Fish Diversity Monitored by Environmental DNA in the Yangtze River Mainstream
Abstract
:1. Introduction
2. Materials and Methods
2.1. Sample Sites and Collection
2.2. DNA Extraction and PCR
2.3. Library Preparation and Sequencing
2.4. Bioinformatic Statistics and Fish Identification
2.5. Seasonal Difference Comparison
3. Results
3.1. Fish Resource Results by eDNA
3.2. Diversity of Fish Communities
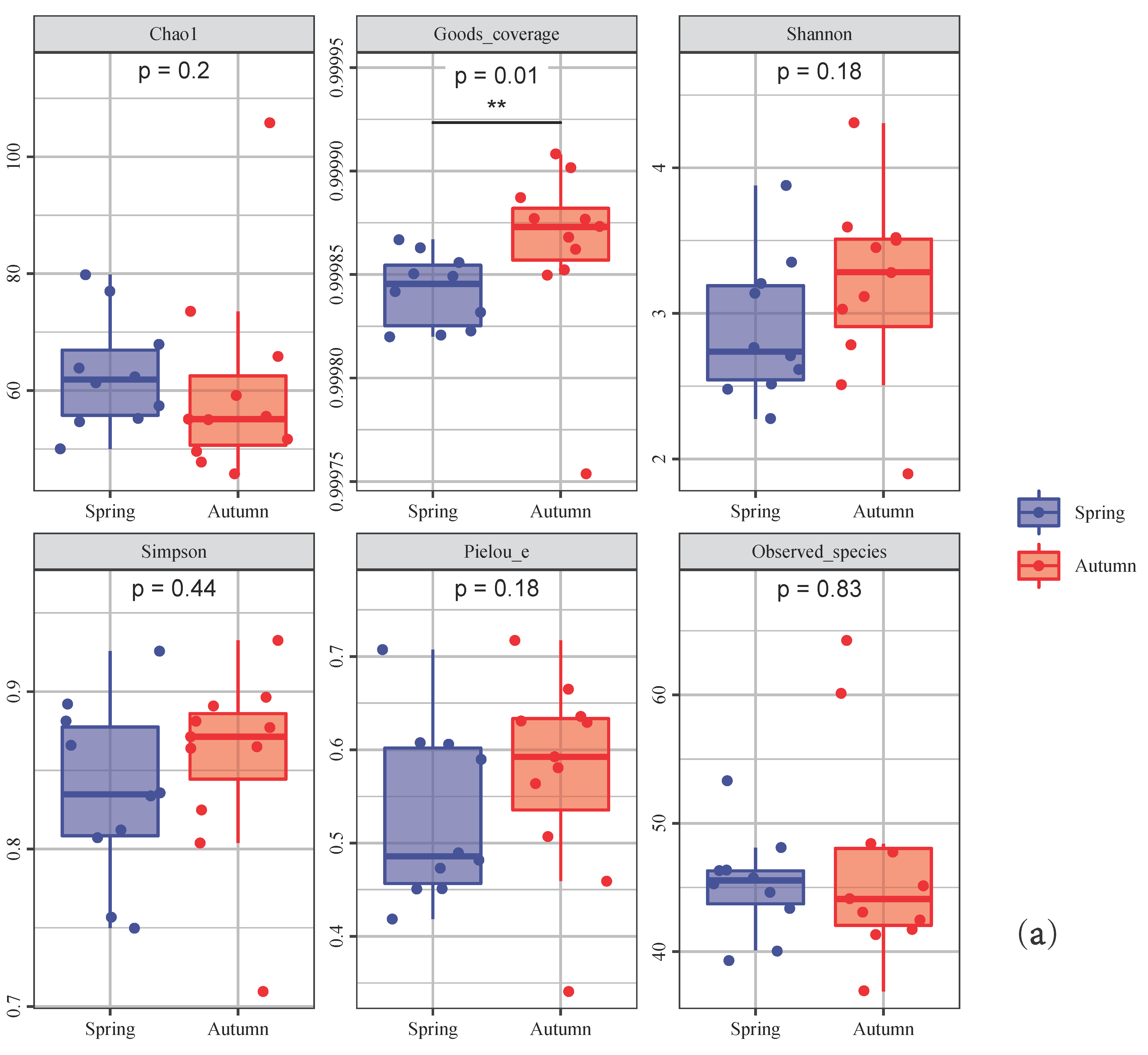
3.3. Seasonal Variation Analysis
3.4. Relationship with Environmental Factors
4. Discussion
4.1. Fish Diversity in the Mainstream of the Yangtze River Based on eDNA
- (1)
- Insufficient sample collection: The fish activities in the Yangtze River mainstream are relatively large; in the present study, we only set 11 sampling points in the Yangtze River mainstream, which may have led to insufficient sampling. Ficetola et al. (2015) pointed out that replication, i.e., the number of water samples collected as opposed to the number of technical replicates run from a single sample, can be adjusted to compensate for false negatives [53]. Therefore, adding sampling points according to the study area would increase the possibility of detecting fish.
- (2)
- Less eDNA preservation: The amount of eDNA secreted by fish is related to itself; studies have shown that different species secrete DNA at different rates and quantities [54]. Moreover, the preservation of eDNA is also very important, as the long transportation after sampling, temperature, time and other factors will affect the preservation effect of the eDNA experiment. In addition, delayed extraction from the Sterivex filter may also result in less DNA being extracted. Thus, special attention should be paid to the preservation of samples, and timely filtration after sampling will reduce the degradation rate of eDNA.
- (3)
- Lack of local reference database information: This study did not establish a more accurate local reference database for fish in the Yangtze River mainstream, which may have reduced the number of native species identified. The effectiveness of eDNA depends on the quality of the reference sequence database and the classification parameters employed [55]. In future investigations and studies, we will consider establishing a local database. At the same time, the results of this survey will also serve as a reference for future surveys of fish in the Yangtze River mainstream.
- (4)
- Inapplicability of 12S primer to freshwater fish: The primers that we used may not be suitable for the identification of fish in the mainstream of the Yangtze River, which could explain the small number of identification results. Many studies have used other primers, such as the mitochondrial cytochrome c oxidase subunit I (COI) sequence and the 16S primer for the fish amplification [56]. In future studies, different primers can be designed to explore primers that are more suitable for Yangtze River fish. In the non-closed fishing season, fish body muscle tissue can be used for primer design, which will be our next research direction.
4.2. Seasonal Variation between Spring and Autumn
4.3. Relationship between Fish Assemblage and Environmental Factors
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Shen, Y.; Hubert, N.; Huang, Y.; Wang, X.; Gan, X.; Peng, Z.; He, S. DNA barcoding the ichthyofauna of the Yangtze River: Insights from the molecular inventory of a mega-diverse temperate fauna. Mol. Ecol. Resour. 2019, 19, 1278–1291. [Google Scholar] [CrossRef]
- Qu, C.; Stewart, K.A.; Clemente-Carvalho, R.; Zheng, J.; Wang, Y.; Gong, C.; Ma, L.; Zhao, J.; Lougheed, S.C. Comparing fish prey diversity for a critically endangered aquatic mammal in a reserve and the wild using eDNA metabarcoding. Sci. Rep. 2020, 10, 16715. [Google Scholar] [CrossRef] [PubMed]
- Eble, J.A.; Daly-Engel, T.S.; DiBattista, J.D.; Koziol, A.; Gaither, M.R. Marine environmental DNA: Approaches, applications, and opportunities. Adv. Mar. Biol. 2020, 86, 141–169. [Google Scholar] [CrossRef]
- Zou, K.; Chen, J.; Ruan, H.; Li, Z.; Guo, W.; Li, M.; Liu, L. eDNA metabarcoding as a promising conservation tool for monitoring fish diversity in a coastal wetland of the Pearl River Estuary compared to bottom trawling. Sci. Total Environ. 2020, 702, 134704. [Google Scholar] [CrossRef] [PubMed]
- Ogram, A.; Sayler, G.S.; Barkay, T. The extraction and purification of microbial DNA from sediments. J. Microbiol. Methods 1987, 7, 57–66. [Google Scholar] [CrossRef]
- Rondon, M.R.; August, P.R.; Bettermann, A.D.; Brady, S.F.; Grossman, T.H.; Liles, M.R.; Loiacono, K.A.; Lynch, B.A.; MacNeil, I.A.; Minor, C.; et al. Cloning the soil metagenome: A strategy for accessing the genetic and functional diversity of uncultured microorganisms. Appl. Environ. Microbiol. 2000, 66, 2541–2547. [Google Scholar] [CrossRef] [Green Version]
- Barnes, M.A.; Turner, C.R. The ecology of environmental DNA and implications for conservation genetics. Conserv. Genet. 2016, 17, 1–17. [Google Scholar] [CrossRef] [Green Version]
- Castañeda, R.A.; Van Nynatten, A.; Crookes, S.; Ellender, B.R.; Heath, D.D.; MacIsaac, H.J.; Mandrak, N.E.; Weyl, O.L.F. Detecting Native Freshwater Fishes Using Novel Non-invasive Methods. Front. Environ. Sci. 2020, 8, 29. [Google Scholar] [CrossRef]
- Hering, D.; Borja, A.; Jones, J.I.; Pont, D.; Boets, P.; Bouchez, A.; Bruce, K.; Drakare, S.; Hänfling, B.; Kahlert, M.; et al. Implementation options for DNA-based identification into ecological status assessment under the European Water Framework Directive. Water Res. 2018, 138, 192–205. [Google Scholar] [CrossRef]
- Ficetola, G.F.; Miaud, C.; Pompanon, F.; Taberlet, P. Species detection using environmental DNA from water samples. Biol. Lett. 2008, 4, 423–425. [Google Scholar] [CrossRef] [Green Version]
- Dejean, T.; Valentini, A.; Miquel, C.; Taberlet, P.; Bellemain, E.; Miaud, C. Improved detection of an alien invasive species through environmental DNA barcoding: The example of the American bullfrogLithobates catesbeianus. J. Appl. Ecol. 2012, 49, 953–959. [Google Scholar] [CrossRef]
- Jerde, C.L.; Mahon, A.R.; Chadderton, W.L.; Lodge, D.M. “Sight-unseen” detection of rare aquatic species using environmental DNA. Conserv. Lett. 2011, 4, 150–157. [Google Scholar] [CrossRef]
- Goldberg, C.S.; Sepulveda, A.; Ray, A.; Baumgardt, J.; Waits, L.P. Environmental DNA as a new method for early detection of New Zealand mudsnails (Potamopyrgus antipodarum). Freshw. Sci. 2013, 32, 792–800. [Google Scholar] [CrossRef] [Green Version]
- Geerts, A.N.; Boets, P.; Van den Heede, S.; Goethals, P.; Van der heyden, C. A search for standardized protocols to detect alien invasive crayfish based on environmental DNA (eDNA): A lab and field evaluation. Ecol. Indic. 2018, 84, 564–572. [Google Scholar] [CrossRef]
- Egan, S.P.; Grey, E.; Olds, B.; Feder, J.L.; Ruggiero, S.T.; Tanner, C.E.; Lodge, D.M. Rapid Molecular Detection of Invasive Species in Ballast and Harbor Water by Integrating Environmental DNA and Light Transmission Spectroscopy. Environ. Sci. Technol. 2015, 49, 4113–4121. [Google Scholar] [CrossRef] [Green Version]
- Xia, Z.; Zhan, A.; Gao, Y.; Zhang, L.; Haffner, G.D.; MacIsaac, H.J. Early detection of a highly invasive bivalve based on environmental DNA (eDNA). Biol. Invasions 2017, 20, 437–447. [Google Scholar] [CrossRef]
- Simpfendorfer, C.A.; Kyne, P.M.; Noble, T.H.; Goldsbury, J.; Basiita, R.K.; Lindsay, R.; Jerry, D.R. Environmental DNA detects critically endangered largetooth sawfish in the wild. Endanger. Species Res. 2016, 30, 109–116. [Google Scholar] [CrossRef] [Green Version]
- Weltz, K.; Lyle, J.M.; Ovenden, J.; Morgan, J.A.T.; Moreno, D.A.; Semmens, J.M. Application of environmental DNA to detect an endangered marine skate species in the wild. PLoS ONE 2017, 12, e0178124. [Google Scholar] [CrossRef]
- Mauvisseau, Q.; Davy-Bowker, J.; Bulling, M.; Brys, R.; Neyrinck, S.; Troth, C.; Sweet, M. Combining ddPCR and environmental DNA to improve detection capabilities of a critically endangered freshwater invertebrate. Sci. Rep. 2019, 9, 14064. [Google Scholar] [CrossRef] [Green Version]
- Nevers, M.B.; Byappanahalli, M.N.; Morris, C.C.; Shively, D.; Przybyla-Kelly, K.; Spoljaric, A.M.; Dickey, J.; Roseman, E.F. Environmental DNA (eDNA): A tool for quantifying the abundant but elusive round goby (Neogobius melanostomus). PLoS ONE 2018, 13, e0191720. [Google Scholar] [CrossRef]
- Yamanaka, H.; Minamoto, T. The use of environmental DNA of fishes as an efficient method of determining habitat connectivity. Ecol. Indic. 2016, 62, 147–153. [Google Scholar] [CrossRef] [Green Version]
- Sigsgaard, E.E.; Nielsen, I.B.; Bach, S.S.; Lorenzen, E.D.; Robinson, D.P.; Knudsen, S.W.; Pedersen, M.W.; Jaidah, M.A.; Orlando, L.; Willerslev, E.; et al. Population characteristics of a large whale shark aggregation inferred from seawater environmental DNA. Nat. Ecol. Evol. 2016, 1, 4. [Google Scholar] [CrossRef]
- Bakker, J.; Wangensteen, O.S.; Chapman, D.D.; Boussarie, G.; Buddo, D.; Guttridge, T.L.; Hertler, H.; Mouillot, D.; Vigliola, L.; Mariani, S. Environmental DNA reveals tropical shark diversity in contrasting levels of anthropogenic impact. Sci. Rep. 2017, 7, 16886. [Google Scholar] [CrossRef]
- Boussarie, G.; Bakker, J.; Wangensteen, O.S.; Mariani, S.; Bonnin, L.; Juhel, J.B.; Kiszka, J.J.; Kulbicki, M.; Manel, S.; Robbins, W.D.; et al. Environmental DNA illuminates the dark diversity of sharks. Sci. Adv. 2018, 4, eaap9661. [Google Scholar] [CrossRef] [Green Version]
- Thalinger, B.; Wolf, E.; Traugott, M.; Wanzenböck, J. Monitoring spawning migrations of potamodromous fish species via eDNA. Sci. Rep. 2019, 9, 15388. [Google Scholar] [CrossRef] [Green Version]
- Cowart, D.A.; Breedveld, K.G.H.; Ellis, M.J.; Hull, J.M.; Larson, E.R. Environmental DNA (eDNA) applications for the conservation of imperiled crayfish (Decapoda: Astacidea) through monitoring of invasive species barriers and relocated populations. J. Crustacean Biol. 2018, 38, 257–266. [Google Scholar] [CrossRef]
- Port, J.A.; O’Donnell, J.L.; Romero-Maraccini, O.C.; Leary, P.R.; Litvin, S.Y.; Nickols, K.J.; Kelly, R.P. Assessing vertebrate biodiversity in a kelp forest ecosystem using environmental DNA. Mol. Ecol. 2015, 25, 527–541. [Google Scholar] [CrossRef]
- Forsström, T.; Vasemägi, A. Can environmental DNA (eDNA) be used for detection and monitoring of introduced crab species in the Baltic Sea? Mar. Pollut. Bull. 2016, 109, 350–355. [Google Scholar] [CrossRef]
- Lanzén, A.; Lekang, K.; Jonassen, I.; Thompson, E.M.; Troedsson, C. High-throughput metabarcoding of eukaryotic diversity for environmental monitoring of offshore oil-drilling activities. Mol. Ecol. 2016, 25, 4392–4406. [Google Scholar] [CrossRef]
- Stat, M.; Huggett, M.J.; Bernasconi, R.; DiBattista, J.D.; Berry, T.E.; Newman, S.J.; Harvey, E.S.; Bunce, M. Ecosystem biomonitoring with eDNA: Metabarcoding across the tree of life in a tropical marine environment. Sci. Rep. 2017, 7, 12240. [Google Scholar] [CrossRef]
- DiBattista, J.D.; Coker, D.J.; Sinclair-Taylor, T.H.; Stat, M.; Berumen, M.L.; Bunce, M. Assessing the utility of eDNA as a tool to survey reef-fish communities in the Red Sea. Coral Reefs 2017, 36, 1245–1252. [Google Scholar] [CrossRef]
- DiBattista, J.D.; Reimer, J.D.; Stat, M.; Masucci, G.D.; Biondi, P.; De Brauwer, M.; Bunce, M. Digging for DNA at depth: Rapid universal metabarcoding surveys (RUMS) as a tool to detect coral reef biodiversity across a depth gradient. PeerJ 2019, 7, e6379. [Google Scholar] [CrossRef] [PubMed]
- Thomsen, P.F.; Willerslev, E. Environmental DNA–An emerging tool in conservation for monitoring past and present biodiversity. Biol. Conserv. 2015, 183, 4–18. [Google Scholar] [CrossRef]
- Jeunen, G.J.; Knapp, M.; Spencer, H.G.; Lamare, M.D.; Taylor, H.R.; Stat, M.; Gemmell, N.J. Environmental DNA (eDNA) metabarcoding reveals strong discrimination among diverse marine habitats connected by water movement. Mol. Ecol. Resour. 2019, 19, 426–438. [Google Scholar] [CrossRef] [PubMed]
- Lacoursière-Roussel, A.; Côté, G.; Leclerc, V.; Bernatchez, L. Quantifying relative fish abundance with eDNA: A promising tool for fisheries management. J. Appl. Ecol. 2016, 53, 1148–1157. [Google Scholar] [CrossRef]
- Kimmerling, N.; Zuqert, O.; Amitai, G.; Gurevich, T.; Armoza-Zvuloni, R.; Kolesnikov, I.; Berenshtein, I.; Melamed, S.; Gilad, S.; Benjamin, S.; et al. Quantitative species-level ecology of reef fish larvae via metabarcoding. Nat. Ecol. Evol. 2018, 2, 306–316. [Google Scholar] [CrossRef] [PubMed]
- West, K.M.; Stat, M.; Harvey, E.S.; Skepper, C.L.; DiBattista, J.D.; Richards, Z.T.; Travers, M.J.; Newman, S.J.; Bunce, M. eDNA metabarcoding survey reveals fine-scale coral reef community variation across a remote, tropical island ecosystem. Mol. Ecol. 2020, 29, 1069–1086. [Google Scholar] [CrossRef]
- Zhang, H.; Yoshizawa, S.; Iwasaki, W.; Xian, W.W. Seasonal Fish Assemblage Structure Using Environmental DNA in the Yangtze Estuary and Its Adjacent Waters. Front. Mar. Sci. 2019, 6, 515. [Google Scholar] [CrossRef]
- Miya, M.; Sato, Y.; Fukunaga, T.; Sado, T.; Poulsen, J.Y.; Sato, K.; Minamoto, T.; Yamamoto, S.; Yamanaka, H.; Araki, H.; et al. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: Detection of more than 230 subtropical marine species. R. Soc. Open Sci. 2015, 2, 150088. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Rognes, T.; Flouri, T.; Nichols, B.; Quince, C.; Mahé, F. VSEARCH: A versatile open source tool for metagenomics. PeerJ 2016, 4, e2584. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt Removes Adapter Sequences from High-Throughput Sequencing Reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Chao, A. Nonparametric Estimation of the Number of Classes in a Population. Scand. J. Stat. 1984, 11, 265–270. [Google Scholar]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 379–656. [Google Scholar] [CrossRef] [Green Version]
- Legovic, T. Statistical ecology. A primer on methods and computing. Ecol. Model. 1991, 54, 143–144. [Google Scholar] [CrossRef]
- Faith, D.P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 1992, 61, 1–10. [Google Scholar] [CrossRef]
- Pielou, E.C. The measurement of diversity in different types of biological collections. J. Theor. Biol. 1966, 13, 131–144. [Google Scholar] [CrossRef]
- Good, I.J. The population frequency of species and theestimation of the population parameters. Biometrics 1958, 40, 237–246. [Google Scholar] [CrossRef]
- Lozupone, C.A.; Hamady, M.; Kelley, S.T.; Knight, R. Quantitative and qualitative beta diversity measures lead to different insights into factors that structure microbial communities. Appl. Environ. Microbiol 2007, 73, 1576–1585. [Google Scholar] [CrossRef] [Green Version]
- Lozupone, C.; Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 2005, 71, 8228–8235. [Google Scholar] [CrossRef] [Green Version]
- Fish Laboratory, Institute of Hydrobiology, Chinese Academy of Sciences. Fishes of Yangtze River; Science Press: Beijing, China, 1976. [Google Scholar]
- Anderson, M.J.; Ellingsen, K.E.; McArdle, B.H. Multivariate dispersion as a measure of beta diversity. Ecol. Lett. 2006, 9, 683–693. [Google Scholar] [CrossRef]
- Ficetola, G.F.; Pansu, J.; Bonin, A.; Coissac, E.; Giguet-Covex, C.; De Barba, M.; Gielly, L.; Lopes, C.M.; Boyer, F.; Pompanon, F.; et al. Replication levels, false presences and the estimation of the presence/absence from eDNA metabarcoding data. Mol. Ecol Resour. 2015, 15, 543–556. [Google Scholar] [CrossRef]
- Russo, T.; Maiello, G.; Talarico, L.; Baillie, C.; Colosimo, G.; D’Andrea, L.; Di Maio, F.; Fiorentino, F.; Franceschini, S.; Garofalo, G.; et al. All is fish that comes to the net: Metabarcoding for rapid fisheries catch assessment. Ecol. Appl. 2021, 31, e02273. [Google Scholar] [CrossRef]
- Gold, Z.; Curd, E.E.; Goodwin, K.D.; Choi, E.S.; Frable, B.W.; Thompson, A.R.; Walker, H.J.; Burton, R.S.; Kacev, D.; Martz, L.D.; et al. Improving metabarcoding taxonomic assignment: A case study of fishes in a large marine ecosystem. Mol. Ecol. Resour. 2021, 21, 2546–2564. [Google Scholar] [CrossRef]
- Zhang, S.; Zhao, J.; Yao, M. A comprehensive and comparative evaluation of primers for metabarcoding eDNA from fish. Methods Ecol. Evol. 2020, 11, 1609–1625. [Google Scholar] [CrossRef]
- Jia, H.; Wang, Y.; Yoshizawa, S.; Iwasaki, W.; Li, Y.; Xian, W.; Zhang, H. Seasonal Variation and Assessment of Fish Resources in the Yangtze Estuary Based on Environmental DNA. Water 2020, 12, 2874. [Google Scholar] [CrossRef]
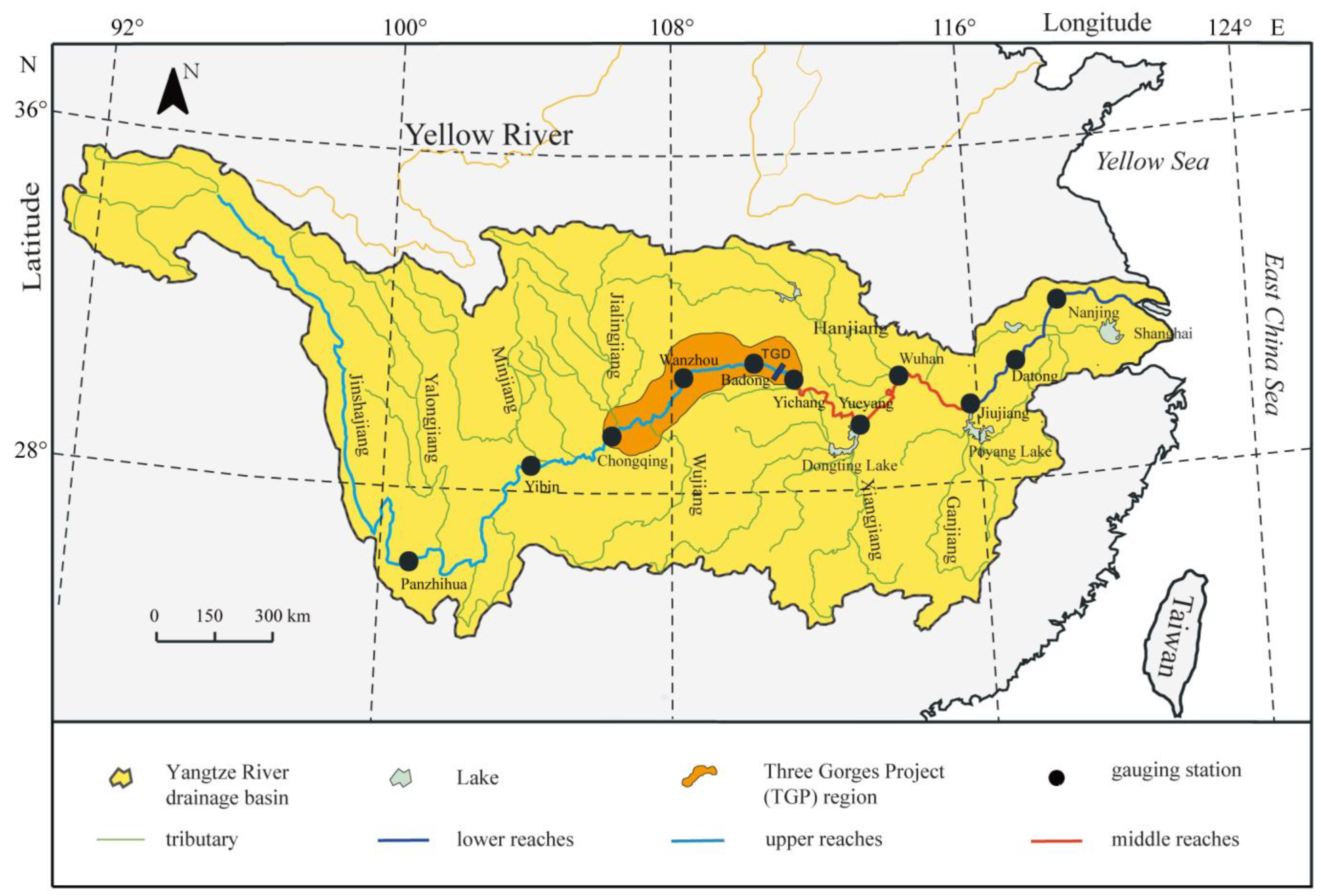




| Number | Station | Longitude | Latitude | Spring | Autumn | ||
|---|---|---|---|---|---|---|---|
| Sampling | Time | Sampling | Time | ||||
| 1 | Panzhihua | 101.70 | 26.56 | C19SE01 | 12 April 2019 | C19AE01 | 13 October 2019 |
| 2 | Yibin | 104.66 | 28.77 | C19SE02 | 14 April 2019 | C19AE02 | 15 October 2019 |
| 3 | Chongqing | 106.57 | 29.55 | C19AE03 | 15 October 2019 | ||
| 4 | Wanzhou | 108.43 | 30.76 | C19SE03 | 15 April 2019 | C19AE04 | 16 October 2019 |
| 5 | Badong | 110.34 | 31.05 | C19SE04 | 16 April 2019 | C19AE05 | 17 October 2019 |
| 6 | Yichang | 111.40 | 30.57 | C19SE05 | 17 April 2019 | C19AE06 | 18 October 2019 |
| 7 | Yuyang | 113.12 | 29.40 | C19SE06 | 17 April 2019 | C19AE07 | 18 October 2019 |
| 8 | Wuhan | 114.30 | 30.55 | C19SE07 | 18 April 2019 | C19AE08 | 19 October 2019 |
| 9 | Jiujiang | 116.03 | 29.75 | C19SE08 | 19 April 2019 | C19AE09 | 20 October 2019 |
| 10 | Datong | 117.74 | 30.86 | C19SE09 | 20 April 2019 | C19AE10 | 21 October 2019 |
| 11 | Nanjing | 118.75 | 32.12 | C19SE10 | 20 April 2019 | C19AE11 | 22 October 2019 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jia, H.; Zhang, H.; Xian, W. Fish Diversity Monitored by Environmental DNA in the Yangtze River Mainstream. Fishes 2022, 7, 1. https://doi.org/10.3390/fishes7010001
Jia H, Zhang H, Xian W. Fish Diversity Monitored by Environmental DNA in the Yangtze River Mainstream. Fishes. 2022; 7(1):1. https://doi.org/10.3390/fishes7010001
Chicago/Turabian StyleJia, Hui, Hui Zhang, and Weiwei Xian. 2022. "Fish Diversity Monitored by Environmental DNA in the Yangtze River Mainstream" Fishes 7, no. 1: 1. https://doi.org/10.3390/fishes7010001
APA StyleJia, H., Zhang, H., & Xian, W. (2022). Fish Diversity Monitored by Environmental DNA in the Yangtze River Mainstream. Fishes, 7(1), 1. https://doi.org/10.3390/fishes7010001







