Expanded Screening of One Million Swedish Babies with R4S and CLIR for Post-Analytical Evaluation of Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
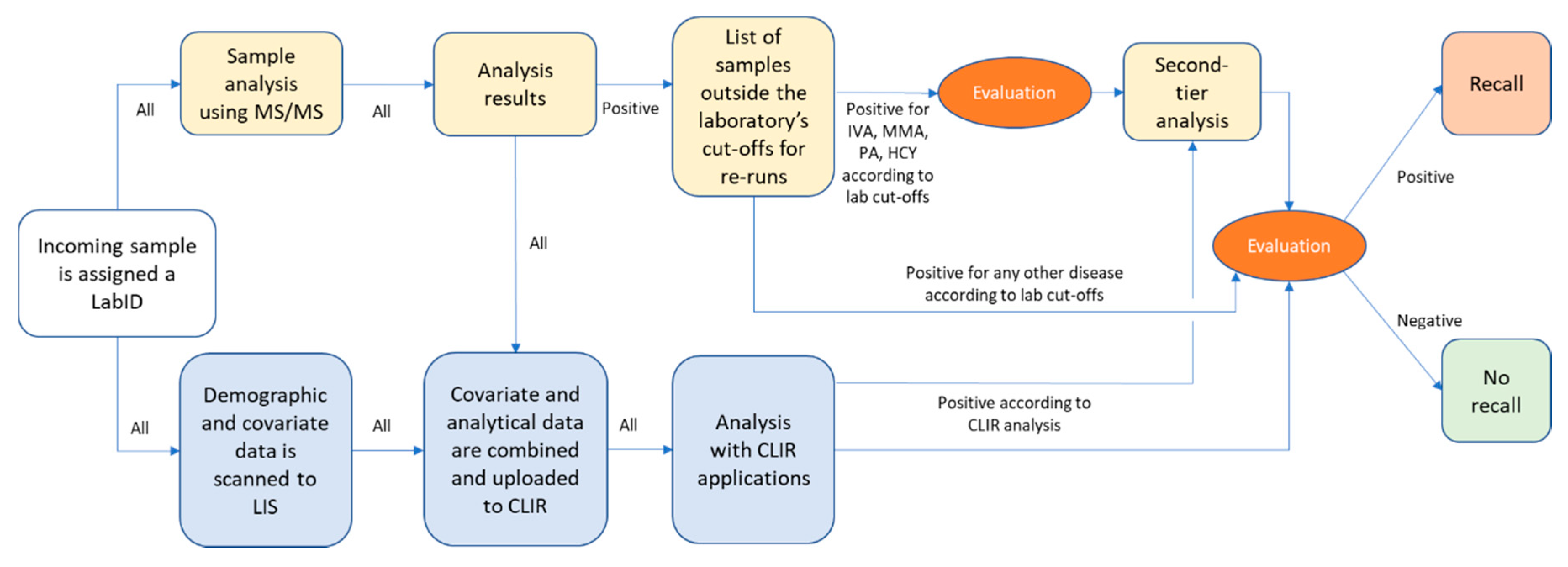
2.2. Screening Laboratory Work-Flow
2.3. MS/MS Screening Method
2.4. Evaluation of Screening Data
2.5. Confirmation of True or False Cases and Feedback on Missed Cases
2.6. Data Collection and Analysis
2.7. Compliance with Ethical Standards
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| ARG | Arginase deficiency |
| ASA | Argininosuccinic aciduria |
| BKT | Betaketothiolase deficiency |
| CACT | Carnitine acylcarnitine translocase deficiency |
| CIT | Citrullinemia type 1 |
| CLIR | Collaborative laboratory integrated reports |
| CPT1 | Carnitine palmitoyl transferase 1 deficiency |
| CPT2 | Carnitine palmitoyl transferase 2 deficiency |
| CUD | Carnitine uptake deficiency |
| DBS | Dried blood spots |
| GA1 | Glutaric aciduria type 1 |
| HCY | Homocystinuria |
| IVA | Isovaleric aciduria |
| LCHAD | Long-chain 3-hydroxyacyl-coenzyme A dehydrogenase deficiency |
| LIS | Laboratory information system |
| MAD | Multiple acyl-coenzyme A dehydrogenase deficiency |
| MCAD | Medium-chain acyl-coenzyme A dehydrogenase deficiency |
| MMA | Methylmalonic aciduria |
| MS/MS | Tandem mass spectrometry |
| MSUD | Maple syrup disease |
| N/A | Not applicable |
| PA | Propionic aciduria |
| PKU | Phenylketonuria |
| PPV | Positive predictive value |
| R4S | Region 4 Stork |
| TYR | Tyrosinemia type 1 |
| VLCAD | Very long-chain acyl-coenzyme A dehydrogenase deficiency |
References
- Millington, D.S.; Kodo, N.; Norwood, D.L.; Roe, C.R. Tandem mass spectrometry: A new method for acylcarnitine profiling with potential for neonatal screening for inborn errors of metabolism. J. Inherit. Metab. Dis. 1990, 13, 321–324. [Google Scholar] [CrossRef] [PubMed]
- Chace, D.H.; Hillman, S.L.; Van Hove, J.L.; Naylor, E.W. Rapid diagnosis of MCAD deficiency: Quantitative analysis of octanoylcarnitine and other acylcarnitines in newborn blood spots by tandem mass spectrometry. Clin. Chem. 1997, 43, 2106–2113. [Google Scholar] [CrossRef] [PubMed]
- Wiley, V.; Carpenter, K.; Wilcken, B. Newborn screening with tandem mass spectrometry: 12 months’ experience in NSW Australia. Acta Paediatr. Suppl. 1999, 88, 48–51. [Google Scholar] [CrossRef]
- Wilcken, B.; Wiley, V.; Hammond, J.; Carpenter, K. Screening newborns for inborn errors of metabolism by tandem mass spectrometry. N. Engl. J. Med. 2003, 348, 2304–2312. [Google Scholar] [CrossRef] [PubMed]
- Schulze, A.; Lindner, M.; Kohlmuller, D.; Olgemoller, K.; Mayatepek, E.; Hoffmann, G.F. Expanded newborn screening for inborn errors of metabolism by electrospray ionization-tandem mass spectrometry: Results, outcome, and implications. Pediatrics 2003, 111, 1399–1406. [Google Scholar] [CrossRef]
- Zytkovicz, T.H.; Fitzgerald, E.F.; Marsden, D.; Larson, C.A.; Shih, V.E.; Johnson, D.M.; Strauss, A.W.; Comeau, A.M.; Eaton, R.B.; Grady, G.F. Tandem mass spectrometric analysis for amino, organic, and fatty acid disorders in newborn dried blood spots: A two-year summary from the New England Newborn Screening Program. Clin. Chem. 2001, 47, 1945–1955. [Google Scholar] [CrossRef]
- Vilarinho, L.; Rocha, H.; Sousa, C.; Marcao, A.; Fonseca, H.; Bogas, M.; Osorio, R.V. Four years of expanded newborn screening in Portugal with tandem mass spectrometry. J. Inherit. Metab. Dis. 2010, 33, S133–S138. [Google Scholar] [CrossRef]
- Frazier, D.M.; Millington, D.S.; McCandless, S.E.; Koeberl, D.D.; Weavil, S.D.; Chaing, S.H.; Muenzer, J. The tandem mass spectrometry newborn screening experience in North Carolina: 1997–2005. J. Inherit. Metab. Dis. 2006, 29, 76–85. [Google Scholar] [CrossRef]
- Lund, A.M.; Hougaard, D.M.; Simonsen, H.; Andresen, B.S.; Christensen, M.; Duno, M.; Skogstrand, K.; Olsen, R.K.; Jensen, U.G.; Cohen, A.; et al. Biochemical screening of 504,049 newborns in Denmark, the Faroe Islands and Greenland—Experience and development of a routine program for expanded newborn screening. Mol. Genet. Metab. 2012, 107, 281–293. [Google Scholar] [CrossRef]
- Alm, J.; Larsson, A. Evaluation of a nation-wide neonatal metabolic screening programme in Sweden 1965–1979. Acta Paediatr. Scand. 1981, 70, 601–607. [Google Scholar] [CrossRef]
- Ohlsson, A. Neonatal Screening in Sweden and Disease-Causing Variants in Phenylketonuria, Galactosaemia and Biotinidase Deficiency. Ph.D. Thesis, Karolinska Institutet, Stockholm, Sweden, 2016. [Google Scholar]
- Ohlsson, A.; Bruhn, H.; Nordenstrom, A.; Zetterstrom, R.H.; Wedell, A.; von Dobeln, U. The spectrum of PAH mutations and increase of milder forms of phenylketonuria in Sweden during 1965–2014. JIMD Rep. 2016. [Google Scholar] [CrossRef]
- Statistics Sweden. Available online: http://www.statistikdatabasen.scb.se/pxweb/en/ssd/ (accessed on 25 May 2020).
- McHugh, D.; Cameron, C.A.; Abdenur, J.E.; Abdulrahman, M.; Adair, O.; Al Nuaimi, S.A.; Ahlman, H.; Allen, J.J.; Antonozzi, I.; Archer, S.; et al. Clinical validation of cutoff target ranges in newborn screening of metabolic disorders by tandem mass spectrometry: A worldwide collaborative project. Genet. Med. 2011, 13, 230–254. [Google Scholar] [CrossRef]
- Hall, P.L.; Marquardt, G.; McHugh, D.M.; Currier, R.J.; Tang, H.; Stoway, S.D.; Rinaldo, P. Postanalytical tools improve performance of newborn screening by tandem mass spectrometry. Genet. Med. 2014, 16, 889–895. [Google Scholar] [CrossRef] [PubMed]
- Marquardt, G.; Currier, R.; McHugh, D.M.; Gavrilov, D.; Magera, M.J.; Matern, D.; Oglesbee, D.; Raymond, K.; Rinaldo, P.; Smith, E.H.; et al. Enhanced interpretation of newborn screening results without analyte cutoff values. Genet. Med. 2012, 14, 648–655. [Google Scholar] [CrossRef] [PubMed]
- CLIR Database. Available online: https://clir.mayo.edu/ (accessed on 25 May 2020).
- Minter Baerg, M.M.; Stoway, S.D.; Hart, J.; Mott, L.; Peck, D.S.; Nett, S.L.; Eckerman, J.S.; Lacey, J.M.; Turgeon, C.T.; Gavrilov, D.; et al. Precision newborn screening for lysosomal disorders. Genet. Med.: Off. J. Am. Coll. Med Genet. 2018, 20, 847–854. [Google Scholar] [CrossRef] [PubMed]
- Turgeon, C.T.; Magera, M.J.; Cuthbert, C.D.; Loken, P.R.; Gavrilov, D.K.; Tortorelli, S.; Raymond, K.M.; Oglesbee, D.; Rinaldo, P.; Matern, D. Determination of total homocysteine, methylmalonic acid, and 2-methylcitric acid in dried blood spots by tandem mass spectrometry. Clin. Chem. 2010, 56, 1686–1695. [Google Scholar] [CrossRef] [PubMed]
- Janzen, N.; Steuerwald, U.; Sander, S.; Terhardt, M.; Peter, M.; Sander, J. UPLC-MS/MS analysis of C5-acylcarnitines in dried blood spots. Clin. Chim. Acta; Int. J. Clin. Chem. 2013, 421, 41–45. [Google Scholar] [CrossRef]
- Matern, D.; Tortorelli, S.; Oglesbee, D.; Gavrilov, D.; Rinaldo, P. Reduction of the false-positive rate in newborn screening by implementation of MS/MS-based second-tier tests: The Mayo Clinic experience (2004–2007). J. Inherit. Metab. Dis. 2007, 30, 585–592. [Google Scholar] [CrossRef]
- Watson, M.S.; Mann, M.Y.; Lloyd-Puryear, M.A.; Rinaldo, P.; Howell, R.R.; American College of Medical Genetics Newborn Screening Expert Group. Newborn screening: Toward a uniform screening panel and system—executive summary. Pediatrics 2006, 117, S296–S307. [Google Scholar] [CrossRef]
- Wilson, J.; Jungner, G. Principles and Practice of Screening for Disease. WHO Papers 1968, WHO Papers No. 34. 1068. Available online: https://apps.who.int/iris/handle/10665/37650 (accessed on 25 May 2020).
- Mudd, S.H.; Skovby, F.; Levy, H.L.; Pettigrew, K.D.; Wilcken, B.; Pyeritz, R.E.; Andria, G.; Boers, G.H.; Bromberg, I.L.; Cerone, R.; et al. The natural history of homocystinuria due to cystathionine beta-synthase deficiency. Am. J. Hum. Genet. 1985, 37, 1–31. [Google Scholar]
- Yap, S.; Naughten, E. Homocystinuria due to cystathionine beta-synthase deficiency in Ireland: 25 Years’ experience of a newborn screened and treated population with reference to clinical outcome and biochemical control. J. Inherit. Metab. Dis. 1998, 21, 738–747. [Google Scholar] [CrossRef] [PubMed]
- Huemer, M.; Kozich, V.; Rinaldo, P.; Baumgartner, M.R.; Merinero, B.; Pasquini, E.; Ribes, A.; Blom, H.J. Newborn screening for homocystinurias and methylation disorders: Systematic review and proposed guidelines. J. Inherit. Metab. Dis. 2015, 38, 1007–1019. [Google Scholar] [CrossRef] [PubMed]
- Therrell, B.L., Jr.; Lloyd-Puryear, M.A.; Camp, K.M.; Mann, M.Y. Inborn errors of metabolism identified via newborn screening: Ten-year incidence data and costs of nutritional interventions for research agenda planning. Mol. Genet. Metab. 2014, 113, 14–26. [Google Scholar] [CrossRef] [PubMed]
- Wilcken, B.; Wiley, V.; Sim, K.G.; Carpenter, K. Carnitine transporter defect diagnosed by newborn screening with electrospray tandem mass spectrometry. J. Pediatr. 2001, 138, 581–584. [Google Scholar] [CrossRef]
- Wilson, C.; Knoll, D.; de Hora, M.; Kyle, C.; Glamuzina, E.; Webster, D. The decision to discontinue screening for carnitine uptake disorder in New Zealand. J. Inherit. Metab. Dis. 2019, 42, 86–92. [Google Scholar] [CrossRef]
- Holme, E.; Greter, J.; Jacobson, C.E.; Lindstedt, S.; Nordin, I.; Kristiansson, B.; Jodal, U. Carnitine deficiency induced by pivampicillin and pivmecillinam therapy. Lancet 1989, 2, 469–473. [Google Scholar] [CrossRef]
- Ahrens-Nicklas, R.C.; Serdaroglu, E.; Muraresku, C.; Ficicioglu, C. Cobalamin C disease missed by newborn screening in a patient with low carnitine level. JIMD Rep. 2015, 23, 71–75. [Google Scholar] [CrossRef]
- Gregersen, N.; Blakemore, A.I.; Winter, V.; Andresen, B.; Kolvraa, S.; Bolund, L.; Curtis, D.; Engel, P.C. Specific diagnosis of medium-chain acyl-CoA dehydrogenase (MCAD) deficiency in dried blood spots by a polymerase chain reaction (PCR) assay detecting a point-mutation (G985) in the MCAD gene. Clin. Chim. Acta 1991, 203, 23–34. [Google Scholar] [CrossRef]
- Johansson, A.; Guthenberg, C.; Ahlman, H.; Von Dobeln, U.; Hagenfeldt, L. Prevalence of the 985A>G mutation in the medium-chain acyl-CoA dehydrogenase (MCAD) gene in Sweden. Scand. J. Clin. Lab. Investig. 1999, 59, 289–291. [Google Scholar] [CrossRef]
- Rocha, H.; Castineiras, D.; Delgado, C.; Egea, J.; Yahyaoui, R.; Gonzalez, Y.; Conde, M.; Gonzalez, I.; Rueda, I.; Rello, L.; et al. Birth prevalence of fatty acid beta-Oxidation disorders in Iberia. JIMD Rep. 2014, 16, 89–94. [Google Scholar] [CrossRef]
- Hesse, J.; Braun, C.; Behringer, S.; Matysiak, U.; Spiekerkoetter, U.; Tucci, S. The diagnostic challenge in very-long chain acyl-CoA dehydrogenase deficiency (VLCADD). J. Inherit. Metab. Dis. 2018, 41, 1169–1178. [Google Scholar] [CrossRef] [PubMed]
- Stranneheim, H.; Engvall, M.; Naess, K.; Lesko, N.; Larsson, P.; Dahlberg, M.; Andeer, R.; Wredenberg, A.; Freyer, C.; Barbaro, M.; et al. Rapid pulsed whole genome sequencing for comprehensive acute diagnostics of inborn errors of metabolism. BMC Genom. 2014, 15, 1090. [Google Scholar] [CrossRef] [PubMed]
- Balasubramaniam, S.; Christodoulou, J.; Rahman, S. Disorders of riboflavin metabolism. J. Inherit. Metab. Dis. 2019, 42, 608–619. [Google Scholar] [CrossRef] [PubMed]
- Fyro, K.; Bodegard, G. Four-year follow-up of psychological reactions to false positive screening tests for congenital hypothyroidism. Acta Paediatr. Scand. 1987, 76, 107–114. [Google Scholar] [CrossRef]
- Gurian, E.A.; Kinnamon, D.D.; Henry, J.J.; Waisbren, S.E. Expanded newborn screening for biochemical disorders: The effect of a false-positive result. Pediatrics 2006, 117, 1915–1921. [Google Scholar] [CrossRef]
| Group | Disease | All Recalls | True Positive Cases | Incidence | Incidence before Screening | False Positive Cases | Missed Cases | PPV |
|---|---|---|---|---|---|---|---|---|
| Amino acidemias | HCY | 15 | 3 | 1:330,000 | 1:265,000 | 12 | - | 20% |
| MSUD | 20 | 9 | 1:110,000 | 1:265,000 | 11 | - | 45% | |
| PKU | 77 | 74 2 | 1:14,000 | N/A | 3 | - | 96% | |
| Carnitine disorders | CACT/CPT2 | 14 | 3 3 | 1:330,000 | N/A | 11 | 2 CPT2 6 | 21% |
| CUD | 107 | 13 | 1:80,000 | N/A | 94 5 | - | 12% | |
| CPT1 | 6 | 5 | 1:200,000 | N/A | 1 | - | 83% | |
| Fatty acid oxidation defects | LCHAD | 14 | 13 | 1:80,000 | 1:92,000 | 1 | - | 93% |
| MAD | 39 | 5 | 1:170,000 | N/A | 34 | - | 15% | |
| MCAD | 67 | 59 | 1:17,000 | 1:235,000 | 8 | - | 88% | |
| VLCAD | 46 | 24 | 1:42,000 | 1:1,060,000 | 22 | - | 52% | |
| Organic acidurias | BKT | 2 | 2 | 1:500,000 | N/A | 0 | - | 100% |
| GA1 | 31 | 9 | 1:110,000 | 1:235,000 | 22 | - | 29% | |
| IVA | 18 | 6 | 1:170,000 | 1:530,000 | 12 | - | 33% | |
| MMA/PA | 165 | 63 (15) 4 | 1:16,000 (1:67,000) | N/A (1:118,000) | 102 (150) | 1 PA 7 | 38% (9%) | |
| TYR | 10 | 9 | 1:110,000 | N/A | 1 | - | 90% | |
| Urea cycle disorders | ARG | 4 | 2 | 1:500,000 | 1:265,000 | 2 | - | 50% |
| ASA 1 | 5 | 3 | 1:330,000 | N/A | 2 | 1 ASA 8 | 60% | |
| CIT 1 | 25 | 9 | 1:110,000 | 1:2,120,000 | 16 | - | 36% | |
| Total | 665 | 311 (263) | 1:3200 (1:3800) | N/A | 354 (402) | 4 | 47% (40%) |
| Before or after Second-Tier | Time Period | All Recalls | True Positive Cases | False Positive Cases | PPV | Number of Screened Newborns in Period | |
|---|---|---|---|---|---|---|---|
| IVA | Before | 15 Nov. 2010–08 Dec. 2010 | 11 | 0 | 11 | 0% | 7334 |
| After | 09 Dec. 2010–1 Jul. 2019 | 7 | 6 | 1 | 86% | 993,000 | |
| HCY | Before | 15 Nov. 2010–31 Dec. 2016 | 13 | 1 | 12 | 8% | 706,543 |
| After | 1 Jan. 2017–1 Jul. 2019 | 2 | 2 | 0 | 100% | 293,791 | |
| MMA/PA | Before | 15 Nov. 2010–31 Dec. 2016 | 139 | 47 (8) | 92 (131) | 34% (6%) | 706,543 |
| After | 1 Jan. 2017–1 Jul. 2019 | 26 | 16 (7) | 10 (19) | 62% (27%) | 293,791 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sörensen, L.; von Döbeln, U.; Åhlman, H.; Ohlsson, A.; Engvall, M.; Naess, K.; Backman-Johansson, C.; Nordqvist, Y.; Wedell, A.; Zetterström, R.H. Expanded Screening of One Million Swedish Babies with R4S and CLIR for Post-Analytical Evaluation of Data. Int. J. Neonatal Screen. 2020, 6, 42. https://doi.org/10.3390/ijns6020042
Sörensen L, von Döbeln U, Åhlman H, Ohlsson A, Engvall M, Naess K, Backman-Johansson C, Nordqvist Y, Wedell A, Zetterström RH. Expanded Screening of One Million Swedish Babies with R4S and CLIR for Post-Analytical Evaluation of Data. International Journal of Neonatal Screening. 2020; 6(2):42. https://doi.org/10.3390/ijns6020042
Chicago/Turabian StyleSörensen, Lene, Ulrika von Döbeln, Henrik Åhlman, Annika Ohlsson, Martin Engvall, Karin Naess, Carolina Backman-Johansson, Yvonne Nordqvist, Anna Wedell, and Rolf H. Zetterström. 2020. "Expanded Screening of One Million Swedish Babies with R4S and CLIR for Post-Analytical Evaluation of Data" International Journal of Neonatal Screening 6, no. 2: 42. https://doi.org/10.3390/ijns6020042
APA StyleSörensen, L., von Döbeln, U., Åhlman, H., Ohlsson, A., Engvall, M., Naess, K., Backman-Johansson, C., Nordqvist, Y., Wedell, A., & Zetterström, R. H. (2020). Expanded Screening of One Million Swedish Babies with R4S and CLIR for Post-Analytical Evaluation of Data. International Journal of Neonatal Screening, 6(2), 42. https://doi.org/10.3390/ijns6020042





