Nonconventional Yeasts Engineered Using the CRISPR-Cas System as Emerging Microbial Cell Factories
Abstract
1. Introduction
2. Industrial Value of NCYs
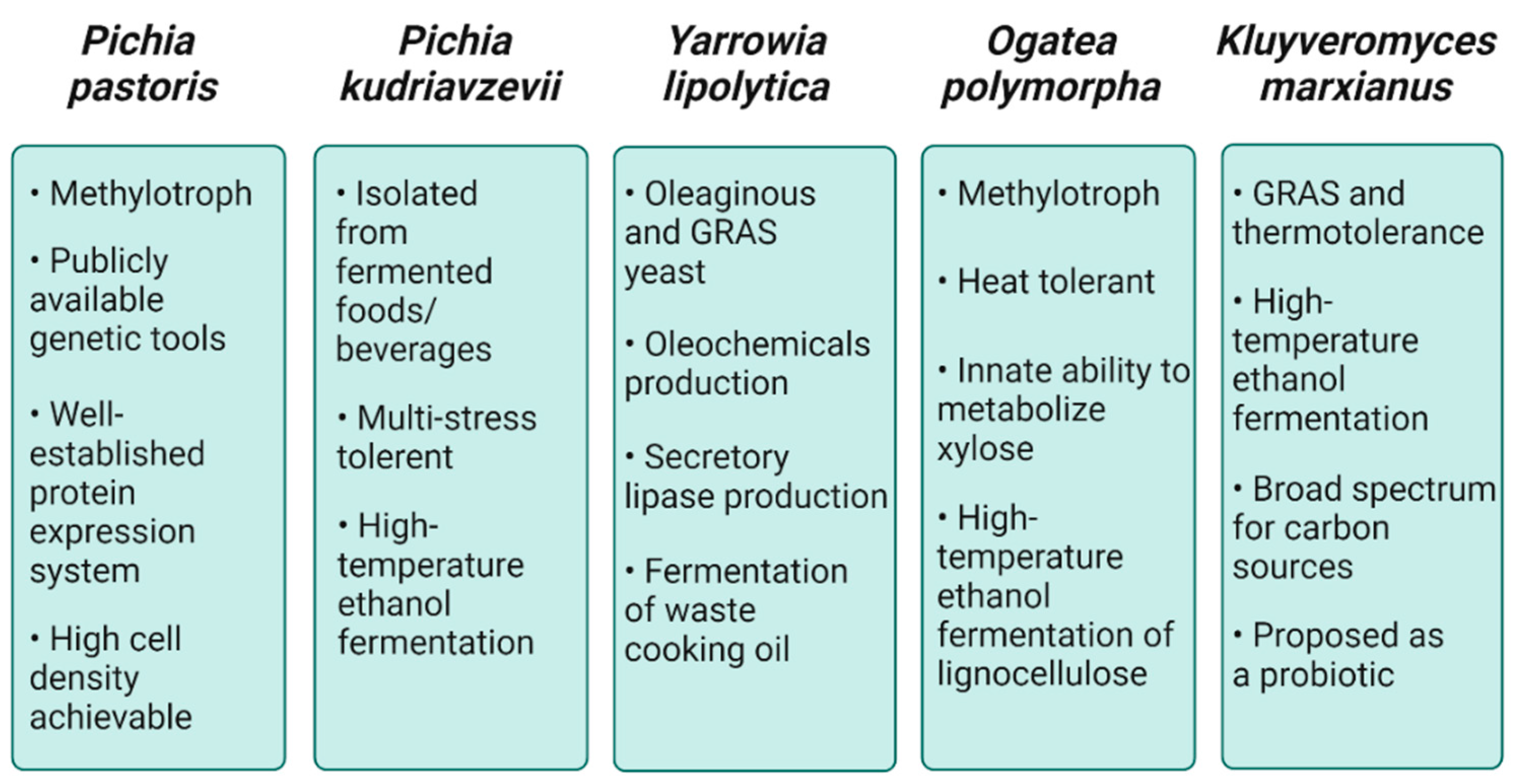
2.1. Pichia pastoris
2.2. Pichia kudriavzevii
2.3. Yarrowia lipolytica
2.4. Ogataea polymorpha
2.5. Kluyveromyces marxianus
3. Genetic Engineering Tools for NCYs
4. CRISPR-Cas System-Guided Metabolic Engineering in NCYs
4.1. CRISPR-Cas System: Classification, Components, and Mechanism
4.2. Challenges and Strategies of CRISPR-Cas9-Guided Genome Editing in NCYs
| Strain | sgRNA Promoter | Plasmid (Backbone) | Cas9 Promoter | Editing Efficiency 1 (%) | Reference |
|---|---|---|---|---|---|
| Y. lipolytica | SCR1′-tRNA2 TEFin (Pol II) 3 | pCRISPRyl pCASyl pGGA | TEF1 | 0–68.9 | [71,72,73,74] |
| O. polymorpha | ScSNR52 (Pol III) ScTDH3 (Pol III) | pCRCT pYTK079 | ScTEF1 AaTEF | 1–75 | [75,76] |
| P. pastoris | HTX1 (Pol II) 3 PFK300 (Pol II) 3 LAT1(Pol II) 3 | pPpT4 p414 BB3cH | HTX1 GAP | 75–93.8 | [77,78,79] |
| K. marxianus | ScTDH3 (Pol II) 3 RPR1′-tRNA (Pol III) 2 | pYTK079 pIW601 | AaTEF 1 ScTEF1 | 10–82 | [76,80] |
| P. kudriavzevii | RPR1 (Pol III) RPR1′-tRNA (Pol III) 2 | pRS416 pRS415 pCast | TEF1 | 64 | [30,81,82] |
4.3. Biotechnological Application of CRISPR-Cas9-Introduced NCYs
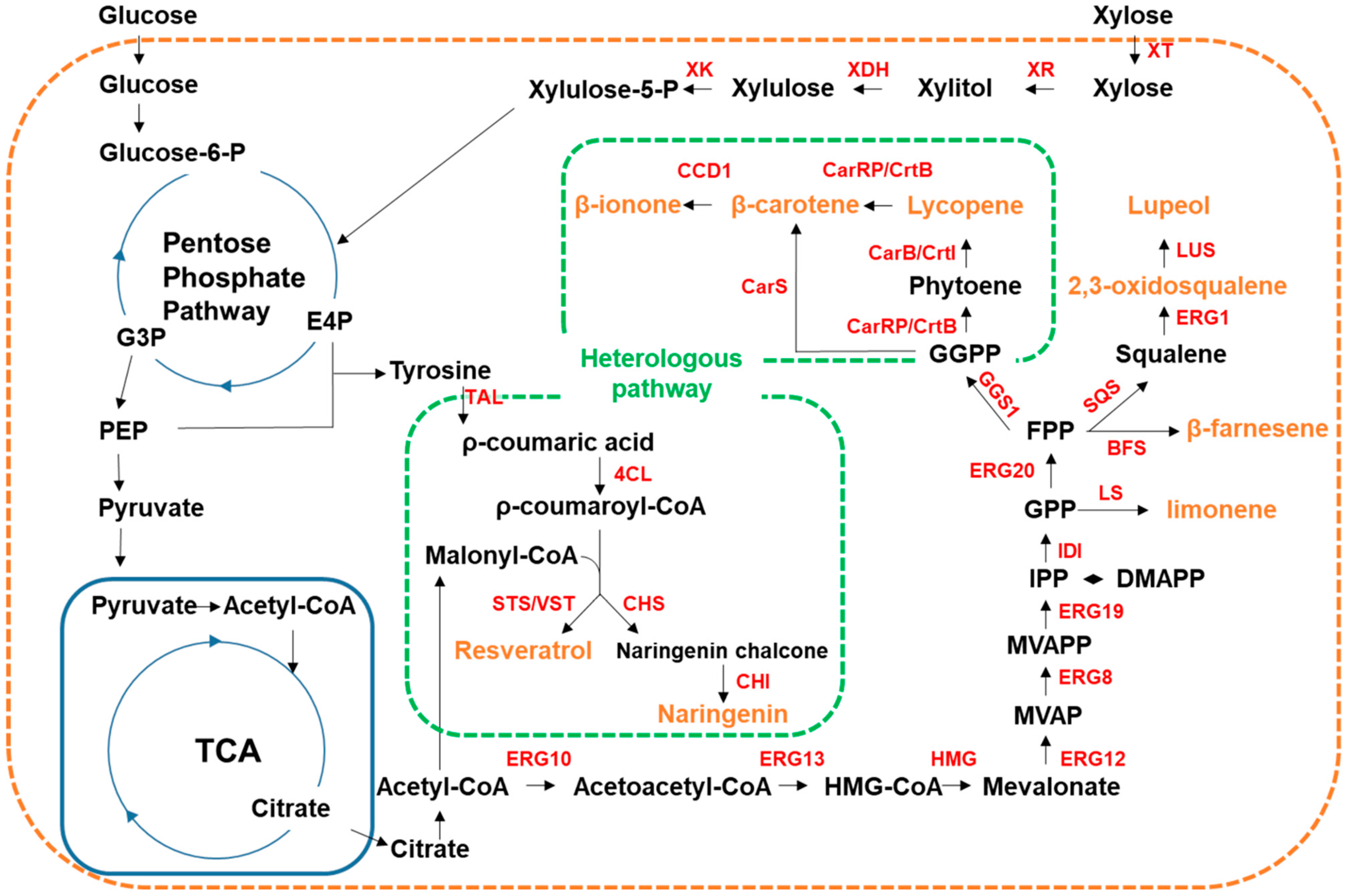
4.3.1. Secondary Plant Metabolites
| Strain | Target Genes | Product | Reference | |
|---|---|---|---|---|
| Endogenous Gene Editing | Heterologous Gene Editing | |||
| Y. lipolytica | HMG1, GGS1 | crtE (Pantoea ananatis), crtI (P. ananatis), crtB (P. ananatis) | Lycopene 3.38 mg/g DCW 2 | [71] |
| GGS1 | carB (Mucor circinelloides), carRP (M. circinelloides), | β-carotene 4.8 mg/g DCW | [72] | |
| GGS1, ERG13, HMG | carB (M. circinelloides), carRP (M. circinelloides) | β-carotene 4.5 g/L | [73] | |
| GGS1, HMG1, ERG8, ERG10, ERG12, ERG13, ERG20, ERG19, IDI | carB (M. circinelloides), carRP (M. circinelloides), CCD1 (Petunia hybrid), PK (Bifidobacterium bifidum), PTA (Bacillus subtilis) | β-ionone 358.4 mg/L 0.98 g/L (fed-batch) | [112] | |
| XK, HMG11, ERG121 | LS (Agastache rugosa) 1, NDPS (Solanum lycopersicum) 1, XR (Scheffersomyces stipitis), XDH (S. stipitis) | Limonene 20.57 mg/L | [113] | |
| HMG, ERG12, IDI, ERG20, SQS | BFS (Artemisia annua), LS (Citrus limon), LS (Perilla frutescens), CnVS (Callitropsis nootkatensis), crtI (Xanthophyllomyces dendrorhous), crtYB (X. dendrohous), acs (Salmonella enterica) | β-farnesene 955 mg/L Limonene 35.9 mg/L Valencene 113.9 mg/L Squalene 402.4 mg/L β-carotene 164 mg/L 2,3-oxidosqualene 22 mg/L | [111] | |
| HMG1, ERG1, ERG9, OLE1, PAH1, DGK1 | LUS (Ricinus communis) | Lupeol 441.72 mg/L | [114] | |
| GGS1 | carS (Schizochytrium sp.) | β-carotene 0.41 mg/g DCW | [74] | |
| XT1, XR1, XDH1, XKS1 | TAL (Rhodotorula glutinis), 4CL (Arabidopsis thaliana), CHS (A. thaliana), CHI (A. thaliana) | Naringenin 715.3 mg/L | [109] | |
| ARO4, ARO7 | TAL (Flavobacterium johnsoniae), VST (Vitis vinifera), 4CL (A. thaliana) | Resveratrol 12.4 g/L | [103] | |
| O. polymorpha | - | TAL (Herpetosiphon aurantiacus), STS (V. vinifera), 4CL (A. thaliana) | Resveratrol 97.23 mg/L | [75] |
4.3.2. Other Industrial Products
| Strain | Target Genes | Product | Reference | |
|---|---|---|---|---|
| Endogenous Gene Editing | Heterologous Gene Editing | |||
| Y. lipolytica | - | CAD (Aspergillus terreus), mttA (A. terreus) 1 | Itaconic acid 22.03 g/L | [116] |
| P. kudriavzevii | ICD, mttA1 | CAD (A. terreus) 1 | Itaconic acid 1.23 g/L | [30] |
| Y. lipolytica | SCT1, OLE1 | FAR (M. aquaeolei) | Fatty alcohol 5.75 g/L | [120] |
| Y. lipolytica | PLA2 | - | Lipid 25 g/L | [119] |
| Y. lipolytica | AXP | celB (Pyrococcus furiosus) | β-glycosidase 187.5 µkatoNPGal/L 2 | [124] |
| K. marxianus | ARO1, ARO2, ARO3, ARO4, ARO7, ARO8, ARO9, PHA2, TAL1, TKL1, RPE1, RKI1, LAC4 | xfpk (Bifidobacterium breve), ppsA (Escherichia coli), pta (Salmonella enterica) | 2-penylethanol 850 mg/L | [48] |
| Y. lipolytica | - | FAP (Chlorella variabilis) | Hydrocarbons 58.7 mg/L | [125] |
| K. marxianus | ACO2b, SDH2, RIP1, MSS51 | - | Ethyl acetate 150 mg/L | [123] |
5. Future Perspectives
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Li, W.; Shen, X.; Wang, J.; Sun, X.; Yuan, Q. Engineering microorganisms for the biosynthesis of dicarboxylic acids. Biotechnol. Adv. 2021, 48, 107710. [Google Scholar] [CrossRef] [PubMed]
- Karim, N.; Shishir, M.R.I.; Gowd, V.; Chen, W. Hesperidin-an emerging bioactive compound against metabolic diseases and its potential biosynthesis pathway in microorganism. Food Rev. Int. 2021. [Google Scholar] [CrossRef]
- Srinivasan, P.; Smolke, C.D. Biosynthesis of medicinal tropane alkaloids in yeast. Nature 2020, 585, 614–619. [Google Scholar] [CrossRef]
- Raschmanová, H.; Weninger, A.; Glieder, A.; Kovar, K.; Vogl, T. Implementing CRISPR-Cas technologies in conventional and non-conventional yeasts: Current state and future prospects. Biotechnol. Adv. 2018, 36, 641–665. [Google Scholar] [CrossRef] [PubMed]
- Parapouli, M.; Vasileiadis, A.; Afendra, A.-S.; Hatziloukas, E. Saccharomyces cerevisiae and its industrial applications. AIMS Microbiol. 2020, 6, 1. [Google Scholar] [CrossRef]
- Rainha, J.; Gomes, D.; Rodrigues, L.R.; Rodrigues, J.L. Synthetic biology approaches to engineer Saccharomyces cerevisiae towards the industrial production of valuable polyphenolic compounds. Life 2020, 10, 56. [Google Scholar] [CrossRef]
- Rainha, J.; Rodrigues, J.L.; Rodrigues, L.R. CRISPR-Cas9: A powerful tool to efficiently engineer Saccharomyces cerevisiae. Life 2020, 11, 13. [Google Scholar] [CrossRef]
- Kręgiel, D.; Pawlikowska, E.; Antolak, H. Nonconventional yeasts in fermentation processes: Potentialities and limitations. In Old Yeasts: New Questions; IntechOpen: London, UK, 2017; p. 87. [Google Scholar]
- Bertels, L.-K.; Fernández Murillo, L.; Heinisch, J.J. The pentose phosphate pathway in yeasts-More than a poor cousin of glycolysis. Biomolecules 2021, 11, 725. [Google Scholar] [CrossRef]
- Patra, P.; Das, M.; Kundu, P.; Ghosh, A. Recent advances in systems and synthetic biology approaches for developing novel cell-factories in non-conventional yeasts. Biotechnol. Adv. 2021, 47, 107695. [Google Scholar] [CrossRef]
- Jensen, M.K.; Keasling, J.D. Recent applications of synthetic biology tools for yeast metabolic engineering. FEMS Yeast Res. 2015, 15, 1–10. [Google Scholar] [CrossRef]
- Barrangou, R.; Fremaux, C.; Deveau, H.; Richards, M.; Boyaval, P.; Moineau, S.; Romero, D.A.; Horvath, P. CRISPR provides acquired resistance against viruses in prokaryotes. Science 2007, 315, 1709–1712. [Google Scholar] [CrossRef] [PubMed]
- Walsh, R.M.; Hochedlinger, K. A variant CRISPR-Cas9 system adds versatility to genome engineering. Proc. Natl. Acad. Sci. USA 2013, 110, 15514–15515. [Google Scholar] [CrossRef] [PubMed]
- Levi, O.; Arava, Y. Expanding the CRISPR/Cas9 toolbox for gene engineering in S. cerevisiae. Curr. Microbiol. 2020, 77, 468–478. [Google Scholar] [CrossRef]
- Mitsui, R.; Yamada, R.; Ogino, H. CRISPR system in the yeast Saccharomyces cerevisiae and its application in the bioproduction of useful chemicals. World J. Microbiol. Biotechnol. 2019, 35, 111. [Google Scholar] [CrossRef] [PubMed]
- Geijer, C.; Ledesma-Amaro, R.; Tomás-Pejó, E. Unraveling the potential of non-conventional yeasts in biotechnology. FEMS Yeast Res. 2022, 22, foab071. [Google Scholar] [CrossRef] [PubMed]
- Cai, P.; Gao, J.; Zhou, Y. CRISPR-mediated genome editing in non-conventional yeasts for biotechnological applications. Microb. Cell Fact. 2019, 18, 63. [Google Scholar] [CrossRef] [PubMed]
- Rebello, S.; Abraham, A.; Madhavan, A.; Sindhu, R.; Binod, P.; Karthika Bahuleyan, A.; Aneesh, E.M.; Pandey, A. Non-conventional yeast cell factories for sustainable bioprocesses. FEMS Microbiol. Lett. 2018, 365, fny222. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, V.; Radecka, D.; Aerts, G.; Verstrepen, K.J.; Lievens, B.; Thevelein, J.M. Phenotypic landscape of non-conventional yeast species for different stress tolerance traits desirable in bioethanol fermentation. Biotechnol. Biofuels 2017, 10, 216. [Google Scholar] [CrossRef]
- Chen, M.-T.; Lin, S.; Shandil, I.; Andrews, D.; Stadheim, T.A.; Choi, B.-K. Generation of diploid Pichia pastoris strains by mating and their application for recombinant protein production. Microb. Cell Fact. 2012, 11, 91. [Google Scholar] [CrossRef]
- Yarimizu, T.; Nonklang, S.; Nakamura, J.; Tokuda, S.; Nakagawa, T.; Lorreungsil, S.; Sutthikhumpha, S.; Pukahuta, C.; Kitagawa, T.; Nakamura, M. Identification of auxotrophic mutants of the yeast Kluyveromyces marxianus by non-homologous end joining-mediated integrative transformation with genes from Saccharomyces cerevisiae. Yeast 2013, 30, 485–500. [Google Scholar] [CrossRef]
- Kao, Y.-T.; Gonzalez, K.L.; Bartel, B. Peroxisome function, biogenesis, and dynamics in plants. Plant Physiol. 2018, 176, 162–177. [Google Scholar] [CrossRef] [PubMed]
- Ahmad, M.; Hirz, M.; Pichler, H.; Schwab, H. Protein expression in Pichia pastoris: Recent achievements and perspectives for heterologous protein production. Appl. Microbiol. Biotechnol. 2014, 98, 5301–5317. [Google Scholar] [CrossRef]
- Kurtzman, C.P. Biotechnological strains of Komagataella (Pichia) pastoris are Komagataella phaffii as determined from multigene sequence analysis. J. Ind. Microbiol. Biotechnol. 2009, 36, 1435–1438. [Google Scholar] [CrossRef] [PubMed]
- Baghban, R.; Farajnia, S.; Ghasemi, Y.; Mortazavi, M.; Zarghami, N.; Samadi, N. New developments in Pichia pastoris expression system, review and update. Curr. Pharm. Biotechnol. 2018, 19, 451–467. [Google Scholar] [CrossRef] [PubMed]
- Murata, Y.; Nwuche, C.O.; Nweze, J.E.; Ndubuisi, I.A.; Ogbonna, J.C. Potentials of multi-stress tolerant yeasts, Saccharomyces cerevisiae and Pichia kudriavzevii for fuel ethanol production from industrial cassava wastes. Process Biochem. 2021, 111, 305–314. [Google Scholar] [CrossRef]
- Greppi, A.; Saubade, F.; Botta, C.; Humblot, C.; Guyot, J.P.; Cocolin, L. Potential probiotic Pichia kudriavzevii strains and their ability to enhance folate content of traditional cereal-based African fermented food. Food Microbiol. 2017, 62, 169–177. [Google Scholar] [CrossRef]
- Choi, D.H.; Park, E.H.; Kim, M.D. Isolation of thermotolerant yeast Pichia kudriavzevii from nuruk. Food Sci. Biotechnol. 2017, 26, 1357–1362. [Google Scholar] [CrossRef]
- Johansen, P.G.; Owusu-Kwarteng, J.; Parkouda, C.; Padonou, S.W.; Jespersen, L. Occurrence and importance of yeasts in indigenous fermented food and beverages produced in sub-Saharan Africa. Front. Microbiol. 2019, 10, 1789. [Google Scholar] [CrossRef]
- Sun, W.; Vila-Santa, A.; Liu, N.; Prozorov, T.; Xie, D.; Faria, N.T.; Ferreira, F.C.; Mira, N.P.; Shao, Z. Metabolic engineering of an acid-tolerant yeast strain Pichia kudriavzevii for itaconic acid production. Metab. Eng. Commun. 2020, 10, e00124. [Google Scholar] [CrossRef]
- Li, C.; Liu, Q.; Wang, Y.; Yang, X.; Chen, S.; Zhao, Y.; Wu, Y.; Li, L. Salt stress improves thermotolerance and high-temperature bioethanol production of multi-stress-tolerant Pichia kudriavzevii by stimulating intracellular metabolism and inhibiting oxidative damage. Biotechnol. Biofuels 2021, 14, 222. [Google Scholar] [CrossRef]
- Isono, N.; Hayakawa, H.; Usami, A.; Mishima, T.; Hisamatsu, M. A comparative study of ethanol production by Issatchenkia orientalis strains under stress conditions. J. Biosci. Bioeng. 2012, 113, 76–78. [Google Scholar] [CrossRef] [PubMed]
- Chan, G.F.; Gan, H.M.; Ling, H.L.; Rashid, N.A.A. Genome sequence of Pichia kudriavzevii M12, a potential producer of bioethanol and phytase. Eukaryot. Cell 2012, 11, 1300–1301. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Zhan, T.; Xu, H.; Chen, J.; Bi, C.; Fan, F.; Zhang, X. Characterization of JEN family carboxylate transporters from the acid-tolerant yeast Pichia kudriavzevii and their applications in succinic acid production. Microb. Biotechnol. 2021, 14, 1130–1147. [Google Scholar] [CrossRef] [PubMed]
- Blazeck, J.; Hill, A.; Liu, L.; Knight, R.; Miller, J.; Pan, A.; Otoupal, P.; Alper, H.S. Harnessing Yarrowia lipolytica lipogenesis to create a platform for lipid and biofuel production. Nat. Commun. 2014, 5, 3131. [Google Scholar] [CrossRef]
- Liu, H.H.; Ji, X.J.; Huang, H. Biotechnological applications of Yarrowia lipolytica: Past, present and future. Biotechnol. Adv. 2015, 33, 1522–1546. [Google Scholar] [CrossRef]
- Cavallo, E.; Charreau, H.; Cerrutti, P.; Foresti, M.L. Yarrowia lipolytica: A model yeast for citric acid production. FEMS Yeast Res. 2017, 17, fox084. [Google Scholar] [CrossRef]
- Wang, N.; Chi, P.; Zou, Y.; Xu, Y.; Xu, S.; Bilal, M.; Fickers, P.; Cheng, H. Metabolic engineering of Yarrowia lipolytica for thermoresistance and enhanced erythritol productivity. Biotechnol. Biofuels 2020, 13, 176. [Google Scholar] [CrossRef]
- Muhammad, A.; Feng, X.; Rasool, A.; Sun, W.; Li, C. Production of plant natural products through engineered Yarrowia lipolytica. Biotechnol. Adv. 2020, 43, 107555. [Google Scholar] [CrossRef]
- Kim, O.C.; Suwannarangsee, S.; Oh, D.B.; Kim, S.; Seo, J.W.; Kim, C.H.; Kang, H.A.; Kim, J.Y.; Kwon, O. Transcriptome analysis of xylose metabolism in the thermotolerant methylotrophic yeast Hansenula polymorpha. Bioprocess Biosyst. Eng. 2013, 36, 1509–1518. [Google Scholar] [CrossRef]
- Choudhary, J.; Singh, S.; Nain, L. Thermotolerant fermenting yeasts for simultaneous saccharification fermentation of lignocellulosic biomass. Electron. J. Biotechnol. 2016, 21, 82–92. [Google Scholar] [CrossRef]
- Fonseca, G.G.; Gombert, A.K.; Heinzle, E.; Wittmann, C. Physiology of the yeast Kluyveromyces marxianus during batch and chemostat cultures with glucose as the sole carbon source. FEMS Yeast Res. 2007, 7, 422–435. [Google Scholar] [CrossRef] [PubMed]
- Banat, I.M.; Nigam, P.; Marchant, R. Isolation of thermotolerant, fermentative yeasts growing at 52 °C and producing ethanol at 45 °C and 50 °C. World J. Microbiol. Biotechnol. 1992, 8, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Álvarez-Cao, M.E.; Rico-Díaz, A.; Cerdán, M.E.; Becerra, M.; González-Siso, M.I. Valuation of agro-industrial wastes as substrates for heterologous production of α-galactosidase. Microb. Cell Fact. 2018, 17, 137. [Google Scholar] [CrossRef] [PubMed]
- Nurcholis, M.; Lertwattanasakul, N.; Rodrussamee, N.; Kosaka, T.; Murata, M.; Yamada, M. Integration of comprehensive data and biotechnological tools for industrial applications of Kluyveromyces marxianus. Appl. Microbiol. Biotechnol. 2020, 104, 475–488. [Google Scholar] [CrossRef]
- Bilal, M.; Ji, L.; Xu, Y.; Xu, S.; Lin, Y.; Iqbal, H.M.N.; Cheng, H. Bioprospecting Kluyveromyces marxianus as a robust host for industrial biotechnology. Front. Bioeng. Biotechnol. 2022, 10, 851768. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Merino, R.A.; Varela, J.A.; Coughlan, A.Y.; Hoshida, H.; Da Silveira, W.B.; Wilde, C.; Kuijpers, N.G.A.; Geertman, J.M.; Wolfe, K.H.; Morrissey, J.P. Ploidy variation in Kluyveromyces marxianus separates dairy and non-dairy isolates. Front. Genet. 2018, 9, 94. [Google Scholar] [CrossRef] [PubMed]
- Rajkumar, A.S.; Morrissey, J.P. Rational engineering of Kluyveromyces marxianus to create a chassis for the production of aromatic products. Microb. Cell Fact. 2020, 19, 207. [Google Scholar] [CrossRef]
- Cernak, P.; Estrela, R.; Poddar, S.; Skerker, J.M.; Cheng, Y.F.; Carlson, A.K.; Chen, B.; Glynn, V.M.; Furlan, M.; Ryan, O.W.; et al. Engineering Kluyveromyces marxianus as a robust synthetic biology platform host. mBio 2018, 9, e01410–e01418. [Google Scholar] [CrossRef]
- Mao, W.; Han, Y.; Wang, X.; Zhao, X.; Chi, Z.; Chi, Z.; Liu, G. A new engineered endo-inulinase with improved activity and thermostability: Application in the production of prebiotic fructo-oligosaccharides from inulin. Food Chem. 2019, 294, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, C.; Wheeldon, I. Genome and metabolic engineering in non-conventional yeasts: Current advances and applications. Synth. Syst. Biotechnol. 2017, 2, 198–207. [Google Scholar]
- Da Silva, N.A.; Srikrishnan, S. Introduction and expression of genes for metabolic engineering applications in Saccharomyces cerevisiae. FEMS Yeast Res. 2012, 12, 197–214. [Google Scholar] [CrossRef]
- Lee, M.E.; DeLoache, W.C.; Cervantes, B.; Dueber, J.E. A highly characterized yeast toolkit for modular, multipart assembly. ACS Synth. Biol. 2015, 4, 975–986. [Google Scholar] [CrossRef]
- Liu, L.; Otoupal, P.; Pan, A.; Alper, H.S. Increasing expression level and copy number of a Yarrowia lipolytica plasmid through regulated centromere function. FEMS Yeast Res. 2014, 14, 1124–1127. [Google Scholar]
- Camattari, A.; Goh, A.; Yip, L.Y.; Tan, A.H.M.; Ng, S.W.; Tran, A.; Liu, G.; Liachko, I.; Dunham, M.J.; Rancati, G. Characterization of a panARS-based episomal vector in the methylotrophic yeast Pichia pastoris for recombinant protein production and synthetic biology applications. Microb. Cell Fact. 2016, 15, 139. [Google Scholar] [CrossRef]
- Nakamura, Y.; Nishi, T.; Noguchi, R.; Ito, Y.; Watanabe, T.; Nishiyama, T.; Aikawa, S.; Hasunuma, T.; Ishii, J.; Okubo, Y. A stable, autonomously replicating plasmid vector containing Pichia pastoris centromeric DNA. Appl. Environ. Microbiol. 2018, 84, e02882. [Google Scholar] [CrossRef]
- Wang, L.; Lin, J.; Zhang, T.; Xu, K.; Ren, C.; Zhang, Z. Simultaneous screening and validation of effective zinc finger nucleases in yeast. PLoS ONE 2013, 8, e64687. [Google Scholar] [CrossRef]
- Gan, Y.; Lin, Y.; Guo, Y.; Qi, X.; Wang, Q. Metabolic and genomic characterisation of stress-tolerant industrial Saccharomyces cerevisiae strains from TALENs-assisted multiplex editing. FEMS Yeast Res. 2018, 18, foy045. [Google Scholar] [CrossRef]
- Song, A.J.; Palmiter, R.D. Detecting and avoiding problems when using the Cre–lox system. Trends Genet. 2018, 34, 333–340. [Google Scholar] [CrossRef]
- Mali, P.; Esvelt, K.M.; Church, G.M. Cas9 as a versatile tool for engineering biology. Nat. Methods 2013, 10, 957–963. [Google Scholar] [CrossRef]
- Mojica, F.J.M.; Montoliu, L. On the origin of CRISPR-Cas technology: From prokaryotes to mammals. Trends Microbiol. 2016, 24, 811–820. [Google Scholar] [CrossRef]
- Makarova, K.S.; Wolf, Y.I.; Alkhnbashi, O.S.; Costa, F.; Shah, S.A.; Saunders, S.J.; Barrangou, R.; Brouns, S.J.; Charpentier, E.; Haft, D.H.; et al. An updated evolutionary classification of CRISPR–Cas systems. Nat. Rev. Microbiol. 2015, 13, 722–736. [Google Scholar] [CrossRef]
- Liu, Z.; Dong, H.; Cui, Y.; Cong, L.; Zhang, D. Application of different types of CRISPR/Cas-based systems in bacteria. Microb. Cell Fact. 2020, 19, 172. [Google Scholar] [CrossRef]
- Anders, C.; Niewoehner, O.; Duerst, A.; Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 2014, 513, 569–573. [Google Scholar] [CrossRef]
- Asmamaw, M.; Zawdie, B. Mechanism and applications of CRISPR/Cas-9-mediated genome editing. Biologics 2021, 15, 353–361. [Google Scholar]
- Palermo, G.; Chen, J.S.; Ricci, C.G.; Rivalta, I.; Jinek, M.; Batista, V.S.; Doudna, J.A.; McCammon, J.A. Key role of the REC lobe during CRISPR–Cas9 activation by ‘sensing’,‘regulating’, and ‘locking’ the catalytic HNH domain. Q. Rev. Biophys. 2018, 51, e91. [Google Scholar] [CrossRef]
- Ming, S.H.; Tian-Rui, X.U.; Ce-Shi, C.H. The big bang of genome editing technology: Development and application of the CRISPR/Cas9 system in disease animal models. Zool. Res. 2016, 37, 191–204. [Google Scholar]
- Daley, J.M.; Palmbos, P.L.; Wu, D.; Wilson, T.E. Nonhomologous end joining in yeast. Annu. Rev. Genet. 2005, 39, 431–451. [Google Scholar] [CrossRef]
- Brandsma, I.; Gent, D.C. Pathway choice in DNA double strand break repair: Observations of a balancing act. Genome Integr. 2012, 3, 9. [Google Scholar] [CrossRef]
- Meng, J.; Qiu, Y.; Shi, S. CRISPR/Cas9 systems for the development of Saccharomyces cerevisiae cell factories. Front. Bioeng. Biotechnol. 2020, 8, 594347. [Google Scholar] [CrossRef]
- Schwartz, C.; Shabbir-Hussain, M.; Frogue, K.; Blenner, M.; Wheeldon, I. Standardized markerless gene integration for pathway engineering in Yarrowia lipolytica. ACS Synth. Biol. 2017, 6, 402–409. [Google Scholar] [CrossRef]
- de Souza, C.P.; Ribeiro, B.D.; Zarur Coelho, M.A.; Almeida, R.V.; Nicaud, J.M. Construction of wild-type Yarrowia lipolytica IMUFRJ 50682 auxotrophic mutants using dual CRISPR/Cas9 strategy for novel biotechnological approaches. Enzyme Microb. Technol. 2020, 140, 109621. [Google Scholar] [CrossRef]
- Zhang, X.K.; Wang, D.N.; Chen, J.; Liu, Z.J.; Wei, L.J.; Hua, Q. Metabolic engineering of β-carotene biosynthesis in Yarrowia lipolytica. Biotechnol. Lett. 2020, 42, 945–956. [Google Scholar] [CrossRef]
- Gao, S.; Tong, Y.; Zhu, L.; Ge, M.; Jiang, Y.; Chen, D.; Yang, S. Production of β-carotene by expressing a heterologous multifunctional carotene synthase in Yarrowia lipolytica. Biotechnol. Lett. 2017, 39, 921–927. [Google Scholar] [CrossRef]
- Wang, L.; Deng, A.; Zhang, Y.; Liu, S.; Liang, Y.; Bai, H.; Cui, D.; Qiu, Q.; Shang, X.; Yang, Z.; et al. Efficient CRISPR–Cas9 mediated multiplex genome editing in yeasts. Biotechnol. Biofuels 2018, 11, 277. [Google Scholar] [CrossRef]
- Juergens, H.; Varela, J.A.; Gorter de Vries, A.R.; Perli, T.; Gast, V.J.; Gyurchev, N.Y.; Rajkumar, A.S.; Mans, R.; Pronk, J.T.; Morrissey, J.P. Genome editing in Kluyveromyces and Ogataea yeasts using a broad-host-range Cas9/gRNA co-expression plasmid. FEMS Yeast Res. 2018, 18, foy012. [Google Scholar] [CrossRef]
- Siripong, W.; Angela, C.; Tanapongpipat, S.; Runguphan, W. Metabolic engineering of Pichia pastoris for production of isopentanol (3-methyl-1-butanol). Enzyme Microb. Technol. 2020, 138, 109557. [Google Scholar] [CrossRef]
- Liu, Q.; Shi, X.; Song, L.; Liu, H.; Zhou, X.; Wang, Q.; Zhang, Y.; Cai, M. CRISPR–Cas9-mediated genomic multiloci integration in Pichia pastoris. Microb. Cell Fact. 2019, 18, 144. [Google Scholar] [CrossRef]
- Gassler, T.; Heistinger, L.; Mattanovich, D.; Gasser, B.; Prielhofer, R. CRISPR/Cas9-mediated homology-directed genome editing in Pichia pastoris. In Recombinant Protein Production in Yeast; Springer: Berlin, Germany, 2019; pp. 211–225. [Google Scholar]
- Löbs, A.-K.; Engel, R.; Schwartz, C.; Flores, A.; Wheeldon, I. CRISPR–Cas9-enabled genetic disruptions for understanding ethanol and ethyl acetate biosynthesis in Kluyveromyces marxianus. Biotechnol. Biofuels 2017, 10, 164. [Google Scholar] [CrossRef]
- Tran, V.G.; Cao, M.; Fatma, Z.; Song, X.; Zhao, H. Development of a CRISPR/Cas9-based tool for gene deletion in Issatchenkia orientalis. mSphere 2019, 4, e00345-19. [Google Scholar] [CrossRef]
- Lee, Y.G.; Kim, C.; Kuanyshev, N.; Kang, N.K.; Fatma, Z.; Wu, Z.Y.; Cheng, M.H.; Singh, V.; Yoshikuni, Y.; Zhao, H.; et al. Cas9-based metabolic engineering of Issatchenkia orientalis for enhanced utilization of cellulosic hydrolysates. J. Agric. Food Chem. 2022, 70, 12085–12094. [Google Scholar] [CrossRef]
- Emerson, C.H.; Bertuch, A.A. Consider the workhorse: Nonhomologous end-joining in budding yeast. Biochem. Cell Biol. 2016, 94, 396–406. [Google Scholar] [CrossRef]
- Yang, H.; Matsumoto, Y.; Trujillo, K.M.; Lees-Miller, S.P.; Osley, M.A.; Tomkinson, A.E. Role of the yeast DNA repair protein Nej1 in end processing during the repair of DNA double strand breaks by non-homologous end joining. DNA Repair 2015, 31, 1–10. [Google Scholar] [CrossRef]
- Zhang, Y.; Hefferin, M.L.; Chen, L.; Shim, E.Y.; Tseng, H.M.; Kwon, Y.; Sung, P.; Lee, S.E.; Tomkinson, A.E. Role of Dnl4–Lif1 in nonhomologous end-joining repair complex assembly and suppression of homologous recombination. Nat. Struct. Mol. Biol. 2007, 14, 639–646. [Google Scholar] [CrossRef]
- Utomo, J.C.; Hodgins, C.L.; Ro, D.K. Multiplex Genome Editing in Yeast by CRISPR/Cas9–A Potent and Agile Tool to Reconstruct Complex Metabolic Pathways. Front. Plant Sci. 2021, 12, 719148. [Google Scholar] [CrossRef]
- Yan, Y.; Finnigan, G.C. Development of a multi-locus CRISPR gene drive system in budding yeast. Sci. Rep. 2018, 8, 17277. [Google Scholar] [CrossRef]
- Rajkumar, A.S.; Varela, J.A.; Juergens, H.; Daran, J.M.G.; Morrissey, J.P. Biological parts for Kluyveromyces marxianus synthetic biology. Front. Bioeng. Biotechnol. 2019, 7, 97. [Google Scholar] [CrossRef] [PubMed]
- Näätsaari, L.; Mistlberger, B.; Ruth, C.; Hajek, T.; Hartner, F.S.; Glieder, A. Deletion of the Pichia pastoris KU70 homologue facilitates platform strain generation for gene expression and synthetic biology. PLoS ONE 2012, 7, e39720. [Google Scholar] [CrossRef]
- Zhang, K.; Duan, X.; Cai, P.; Gao, L.; Wu, X.; Yao, L.; Zhou, Y.J. Fusing an exonuclease with Cas9 enhances homologous recombination in Pichia pastoris. Microb. Cell Fact. 2022, 21, 182. [Google Scholar] [CrossRef]
- Paffett, K.S.; Clikeman, J.A.; Palmer, S.; Nickoloff, J.A. Overexpression of Rad51 inhibits double-strand break-induced homologous recombination but does not affect gene conversion tract lengths. DNA Repair 2005, 4, 687–698. [Google Scholar] [CrossRef]
- Cai, P.; Duan, X.; Wu, X.; Gao, L.; Ye, M.; Zhou, Y.J. Recombination machinery engineering facilitates metabolic engineering of the industrial yeast Pichia pastoris. Nucleic Acids Res. 2021, 49, 7791–7805. [Google Scholar] [CrossRef]
- Ji, Q.; Mai, J.; Ding, Y.; Wei, Y.; Ledesma-Amaro, R.; Ji, X.-J. Improving the homologous recombination efficiency of Yarrowia lipolytica by grafting heterologous component from Saccharomyces cerevisiae. Metab. Eng. Commun. 2020, 11, e00152. [Google Scholar] [CrossRef]
- Weninger, A.; Hatzl, A.M.; Schmid, C.; Vogl, T.; Glieder, A. Combinatorial optimization of CRISPR/Cas9 expression enables precision genome engineering in the methylotrophic yeast Pichia pastoris. J. Biotechnol. 2016, 235, 139–149. [Google Scholar] [CrossRef]
- Gao, Y.; Zhao, Y. Self-processing of ribozyme-flanked RNAs into guide RNAs in vitro and in vivo for CRISPR-mediated genome editing. J. Integr. Plant Biol. 2014, 56, 343–349. [Google Scholar] [CrossRef] [PubMed]
- Ryan, O.W.; Skerker, J.M.; Maurer, M.J.; Li, X.; Tsai, J.C.; Poddar, S.; Lee, M.E.; DeLoache, W.; Dueber, J.E.; Arkin, A.P.; et al. Selection of chromosomal DNA libraries using a multiplex CRISPR system. eLife 2014, 3, e03703. [Google Scholar] [CrossRef] [PubMed]
- Naito, Y.; Hino, K.; Bono, H.; Ui-Tei, K. CRISPRdirect: Software for designing CRISPR/Cas guide RNA with reduced off-target sites. Bioinformatics 2015, 31, 1120–1123. [Google Scholar] [CrossRef] [PubMed]
- Kleinstiver, B.P.; Prew, M.S.; Tsai, S.Q.; Nguyen, N.T.; Topkar, V.V.; Zheng, Z.; Joung, J.K. Broadening the targeting range of Staphylococcus aureus CRISPR-Cas9 by modifying PAM recognition. Nat. Biotechnol. 2015, 33, 1293–1298. [Google Scholar] [CrossRef]
- Rees, H.A.; Yeh, W.-H.; Liu, D.R. Development of hRad51–Cas9 nickase fusions that mediate HDR without double-stranded breaks. Nat. Commun. 2019, 10, 2212. [Google Scholar] [CrossRef]
- Schusterbauer, V.; Fischer, J.E.; Gangl, S.; Schenzle, L.; Rinnofner, C.; Geier, M.; Sailer, C.; Glieder, A.; Thallinger, G.G. Whole Genome Sequencing Analysis of Effects of CRISPR/Cas9 in Komagataella phaffii: A Budding Yeast in Distress. J. Fungi 2022, 8, 992. [Google Scholar] [CrossRef]
- Butelli, E.; Titta, L.; Giorgio, M.; Mock, H.P.; Matros, A.; Peterek, S.; Schijlen, E.G.; Hall, R.D.; Bovy, A.G.; Luo, J.; et al. Enrichment of tomato fruit with health-promoting anthocyanins by expression of select transcription factors. Nat. Biotechnol. 2008, 26, 1301–1308. [Google Scholar] [CrossRef]
- Kawser Hossain, M.; Abdal Dayem, A.; Han, J.; Yin, Y.; Kim, K.; Kumar Saha, S.; Yang, G.M.; Choi, H.Y.; Cho, S.G. Molecular mechanisms of the anti-obesity and anti-diabetic properties of flavonoids. Int. J. Mol. Sci. 2016, 17, 569. [Google Scholar] [CrossRef]
- Sáez-Sáez, J.; Wang, G.; Marella, E.R.; Sudarsan, S.; Cernuda Pastor, M.; Borodina, I. Engineering the oleaginous yeast Yarrowia lipolytica for high-level resveratrol production. Metab. Eng. 2020, 62, 51–61. [Google Scholar] [CrossRef] [PubMed]
- Pyne, M.E.; Narcross, L.; Martin, V.J.J. Engineering plant secondary metabolism in microbial systems. Plant Physiol. 2019, 179, 844–861. [Google Scholar] [CrossRef] [PubMed]
- Zha, J.; Wu, X.; Gong, G.; Koffas, M.A.G. Pathway enzyme engineering for flavonoid production in recombinant microbes. Metab. Eng. Commun. 2019, 9, e00104. [Google Scholar] [CrossRef] [PubMed]
- Sheng, H.; Sun, X.; Yan, Y.; Yuan, Q.; Wang, J.; Shen, X. Metabolic engineering of microorganisms for the production of flavonoids. Front. Bioeng. Biotechnol. 2020, 8, 589069. [Google Scholar] [CrossRef]
- Soreanu, I.; Hendler, A.; Dahan, D.; Dovrat, D.; Aharoni, A. Marker-free genetic manipulations in yeast using CRISPR/CAS9 system. Curr. Genet. 2018, 64, 1129–1139. [Google Scholar] [CrossRef]
- Lv, Y.; Marsafari, M.; Koffas, M.; Zhou, J.; Xu, P. Optimizing oleaginous yeast cell factories for flavonoids and hydroxylated flavonoids biosynthesis. ACS Synth. Biol. 2019, 8, 2514–2523. [Google Scholar] [CrossRef] [PubMed]
- Wei, W.; Zhang, P.; Shang, Y.; Zhou, Y.; Ye, B.C. Metabolically engineering of Yarrowia lipolytica for the biosynthesis of naringenin from a mixture of glucose and xylose. Bioresour. Technol. 2020, 314, 123726. [Google Scholar] [CrossRef]
- Toti, E.; Chen, C.Y.O.; Palmery, M.; Villaño Valencia, D.; Peluso, I. Non-provitamin A and provitamin A carotenoids as immunomodulators: Recommended dietary allowance, therapeutic index, or personalized nutrition? Oxid. Med. Cell. Longev. 2018, 2018. [Google Scholar] [CrossRef] [PubMed]
- Arnesen, J.A.; Kildegaard, K.R.; Cernuda Pastor, M.; Jayachandran, S.; Kristensen, M.; Borodina, I. Yarrowia lipolytica strains engineered for the production of terpenoids. Front. Bioeng. Biotechnol. 2020, 8, 945. [Google Scholar] [CrossRef]
- Lu, Y.; Yang, Q.; Lin, Z.; Yang, X. A modular pathway engineering strategy for the high-level production of β-ionone in Yarrowia lipolytica. Microb. Cell Fact. 2020, 19, 49. [Google Scholar] [CrossRef]
- Yao, F.; Liu, S.C.; Wang, D.N.; Liu, Z.J.; Hua, Q.; Wei, L.J. Engineering oleaginous yeast Yarrowia lipolytica for enhanced limonene production from xylose and lignocellulosic hydrolysate. FEMS Yeast Res. 2020, 20, foaa046. [Google Scholar] [CrossRef]
- Zhang, J.L.; Bai, Q.Y.; Peng, Y.Z.; Fan, J.; Jin, C.C.; Cao, Y.X.; Yuan, Y.J. High production of triterpenoids in Yarrowia lipolytica through manipulation of lipid components. Biotechnol. Biofuels 2020, 13, 133. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Lu, X.; Zong, H.; Li, J.; Zhuge, B. Itaconic acid production in microorganisms. Biotechnol. Lett. 2018, 40, 455–464. [Google Scholar] [CrossRef] [PubMed]
- Zhao, C.; Cui, Z.; Zhao, X.; Zhang, J.; Zhang, L.; Tian, Y.; Qi, Q.; Liu, J. Enhanced itaconic acid production in Yarrowia lipolytica via heterologous expression of a mitochondrial transporter MTT. Appl. Microbiol. Biotechnol. 2019, 103, 2181–2192. [Google Scholar] [CrossRef]
- Blazeck, J.; Miller, J.; Pan, A.; Gengler, J.; Holden, C.; Jamoussi, M.; Alper, H.S. Metabolic engineering of Saccharomyces cerevisiae for itaconic acid production. Appl. Microbiol. Biotechnol. 2014, 98, 8155–8164. [Google Scholar] [CrossRef] [PubMed]
- Yook, S.D.; Kim, J.; Gong, G.; Ko, J.K.; Um, Y.; Han, S.O.; Lee, S.M. High-yield lipid production from lignocellulosic biomass using engineered xylose-utilizing Yarrowia lipolytica. Glob. Change Biol. Bioenergy 2020, 12, 670–679. [Google Scholar] [CrossRef]
- Li, J.X.; Xu, J.; Ruan, J.C.; Meng, H.M.; Su, H.; Han, X.F.; Lu, M.; Li, F.L.; Wang, S.A. Disrupting a phospholipase A2 gene increasing lipid accumulation in the oleaginous yeast Yarrowia lipolytica. J. Appl. Microbiol. 2021, 130, 100–108. [Google Scholar] [CrossRef]
- Zhang, J.L.; Cao, Y.X.; Peng, Y.Z.; Jin, C.C.; Bai, Q.Y.; Zhang, R.S.; Liu, D.; Yuan, Y.J. High production of fatty alcohols in Yarrowia lipolytica by coordination with glycolysis. Sci. China Chem. 2019, 62, 1007–1016. [Google Scholar] [CrossRef]
- Reyes-Sánchez, F.J.; Páez-Lerma, J.B.; Rojas-Contreras, J.A.; López-Miranda, J.; Soto-Cruz, N.Ó.; Reinhart-Kirchmayr, M. Study of the enzymatic capacity of Kluyveromyces marxianus for the synthesis of esters. J. Mol. Microbiol. Biotechnol. 2019, 29, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Lang, X.; Moran Cabrera, M.; De Keyser, S.; Sun, X.; Da Silva, N.; Wheeldon, I. CRISPR-mediated multigene integration enables Shikimate pathway refactoring for enhanced 2-phenylethanol biosynthesis in Kluyveromyces marxianus. Biotechnol. Biofuels 2021, 14, 3. [Google Scholar] [CrossRef]
- Löbs, A.K.; Schwartz, C.; Thorwall, S.; Wheeldon, I. Highly multiplexed CRISPRi repression of respiratory functions enhances mitochondrial localized ethyl acetate biosynthesis in Kluyveromyces marxianus. ACS Synth. Biol. 2018, 7, 2647–2655. [Google Scholar] [CrossRef] [PubMed]
- Swietalski, P.; Hetzel, F.; Seitl, I.; Fischer, L. Secretion of a low and high molecular weight β-glycosidase by Yarrowia lipolytica. Microb. Cell Fact. 2020, 19, 100. [Google Scholar] [CrossRef] [PubMed]
- Bruder, S.; Moldenhauer, E.J.; Lemke, R.D.; Ledesma-Amaro, R.; Kabisch, J. Drop-in biofuel production using fatty acid photodecarboxylase from Chlorella variabilis in the oleaginous yeast Yarrowia lipolytica. Biotechnol. Biofuels 2019, 12, 202. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Park, J.; Kim, I.J.; Kim, S.R. Nonconventional Yeasts Engineered Using the CRISPR-Cas System as Emerging Microbial Cell Factories. Fermentation 2022, 8, 656. https://doi.org/10.3390/fermentation8110656
Park J, Kim IJ, Kim SR. Nonconventional Yeasts Engineered Using the CRISPR-Cas System as Emerging Microbial Cell Factories. Fermentation. 2022; 8(11):656. https://doi.org/10.3390/fermentation8110656
Chicago/Turabian StylePark, Jongbeom, In Jung Kim, and Soo Rin Kim. 2022. "Nonconventional Yeasts Engineered Using the CRISPR-Cas System as Emerging Microbial Cell Factories" Fermentation 8, no. 11: 656. https://doi.org/10.3390/fermentation8110656
APA StylePark, J., Kim, I. J., & Kim, S. R. (2022). Nonconventional Yeasts Engineered Using the CRISPR-Cas System as Emerging Microbial Cell Factories. Fermentation, 8(11), 656. https://doi.org/10.3390/fermentation8110656








