An Inter-Laboratory Comparative Study on the Influence of Reagents to Perform the Identification of the Xylella fastidiosa Subspecies Using Tetraplex Real Time PCR
Abstract
:1. Introduction
- (1)
- A test performance study (TPS), with the participation of 18 official laboratories (OLs) of the Italian (NPPO), was used to evaluate the influence of different master mixes on the performance of Set N° 2 of TqPCR [16]. TqPCR was compared with the real-time PCR method developed by Harper et al. [18], which is known as one of the most appropriate tests for the detection of Xf, with high diagnostic sensitivity and specificity [19].
- (2)
- Intra-laboratory study (ITS), within the Research Centre for Plant Protection and Certification of the Council for Agricultural Research and Economics (CREA-DC), was used to evaluate the analytical sensitivity (ASE) of TqPCR [16] with respect to Harper et al. [18] and Hodgetts et al. [20] assays by testing spiked plant matrices. Additionally, analyses of naturally infected samples to determine the subspecies and STs via MLST [14] and TqPCR [16] were performed.
2. Materials and Methods
2.1. Test Performance Study (TPS)
2.1.1. Plant Material
2.1.2. Bacterial Strains
2.1.3. Sample Preparation and DNA Extraction
2.1.4. Sample Homogeneity and Stability Test
2.1.5. Tetraplex Real-Time PCR (TqPCR)
2.1.6. Performance Criteria Evaluation
2.1.7. Outliers
2.2. Intra-Laboratory Study (ITS)
3. Results
3.1. Test Performance Study (TPS)
3.1.1. Participants
3.1.2. Sample Homogeneity and Stability Test
3.1.3. Evaluation of Performance Criteria
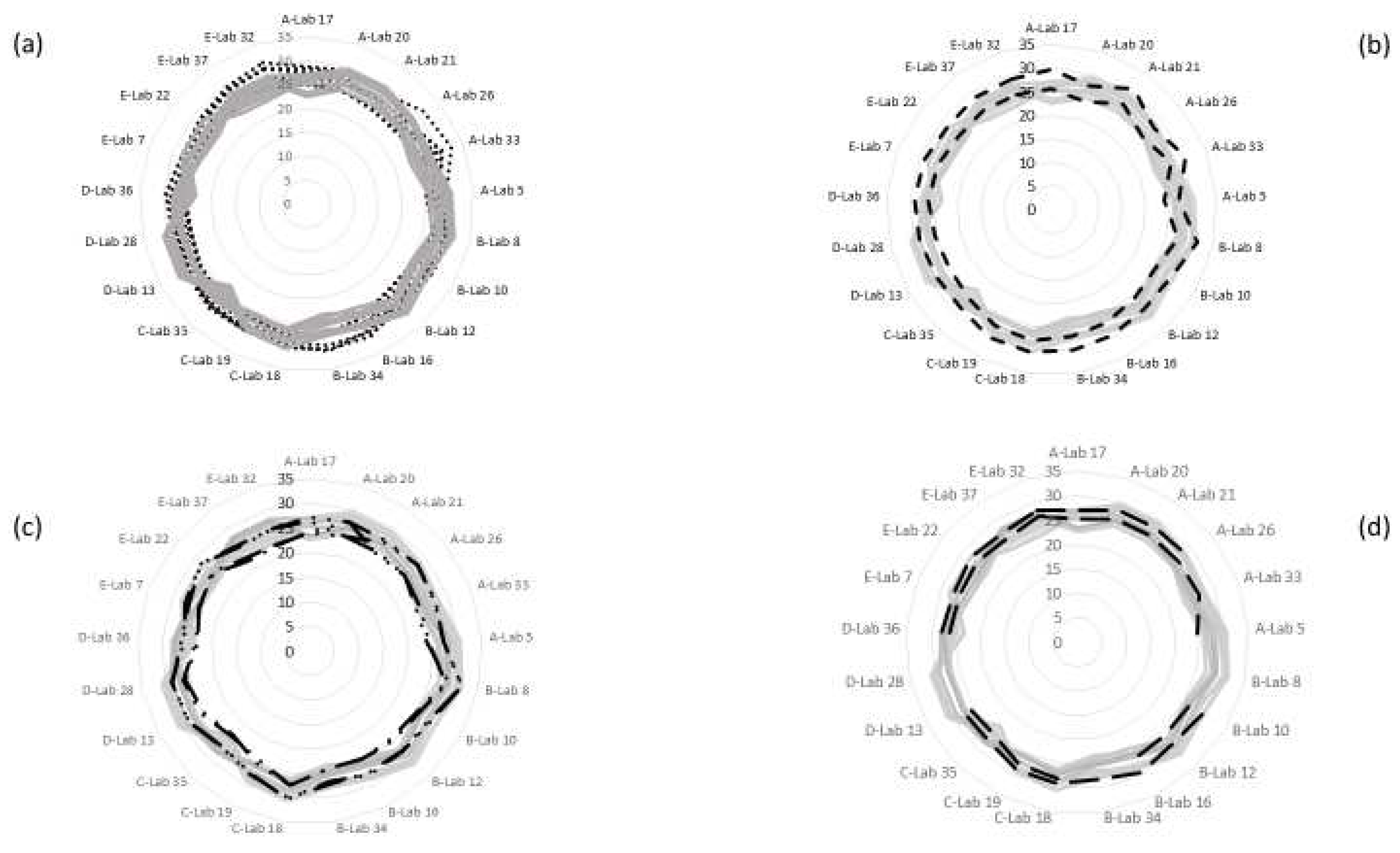
3.1.4. Comparison of Real-Time PCR and Tetraplex Real-Time PCR
3.2. Intralaboratory Study (ITS)
4. Discussion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Delbianco, A.; Gibin, D.; Pasinato, L.; Boscia, D.; Morelli, M. Update of the Xylella spp. host plant database–systematic literature search up to 30 June 2022. EFSA J. 2023, 21, 7726. [Google Scholar] [CrossRef]
- Burbank, L.P.; Roger, M.C. Microbe Profile: Xylella fastidiosa—A devasting agricultural pathogen with an endophytic lifestyle. Microbiology 2021, 167, 001091. [Google Scholar] [CrossRef] [PubMed]
- Commission Implementing Regulation (EU) 2019/2072 of 28 November 2019. Available online: https://www.legislation.gov.uk/eur/2019/2072/2020-12-31 (accessed on 17 January 2023).
- Saponari, M.; Boscia, D.; Nigro, F.; Martelli, G.P. Identification of DNA sequences related to Xylella fastidiosa in oleander, almond and olive trees exhibiting leaf scorch symptoms in Apulia (Southern Italy). J. Plant Pathol. 2013, 95, 659–668. [Google Scholar]
- Trkulja, V.; Tomic, A.; Iličić, R.; Nožinić, M.; Popović Milovanović, T. Xylella fastidiosa in Europe: From the Introduction to the Current Status. Plant Pathol. J. 2022, 38, 551–571. [Google Scholar] [CrossRef] [PubMed]
- EPPO 2020 Global Database. Available online: https://gd.eppo.int/taxon/XYLEFA/datasheet (accessed on 28 June 2023).
- Bosso, L.; Russo, D.; Di Febbraro, M.; Cristinzio, G.; Zoina, A. Potential distribution of Xylella fastidiosa in Italy: A maximum entropy model. Phytopathol. Mediterr. 2016, 55, 62–72. [Google Scholar] [CrossRef]
- Marcelletti, S.; Scortichini, M. Xylella fastidiosa CoDiRO strain associated with the olive quick decline syndrome in southern Italy belongs to a clonal complex of the subspecies pauca that evolved in Central America. Microbiology 2016, 162, 2087–2098. [Google Scholar] [CrossRef]
- Denancé, N.; Briand, M.; Gaborieau, R.; Gaillard, S.; Jacques, M.A. Identification of genetic relationships and subspecies signatures in Xylella fastidiosa. BMC Genom. 2019, 20, 239. [Google Scholar] [CrossRef]
- Commission Implementing Regulation (EU) 2020/1201 of 14 August 2020 as regards measures to prevent the introduction into and the spread within the Union of Xylella fastidiosa (Wells et al.). Off. J. Eur. Union, 2020; L 269/2. Available online: http://data.europa.eu/eli/reg_impl/2020/1201/oj(accessed on 28 June 2023).
- European and Mediterranean Plant Protection Organization. PM 7/24 (5) Xylella fastidiosa. EPPO Bull. 2023, 53, 205–276. [Google Scholar] [CrossRef]
- Yuan, X.; Morano, L.; Bromley, R.; Spring-Pearson, S.; Stouthamer, R.; Nunney, L. Multilocus sequence typing of Xylella fastidiosa causing Pierce’s disease and oleander leaf scorch in the United States. Phytopathology 2010, 100, 601–611. [Google Scholar] [CrossRef]
- Pooler, M.R.; Hartung, J.S. Specific PCR detection and identification of Xylella fastidiosa strains causing citrus variegated chlorosis. Curr. Microbiol. 1995, 31, 377–381. [Google Scholar] [CrossRef]
- Hernandez-Martinez, R.; Costa, H.S.; Dumenyo, C.K.; Cooksey, D.A. Differentiation of strains of Xylella fastidiosa infecting grape, almonds, and oleander using a multiprimer PCR assay. Plant Dis. 2006, 90, 1382–1388. [Google Scholar] [CrossRef]
- Denancé, N.; Legendre, B.; Briand, M.; Olivier, V.; de Boisseson, C.; Poliakoff, F.; Jacques, M.-A. Several subspecies and sequence types are associated with the emergence of Xylella fastidiosa in natural settings in France. Plant Pathol. 2017, 66, 1054–1064. [Google Scholar] [CrossRef]
- Dupas, E.; Briand, M.; Jacques, M.-A.; Cesbron, S. Novel Tetraplex Quantitative PCR Assays for Simultaneous Detection and Identification of Xylella fastidiosa Subspecies in Plant Tissues. Front. Plant Sci. 2019, 10, 1732. [Google Scholar] [CrossRef] [PubMed]
- Bergsma-Vlami, M.; van de Bilt, J.L.J.; Tjou-Tam-Sin, N.N.A.; Helderman, C.M.; Gorkink-Smits, P.P.M.A.; Landman, N.M.; van Nieuwburg, J.G.W.; van Veen, E.J.; Westenberg, M. Assessment of the genetic diversity of Xylella fastidiosa in imported ornamental Coffea arabica plants. Plant Pathol. 2017, 66, 1065–1074. [Google Scholar] [CrossRef]
- Harper, S.; Ward, L.; Clover, G. Development of LAMP and qPCR methods for the rapid detection of Xylella fastidiosa for quarantine and field applications. Phytopathology 2010, 100, 1282–1288, (erratum 2013). [Google Scholar] [CrossRef] [PubMed]
- Modesti, V.; Pucci, N.; Lucchesi, S.; Campus, L.; Loreti, S. Experience of the Latium region (Central Italy) as a pest-free area for monitoring of Xylella fastidiosa: Distinctive features of molecular diagnostic methods. Eur. J. Plant Pathol. 2016, 148, 557–566. [Google Scholar] [CrossRef]
- Hodgetts, J.; Glover, R.; Cole, J.; Hall, J.; Boonham, N. Genomics informed design of a suite of real-time PCR assays for the specific detection of each Xylella fastidiosa subspecies. J. Appl. Microbiol. 2020, 131, 855–872. [Google Scholar] [CrossRef] [PubMed]
- European and Mediterranean Plant Protection Organization. PM7/122 (2). EPPO Bull. 2022, 52, 604–618. [Google Scholar] [CrossRef]
- European and Mediterranean Plant Protection Organization. PM7/98 (5). EPPO Bull. 2021, 51, 468–498. [Google Scholar] [CrossRef]
- Langton, S.D.; Chevennement, R.; Nagelkerke, N.; Lombard, B. Analysis collaborative trials for qualitative microbiological methods. Int. J. Food Microbiol. 2002, 79, 175–181. [Google Scholar] [CrossRef]
- Tatulli, G.; Modesti, V.; Pucci, N.; Scala, V.; L’Aurora, A.; Lucchesi, S.; Salustri, M.; Scortichini, M.; Loreti, S. Further In Vitro Assessment and Mid-Term Evaluation of Control Strategy of Xylella fastidiosa subsp. pauca in Olive Groves of Salento (Apulia, Italy). Pathogens 2021, 10, 85. [Google Scholar] [CrossRef]
- Cesbron, S.; Dupas, E.; Beaurepère, Q.; Briand, M.; Montes-Borrego, M.; Velasco-Amo, M.d.P.; Landa, B.B.; Jacques, M.-A. Development of A Nested-MultiLocus Sequence Typing Approach for A Highly Sensitive and Specific Identification of Xylella fastidiosa Subspecies Directly from Plant Samples. Agronomy 2020, 10, 1099. [Google Scholar] [CrossRef]
- Reisenzein, H. PCR assays for the detection of Xylella fastidiosa. Review and comparison of published protocols. In Xylella fastidiosa the Olive Quick Decline Syndrome (OQDS). A Serious Worldwide Challenge for the Safeguard of Olive Trees; D’Onghia, A.M., Brunel, S., Valentini, F., Eds.; Options Méditerranéennes: Série A. Séminaires Méditerranéens; n. 121; CIHEAM: Bari, Italy, 2017; pp. 57–60. [Google Scholar]
- Marchi, G.; Rizzo, D.; Ranaldi, F.; Ghelardini, L.; Ricciolini, M.; Scarpelli, I.; Drosera, L.; Goti, E.; Capretti, P.; Surico, G. First detection of Xylella fastidiosa subsp. multiplex DNA in Tuscany (Italy). Phytopathol. Mediterr. 2018, 57, 363–364. [Google Scholar] [CrossRef]
- Saponari, M.; D’Attoma, G.; Abou Kubaa, R.; Loconsole, G.; Altamura, G.; Zicca, S.; Rizzo, D.; Boscia, D. A new variant of Xylella fastidiosa subspecies multiplex detected in different host plants in the recently emerged outbreak in the region of Tuscany, Italy. Eur. J. Plant Pathol. 2019, 154, 1195–1200. [Google Scholar] [CrossRef]
- Giampetruzzi, A.; Loconsole, G.; Zicca, S.; Boscia, D.; Balestra, G.M.; Saponari, M. Draft Genome Sequence Resource of Xylella fastidiosa Strain Alm_Lz_1 Associated with a New Outbreak in Lazio, Italy. Phytopathology 2023, 113, 108–111. [Google Scholar] [CrossRef] [PubMed]
- Olmo, D.; Nieto, A.; Borràs, D.; Montesinos, M.; Adrover, F.; Pascual, A.; Gost, P.A.; Quetglas, B.; Urbano, A.; de Dios García, J.; et al. Landscape epidemiology of Xylella fastidiosa in the Balearic Islands. Agronomy 2021, 11, 473. [Google Scholar] [CrossRef]
- Nunney, L.R.; Hopkins, D.L.; Morano, L.D.; Russell, S.E.; Stouthamer, R. Intersubspecific recombination in Xylella fastidiosa strains native to the United States: Infection of novel hosts associated with an unsuccessful invasion. Appl. Environ. Microbiol. 2014, 80, 1159–1169. [Google Scholar] [CrossRef]
| Sample ID | Sample Type (DNA Extract) |
Phyto- Sanitary Status | Host |
|---|---|---|---|
| S1 | Healthy | Negative | Petioles and vine leaves (Vitis vinifera) |
| S2 | Healthy | Negative | Petioles and olive leaves (Olea europaea) |
| S3 | Artificially contaminated (Xfm-6 pg/µL) | Positive | Petioles and lavender leaves (Lavandula spp.) |
| S4 | Artificially contaminated (Xfm-0.6 pg/µL) | Positive | Petioles and rosemary leaves (Rosmarinus officinalis) |
| S5 | Artificially contaminated (Xff-0.6 pg/µL) | Positive | Petioles and vine leaves |
| S6 | Artificially contaminated (Xfp-0.6 pg/µL) | Positive | Petioles and lavender leaves |
| S7 | Healthy | Negative | Petioles and lavender leaves |
| S8 | Healthy | Negative | Petioles and rosemary leaves |
| S9 | Artificially contaminated (Xfm-0.6 pg/µL) | Positive | Petioles and lavender leaves |
| S10 | Naturally infected (Xfm) | Positive | Petioles and almond leaves (Prunus dulcis) |
| S11 | Naturally infected (Xfp) | Positive | Petioles and olive leaves |
| S12 | Artificially contaminated (Xff-6 pg/µL) | Positive | Petioles and vine leaves |
| PAC1 | Bacterial DNA (Xff-60 pg/µL) | Positive | Bacterial strain |
| PAC2 | Bacterial DNA (Xfm-60 pg/µL | Positive | Bacterial strain |
| PAC3 | Bacterial DNA (Xfp-60 pg/µL) | Positive | Bacterial strain |
| NAC | Water DEPC (Diethyl pyrocarbonate) | Negative | - |
| CREA-DC Centro difesa e certificazione—Roma * |
| Agenzia Settore Agroalimentare delle Marche—SFR Regione Marche, Osimo (AN) |
| Agenzia Agris, Ussana (SU) |
| CNR—Istituto per la Protezione Sostenibile delle Piante, Bari (BA) |
| CRSFA, Centro di Ricerca, Sperimentazione e Formazione in Agricoltura “Basile Caramia”, Locorotondo (BA) |
| DAFNE, Dipartimento di Scienze Agrarie degli Alimenti, Risorse Naturali e Ingegneria—Università degli Studi di Foggia, Foggia (FG) |
| DAFNE, Università degli Studi della Tuscia, Dipartimento di Scienze agrarie e forestali, Viterbo (VT) |
| Dipartimento di Scienze Agrarie, Alimentari ed Ambientali, Università Politecnica delle Marche, Ancona (AN) |
| Enocontrol S.c.a.r.l. Centro d’analisi e ricerca, Cuneo (CN) |
| ERSA, Laboratorio di Fitopatologia e Biotecnologie, Udine (UD) |
| Fondazione Edmund Mach, San Michele all’Adige (TN) |
| Laboratorio Fitopatologico Regione Emilia-Romagna, Bologna (BO) |
| Laboratorio Fitopatologico Regione Liguria, Genova (GE) |
| Laboratorio Fitopatologico Regione Campania, Napoli, (NA) |
| Laboratorio fitosanitario e settore fitosanitario e servizi tecnico-scientifici, Direzione Agricoltura e cibo, Regione Piemonte, Torino (TO) |
| Laboratorio SFR Regione Lombardia, Vertemate con Minoprio (MI) |
| Regione Toscana SFR e di vigilanza del controllo agroforestale—Laboratorio Fitopatologico Regionale, Pistoia (PT) |
| SELGE—Dipartimento di scienze del suolo, della pianta e degli alimenti—Università di Bari (BA) |
|
Group Code and Total Number of OLs | Enzyme |
Step of Amplification Protocol | Cycles | Time |
Temperature °C |
|---|---|---|---|---|---|
| A 6 OLs | SsoAdvanced™ Universal Probes Supermix (Bio-Rad) | Denaturation | 1× | 3 min | 95 |
| Amplification/ Fluorescence detection | 40× | 15 s | 95 | ||
| 30 s | 60 | ||||
| B 5 OLs | SsoAdvanced™ Universal Probes Supermix (Bio-Rad) | Denaturation | 1× | 3 min | 95 |
| Amplification/ Fluorescence detection | 40× | 15 s | 95 | ||
| 30 s | 63 | ||||
| C 4 OLs | QuantiNova Pathogen + IC kit (Qiagen) | Denaturation | 1× | 2 min | 95 |
| Amplification/ Fluorescence detection | 40× | 15 s | 95 | ||
| 30 s | 60 | ||||
| D 4 OLs | TaqManTM Fast Universal PCR Master Mix (Applied Biosystems™) | Enzyme activation | 1× | 2 min | 50 |
| Denaturation | 1× | 10 min | 95 | ||
| Amplification/ Fluorescence detection | 40× | 15 s | 95 | ||
| 30 s | 60 | ||||
| E 4 OLs | Brilliant Multiplex qPCR Master Mix (Agilent) | Denaturation | 1× | 10 min | 95 |
| Amplification/ Fluorescence detection | 40× | 15 s | 95 | ||
| 1 min | 60 |
| Target Species | Primers and Probe | Sequence |
|---|---|---|
| X. fastidiosa | 1-XF-F | 5′-AAC CTG CGT GAC TCT GGT TT-3′ |
| 1-XF-R | 5′-CAT GTT TCG CTG CTT GGT CC-3′ | |
| 1-XF-Probe | 5′-FAM-GCT CAG GCT GAC GGT TTC ACA GTG CA-BHQ1-3′ | |
| X. fastidiosa subspecies fastidiosa | 2-XFF-F | 5′-TTA CAT CGT TTT CGC GCA CG-3′ |
| 2-XFF-R | 5′-TCG GTT GAT CGC AAT ACC CA-3′ | |
| 2-XFF-Probe | 5′-HEX-CCC GAC TCG GCG CGG TTC CA-BHQ1-3′ | |
| X. fastidiosa subspecies multiplex | 3-XFM-F | 5′-ACG ATG TTT GAG CCG TTT GC-3′ |
| 3-XFM-R | 5′-TGT CAC CCA CTA CGA AAC GG-3′ | |
| 3-XFM-Probe | 5′-ROX- ACG CAG CCC ACC ACG ATT TAG CCG-BHQ2-3′ | |
| X. fastidiosa subspecies pauca | 4-XFP-F | 5′-TGC GTT TTC CTA GGT GGC AT-3′ |
| 4-XFP-R | 5′-GTT GGA ACC TTG AAT GCG CA-3′ | |
| 4-XFP-Probe | 5′-CY5-CCA AAG GGC GGC CAC CTC GC-BHQ2-3′ |
|
TqPCR Mastermix |
Performance Criteria (%) | Xf | Xff | Xfm | Xfp |
|---|---|---|---|---|---|
| A | DSE | 97.91 | 100 | 100 | 91.66 |
| DSP | 100 | 100 | 100 | 93.5 | |
| ACC | 98.61 | 100 | 100 | 91.66 | |
| B | DSE | 95 | 100 | 100 | 87.5 |
| DSP | 100 | 100 | 100 | 100 | |
| ACC | 96.6 | 100 | 100 | 98 | |
| C | DSE | 87.5 | 87.5 | 87.5 | 100 |
| DSP | 93.7 | 100 | 100 | 100 | |
| ACC | 89.6 | 97.9 | 95.8 | 100 | |
| D | DSE | 100 | 100 | 100 | 100 |
| DSP | 100 | 100 | 100 | 100 | |
| ACC | 100 | 100 | 100 | 100 | |
| E | DSE | 100 | 100 | 100 | 100 |
| DSP | 100 | 100 | 100 | 100 | |
| ACC | 100 | 100 | 100 | 100 |
|
TqPCR Master Mix | Xf | Xff | Xfm | Xfp | ||||
|---|---|---|---|---|---|---|---|---|
| FP | FN | FP | FN | FP | FN | FP | FN | |
| A | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 |
| B | 0 | 2 | 0 | 0 | 0 | 0 | 0 | 1 |
| C | 1 | 4 | 0 | 1 | 0 | 2 | 0 | 0 |
| D | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| E | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 |
| TqPCR Master Mix | Xf | Xff | Xfm | Xfp |
|---|---|---|---|---|
| A | 95.83 | 100 | 100 | 95.83 |
| B | 95 | 100 | 100 | 100 |
| C | 95 | 93.75 | 87.5 | 100 |
| D | 100 | 100 | 100 | 100 |
| E | 100 | 100 | 100 | 100 |
|
TqPCR Master Mix | Xf | Xff | Xfm | Xfp | ||||
|---|---|---|---|---|---|---|---|---|
| FP | FN | FP | FN | FP | FN | FP | FN | |
| A | - | 1 | - | - | - | - | - | 1 |
| B | - | 1 | - | - | - | - | - | - |
| C | 1 | 3 | - | 1 | - | 1 | - | - |
| D | - | - | - | - | - | - | - | - |
| E | - | - | - | - | - | - | - | - |
| Real-Time Test | Plant Matrix/ Xf Subspecies | Sub Species | Bacterial Colony-Forming Units mL−1 | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 106 | 105 | 104 | 103 | 102 | 101 | ||||||||||||||||
| N | Cq | DS | N | Cq | DS | N | Cq | DS | N | Cq | DS | N | Cq | DS | N | Cq | DS | ||||
| Harper et al. [18] | O/Xfp | Xf | 9/9 | 24.97 | 0.44 | 9/9 | 29.02 | 0.54 | 9/9 | 32.08 | 0.16 | 9/9 | 35.48 | 0.74 | 8/9 | 37.86 | 0.82 | 4/9 | 38.95 | 0.44 | |
| Vv/Xff | 9/9 | 23.58 | 0.46 | 9/9 | 26.28 | 0.45 | 9/9 | 28.47 | 1.22 | 9/9 | 31.76 | 1.40 | 9/9 | 34.82 | 1.38 | 7/9 | 36.51 | 1.32 | |||
| Pd/Xfm | 9/9 | 25.75 | 0.48 | 9/9 | 29.74 | 0.20 | 9/9 | 32.90 | 1.19 | 9/9 | 35.60 | 0.80 | 9/9 | 36.52 | 0.98 | 3/9 | 36.03 | 1.57 | |||
| Hodgetts et al. [20] | O/Xfp | Xf | 9/9 | 24.13 | 0.46 | 9/9 | 27.46 | 0.46 | 9/9 | 30.22 | 0.43 | 9/9 | 33.28 | 0.39 | 8/9 | 35.94 | 0.78 | 6/9 | 37.01 | 0.44 | |
| Vv/Xff | Xff | 9/9 | 23.12 | 0.47 | 9/9 | 26.06 | 0.41 | 9/9 | 29.19 | 0.32 | 9/9 | 32.40 | 0.47 | 9/9 | 35.06 | 0.55 | 6/9 | 36.91 | 0.62 | ||
| Pd/Xfm | Xfm | 9/9 | 24.09 | 0.38 | 9/9 | 28.34 | 0.19 | 9/9 | 31.28 | 0.78 | 9/9 | 34.04 | 0.46 | 9/9 | 36.10 | 1.60 | 8/9 | 35.24 | 1.98 | ||
| Dupas et al. [16] | Bio-Rad | O/Xfp | Xf | 9/9 | 28.91 | 0.51 | 9/9 | 31.71 | 0.60 | 9/9 | 36.06 | 1.05 | 8/9 | NA | NA | 0/9 | NA | 0/9 | 0/9 | NA | NA |
| Xfp | 9/9 | 25.57 | 0.54 | 9/9 | 28.64 | 0.43 | 9/9 | 31.70 | 0.55 | 9/9 | 36.23 | 0.86 | 3/9 | NA | 0/9 | 0/9 | NA | NA | |||
| Vv/Xff | Xf | 9/9 | 28.06 | 0.73 | 9/9 | 31.96 | 0.82 | 9/9 | 38.39 | 0.78 | 0/9 | NA | NA | 0/9 | NA | 0/9 | 0/9 | NA | NA | ||
| Xff | 9/9 | 25.04 | 0.49 | 9/9 | 28.68 | 0.20 | 9/9 | 33.49 | 1.90 | 6/9 | NA | NA | 0/9 | NA | 0/9 | 0/9 | NA | NA | |||
| Pd/Xfm | Xf | 9/9 | 29.65 | 0.72 | 9/9 | 36.04 | 1.46 | 9/9 | NA | NA | 0/9 | NA | NA | 0/9 | NA | 0/9 | 0/9 | NA | NA | ||
| Xfm | 9/9 | 26.04 | 0.47 | 9/9 | 30.05 | 0.22 | 9/9 | 34.29 | 1.67 | 6/9 | NA | NA | 0/9 | NA | 0/9 | 0/9 | NA | NA | |||
| Agilent | O/Xfp | Xf | 9/9 | 26.30 | 0.45 | 9/9 | 29.48 | 0.50 | 9/9 | 31.91 | 0.32 | 5/9 | 35.60 | 0.66 | 0/9 | NA | 0/9 | 0/9 | NA | NA | |
| Xfp | 9/9 | 23.99 | 0.47 | 9/9 | 27.28 | 0.46 | 9/9 | 29.78 | 0.46 | 9/9 | 33.07 | 1.05 | 1/9 | 38.94 | 0/9 | 0/9 | NA | NA | |||
| Vv/Xff | Xf | 9/9 | 25.46 | 0.45 | 9/9 | 28.15 | 0.33 | 9/9 | 31.20 | 0.29 | 9/9 | 34.96 | 1.84 | 9/9 | NA | 0/9 | 0/9 | NA | NA | ||
| Xff | 9/9 | 23.74 | 0.42 | 9/9 | 26.45 | 0.36 | 9/9 | 29.59 | 0.41 | 9/9 | 32.75 | 0.29 | 9/9 | 35.77 | 2/9 | 2/9 | NA | NA | |||
| Pd/Xfm | Xf | 9/9 | 27.32 | 0.33 | 9/9 | 31.23 | 0.14 | 9/9 | 34.22 | 1.10 | 9/9 | 35.23 | NA | 1/9 | NA | 0/9 | 0/9 | NA | NA | ||
| Xfm | 9/9 | 24.26 | 0.37 | 9/9 | 28.16 | 0.13 | 9/9 | 31.12 | 0.87 | 9/9 | 34.51 | 1.08 | 6/9 | 36.02 | 0/9 | 0/9 | 35.64 | 1.68 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pucci, N.; Scala, V.; Cesari, E.; Crosara, V.; Fiorani, R.; L’Aurora, A.; Lucchesi, S.; Tatulli, G.; Barra, E.; Ciarroni, S.; et al. An Inter-Laboratory Comparative Study on the Influence of Reagents to Perform the Identification of the Xylella fastidiosa Subspecies Using Tetraplex Real Time PCR. Horticulturae 2023, 9, 1053. https://doi.org/10.3390/horticulturae9091053
Pucci N, Scala V, Cesari E, Crosara V, Fiorani R, L’Aurora A, Lucchesi S, Tatulli G, Barra E, Ciarroni S, et al. An Inter-Laboratory Comparative Study on the Influence of Reagents to Perform the Identification of the Xylella fastidiosa Subspecies Using Tetraplex Real Time PCR. Horticulturae. 2023; 9(9):1053. https://doi.org/10.3390/horticulturae9091053
Chicago/Turabian StylePucci, Nicoletta, Valeria Scala, Erica Cesari, Valeria Crosara, Riccardo Fiorani, Alessia L’Aurora, Simone Lucchesi, Giuseppe Tatulli, Eleonora Barra, Serena Ciarroni, and et al. 2023. "An Inter-Laboratory Comparative Study on the Influence of Reagents to Perform the Identification of the Xylella fastidiosa Subspecies Using Tetraplex Real Time PCR" Horticulturae 9, no. 9: 1053. https://doi.org/10.3390/horticulturae9091053
APA StylePucci, N., Scala, V., Cesari, E., Crosara, V., Fiorani, R., L’Aurora, A., Lucchesi, S., Tatulli, G., Barra, E., Ciarroni, S., De Amicis, F., Fascella, S., Giacobbi, F., Gaffuri, F., Gualandri, V., Landi, L., Loconsole, G., Molinatto, G., Pollastro, S., ... Loreti, S. (2023). An Inter-Laboratory Comparative Study on the Influence of Reagents to Perform the Identification of the Xylella fastidiosa Subspecies Using Tetraplex Real Time PCR. Horticulturae, 9(9), 1053. https://doi.org/10.3390/horticulturae9091053







