Cucumis melo L. Germplasm in Tunisia: Unexploited Sources of Resistance to Fusarium Wilt
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Material
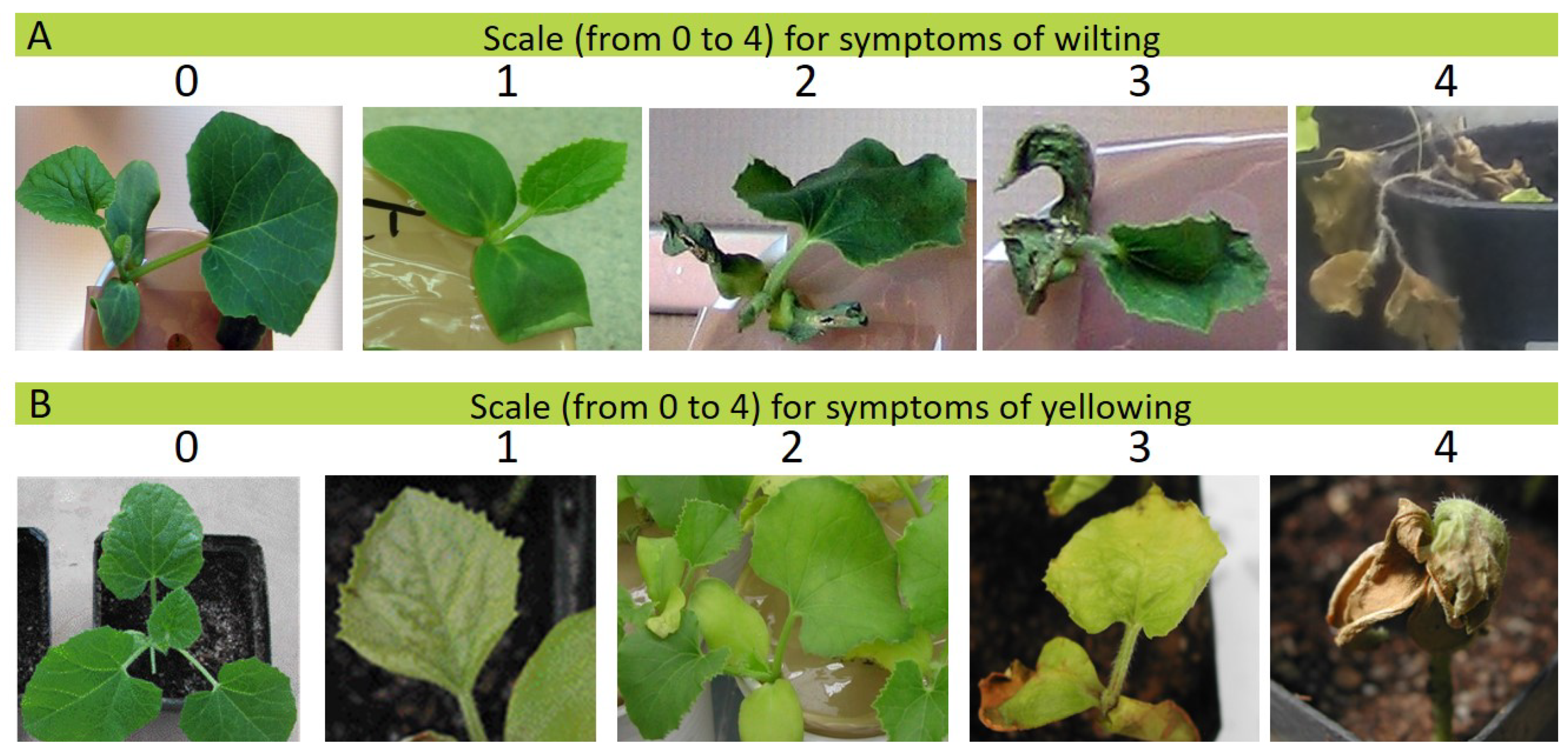
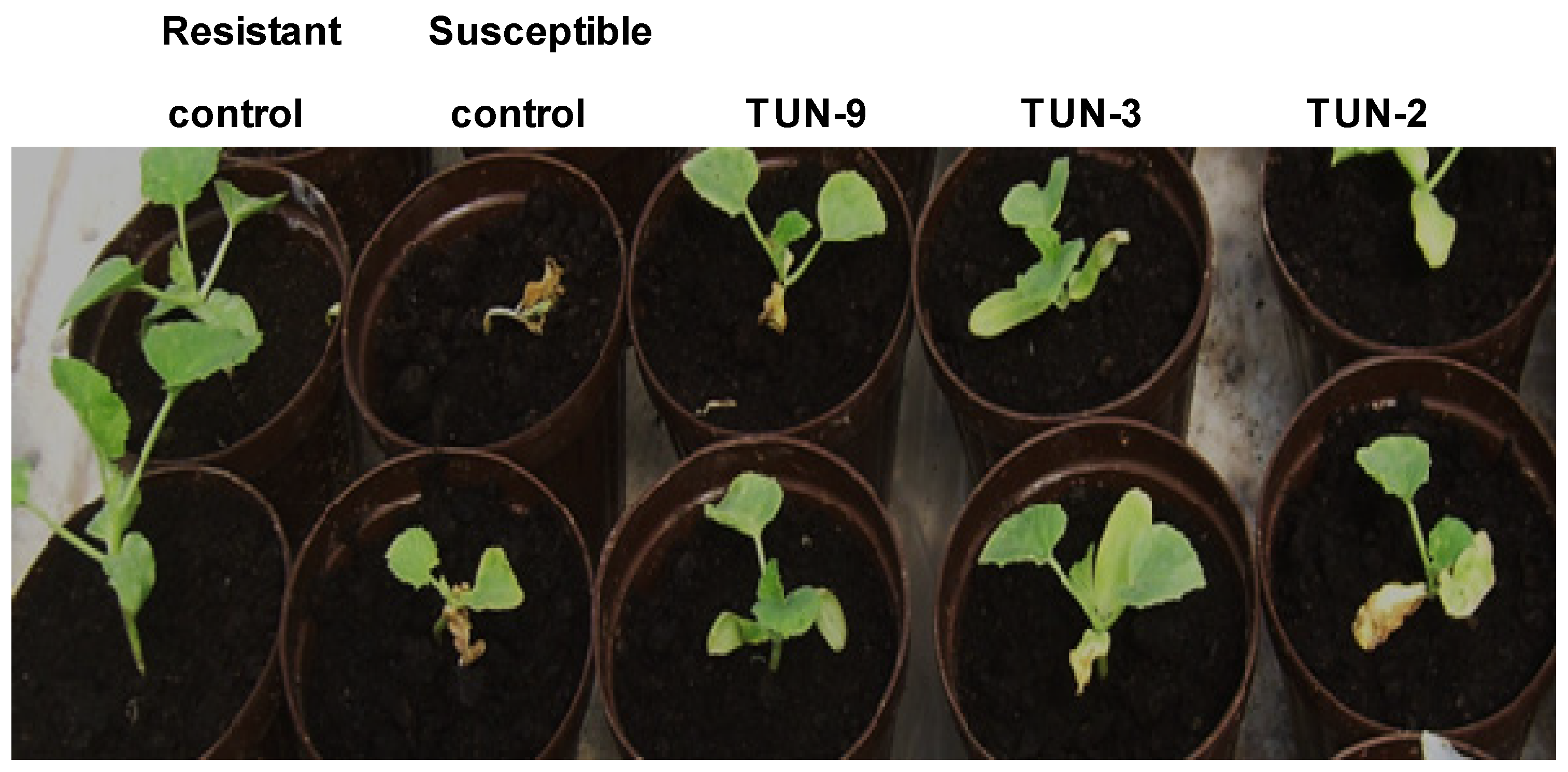
2.2. Phenotypic Evaluation of the Resistance to Fom
2.3. Analyses of Molecular Markers for Fom Races 0, 1, and 2
3. Results
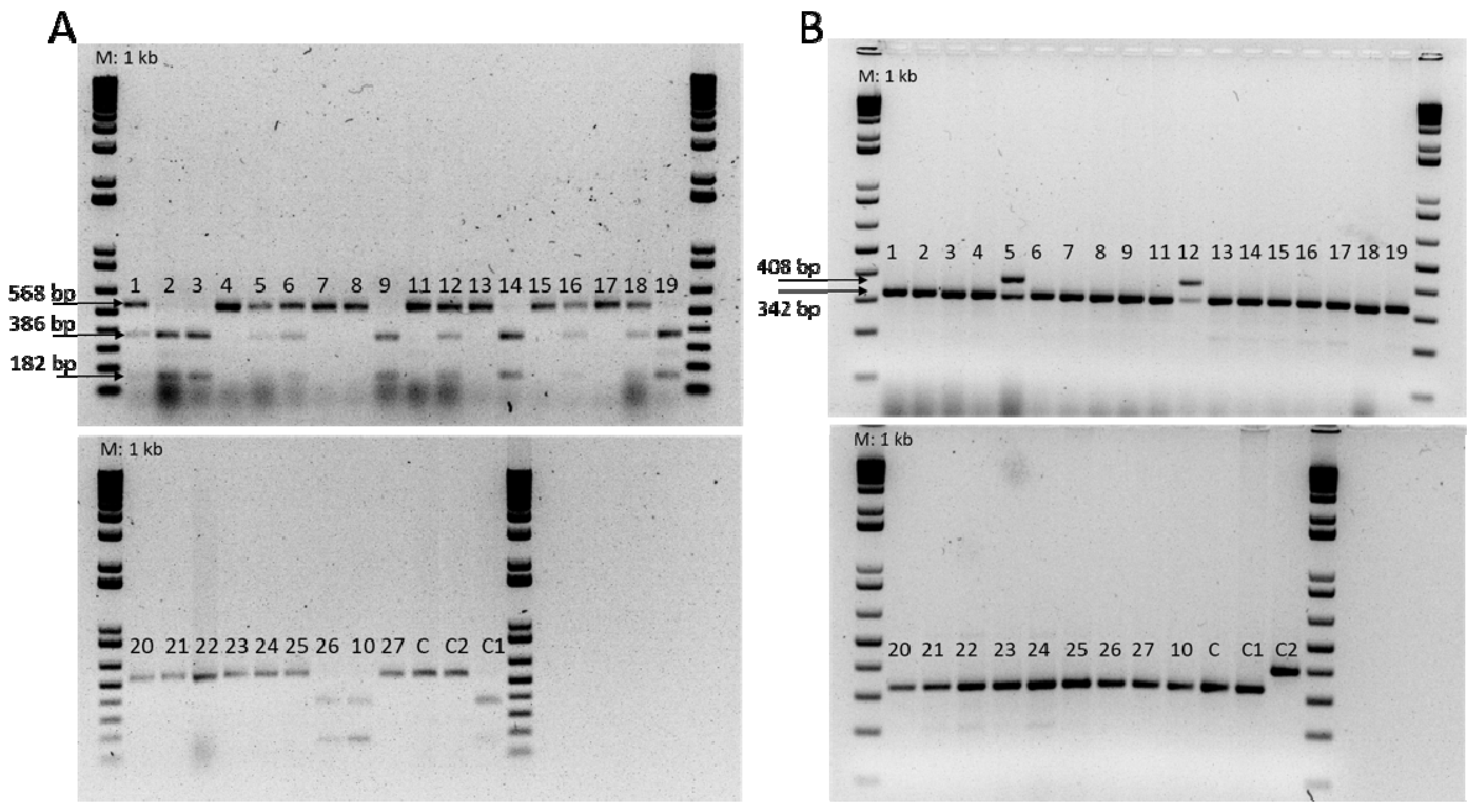
3.1. Phenotypic and Molecular Evaluation of the Resistance to Fom Races 0, 1, and 2
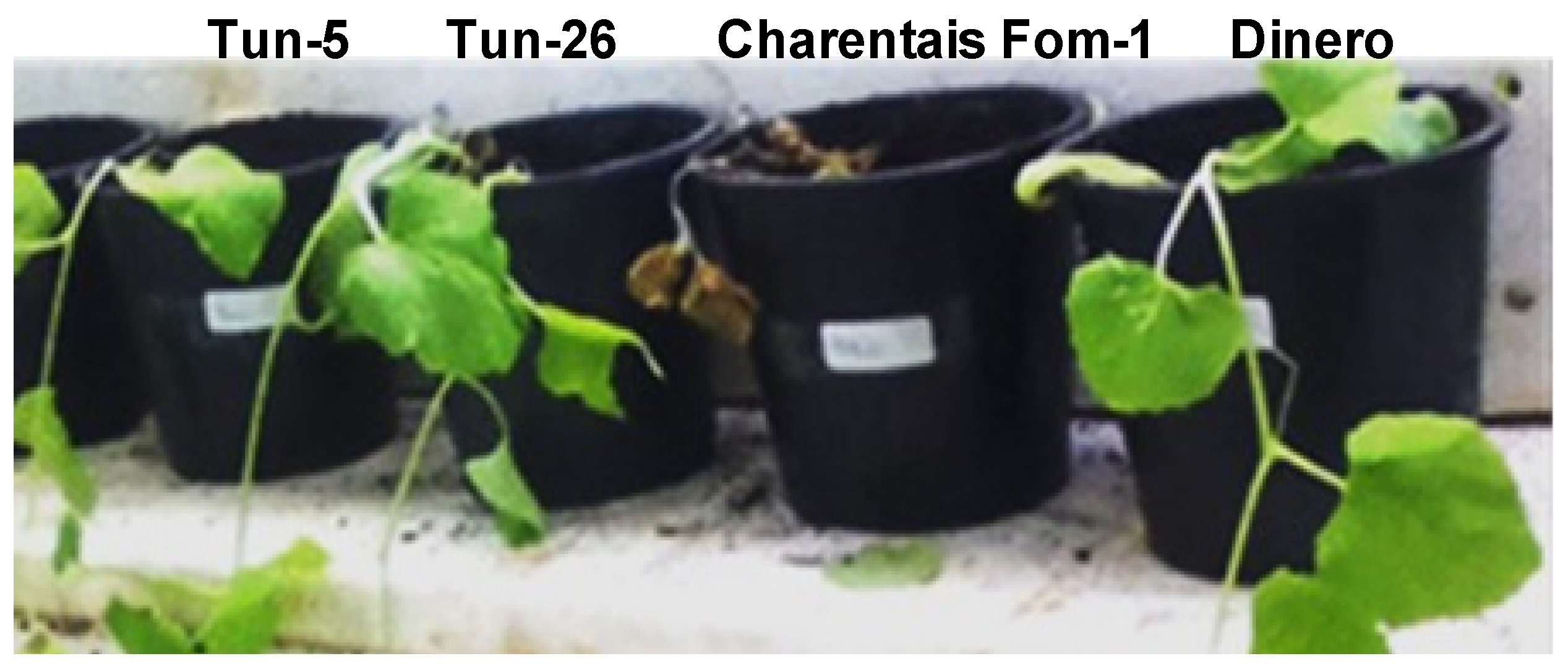
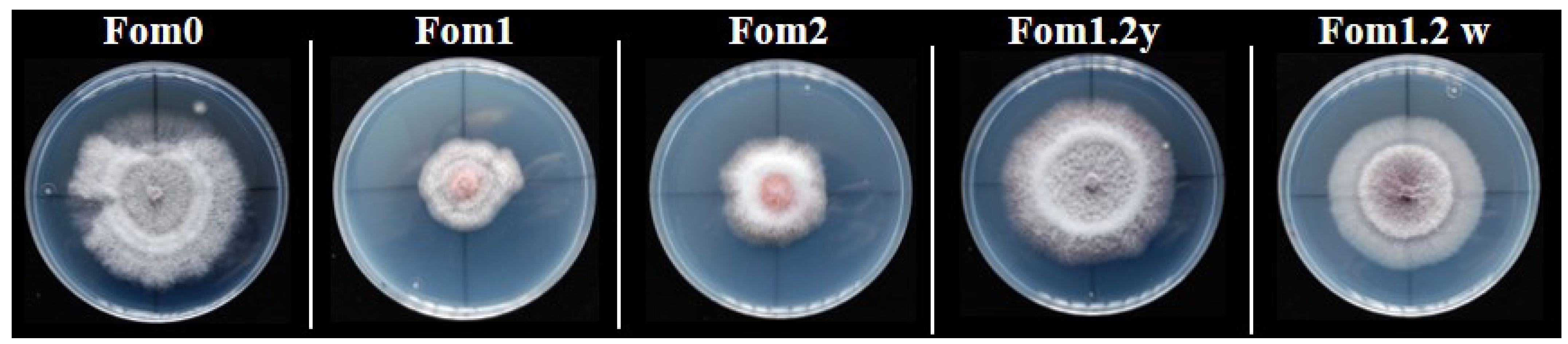
3.2. Phenotypic Evaluations of the Resistance to Fom Pathotypes 1.2y and 1.2w
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Luongo, L.; Ferrarini, A.; Haegi, A.; Vitale, S.; Polverari, A.; Belisario, A. Genetic diversity and pathogenicity of Fusarium oxysporum f.sp melonis races from different areas of Italy. J. Phytopathol. 2015, 163, 73–83. [Google Scholar] [CrossRef]
- González, V.; Armengol, J.; Garcés-Claver, A. First Report of Fusarium petroliphilum Causing Fruit Rot of Butternut Squash in Spain. Plant Dis. 2018, 102, 1662. [Google Scholar] [CrossRef]
- González, V.; Armijos, E.; Garcés-Claver, A. FungalEndophytes as Biocontrol Agents against the Main Soil-Borne Diseases of Melon and Watermelon in Spain. Agronomy 2020, 10, 820. [Google Scholar] [CrossRef]
- González, V.; García-Martínez, S.; Flores-León, A.; Ruiz, J.J.; Pico, M.B.; Garces-Claver, A. Neocosmospora keratoplastica, a relevant human fusarial pathogenis found to be associated with wilt and root rot of Muskmelon and Watermelon crops in Spain: Epidemiological and molecular evidences. Eur. J. Plant. Pathol. 2020, 156, 1189–1196. [Google Scholar] [CrossRef]
- González, V.; García-Martínez, S.; Ruiz, J.J.; Flores-León, A.; Picó, B.; Garcés-Claver, A. First Report of Neocosmospora falciformis Causing Wilt and Root Rot of Muskmelon in Spain. Plant Dis. 2020, 104, 1256. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; Sta-Baba, R.; Ayed, C.; Belgacem, S.; Boughalleb, N.; Cherif, M. Physiological races of Fusarium oxysporum f.sp. melonis in Tunisia. Phytoparasitica 2013, 41, 593–596. [Google Scholar] [CrossRef]
- Flores-León, A.; García-Martínez, S.; González, V.; Garcés-Claver, A.; Martí, R.; Julián, C.; Sifres, A.; Pérez-De-Castro, A.; Díez, M.J.; López, C.; et al. Grafting Snake Melon [Cucumis melo L. subsp. melo Var. flexuosus (L.) Naudin] in Organic Farming: Effects on Agronomic Performance; Resistance to Pathogens; Sugar, Acid, and VOC Profiles; and Consumer Acceptance. Front. Plant Sci. 2021, 12, 613845. [Google Scholar] [CrossRef] [PubMed]
- Rhouma, A.; Salem, I.B.; M’hamdi, M.; Boughalleb, N. Relationship study among soils physico-chemical properties and Monosporascus cannonballus ascospores densities for cucurbit fields in Tunisia. Eur. J. Plant Pathol. 2019, 153, 65–78. [Google Scholar] [CrossRef]
- Risser, G.; Banihashemi, Z.; Davis, D.W. A proposed nomenclature of Fusarium oxysporum f.sp. melonis races and resistancegenes in Cucumis melo. Phytopathology 1976, 66, 1105–1106. [Google Scholar] [CrossRef]
- Tamietti, G.; Valentino, D. Soil solarization as an ecological method for the control of Fusarium wilt of melon in Italy. Crop Prot. 2006, 25, 389–397. [Google Scholar] [CrossRef]
- Oumouloud, A.; Arnedo-Andrés, M.S.; González-Torres, R.; Álvarez, J.M. Inheritance of resistance to Fusarium oxysporum f. sp. melonis races 0 and 2 in melon accession Tortuga. Euphytica 2010, 176, 183–189. [Google Scholar] [CrossRef]
- Oumouloud, A.; El-Otmani, M.; Chikh-Rouhou, H.; Garcés-Claver, A.; Torres, R.G.; Perltreves, R.; Alvarez, J.M. Breeding melon for resistance to Fusarium wilt: Recent developments. Euphytica 2013, 192, 155–169. [Google Scholar] [CrossRef] [Green Version]
- Chikh-Rouhou, H.; González-Torres, R.; Oumouloud, A.; Alvarez, J.M. Inheritance of race 1.2 Fusarium wilt resistance in four melon cultivars. Euphytica 2011, 182, 177–186. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; González-Torres, R.; Álvarez, J.M. Characterization of the resistance to Fusarium oxysporum f.sp melonis race 1.2 in Cucumis melo ‘BG-5384’. In Proceedings of the Cucurbitaceae 2008, IXth EUCARPIA Meeting on Genetics and Breeding of Cucurbitaceae, Avignon, France, 21–24 May 2008; pp. 419–422. [Google Scholar]
- Perchepied, L.; Pitrat, M. Polygenic Inheritance of Partial Resistance to Fusarium oxysporum f. sp. melonis Race 1.2 in Melon. Phytopathology 2004, 94, 1331–1336. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Herman, R.; Perl-Treves, R. Characterization and Inheritance of a New Source of Resistance to Fusarium oxysporum f. sp. melonis Race 1.2 in Cucumis melo. Plant Dis. 2007, 91, 1180–1186. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; González-Torres, R.; Alvarez, J.M.; Oumouloud, A. Screening and Morphological Characterization of Melons for Resistance to Fusarium oxysporum f.sp. melonis Race 1.2. HortScience 2010, 45, 1021–1025. [Google Scholar] [CrossRef] [Green Version]
- Chikh-Rouhou, H.; Alvarez, J.M.; González-Torres, R. Differential interaction between melon cultivars and race 1.2 of Fusarium oxysporum f.sp. melonis. Commun. Agric. Appl. Boil. Sci. 2007, 72, 825–829. [Google Scholar]
- Tezuka, T.; Waki, K.; Yashiro, K.; Kuzuya, M.; Ishikawa, T.; Takatsu, Y.; Miyagi, M. Construction of a linkage map and identification of DNA markers linked to Fom-1, a gene conferring resistance to Fusarium oxysporum f.sp. melonis race 2 in melon. Euphytica 2009, 168, 177–188. [Google Scholar] [CrossRef]
- Oumouloud, A.; Arnedo-Andres, M.S.; Gonzalez-Torres, R.; Alvarez, J.M. Development of molecular markers linked to the Fom-1 locus for resistance to Fusarium race 2 in melon. Euphytica 2008, 164, 347–356. [Google Scholar] [CrossRef]
- Brotman, Y.; Kovalski, I.; Dogimont, C.; Pitrat, M.; Portnoy, V.; Katzir, N.; Perl-Treves, R. Molecular markers linked to papaya ring spot virus resistance and Fusarium race 2 resistance in melon. Theor. Appl. Genet. 2004, 110, 337–345. [Google Scholar] [CrossRef]
- Wang, Y.-H.; Thomas, C.E.; Dean, R.A. Genetic mapping of a Fusarium wilt resistance gene (Fom-2) in melon (Cucumis melo L.). Mol. Breed. 2000, 6, 379–389. [Google Scholar] [CrossRef]
- Wechter, W.P.; Whitehead, M.P.; Thomas, C.E.; Dean, R.A. Identification of a randomly amplified polymorphic DNA marker linked to the Fom-2 Fusarium wilt resistance gene in muskmelon MR-1. Phytopathology 1995, 85, 1245–1249. [Google Scholar] [CrossRef]
- Oumouloud, A.; El Otmani, M.; Alvarez, J.M.A. Molecular characterization of Fom-1 gene and development of functional markers for molecular breeding of resistance to Fusarium race 2 in melon. Euphytica 2015, 205, 491–501. [Google Scholar] [CrossRef] [Green Version]
- Oumouloud, A.; Mokhtari, M.; Chikh-Rouhou, H.; Arnedo-Andrés, M.S.; González-Torres, R.; Álvarez, J.M. Characterization of the Fusarium wilt resistance Fom-2 gene in melon. Mol. Breed. 2012, 30, 325–334. [Google Scholar] [CrossRef]
- Andersen, J.R.; Lübberstedt, T. Functional markers in plants. Trends Plant Sci. 2003, 8, 554–560. [Google Scholar] [CrossRef] [PubMed]
- Perchepied, L.; Dogimont, C.; Pitrat, M. Strain-specific and recessive QTLs involved in the control of partial resistance to Fusarium oxysporum f. sp. melonis race 1.2 in a recombinant inbred line population of melon. Theor. Appl. Genet. 2005, 111, 65–74. [Google Scholar] [CrossRef]
- Herman, R.; Zvirin, Z.; Kovalski, I. Characterization of Fusarium race 1.2 resistance in melon and mapping of a major QTL for this trait near a fruit netting locus. In Proceedings of the IXth Eucarpia Meeting on Genetics and Breeding of Cucurbitaceae 2008, Avignon, France, 21–24 May 2008; pp. 149–151. [Google Scholar]
- Chikh-Rouhou, H.; Tlili, I.; Ilahy, R.; R’Him, T.; Sta-Baba, R. Fruit quality assessment and characterization of melon genotypes. Int. J. Veg. Sci. 2021, 27, 3–19. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; Ben Belgacem, A.M.; Sta-Baba, R.; Tarchoun, N.; Gómez-Guillamón, M.L. New source of resistance to Aphis gossypii in Tunisian melon accessions using phenotypic and molecular marker approaches. Phytoparasitica 2019, 47, 405–413. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; Mezghani, N.; Mnasri, S.; Mezghani, N.; Garcés-Claver, A. Assessing the Genetic Diversity and Population Structure of a Tunisian Melon (Cucumis melo L.) Collection Using Phenotypic Traits and SSR Molecular Markers. Agronomy 2021, 11, 1121. [Google Scholar] [CrossRef]
- Pitrat, M. Melon Genetic Resources: Phenotypic Diversity and Horticultural Taxonomy. In Genetics and Genomics of Cucurbitaceae; Grumet, R., Katzir, N., Garcia-Mas, J., Eds.; Springer: Cham, Switzerland, 2016; Volume 20, pp. 25–60. [Google Scholar]
- Álvarez, J.M.; González-Torres, R.; Mallor, C.; Gómez-Guillamón, M.L. Potential sources of resistanse to Fusarium wilt and powdery mildew in melons. HortScience 2005, 40, 1657–1660. [Google Scholar] [CrossRef] [Green Version]
- Mas, P.; Risser, G. Caracterisation, Symptômes et Virulence de Diverses Races de Fusarium oxysporum Schl. f. sp. melonis Sn. et Hans. In Proceedings of the Actes du Premier Congres de l’Union Phytopathologique Mediterraneenne 1966, Bari-Naples, Italy, 23 September 1966. [Google Scholar]
- Rodríguez-Maza, M.J.; Garcés-Claver, A.; Park, S.-W.; Kang, B.-C.; Arnedo-Andrés, M.S. A versatile PCR marker for pungency in Capsicum spp. Mol. Breed. 2012, 30, 889–898. [Google Scholar] [CrossRef]
- Meighan, A.M.; Rabiei, B.; Khodaparast, S.A. Identification of Fusarium wilt resistance sources in melon (Cucumis melo L.) landraces of Iran using marker-assisted selection technique. Australas. Plant Pathol. 2020, 49, 413–423. [Google Scholar] [CrossRef]
- Chikh-Rouhou, H.; Garcés-Claver, A.; Sta-Baba, R.; González, V.; Daami-Remadi, M. Screening for resistance to race 1 of Fusarium oxysporum f.sp melonis in Tunisian melon cultivars using molecular markers. Comm. Agric. Appl. Biol. Sci. 2018, 83, 87–92. [Google Scholar]
- Palomares-Rius, F.; Garcés-Claver, A.; Picó, B.; Esteras, C.; Yuste-Lisbona, F.; Gómez-Guillamón, M.L. ‘Carmen’, a Yellow-Canary melon resistant to Podosphaera xanthii, Aphis gossypii and Cucurbit Yellow Stunting Disorder Virus. HortScience 2018, 53, 1072–1075. [Google Scholar] [CrossRef] [Green Version]
- Oumouloud, A.; Torres, R.G.; Garcés-Claver, A.; Rouhou, H.C.; Álvarez, J. Differential response of Cucumis melo to Fusarium oxysporum f. sp. melonis race 1.2 isolates. Crop Prot. 2013, 44, 91–94. [Google Scholar] [CrossRef]
- Dhillon, N.P.S.; Monforte, A.J.; Pitrat, M.; Pandey, S.; Singh, P.K.; Reitsma, K.R.; Garcia-Mas, J.; Sharma, A.; McCreight, J.D. Melon landraces of India: Contributions and importance. In Plant Breeding Reviews; Janick, J., Ed.; Wiley: Hoboken, NJ, USA, 2012; Volume 35, pp. 85–150. [Google Scholar]



| ID | Accessions | Horticultural Group * |
|---|---|---|
| TUN-1 | Maazoun Chott-Mariem | inodorus |
| TUN-2 | Maazoun Menzel Chaker | inodorus |
| TUN-3 | Maazoun Mehdia (MM2009) | inodorus |
| TUN-4 | Maazoun Fethi | inodorus |
| TUN-5 | Fakous (FL) | flexuosus |
| TUN-6 | Fakous Salem 2014 | flexuosus |
| TUN-7 | Trabelsi | inodorus |
| TUN-8 | Galaoui | reticulatus |
| TUN-9 | Dziri (DZ P5 2011) | inodorus |
| TUN-10 | Lobneni | reticulatus |
| TUN-11 | Arbi | inodorus |
| TUN-12 | Horchay | chate |
| TUN-13 | Arbi 1 | inodorus |
| TUN-14 | Arbi 2 | inodorus |
| TUN-15 | Arbi 3 | inodorus |
| TUN-16 | Sarachika | inodorus |
| TUN-17 | RD | cantalupensis |
| TUN-18 | Rupa | cantalupensis |
| TUN-19 | Chamem (Ananastype) | reticulatus |
| TUN-20 | HTM Kairouan | reticulatus |
| TUN-21 | Acc Jendouba | inodorus |
| TUN-22 | Dziri (Menzel Kamel) | inodorus |
| TUN-23 | Ecotype arbi Dz | inodorus |
| TUN-24 | Maazoun (Kairouan) | inodorus |
| TUN-25 | Asli | inodorus |
| TUN-26 | Stambouli | inodorus |
| TUN-27 | V4 autoféc | inodorus |
| Primer | Primer Sequences | Fragment Size (bp) | Expected Genotype | Reference |
|---|---|---|---|---|
| FOM-1R | F: ATGAGTTTTGATAGTTTCATAAG R: GAACACTCCCTTAGATACTT | 182 + 386 568 | Fom-1/Fom-1 fom-1/fom-1 | Oumouloud et al. [24] |
| FOM2-R | F: GAGAAATTTGCAATGGGTGG R: TTACACTATTATTGCTCAACTTGC | 408 | Fom-2/Fom-2 | Oumouloud et al. [25] |
| FOM2-S | F: ATGAAAAGAAAAGATAACGACGA R: ATTGCTCTAAGTTGATCATATTCTG | 342 | fom-2/fom-2 |
| Melon Accessions | Reaction to Fom | Marker-Assisted Screening | |||
|---|---|---|---|---|---|
| Race 0 | Race 1 | Race 2 | Fom-1 Alleles | Fom-2 Alleles | |
| TUN-1 | R | S | R | R (Fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-2 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-3 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-4 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-5 | R | R | R | R (Fom-1/fom-1) | R (Fom-2/fom-2) |
| TUN-6 | R | S | R | R (Fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-7 | S | R | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-8 | R | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-9 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-10 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-11 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-12 | R | R | R(H) * | R (Fom-1/fom-1) | R (Fom-2/fom-2) |
| TUN-13 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-14 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-15 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-16 | R | S | R | R (Fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-17 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-18 | R | S | R | R (Fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-19 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-20 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-21 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-22 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-23 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-24 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-25 | R | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| TUN-26 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| TUN-27 | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| Control genotypes | |||||
| Charentais T | S | S | S | S (fom-1/fom-1) | S (fom-2/fom-2) |
| Charentais Fom-1 | R | S | R | R (Fom-1/Fom-1) | S (fom-2/fom-2) |
| Charentais Fom-2 | R | R | S | S (fom-1/fom-1) | R (Fom-2/Fom-2) |
| 1st Assay | 2nd Assay | |||
|---|---|---|---|---|
| Accession | 1.2y | 1.2w | 1.2y | 1.2w |
| TUN-1 | 45.5 b ± 8.17 | 47.8 b ± 5.04 | 38.5 c ± 11.50 | 45.5 b ± 3.83 |
| TUN-2 | 39.1 c ± 9.88 | 42.0 b ± 12.30 | 31.5 c ± 8.57 | 37.3 c ± 8.47 |
| TUN-3 | 38.5 c ± 10.96 | 42.3 b ± 10.80 | 35.0 c ± 10.84 | 32.7 c ± 7.22 |
| TUN-4 | 46.9 b ± 4.45 | 55.1 a ± 2.11 | 57.8 a ± 1.91 | 51.9 a ± 6.79 |
| TUN-5 | 10.5 ef ± 8.51 | 4.4 e ± 1.31 | 8.2 e ± 5.71 | 4.1 e ± 2.15 |
| TUN-6 | 47.3 b ± 3.16 | 44.6 b ± 4.97 | 44.9 b ± 4.65 | 54.3 a ± 1.91 |
| TUN-7 | 51.3 a ± 4.65 | 54.0 a ± 4.07 | 48.9 ab ± 2.63 | 51.3 a ± 5.26 |
| TUN-8 | 42.5 b ± 10.54 | 51.6 a ± 4.89 | 41.9 b ± 9.49 | 40.8 b ± 5.75 |
| TUN-9 | 38.5 c ± 10.96 | 43.5 b ± 11.43 | 37.3 c ± 10.53 | 36.2 c ± 12.05 |
| TUN-10 | 35.9 c ± 12.49 | 40.0 b ± 10.6 | 31.5 c ± 8.57 | 38.5 c ± 11.50 |
| TUN-11 | 51.8 a ± 2.72 | 54.3 a ± 2.78 | 51.3 a ± 1.91 | 51.9 a ± 5.60 |
| TUN-12 | 25.3 d ± 5.07 | 28.7 d ± 10.89 | 23.5 d ± 8.57 | 28.3 d ± 7.22 |
| TUN-13 | 51.8 a ± 6.56 | 53.4 a ± 2.63 | 50.1 a ± 5.15 | 49.6 b ± 7.47 |
| TUN-14 | 36.8 c ± 10.81 | 38.2 c ± 10.28 | 36.8 c ± 10.32 | 33.3 c ± 6.91 |
| TUN-15 | 53.6 a ± 3.60 | 55.4 a ± 2.20 | 50.1 a ± 8.69 | 49.6 b ± 4.09 |
| TUN-16 | 48.4 b ± 7.58 | 38.2 c ± 7.07 | 43.8 b ± 4.82 | 38.5 c ± 8.57 |
| TUN-17 | 47.2 b ± 6.56 | 59.5 a±3.65 | 41.3 b ± 1.91 | 51.8 a ± 5.26 |
| TUN-18 | 37.3 c ± 9.61 | 37.0 c ± 7.82 | 49.6 b ± 6.79 | 43.8 b ± 5.75 |
| TUN-19 | 36.5 c ± 11.69 | 36.2 c ± 7.51 | 32.7 c ± 7.87 | 36.2 c ± 9.03 |
| TUN-20 | 48.3 b ± 3.44 | 52.8 a ± 4.07 | 51.3 a ± 1.91 | 47.8 b ± 4.78 |
| TUN-21 | 48.3 b ± 6.56 | 53.1 a ± 5.36 | 35.9 c ± 4.92 | 53.1 a ± 7.47 |
| TUN-22 | 56.5 a ± 3.65 | 51.9 a ± 5.74 | 51.3 a ± 4.07 | 48.4 b ± 4.09 |
| TUN-23 | 51.3 a ± 10.91 | 54.3 a ± 5.06 | 50.1 a ± 3.65 | 54.3 a ± 3.67 |
| TUN-24 | 50.1 a ± 4.91 | 53.1 a ± 3.60 | 51.3 a ± 5.84 | 51.3 a ± 5.71 |
| TUN-25 | 51.8 a ± 2.72 | 52.5 a ± 3.65 | 50.1 a ± 3.60 | 52.5 a ± 7.53 |
| TUN-26 | 11.1 e ± 6.12 | 8.2 e ± 5.84 | 10.5 e ± 8.57 | 8.2 e ± 6.88 |
| TUN-27 | 59.8 a ± 7.07 | 56.3 a ± 1.01 | 48.9 ab ± 4.09 | 43.7 b ± 3.61 |
| Control genotypes | ||||
| Charentais Fom-1 | 60.2 a ± 2.44 | 58.3 a ± 1.72 | 49.6 ab ± 4.52 | 56.6 a ± 1.42 |
| Charentais Fom-2 | 59.2 a ± 4.34 | 55.4 a ± 2.34 | 50.2 a ± 2.72 | 50.8 a ± 6.52 |
| Dinero | 5.5 f ± 4.23 | 2.6 e ± 1.74 | 5.3 e ± 2.25 | 3.5 e ± 2.42 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chikh-Rouhou, H.; Gómez-Guillamón, M.L.; González, V.; Sta-Baba, R.; Garcés-Claver, A. Cucumis melo L. Germplasm in Tunisia: Unexploited Sources of Resistance to Fusarium Wilt. Horticulturae 2021, 7, 208. https://doi.org/10.3390/horticulturae7080208
Chikh-Rouhou H, Gómez-Guillamón ML, González V, Sta-Baba R, Garcés-Claver A. Cucumis melo L. Germplasm in Tunisia: Unexploited Sources of Resistance to Fusarium Wilt. Horticulturae. 2021; 7(8):208. https://doi.org/10.3390/horticulturae7080208
Chicago/Turabian StyleChikh-Rouhou, Hela, Maria Luisa Gómez-Guillamón, Vicente González, Rafika Sta-Baba, and Ana Garcés-Claver. 2021. "Cucumis melo L. Germplasm in Tunisia: Unexploited Sources of Resistance to Fusarium Wilt" Horticulturae 7, no. 8: 208. https://doi.org/10.3390/horticulturae7080208
APA StyleChikh-Rouhou, H., Gómez-Guillamón, M. L., González, V., Sta-Baba, R., & Garcés-Claver, A. (2021). Cucumis melo L. Germplasm in Tunisia: Unexploited Sources of Resistance to Fusarium Wilt. Horticulturae, 7(8), 208. https://doi.org/10.3390/horticulturae7080208








