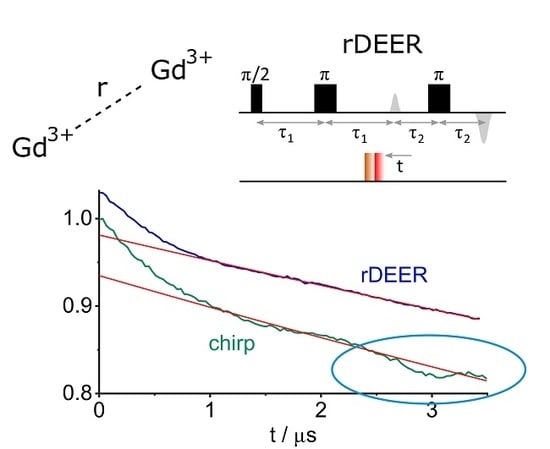
rDEER: A Modified DEER Sequence for Distance Measurements Using Shaped Pulses
Abstract
:1. Introduction
2. Results
3. Discussion
4. Materials and Methods
4.1. Sample Preparation
4.2. Spectroscopy
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Jeschke, G. DEER Distance Measurements on Proteins. Annu. Rev. Phys. Chem. 2012, 63, 419–446. [Google Scholar] [CrossRef] [PubMed]
- Hubbell, W.L.; Altenbach, C. Investigation of structure and dynamics in membrane proteins using site-directed spin labeling. Curr. Opin. Struc. Biol. 1994, 4, 566–573. [Google Scholar] [CrossRef]
- Hubbell, W.L.; Gross, A.; Langen, R.; Lietzow, M.A. Recent advances in site-directed spin labeling of proteins. Curr. Opin. Struc. Biol. 1998, 8, 649–656. [Google Scholar] [CrossRef]
- Klare, J.P.; Steinhoff, H.-J. Spin labeling EPR. Photosynth. Res. 2009, 102, 377–390. [Google Scholar] [CrossRef] [PubMed]
- Shevelev, G.Y.; Krumkacheva, O.A.; Lomzov, A.A.; Kuzhelev, A.A.; Rogozhnikova, O.Y.; Trukhin, D.V.; Troitskaya, T.I.; Tormyshev, V.M.; Fedin, M.V.; Pyshnyi, D.V.; et al. Physiological-temperature distance measurement in nucleic acid using triarylmethyl-based spin labels and pulsed dipolar EPR spectroscopy. J. Am. Chem. Soc. 2014, 136, 9874–9877. [Google Scholar] [CrossRef] [PubMed]
- Giannoulis, A.; Yang, Y.; Gong, Y.-J.; Tan, X.; Feintuch, A.; Carmieli, R.; Bahrenberg, T.; Liu, Y.; Su, X.-C.; Goldfarb, D. DEER distance measurements on trityl/trityl and Gd(III)/trityl labelled proteins. Phys. Chem. Chem. Phys. 2019. [Google Scholar] [CrossRef]
- Yang, Z.; Becker, J.; Saxena, S. On Cu(II)–Cu(II) distance measurements using pulsed electron electron double resonance. J. Magn. Reson. 2007, 188, 337–343. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Kise, D.; Saxena, S. An Approach towards the Measurement of Nanometer Range Distances Based on Cu2+ Ions and ESR. J. Phys. Chem. B 2010, 114, 6165–6174. [Google Scholar] [CrossRef] [PubMed]
- Kaminker, I.; Bye, M.; Mendelman, N.; Gislason, K.; Sigurdsson, S.T.; Goldfarb, D. Distance measurements between manganese(ii) and nitroxide spin-labels by DEER determine a binding site of Mn2+ in the HP92 loop of ribosomal RNA. Phys. Chem. Chem. Phys. 2015, 17, 15098–15102. [Google Scholar] [CrossRef] [PubMed]
- Martorana, A.; Yang, Y.; Zhao, Y.; Li, Q.-F.; Su, X.-C.; Goldfarb, D. Mn(ii) tags for DEER distance measurements in proteins via C-S attachment. Dalt. Trans. 2015, 44, 20812–20816. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Gong, Y.-J.; Litvinov, A.; Liu, H.-K.; Yang, F.; Su, X.-C.; Goldfarb, D. Generic tags for Mn(ii) and Gd(iii) spin labels for distance measurements in proteins. Phys. Chem. Chem. Phys. 2017, 19, 26944–26956. [Google Scholar] [CrossRef] [PubMed]
- Akhmetzyanov, D.; Ching, H.Y.V.; Denysenkov, V.; Demay-Drouhard, P.; Bertrand, H.C.; Tabares, L.C.; Policar, C.; Prisner, T.F.; Un, S. RIDME spectroscopy on high-spin Mn2+ centers. Phys. Chem. Chem. Phys. 2016, 18, 30857–30866. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, D.; Yagi, H.; Huber, T.; Otting, G.; Goldfarb, D. Nanometer-Range Distance Measurement in a Protein Using Mn2+ Tags. J. Phys. Chem. Lett. 2012, 3, 157–160. [Google Scholar] [CrossRef]
- Prokopiou, G.; Lee, M.D.; Collauto, A.; Abdelkader, E.H.; Bahrenberg, T.; Feintuch, A.; Ramirez-Cohen, M.; Clayton, J.; Swarbrick, J.D.; Graham, B.; et al. Small Gd(III) Tags for Gd(III)-Gd(III) Distance Measurements in Proteins by EPR Spectroscopy. Inorg. Chem. 2018, 57. [Google Scholar] [CrossRef] [PubMed]
- Feintuch, A.; Otting, G.; Goldfarb, D. Gd3 + Spin Labeling for Measuring Distances in Biomacromolecules: Why and How? Methods Enzymol. 2015, 563, 415–457. [Google Scholar] [PubMed]
- Potapov, A.; Yagi, H.; Huber, T.; Jergic, S.; Dixon, N.E.; Otting, G.; Goldfarb, D. Nanometer-Scale Distance Measurements in Proteins Using Gd3+ Spin Labeling. J. Am. Chem. Soc. 2010, 132, 9040–9048. [Google Scholar] [CrossRef] [PubMed]
- Lueders, P.; Jeschke, G.; Yulikov, M. Double electron-electron resonance measured between Gd3+ ions and nitroxide radicals. J. Phys. Chem. Lett. 2011, 2, 604–609. [Google Scholar] [CrossRef]
- Mahawaththa, M.C.; Lee, M.D.; Giannoulis, A.; Adams, L.A.; Feintuch, A.; Swarbrick, J.D.; Graham, B.; Nitsche, C.; Goldfarb, D.; Otting, G. Small neutral Gd(iii) tags for distance measurements in proteins by double electron–electron resonance experiments. Phys. Chem. Chem. Phys. 2018, 20, 23535–23545. [Google Scholar] [CrossRef] [PubMed]
- Abdelkader, E.H.; Lee, M.D.; Feintuch, A.; Cohen, M.R.; Swarbrick, J.D.; Otting, G.; Graham, B.; Goldfarb, D. A New Gd3+ Spin Label for Gd3+–Gd3+ Distance Measurements in Proteins Produces Narrow Distance Distributions. J. Phys. Chem. Lett. 2015, 6, 5016–5021. [Google Scholar] [CrossRef] [PubMed]
- Yagi, H.; Banerjee, D.; Graham, B.; Huber, T.; Goldfarb, D.; Otting, G. Gadolinium Tagging for High-Precision Measurements of 6 nm Distances in Protein Assemblies by EPR. J. Am. Chem. Soc. 2011, 133, 10418–10421. [Google Scholar] [CrossRef] [PubMed]
- Sajid, M.; Jeschke, G.; Wiebcke, M.; Godt, A. Conformationally unambiguous spin labeling for distance measurements. Chemistry 2009, 15, 12960–12962. [Google Scholar] [CrossRef] [PubMed]
- Qi, M.; Groß, A.; Jeschke, G.; Godt, A.; Drescher, M. Gd(III)-PyMTA Label Is Suitable for In-Cell EPR. J. Am. Chem. Soc. 2014, 136, 15366–15378. [Google Scholar] [CrossRef] [PubMed]
- Cunningham, T.F.; Putterman, M.R.; Desai, A.; Horne, W.S.; Saxena, S. The Double-Histidine Cu2+-Binding Motif: A Highly Rigid, Site-Specific Spin Probe for Electron Spin Resonance Distance Measurements. Angew. Chemie Int. Ed. 2015, 54, 6330–6334. [Google Scholar] [CrossRef] [PubMed]
- Mascali, F.C.; Ching, H.Y.V.; Rasia, R.M.; Un, S.; Tabares, L.C. Using Genetically Encodable Self-Assembling Gd III Spin Labels to Make In-cell Nanometric Distance Measurements. Angew. Chemie Int. Ed. 2016, 1–4. [Google Scholar]
- Yang, Y.; Yang, F.; Gong, Y.-J.; Chen, J.-L.; Goldfarb, D.; Su, X.-C. A Reactive, Rigid Gd III Labeling Tag for In-Cell EPR Distance Measurements in Proteins. Angew. Chemie Int. Ed. 2017, 56, 2914–2918. [Google Scholar] [CrossRef] [PubMed]
- Martorana, A.; Bellapadrona, G.; Feintuch, A.; Di Gregorio, E.; Aime, S.; Goldfarb, D. Probing Protein Conformation in Cells by EPR Distance Measurements using Gd3+ Spin Labeling. J. Am. Chem. Soc. 2014, 136, 13458–13465. [Google Scholar] [CrossRef] [PubMed]
- Theillet, F.-X.; Binolfi, A.; Bekei, B.; Martorana, A.; Rose, H.M.; Stuiver, M.; Verzini, S.; Lorenz, D.; van Rossum, M.; Goldfarb, D.; et al. Structural disorder of monomeric α-synuclein persists in mammalian cells. Nature 2016, 530, 45–50. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.; Yang, F.; Gong, Y.-J.; Bahrenberg, T.; Feintuch, A.; Su, X.-C.; Goldfarb, D. High Sensitivity In-Cell EPR Distance Measurements on Proteins Using an Optimized Gd(III) Spin Label. J. Phys. Chem. Lett. 2018. [Google Scholar] [CrossRef] [PubMed]
- Dalaloyan, A.; Qi, M.; Ruthstein, S.; Vega, S.; Godt, A.; Feintuch, A.; Goldfarb, D. Gd( III)–Gd(III) EPR distance measurements – the range of accessible distances and the impact of zero field splitting. Phys. Chem. Chem. Phys. 2015, 17, 18464–18476. [Google Scholar] [CrossRef] [PubMed]
- Goldfarb, D. Gd3+ Spin Labeling For Distance Measurements by Pulse Epr Spectroscopy. PCCP 2014, 16, 9685–9699. [Google Scholar] [CrossRef] [PubMed]
- Demay-Drouhard, P.; Ching, H.Y.V.; Akhmetzyanov, D.; Guillot, R.; Tabares, L.C.; Bertrand, H.C.; Policar, C. A Bis-Manganese(II)–DOTA Complex for Pulsed Dipolar Spectroscopy. ChemPhysChem 2016, 17, 2066–2078. [Google Scholar] [CrossRef] [PubMed]
- van Amsterdam, I.M.C.; Ubbink, M.; Canters, G.W.; Huber, M. Measurement of a Cu–Cu Distance of 26 Å by a Pulsed EPR Method. Angew. Chemie Int. Ed. 2003, 42, 62–64. [Google Scholar] [CrossRef]
- Polyhach, Y.; Bordignon, E.; Tschaggelar, R.; Gandra, S.; Godt, A.; Jeschke, G. High sensitivity and versatility of the DEER experiment on nitroxide radical pairs at Q-band frequencies. Phys. Chem. Chem. Phys. 2012, 14, 10762–10773. [Google Scholar] [CrossRef] [PubMed]
- Doll, A.; Pribitzer, S.; Tschaggelar, R.; Jeschke, G. Adiabatic and fast passage ultra-wideband inversion in pulsed EPR. J. Magn. Reson. 2013, 230, 27–39. [Google Scholar] [CrossRef] [PubMed]
- Doll, A.; Jeschke, G. Fourier-transform electron spin resonance with bandwidth-compensated chirp pulses. J. Magn. Reson. 2014, 246, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Doll, A.; Qi, M.; Wili, N.; Pribitzer, S.; Godt, A.; Jeschke, G. Gd(III)-Gd(III) distance measurements with chirp pump pulses. J. Magn. Reson. 2015, 259, 153–162. [Google Scholar] [CrossRef] [PubMed]
- Spindler, P.E.; Zhang, Y.; Endeward, B.; Gershernzon, N.; Skinner, T.E.; Glaser, S.J.; Prisner, T.F. Shaped optimal control pulses for increased excitation bandwidth in EPR. J. Magn. Reson. 2012, 218, 49–58. [Google Scholar] [CrossRef] [PubMed]
- Schöps, P.; Spindler, P.E.; Marko, A.; Prisner, T.F. Broadband spin echoes and broadband SIFTER in EPR. J. Magn. Reson. 2015, 250, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Spindler, P.E.; Glaser, S.J.; Skinner, T.E.; Prisner, T.F. Broadband Inversion PELDOR Spectroscopy with Partially Adiabatic Shaped Pulses. Angew. Chemie Int. Ed. 2013, 52, 3425–3429. [Google Scholar] [CrossRef] [PubMed]
- Giannoulis, A.; Ackermann, K.; Spindler, P.E.; Higgins, C.; Cordes, D.B.; Slawin, A.M.Z.; Prisner, T.F.; Bode, B.E. Nitroxide–nitroxide and nitroxide–metal distance measurements in transition metal complexes with two or three paramagnetic centres give access to thermodynamic and kinetic stabilities. Phys. Chem. Chem. Phys. 2018, 20, 11196–11205. [Google Scholar] [CrossRef] [PubMed]
- Teucher, M.; Bordignon, E. Improved signal fidelity in 4-pulse DEER with Gaussian pulses. J. Magn. Reson. 2018, 296, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Kaminker, I.; Barnes, R.; Han, S. Arbitrary waveform modulated pulse EPR at 200GHz. J. Magn. Reson. 2017, 279, 81–90. [Google Scholar] [CrossRef] [PubMed]
- Kuzhelev, A.; Akhmetzyanov, D.; Denysenkov, V.; Shevelev, G.; Krumkacheva, O.; Bagryanskaya, E.; Prisner, T. High-frequency pulsed electron–electron double resonance spectroscopy on DNA duplexes using trityl tags and shaped microwave pulses. Phys. Chem. Chem. Phys. 2018, 20, 26140–26144. [Google Scholar] [CrossRef] [PubMed]
- Pannier, M.; Veit, S.; Godt, A.; Jeschke, G.; Spiess, H. Dead-Time Free Measurement of Dipole–Dipole Interactions between Electron Spins. J. Magn. Reson. 2000, 142, 331–340. [Google Scholar] [CrossRef] [PubMed]
- Bahrenberg, T.; Rosenski, Y.; Carmieli, R.; Zibzener, K.; Qi, M.; Frydman, V.; Godt, A.; Goldfarb, D.; Feintuch, A. Improved sensitivity for W-band Gd(III)-Gd(III) and nitroxide-nitroxide DEER measurements with shaped pulses. J. Magn. Reson. 2017, 283, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.; Bibi, E. MdfA, an Escherichia coli multidrug resistance protein with an extraordinarily broad spectrum of drug recognition. J. Bacteriol. 1998, 179, 2274–2280. [Google Scholar] [CrossRef]
- Sigal, N.; Cohen-Karni, D.; Siemion, S.; Bibi, E. MdfA from Escherichia coli, a Model Protein for Studying Secondary Multidrug Transport. J. Mol. Microbiol. Biotechnol. 2006, 11, 308–317. [Google Scholar] [CrossRef] [PubMed]
- Yardeni, E.H.; Zomot, E.; Bibi, E. The fascinating but mysterious mechanistic aspects of multidrug transport by MdfA from Escherichia coli. Res. Microbiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- de la Cruz, L.; Nguyen, T.H.D.; Ozawa, K.; Shin, J.; Graham, B.; Huber, T.; Otting, G. Binding of Low Molecular Weight Inhibitors Promotes Large Conformational Changes in the Dengue Virus NS2B-NS3 Protease: Fold Analysis by Pseudocontact Shifts. J. Am. Chem. Soc. 2011, 133, 19205–19215. [Google Scholar] [CrossRef] [PubMed]
- Jeschke, G.; Chechik, V.; Ionita, P.; Godt, A.; Zimmermann, H.; Banham, J.; Timmel, C.R.; Hilger, D.; Jung, H. DeerAnalysis2006—A comprehensive software package for analyzing pulsed ELDOR data. Appl. Magn. Reson. 2006, 30, 473–498. [Google Scholar] [CrossRef]
- Tait, C.E.; Stoll, S. Coherent pump pulses in Double Electron Electron Resonance spectroscopy. Phys. Chem. Chem. Phys. 2016, 18, 18470–18485. [Google Scholar] [CrossRef] [PubMed]
- Goldfarb, D.; Lipkin, Y.; Potapov, A.; Gorodetsky, Y.; Epel, B.; Raitsimring, A.M.; Radoul, M.; Kaminker, I. HYSCORE and DEER with an upgraded 95 GHz pulse EPR spectrometer. J. Magn. Reson. 2008, 194, 8–15. [Google Scholar] [CrossRef] [PubMed]
- Mentink-Vigier, F.; Collauto, A.; Feintuch, A.; Kaminker, I.; Tarle, V.; Goldfarb, D. Increasing sensitivity of pulse EPR experiments using echo train detection schemes. J. Magn. Reson. 2013, 236, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Epel, B.; Gromov, I.; Stoll, S.; Schweiger, A.; Goldfarb, D. Spectrometer manager: A versatile control software for pulse EPR spectrometers. Concepts Magn. Reson. Part B Magn. Reson. Eng. 2005, 26B, 36–45. [Google Scholar] [CrossRef]
- Stoll, S.; Kasumaj, B. Phase Cycling in Electron Spin Echo Envelope Modulation. Appl. Magn. Reson. 2008, 35, 15–32. [Google Scholar] [CrossRef]

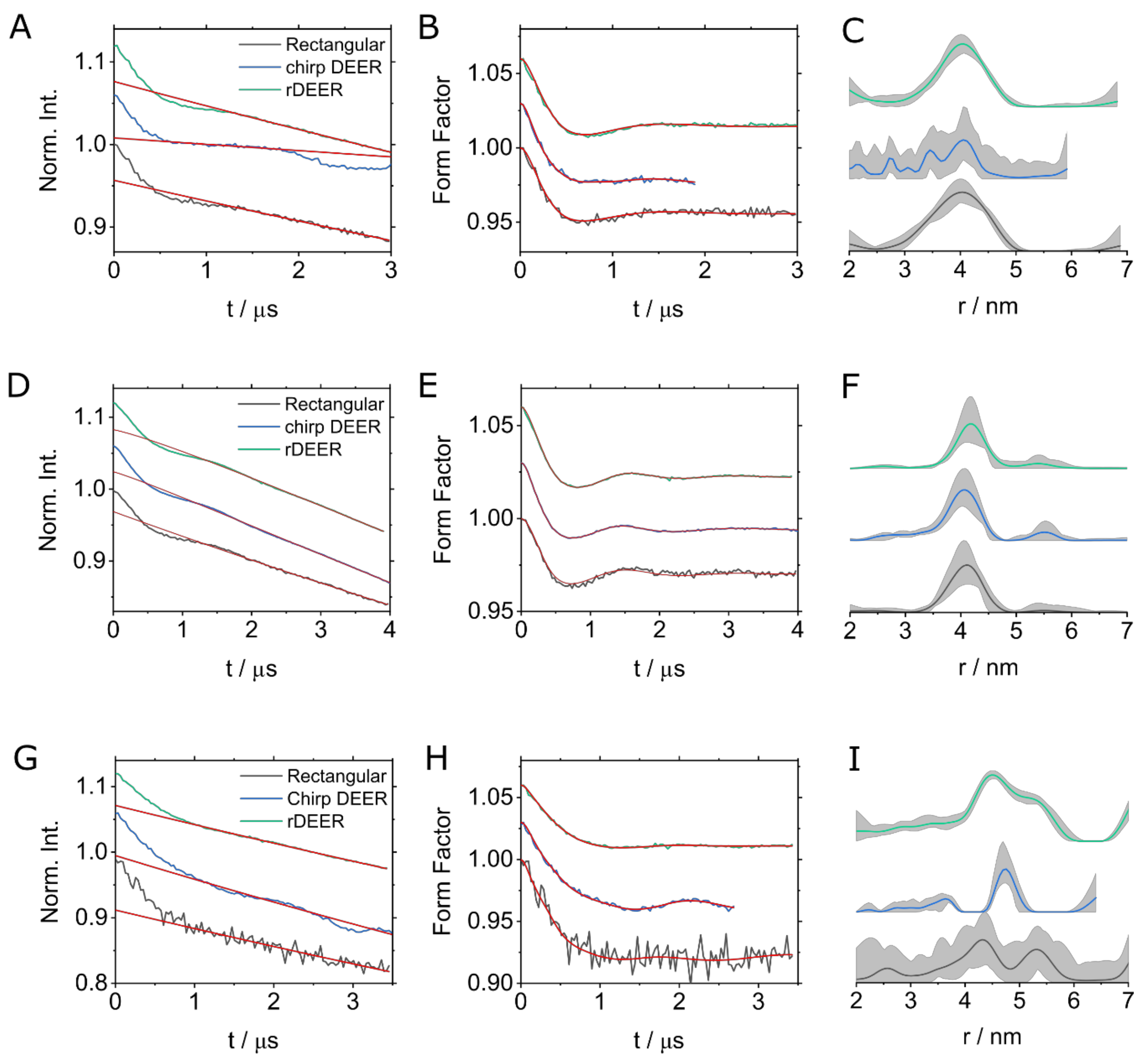
| Experiment | Run-Time/min | SNR | Modulation Depth | Number of Scans ** |
|---|---|---|---|---|
| Ubi-DOTA-M-Gd | ||||
| Rectangular | 15:50 | 7.2 * | 4.3% | 20 |
| chirp DEER | 08:00 | 15.0 * | 5.2% | 10 |
| rDEER | 15:55 | 18.4 * | 4.5% | 20 |
| Ubi-DO3A-Gd | ||||
| Rectangular | 51:44 | 5.6 | 3% | 50 |
| chirp DEER | 52:22 | 31.2 | 3.7% | 50 |
| rDEER | 52:19 | 40.7 | 3.8% | 50 |
| MdfA-C2-Gd | ||||
| Rectangular | 18:20 | 2.2 | 8% | 20 |
| chirp DEER | 18:22 | 15.6 | 6.5% | 20 |
| rDEER | 18:43 | 13.9 | 5% | 20 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bahrenberg, T.; Yang, Y.; Goldfarb, D.; Feintuch, A. rDEER: A Modified DEER Sequence for Distance Measurements Using Shaped Pulses. Magnetochemistry 2019, 5, 20. https://doi.org/10.3390/magnetochemistry5010020
Bahrenberg T, Yang Y, Goldfarb D, Feintuch A. rDEER: A Modified DEER Sequence for Distance Measurements Using Shaped Pulses. Magnetochemistry. 2019; 5(1):20. https://doi.org/10.3390/magnetochemistry5010020
Chicago/Turabian StyleBahrenberg, Thorsten, Yin Yang, Daniella Goldfarb, and Akiva Feintuch. 2019. "rDEER: A Modified DEER Sequence for Distance Measurements Using Shaped Pulses" Magnetochemistry 5, no. 1: 20. https://doi.org/10.3390/magnetochemistry5010020
APA StyleBahrenberg, T., Yang, Y., Goldfarb, D., & Feintuch, A. (2019). rDEER: A Modified DEER Sequence for Distance Measurements Using Shaped Pulses. Magnetochemistry, 5(1), 20. https://doi.org/10.3390/magnetochemistry5010020