Implementation of SCID Screening in Denmark
Abstract
:1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. Samples
2.3. Realtime PCR
2.4. Data Analysis and Cut-Offs
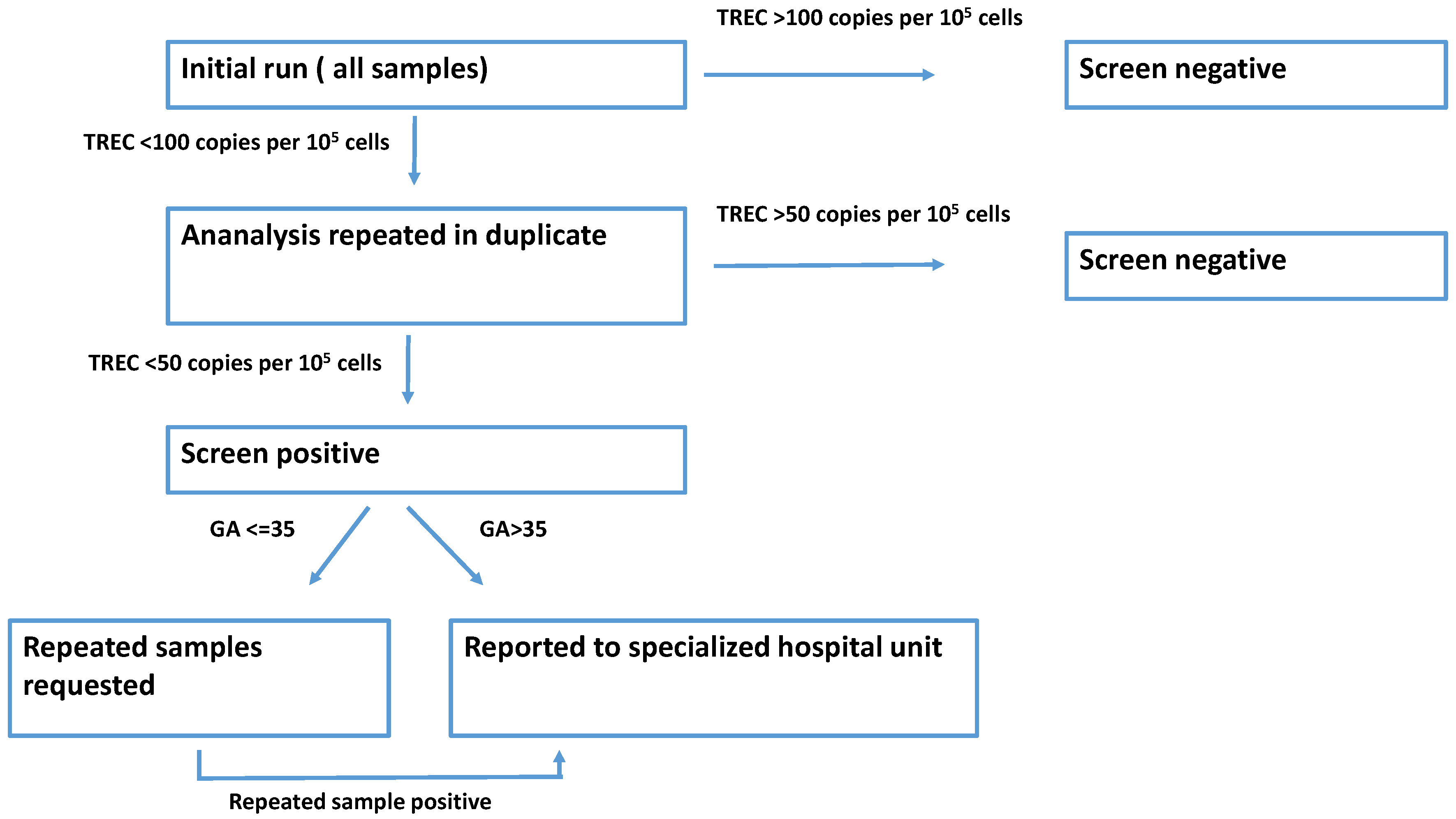
2.5. Screening Algorithm
3. Results
3.1. Validation Study
3.2. Screening
4. Discussion
Author Contributions
Funding
Conflicts of Interest
References
- The Danish Neonatal Screening Webpage. Available online: https://nyfoedte.ssi.dk/ (accessed on 29 July 2021).
- Myers, L.A.; Patel, D.D.; Puck, J.M.; Buckley, R.H. Hematopoietic stem cell transplantation for severe combined immunodeficiency in the neonatal period leads to superior thymic output and improved survival. Blood 2002, 99, 872–878. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- King, J.R.; Hammarström, L. Newborn Screening for Primary Immunodeficiency Diseases: History, Current and Future Practice. J. Clin. Immunol. 2018, 38, 56–66. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buelow, B.J.; Routes, J.M.; Verbsky, J.W. Newborn screening for SCID: Where are we now? Expert Rev. Clin. Immunol. 2014, 10, 1649–1657. [Google Scholar] [CrossRef] [PubMed]
- Lund, A.M.; Hougaard, D.M.; Simonsen, H.; Andresen, B.S.; Christensen, M.; Dunø, M.; Skogstrand, K.; Olsen, R.K.J.; Jensen, U.G.; Cohen, A.; et al. Biochemical screening of 504,049 newborns in Denmark, the Faroe Islands and Greenland—Experience and development of a routine program for expanded newborn screening. Mol. Genet. Metab. 2012, 107, 281–293. [Google Scholar] [CrossRef] [PubMed]
- Nørgaard-Pedersen, B.; Simonsen, H. Biological specimen banks in neonatal screening. Acta Paediatr. Suppl. 1999, 88, 106–109. [Google Scholar] [CrossRef] [PubMed]
- Gutierrez-Mateo, C.; Timonen, A.; Vaahtera, K.; Jaakkola, M.; Hougaard, D.M.; Bybjerg-Grauholm, J.; Baekvad-Hansen, M.; Adamsen, D.; Filippov, G.; Dallaire, S.; et al. Development of a Multiplex real-time PCR Assay for the Newborn Screening of SCID, SMA, and XLA. Int. J. Neonatal Screen. 2019, 5, 39. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Van der Burg, M.; Mahlaoui, N.; Gaspar, H.B.; Pai, S.Y. Universal Newborn Screening for Severe Combined Immunodeficiency (SCID). Front. Pediatr. 2019, 7, 373. [Google Scholar] [CrossRef] [PubMed]
- Routes, J.M. Statewide Newborn Screening for Severe T-Cell Lymphopenia. JAMA 2009, 302, 2465–2470. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Verbsky, J.W.; Baker, M.W.; Grossman, W.J.; Hintermeyer, M.; Dasu, T.; Bonacci, B.; Reddy, S.; Margolis, D.; Casper, J.; Gries, M.; et al. Newborn Screening for Severe Combined Immunodeficiency; The Wisconsin Experience (2008–2011). J. Clin. Immunol. 2012, 32, 82–88. [Google Scholar] [CrossRef] [PubMed]
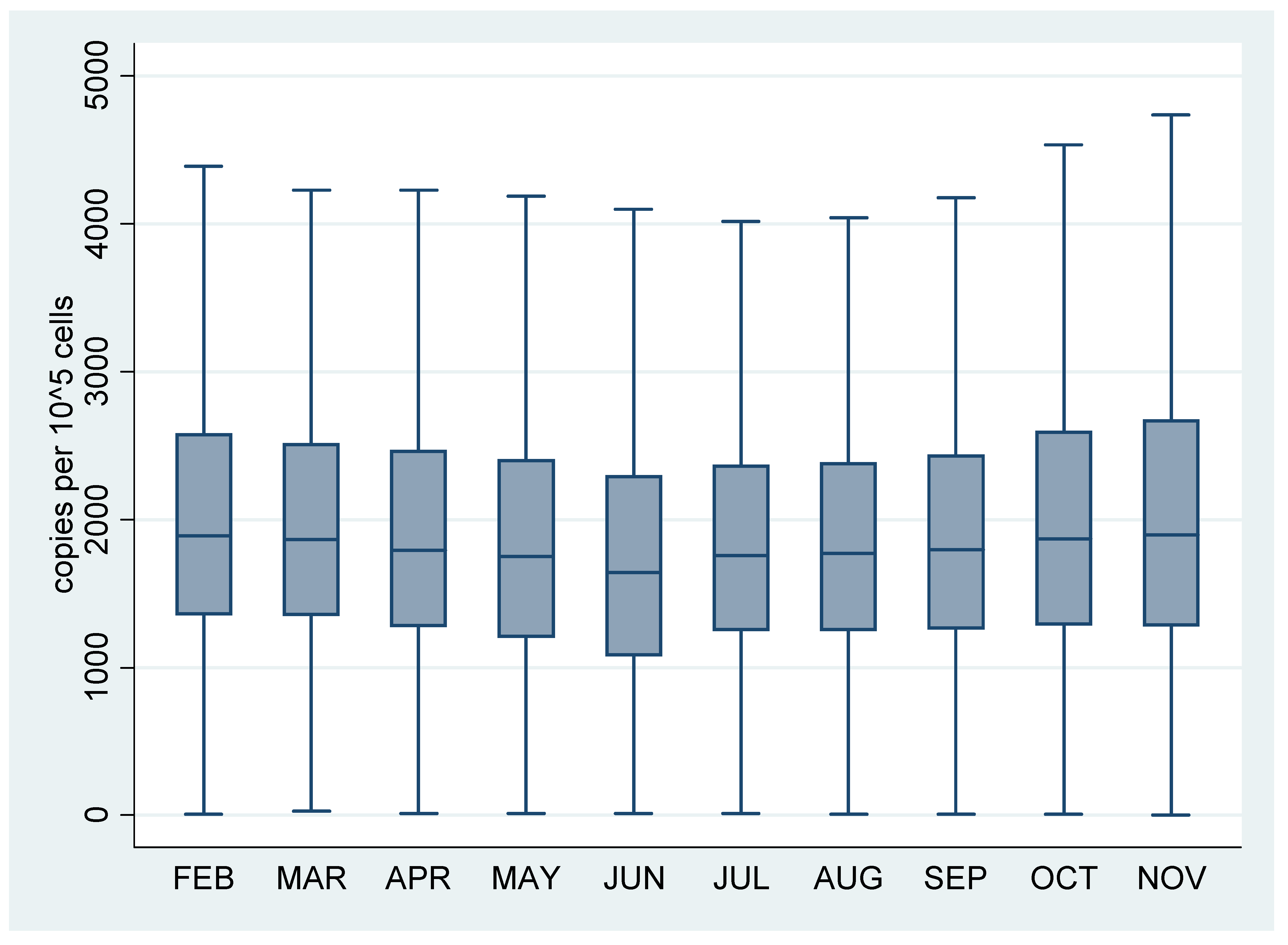
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bækvad-Hansen, M.; Adamsen, D.; Bybjerg-Grauholm, J.; Hougaard, D.M. Implementation of SCID Screening in Denmark. Int. J. Neonatal Screen. 2021, 7, 54. https://doi.org/10.3390/ijns7030054
Bækvad-Hansen M, Adamsen D, Bybjerg-Grauholm J, Hougaard DM. Implementation of SCID Screening in Denmark. International Journal of Neonatal Screening. 2021; 7(3):54. https://doi.org/10.3390/ijns7030054
Chicago/Turabian StyleBækvad-Hansen, Marie, Dea Adamsen, Jonas Bybjerg-Grauholm, and David Michael Hougaard. 2021. "Implementation of SCID Screening in Denmark" International Journal of Neonatal Screening 7, no. 3: 54. https://doi.org/10.3390/ijns7030054
APA StyleBækvad-Hansen, M., Adamsen, D., Bybjerg-Grauholm, J., & Hougaard, D. M. (2021). Implementation of SCID Screening in Denmark. International Journal of Neonatal Screening, 7(3), 54. https://doi.org/10.3390/ijns7030054






