time-lapse | Not given | Area, eccentricity, extent, finger (filopodia), major axis, minor axis | [27] |
| MSCs | Hu | Expansion and differentiation media | Osteo-, adipo- and chondrogenic differentiation, population doubling time | Phase-contrast microscopy | Meta-Morph | Area without holes, breadth, elliptical form factor, fiber breadth, fiber length, hole area, inner radius, length, perimeter, relative hole area, shape factor, total area | [23] |
| MSCs | Hu | Expansion and differentiation media | Osteo- and adipogenic differentiation (classification of stem cell subpopulations) | IF staining of NuMA protein and phalloidin staining of F-actin | Image Pro Plus | Angle, area, area/box, aspect, box height, box ratio, box width, cell area/total area, convex perimeter, dendrites, dendritic length, elliptical perimeter, end points, fractal dimension, length, major axis, maximum diameter, maximum Feret length, maximum radius, mean diameter, mean Feret length, minimum diameter, minimum Feret length, minimum radius, minor axis, perimeter ratio, perimeter, polygonal area, radius ratio, roundness, width. | [33] |
| MSCs, Mu CD8+ T-cells | Hu/Mu | Biomaterial type, beads, expansion and differentiation media | Osteogenic differentiation of hMSCs, naive vs. stimulated phenotype of murine T-cells | Phase-contrast microscopy using live-cell imaging system | MAT-LAB | Area, aspect ratio, cell centroid, Δcell aspect ratio, Δcellular area, migration speed, perimeter | [28] |
| MSCs, immort. MSCs, Rat OPCs | Hu/Rat | Biomaterial type, treatment with nickel(II) sulfate, differen-tiation media | Osteo- and adipogenic differentiation of MSCs, OPC differentiation to astrocytes, cancer-mitigation of biomaterials | CellLight Nucleus-GFP, DAPI staining and IF staining of NuMA for nuclear morphology | Image Pro Plus | Same as in ref. [33] | [34] |
| MSCs, ESCs, iPSCs | Hu | Biomaterial type, self-assembled monolayers (SAM), differentiation media | Osteo-, adipogenic and neural differentiation | DAPI staining and IF staining of NuMA for nuclear morphology | MAT-LAB | Same as in ref. [33] | [35] |
| MSCs, iPSCs | Hu | Biomaterial type, biomaterial stiffness, micropatterning, intracellular magnetic beads | Homeostatic mechanical counterbalance between nuclear shape and perinuclear cytoskeleton architecture | eGFP-actin fusion protein for cell morphology, AlexaFluor 568-tagged beads, Hoechst staining for nuclear morphology and phalloidin staining of F-actin for stress fiber localization | Not given | Angle to bead from the major axis, distance to bead from the nuclear centroid, nuclear elliptical shape (nuclear major axis, nuclear minor axis) | [32] |
| APCs | Hu | Biomaterial topography | Adipo- and osteogenic differentiation | APCs stably express LifeAct-GFP and CAAX-mCherry | Cell-Profiler | Area, circularity, major axis, minor axis | [64] |
| PSC lines, ESC line | Hu | Cultivation/ expansion | Colony morphology for single-colony selection | Phase-contrast microscopy using live-cell imaging system | CL-Quant | Area, compactness, equivalent radius, Fourier descriptors 0–19, inner radius, perimeter, rod-like width, shape factor | [22] |
| Brain: microglia, macro-phages | Mu | Transient and permanent occlusion of middle cerebral artery, traumatic brain injury | Acute brain injury | IF staining of CD11b and CD45 | Fiji | Area, aspect ratio, circularity, Feret’s diameter (caliper), perimeter, solidity | [29] |
| Primary myoblasts | Mu | Differentiation medium | Muscular fiber health | DAPI staining for nuclear morphology and IF staining of myosin heavy chain | Not given | Fiber density, fraction of fiber area, nuclei density, nuclei per fiber area, total fiber length | [30] |
| Primary myoblasts | Hu | Cultivation/expansion | Cellular physiology | Phase-contrast microscopy using live-cell imaging system/ MetaMorph | MAT-LAB (UWT-MIA) | Major axis, minor axis, orientation, roundness | [60] |
| J2-3T3 fibroblasts, primary hepato-cytes | Hu/Mu | Co-culturing of fibroblasts and hepatocytes | Simulation of native microenviron-ment | Hoechst staining for nuclear morphology | Cell-Profiler (Analyst) ilastik | Eccentricity, major axis, perimeter | [57] |
| Hu/Mu | Functional proliferation | Hoechst staining for nuclear morphology | Cell-Profiler (Analyst) | Eccentricity, major axis, perimeter | [58] | ||
| HUVECs | Hu | Hypergravity | Cytoskeleton organization | IF staining of β-tubulin, Phalloidin staining of F-actin | ImageJ | Area, circularity, roundness, solidity | [66] |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hart, M.L.; Lauer, J.C.; Selig, M.; Hanak, M.; Walters, B.; Rolauffs, B. Shaping the Cell and the Future: Recent Advancements in Biophysical Aspects Relevant to Regenerative Medicine. J. Funct. Morphol. Kinesiol. 2018, 3, 2. https://doi.org/10.3390/jfmk3010002
Hart ML, Lauer JC, Selig M, Hanak M, Walters B, Rolauffs B. Shaping the Cell and the Future: Recent Advancements in Biophysical Aspects Relevant to Regenerative Medicine. Journal of Functional Morphology and Kinesiology. 2018; 3(1):2. https://doi.org/10.3390/jfmk3010002
Chicago/Turabian StyleHart, Melanie L., Jasmin C. Lauer, Mischa Selig, Martha Hanak, Brandan Walters, and Bernd Rolauffs. 2018. "Shaping the Cell and the Future: Recent Advancements in Biophysical Aspects Relevant to Regenerative Medicine" Journal of Functional Morphology and Kinesiology 3, no. 1: 2. https://doi.org/10.3390/jfmk3010002
APA StyleHart, M. L., Lauer, J. C., Selig, M., Hanak, M., Walters, B., & Rolauffs, B. (2018). Shaping the Cell and the Future: Recent Advancements in Biophysical Aspects Relevant to Regenerative Medicine. Journal of Functional Morphology and Kinesiology, 3(1), 2. https://doi.org/10.3390/jfmk3010002