Enterococci Isolated from Cypriot Green Table Olives as a New Source of Technological and Probiotic Properties
Abstract
1. Introduction
2. Materials and Methods
2.1. Samples Collection
2.2. Enumeration of Microorganisms
2.3. Isolation of LAB
2.4. Physiological and Molecular Characterization of LAB Strains
2.5. Technological Characteristics
2.5.1. Acidification Activity
2.5.2. Proteolytic Activity
2.5.3. Lipolytic Activity
2.5.4. Exopolysaccharide Production (EPS)
2.5.5. β-Glucosidase Activity
2.5.6. Catabolism of Citric Acid
2.6. Pathogenicity
2.6.1. Hemolytic Activity
2.6.2. DNAse Production
2.6.3. Virulence Activity Using Genotypic Tests
2.7. Screening for Probiotics Characteristics
2.7.1. Resistance to Low pH
2.7.2. Resistance to Bile Salts
2.8. Statistical Analysis
3. Results and Discussion
3.1. Microbial Enumeration
3.2. Isolation and Identification of LAB
3.3. Technological Properties
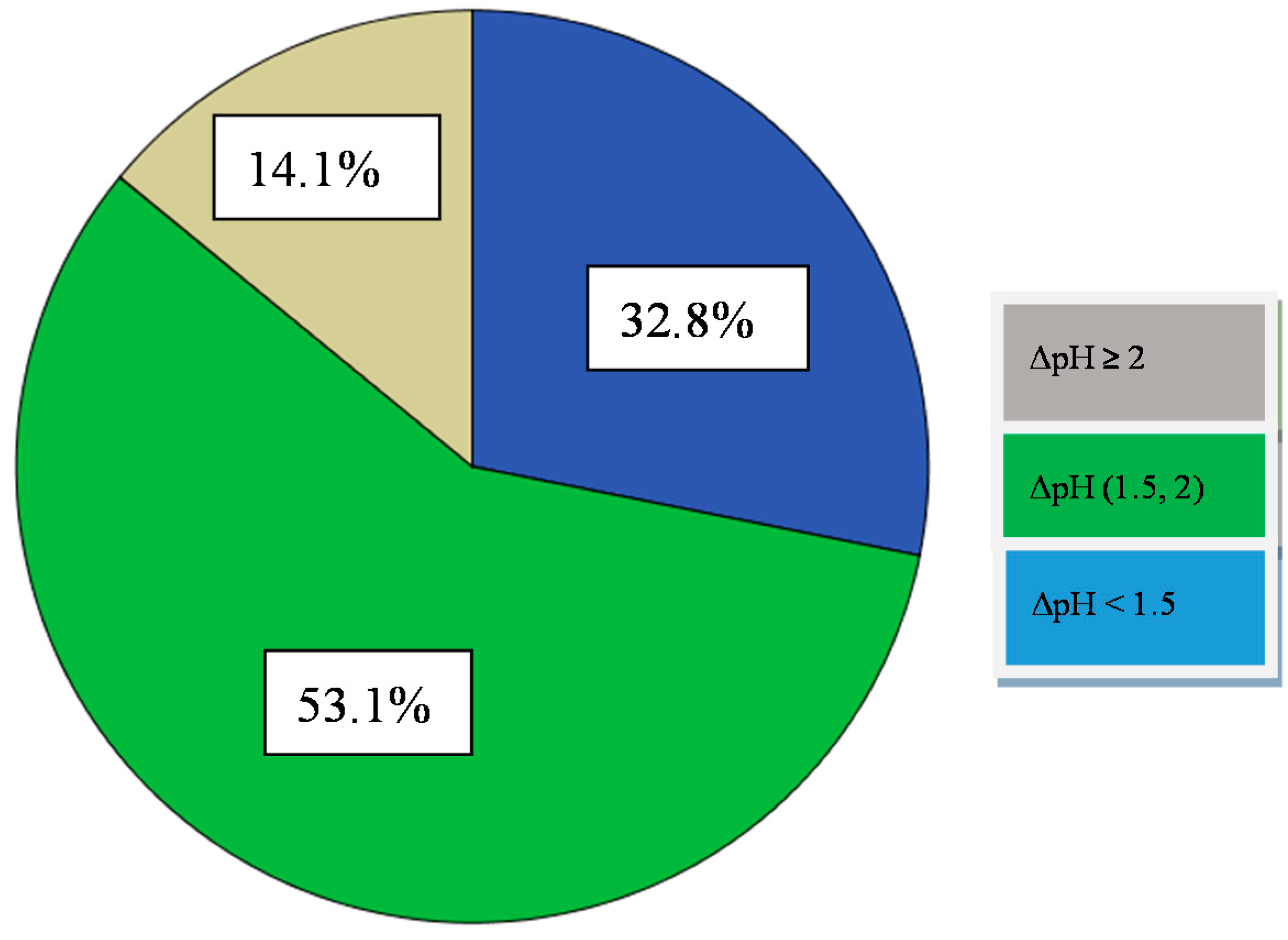
3.3.1. Acidification Activity
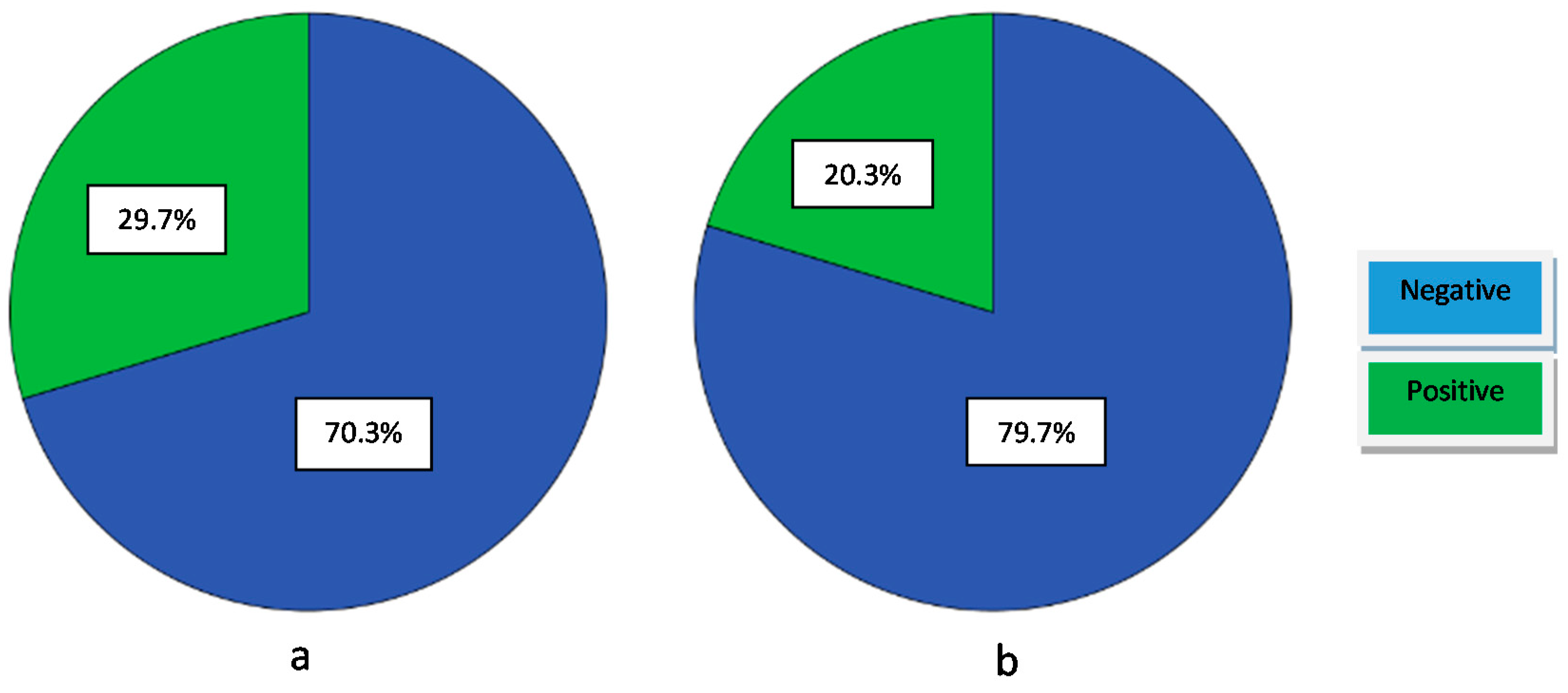
3.3.2. Proteolytic Activity
3.3.3. Lipolytic Activity
3.3.4. Exopolysaccharide Production (EPS)
3.3.5. β-Glucosidase Activity
3.3.6. Catabolism of Citric Acid
3.4. Pathogenicity
3.5. Screening for Probiotic Potential
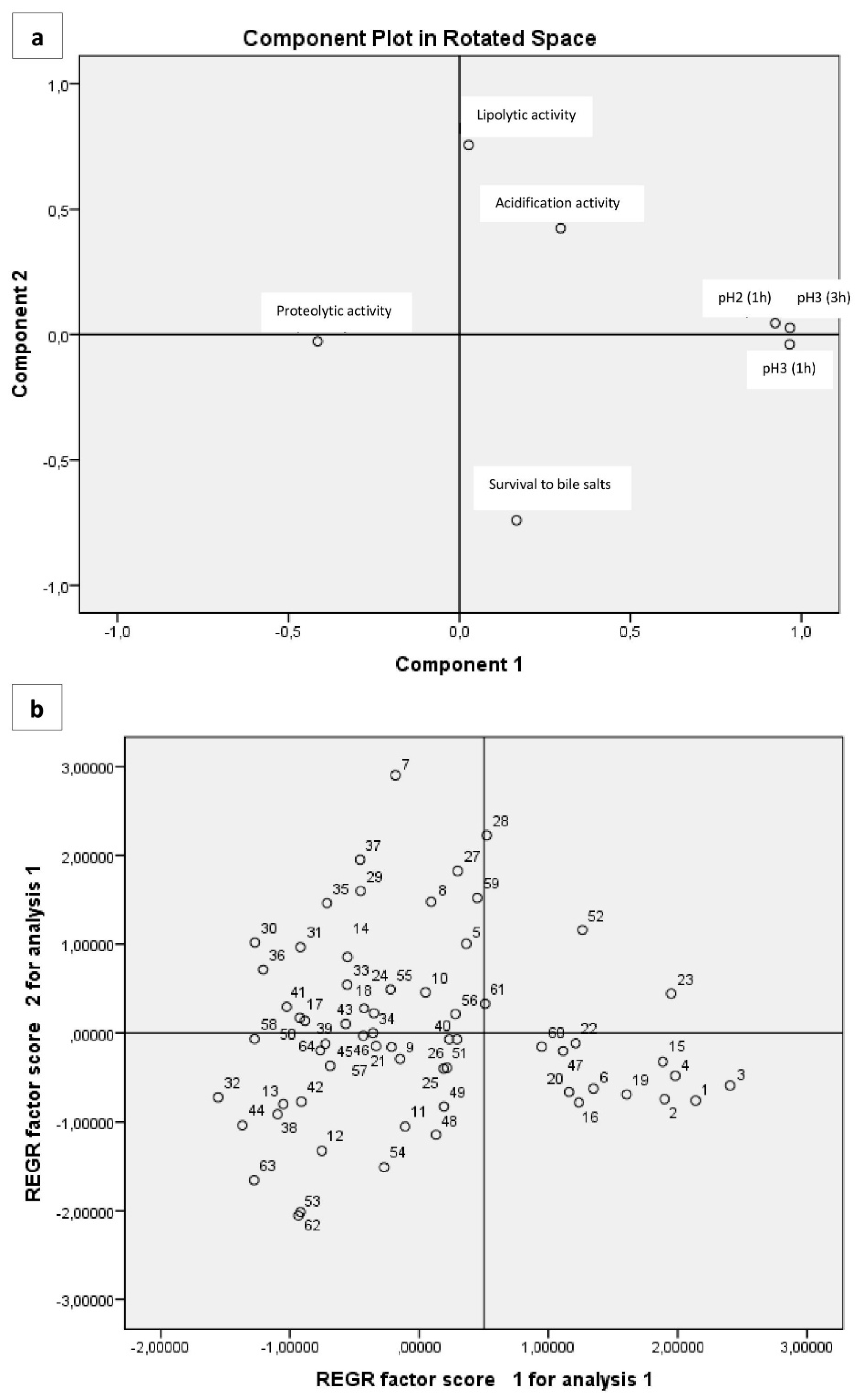
3.6. Multivariate Analysis of Phenotypic Characteristics Related to Probiotic Potential
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Olives, T. No 66—November 2012; International Olive Council: Madrid, Spain, 2012. [Google Scholar]
- López, F.N.A.; Romero, C.; Quintana, M.D.C.D.; López, A.L.; García, P.G.; Fernández, A.G. Kinetic study of the physicochemical and microbiological changes in ‘seasoned’ olives during the shelf-life period. J. Agric. Food Chem. 2005, 53, 5285–5292. [Google Scholar] [CrossRef] [PubMed]
- Bautista-Gallego, J.; Arroyo-López, F.N.; Rantsiou, K.; Jiménez-Díaz, R.; Garrido-Fernández, A.; Cocolin, L. Screening of lactic acid bacteria isolated from fermented table olives with probiotic potential. Food Res. Int. 2013, 50, 135–142. [Google Scholar] [CrossRef]
- Montet, D.; Ray, R.C.; Zakhia-Rozis, N. Lactic Acid Fermentation of Vegetables and Fruits. In Microorganisms and Fermentation of Traditional Foods; CRC Press: Boca Raton, FL, USA, 2014; pp. 108–140. [Google Scholar]
- Stiles, M.E.; Holzapfel, W.H. Lactic acid bacteria of foods and their current taxonomy. Int. J. Food Microbiol. 1997, 36, 1–29. [Google Scholar] [CrossRef]
- Fisher, K.; Phillips, C. The ecology, epidemiology and virulence of Enterococcus. Microbiology 2009, 155, 1749–1757. [Google Scholar] [CrossRef] [PubMed]
- Lebreton, F.; Willems, R.J.L.; Gilmore, M.S. Enterococcus Diversity, Origins in Nature, and Gut Colonization. In Enterococci: From Commensals to Lead. Causes Drug Resistant Infection; Massachusetts Eye and Ear Infirmary: Boston, MA, USA, 2014; pp. 1–56. [Google Scholar]
- Franz, C.M.A.P.; Huch, M.; Abriouel, H.; Holzapfel, W.; Gálvez, A. Enterococci as probiotics and their implications in food safety. Int. J. Food Microbiol. 2011, 151, 125–140. [Google Scholar] [CrossRef] [PubMed]
- Gomes, B.C.; Dora, B.; de Melo, G.; De Martinis, E.C.P.; Paulo, S. Dualistic aspects of Enterococcus spp. in foods. Curr. Res. Technol. Educ. Top. Appl. Microbiol. Microb. Biotechnol. 2010, 1119–1125. [Google Scholar]
- Omar, N.B.; Castro, A.; Lucas, R.; Abriouel, H.; Yousif, N.M.K.; Franz, C.M.A.P.; Holzapfel, W.H.; Ruben, P.P.; Martínez-Canãmero, M.; Gálvez, A. Functional and Safety Aspects of Enterococci Isolated from Different Spanish Foods. Syst. Appl. Microbiol. 2004, 27, 118–130. [Google Scholar] [CrossRef] [PubMed]
- Randazzo, C.L.; Restuccia, C.; Romano, A.D.; Caggia, C. Lactobacillus casei, dominant species in naturally fermented Sicilian green olives. Int. J. Food Microbiol. 2004, 90, 9–14. [Google Scholar] [CrossRef]
- Gonza, J.M.; Lucena-padro, H.; Ruiz-barba, L.; Maldonado-Barraga, A. Enterococcus olivae sp. nov., isolated from Spanish-style green-olive fermentations. Int. J. Syst. Evol. Microbiol. 2014, 64, 2534–2539. [Google Scholar]
- Shinozaki-Kuwahara, N.; Saito, M.; Hirasawa, M.; Hirasawa, M.; Takada, K. Streptococcus dentiloxodontae sp. nov., isolated from the oral cavity of elephants. Int. J. Syst. Evol. Microbiol. 2016, 66, 3878–3883. [Google Scholar] [PubMed]
- Cogan, T.M. Characterization of the lactic acid bacteria in artisanal dairy products. J. Dairy Res. 1997, 64, 409–421. [Google Scholar] [CrossRef]
- Estifanos, H. Isolation and identification of probiotic lactic acid bacteria from curd and in vitro evaluation of its growth inhibition activities against pathogenic bacteria. Afr. J. Microbiol. Res. 2014, 8, 1419–1425. [Google Scholar] [CrossRef]
- Jackson, C.R.; Fedorka-Cray, P.J.; Barrett, J.B. Use of a Genus- and Species-Specific Multiplex PCR for Identification of Enterococci Use of a Genus- and Species-Specific Multiplex PCR for Identification of Enterococci. J. Clin. Microbiol. 2004, 42, 3558. [Google Scholar] [CrossRef] [PubMed]
- Fuka, M.M.; Maksimovic, A.Z.; Tanuwidjaja, I.; Hulak, N.; Schloter, M. Characterization of enterococcal community isolated from an Artisan Istrian raw milk cheese: Biotechnological and safety aspects. Food Technol. Biotechnol. 2017, 55, 368–380. [Google Scholar]
- Franciosi, E.; Settanni, L.; Cavazza, A.; Poznanski, E. Biodiversity and technological potential of wild lactic acid bacteria from raw cows’ milk. Int. Dairy J. 2009, 19, 3–11. [Google Scholar] [CrossRef]
- Dinçer, E.; Kıvanç, M. Lipolytic Activity of Lactic Acid Bacteria Isolated from Turkish Pastırma. Anadolu Univ. J. Sci. Technol. C Life Sci. Biotechnol. 2018, 7, 12–19. [Google Scholar] [CrossRef]
- Imène, K.; Halima, Z.-K.; Nour-Eddine, K. Screening of exopolysaccharide-producing coccal lactic acid bacteria isolated from camel milk and red meat of Algeria. Afr. J. Biotechnol. 2017, 16, 1078–1084. [Google Scholar] [CrossRef]
- Ghabbour, N.; Lamzira, Z.; Thonart, P.; Cidalia, P.; Markaoui, M.; Asehraou, A. Selection of oleuropein-degrading lactic acid bacteria strains isolated from fermenting Moroccan green olives. Grasas Aceites 2011, 62, 84–89. [Google Scholar]
- Ribeiro, S.C.; Coelho, M.C.; Todorov, S.D.; Franco, B.D.G.M.; Dapkevicius, M.L.E.; Silva, C.C.G. Technological properties of bacteriocin-producing lactic acid bacteria isolated from Pico cheese an artisanal cow’s milk cheese. J. Appl. Microbiol. 2014, 116, 573–585. [Google Scholar] [CrossRef] [PubMed]
- Chajęcka-Wierzchowska, W.; Zadernowska, A.; Łaniewska-Trokenheim, Ł. Virulence factors of Enterococcus spp. presented in food. LWT Food Sci. Technol. 2017, 75, 670–676. [Google Scholar] [CrossRef]
- Aspri, M.; Bozoudi, D.; Tsaltas, D.; Hill, C.; Papademas, P. Raw donkey milk as a source of Enterococcus diversity: Assessment of their technological properties and safety characteristics. Food Control 2017, 73, 81–90. [Google Scholar] [CrossRef]
- Vankerckhoven, V.; van Autgaerden, T.; Vael, C.; Lammens, C.; Chapelle, S.; Rossi, R.; Jabes, D.; Goossens, H. Development of a multiplex PCR for the detection of asaI, gelE, cylA, esp, and hyl genes in enterococci and survey for virulence determinants among european hospital isolates of Enterococcus faecium. J. Clin. Microbiol. 2004, 42, 4473–4479. [Google Scholar] [CrossRef] [PubMed]
- Martín-Platero, A.M.; Valdivia, E.; Maqueda, M.; Martínez-Bueno, M. Characterization and safety evaluation of enterococci isolated from Spanish goats’ milk cheeses. Int. J. Food Microbiol. 2009, 132, 24–32. [Google Scholar] [CrossRef] [PubMed]
- De las Rivas, B.; Marcobal, Á.; Muñoz, R. Improved multiplex-PCR method for the simultaneous detection of food bacteria producing biogenic amines. FEMS Microbiol. Lett. 2005, 244, 367–372. [Google Scholar] [CrossRef] [PubMed]
- Ilavenil, S.; Park, H.S.; Vijayakumar, M.; Arasu, M.V.; Kim, D.H.; Ravikumar, S.; Choi, K.C. Probiotic Potential of Lactobacillus Strains with Antifungal Activity Isolated from Animal Manure. Sci. World J. 2015, 2015, 802570. [Google Scholar] [CrossRef] [PubMed]
- Song, M.; Yun, B.; Moon, J.-H.; Park, D.-J.; Lim, K.; Oh, S. Characterization of Selected Lactobacillus Strains for Use as Probiotics. Korean J. Food Sci. Anim. Resour. 2015, 35, 551–556. [Google Scholar] [CrossRef] [PubMed]
- Campaniello, D.; Bevilacqua, A.; D’Amato, D.; Corbo, M.R.; Altieri, C.; Sinigaglia, M. Microbial characterization of table olives processed according to Spanish and natural styles. Food Technol. Biotechnol. 2005, 43, 289–294. [Google Scholar]
- Aponte, M.; Ventorino, V.; Blaiotta, G.; Volpe, G.; Farina, V.; Avellone, G.; Lanza, C.M.; Moschetti, G. Study of green Sicilian table olive fermentations through microbiological, chemical and sensory analyses. Food Microbiol. 2010, 27, 162–170. [Google Scholar] [CrossRef] [PubMed]
- Panagou, E.Z.; Tassou, C.C.; Katsaboxakis, C.Z. Induced lactic acid fermentation of untreated green olives of the Conservolea cultivar by Lactobacillus pentosus. J. Sci. Food Agric. 2003, 83, 667–674. [Google Scholar] [CrossRef]
- De Castro, A.; Montano, A.; Casado, F.-J.; Sanchez, A.-H.; Rejano, L. Utilization of Enterococcus casseliflavus and Lactobacillus pentosus as starter cultures for Spanish-style green olive fermentation. Food Microbiol. 2002, 19, 637–644. [Google Scholar] [CrossRef]
- Sarantinopoulos, P.; Andrighetto, C.; Georgalaki, M.D.; Rea, M.C.; Lombardi, A.; Cogan, T.M.; Kalantzopoulos, G.; Tsakalidou, E. Biochemical properties of enterococci relevant to their technological performance. Int. Dairy J. 2001, 11, 621–647. [Google Scholar] [CrossRef]
- Morea, M.; Baruzzi, F.; Cocconcelli, P.S. Molecular and physiological characterization of dominant bacterial populations in traditional Mozzarella cheese processing. J. Appl. Microbiol. 1999, 87, 574–582. [Google Scholar] [CrossRef] [PubMed]
- Suzzi, G.; Caruso, M.; Gardini, F.; Lombardi, A.; Vannini, L.; Guerzoni, M.E.; Andrighetto, C.; Lanorte, M.T. A survey of the enterococci isolated from an artisanal Italian goat’s cheese (semicotto caprino). J. Appl. Microbiol. 2000, 89, 267–274. [Google Scholar] [CrossRef] [PubMed]
- Durlu özkaya, F.; Xanthopoulos, V.; Tunail, N.; Litopoulou-Tzanetaki, E. Technologically important properties of lactic acid bacteria isolates from Beyaz cheese made from raw ewes’ milk. J. Appl. Microbiol. 2001, 91, 861–870. [Google Scholar] [CrossRef] [PubMed]
- Kanmani, P.; Suganya, K.; Kumar, R.S.; Yuvaraj, N.; Pattukumar, V.; Paari, K.A.; Arul, V. Synthesis and functional characterization of antibiofilm exopolysaccharide produced by Enterococcus faecium mc13 isolated from the gut of fish. Appl. Biochem. Biotechnol. 2013, 169, 1001–1015. [Google Scholar] [CrossRef] [PubMed]
- Charoenprasert, S.; Mitchell, A. Factors influencing phenolic compounds in table olives (Olea europaea). J. Agric. Food Chem. 2012, 60, 7081–7095. [Google Scholar] [CrossRef] [PubMed]
- Laëtitia, G.; Pascal, D.; Yann, D. The Citrate Metabolism in Homo- and Heterofermentative LAB: A Selective Means of Becoming Dominant over Other Microorganisms in Complex Ecosystems. Food Nutr. Sci. 2014, 5, 953–969. [Google Scholar] [CrossRef]
- González, L.; Sacristán, N.; Arenas, R.; Fresno, J.M.; Tornadijo, M.E. Enzymatic activity of lactic acid bacteria (with antimicrobial properties) isolated from a traditional Spanish cheese. Food Microbiol. 2010, 27, 592–597. [Google Scholar] [CrossRef] [PubMed]
- Creti, R.; Imperi, M.; Bertuccini, L.; Fabretti, F.; Orefici, G.; Rosa, R.D.; Baldassarri, L. Survey for virulence determinants among Enterococcus faecalis isolated from different sources. J. Med. Microbiol. 2004, 53, 13–20. [Google Scholar] [CrossRef] [PubMed]
- Franz, C.; Muscholl-Silberhorn, A.; Yousif, N.; Vancanneyt, M.; Swings, J.; Holzapfel, W. Incidence of virulence factors and antibiotic resistance among enterococci isolated from food. Appl. Environ. Microbiol. 2001, 67, 4385–4389. [Google Scholar] [CrossRef] [PubMed]
- Rosado, D.; Brito, J.C.; Harris, D.J. Molecular screening of Hepatozoon (Apicomplexa: Adeleorina) infections in Python sebae from West Africa using 18S rRNA gene sequences. Herpetol. Notes 2015, 8, 461–463. [Google Scholar]
- Klein, G. Taxonomy, ecology and antibiotic resistance of enterococci from food and the gastro-intestinal tract. Int. J. Food Microbiol. 2003, 88, 123–131. [Google Scholar] [CrossRef]
- Strompfová, V.; Lauková, A.; Ouwehand, A.C. Selection of enterococci for potential canine probiotic additives. Vet. Microbiol. 2004, 100, 107–114. [Google Scholar] [CrossRef] [PubMed]
- Martin, B.; Garriga, M.; Hugas, M.; Aymerich, T. Genetic diversity and safety aspects of enterococci from slightly fermented sausages. J. Appl. Microbiol. 2005, 98, 1177–1190. [Google Scholar] [CrossRef] [PubMed]
| Growth Media | Microorganisms | Method | Incubation Conditions |
|---|---|---|---|
| Plate Count Agar (PCA) (Merck, Darmstadt, Germany) | Total viable count | Spread plate | 30 °C/72 h |
| De Man-Rogosa-Sharpe Agar (MRS) (Oxoid, Basingstoke, UK) + natamycin 4% | Lactic acid bacteria | Pour plate/Overlay | 30 °C/72 h |
| M17 (Oxoid, Basingstoke, UK) | Lactic acid bacteria | Pour plate | 37 °C/72 h |
| Sabouraud Agar (Oxoid, Basingstoke, UK) | Yeast and molds | Spread plate | 25 °C/3 day |
| Violet Red Bile Glycose Agar (VRBGA) (BD, Sparks, MD, USA) | Enterobacteriaceae | Pour plate/Overlay | 37 °C/24 h |
| Violet Red Bile Lactose Agar (VRBL) (Oxoid, Basingstoke, UK) | Coliforms | Pour plate/Overlay | 30 °C/24 h |
| Baird Parker egg yolk tellurite (BPM) (Oxoid, Basingstoke, UK) | Micrococcaceae | Spread plate | 37 °C/48 h |
| Nutrient Agar Crystal Violet (NACV) (Oxoid, Basingstoke, UK) | Gram- cocci | Spread plate | 21 °C/48 h |
| Mannitol Salt Agar (MSA) (Oxoid, Basingstoke, UK) | Salt resistant | Spread plate | 30 °C/48 h |
| Strain | Primer | Sequence (5′–3′) | Product Size (bp) |
|---|---|---|---|
| E. durans ATCC19432 | DU1 | CCTACTGATATTAAGACAGCG | 295 |
| DU2 | TAATCCTAAGATAGGTGTTTG | ||
| E. faecalis ATCC19433 | FL1 | ACTTATGTGACTAACTTAACC | 360 |
| FL2 | TAATGGTGAATCTTGGTTTGG | ||
| E. faecium ATCC19434 | FM1 | GAAAAAACAATAGAAGAATTAT | 215 |
| FM2 | TGCTTTTTTGAATTCTTCTTTA | ||
| E. casseliflavus ATCC25788 | CA1 | TCCTGAATTAGGTGAAAAAAC | 288 |
| CA2 | GCTAGTTTACCGTCTTTAACG | ||
| E. gallinarum ATCC49673 | GA1 | TTACTTGCTGATTTTGATTCG | 173 |
| GA2 | TGAATTCTTCTTTGAAATCAG | ||
| E. hirae ATCC 8043 | HI1 | CTTTCTGATATGGATGCTGTC | 187 |
| HI2 | TAAATTCTTCCTTAAATGTTG |
| Target Gene | Primer Sequence | Annealing Temperature (°C) | Fragment Size (bp) | Reference |
|---|---|---|---|---|
| Aggregationsubstance (asa1) | GCACGCTATTACGAACTATGA TAAGAAAGAACATCACCACGA | 50 | 375 | [25] |
| Adhesion of collagen protein (ace) | GAATTGAGCAAAAGTTCAATCG GTCTGTCTTTTCACTTGTTTC | 48 | 1008 | [26] |
| Cytolysin (cylA) | ACTCGGGGATTGATAGGC GCTGCTAAAGCTGCGCTT | 52 | 688 | [25] |
| Endocartidis antigen (efaA) | GCCAATTGGGACAGACCCTC CGCCTTCTGTTCCTTCTTTGGC | 57 | 688 | [26] |
| Enterococcal surface protein (esp) | AGATTTCATCTTTGATTCTTG AATTGATTCTTTAGCATCTGG | 50 | 510 | [25] |
| Gelatinase (gelE) | TATGACAATGCTTTTTGGGAT AGATGCACCCGAAATAATATA | 47 | 213 | [25] |
| Hyluronidase (hyl) | ACAGAAGAGCTGCAGGAAATG GACTGACGTCCAAGTTTCCAA | 53 | 276 | [25] |
| Vancomycin Resistance (vanA) | TCTGCAATAGAGATAGCCGC GGAGTAGCTATCCCAGCATT | 52 | 377 | [26] |
| Vancomycin Resistance (vanB) | GCTCCGCAGCCTGCATGGACA ACGATGCCGCCATCCTCCTGC | 60 | 529 | [26] |
| Histidine decarboxylase (hdc1) | AGATGGTATTGTTTCTTATG AGACCATACACCATAACCTT | 46 | 367 | [27] |
| Histidine decarboxylase (hdc2) | AAYTCNTTYGAYTTYGARAARGARG ATNGGNGANCCDATCATYTTRTGNCC | 50 | 534 | [27] |
| Tyrosine decarboxylase (tdc) | GAYATNATNGGNATNGGNYTNGAYCARG CCRTARTCNGGNATAGCRAARTCNGTRTG | 55 | 924 | [27] |
| Ornithinedecarboxylase (odc) | GTNTTYAAYGCNGAYAARCANTAYTTYGT ATNGARTTNAGTTCRCAYTTYTCNGG | 54 | 1446 | [27] |
| Medium | Log cfu mL−1 |
|---|---|
| TAC a | 9.18 ± 0.07 |
| Yeasts and Molds | 3.48 ± 0.06 |
| LAB on MRS agar | 8.04 ± 0.04 |
| Coliforms | 4.66 ± 0.02 |
| Enterobacteriaceae | 3.83 ± 0.01 |
| Micrococacceae | nd * |
| LAB on M17 agar | 4.86 ± 0.06 |
| Salt Resistant Bacteria | 7.81 ± 0.08 |
| Gram -ve | 5.37 ± 0.03 |
| pH | 3.89 ± 0.01 |
| Proteolytic Activity (Positives) | Proteolytic Activity (Negatives) | Lipolytic Activity (Positives) | Lipolytic Activity (Negatives) | Acidification Activity (High) | Acidification Activity (Medium) | Acidification Activity (Low) | |
|---|---|---|---|---|---|---|---|
| Isolates (Enterococcus) | 7,11–14,16–18,27–37 | 1–6,8–10,15,19–26,38–64 | 11–13,16–18,32,36,42,53,54,62,63 | 1–10,14,15,19–31,33,34,35,37–41,43–52,55–61,64 | 3,23,26,27,28,42,47,52,59 | 1,2,4–10,13,14,18,21,22,24,25,29–33,35,37,39,40,41,43,50,51,55,56,57,60–64 | 11,12,15,16,17,19,20,34,36,38,44,45,46,48,49,53,54,58 |
| Survival Rate | pH 2 (1 h) | pH 3 (1 h) | pH 3 (3 h) | Bile Salts 0.3% |
|---|---|---|---|---|
| >70% | 89% | 57.8% | 82.7% | 10.9% |
| (70%, 80%) | 11% | 20.3% | 9.5% | 20.3% |
| (80%, 90%) | 0% | 12.5% | 7.8% | 34.4% |
| <90% | 0% | 9.4% | 0% | 34.4% |
| Survival Rate | pH 2 (1 h) | pH 3 (1 h) | pH 3 (3 h) | Bile Salts 0.3% |
|---|---|---|---|---|
| >70% | 2–14,17,18,20,21,22,24–64 | 7,8,9,10,12,13,14,17,18,21,24,25,26,29–39,41–46,50,53,57,58,62,63,64 | 5,7,8,9,10,11,12,13,14,17,18,20,21,22,24–51,53,54,55,56,57,58,59,61,62,63,64 | 5,7,8,17,36,37,59 |
| (70%, 80%) | 1,3,4,15,16,19,23 | 5,11,27,28,40,48,49,51,54,55,56,59,61 | 1,2,3,4,6,16,19,23,52,60 | 10,18,23,24,28,29,35,43,45,46,52,55,58 |
| (80%, 90%) | - | 6,16,19,20,22,47,52,60 | 1,2,3,4,15, | 9,14,15,16,20,21,22,27,30,31,39,40,41,42,50,51,53,54,56,57,60,61 |
| ≤90% | - | 1,2,3,4,15,23 | - | 1,2,3,4,6,11,12,13,19,25,26,32,33,34,38,44,47,48,49,62,63,64 |
| Variable | Factor 1 | Factor 2 | Factor 3 |
|---|---|---|---|
| Proteolytic activity | −0.416 | <0.01 | 0.683 |
| Lipolytic activity | 0.055 | 0.753 | 0.250 |
| Acidification activity | 0.311 | 0.413 | −0.545 |
| Survival to pH2 1h | 0.925 | 0.011 | 0.240 |
| Survival to pH3 1h | 0.967 | <0.01 | 0.117 |
| Survival to pH3 3h | 0.964 | <0.01 | 0.119 |
| Survival to bile salts | 0.139 | −0.747 | <0.01 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Anagnostopoulos, D.A.; Bozoudi, D.; Tsaltas, D. Enterococci Isolated from Cypriot Green Table Olives as a New Source of Technological and Probiotic Properties. Fermentation 2018, 4, 48. https://doi.org/10.3390/fermentation4020048
Anagnostopoulos DA, Bozoudi D, Tsaltas D. Enterococci Isolated from Cypriot Green Table Olives as a New Source of Technological and Probiotic Properties. Fermentation. 2018; 4(2):48. https://doi.org/10.3390/fermentation4020048
Chicago/Turabian StyleAnagnostopoulos, Dimitrios A., Despina Bozoudi, and Dimitrios Tsaltas. 2018. "Enterococci Isolated from Cypriot Green Table Olives as a New Source of Technological and Probiotic Properties" Fermentation 4, no. 2: 48. https://doi.org/10.3390/fermentation4020048
APA StyleAnagnostopoulos, D. A., Bozoudi, D., & Tsaltas, D. (2018). Enterococci Isolated from Cypriot Green Table Olives as a New Source of Technological and Probiotic Properties. Fermentation, 4(2), 48. https://doi.org/10.3390/fermentation4020048








