Preliminary Screening of Non-Conventional Yeasts for Olive Mill Wastewater Valorization
Abstract
1. Introduction
2. Materials and Methods
2.1. Yeast Strains
2.2. Primary Screening for Lipid Accumulation Capacity
2.3. Evaluation of Lipase Activity
2.4. Accumulation and Extraction of SCOs from Yeast Cells in Synthetic Medium
2.5. Evaluation of Selected Strain Performance in Olive Mill Wastewater-Supplemented Medium
2.6. Analytical Methods
2.7. Data Analysis
3. Results and Discussion
3.1. Chemical Composition of Olive Mill Wastewater
3.2. Qualitative Screening of Yeast Strains for SCOs Accumulation
- −
- Group 1, characterized by lipid droplets inside some cells, which include the following strains: K (Y. lipolytica); AII176 (Cut. curvatus); 1RP9; 2-13; 4-2; 4-15; 4R11 (M. pulcherrima); 4-22 (P. kudriavzevii); and RB1 (P. nakasei).
- −
- Group 2, composed of strains containing oil droplets in all the cells. These strains were the following: 1P1, 1RP2, 1SPR1, 1SPR14, 2-1, 2-7, 2R1, 4R1, AII 37, AII 68, AII 136, AII 181, AII 49 (M. pulcherrima); M, C, A, RB3 (Y. lipolytica); PD 36-2 (C. zeylanoides); AII 122 (C. tropicalis; RB5, PD 12-1, PD 49-1 (D. hansenii); ND1 (H. osmophila); AII184 (L. kluyveri); AII 40 (Mon. pollinis); AII171 (Mey. caribbica); 2-21, 4-16, 2R-17 (P. kudriavzevii); AII110 (P. kluyveri); ML-3 (P. manshurica); and D (P. nakasei). As expected, the control strains (L. tetrasporus and Y. lipolytica) showed a high intracellular lipid accumulation ability. The strains included in group 2 can be considered as potential lipid producers.
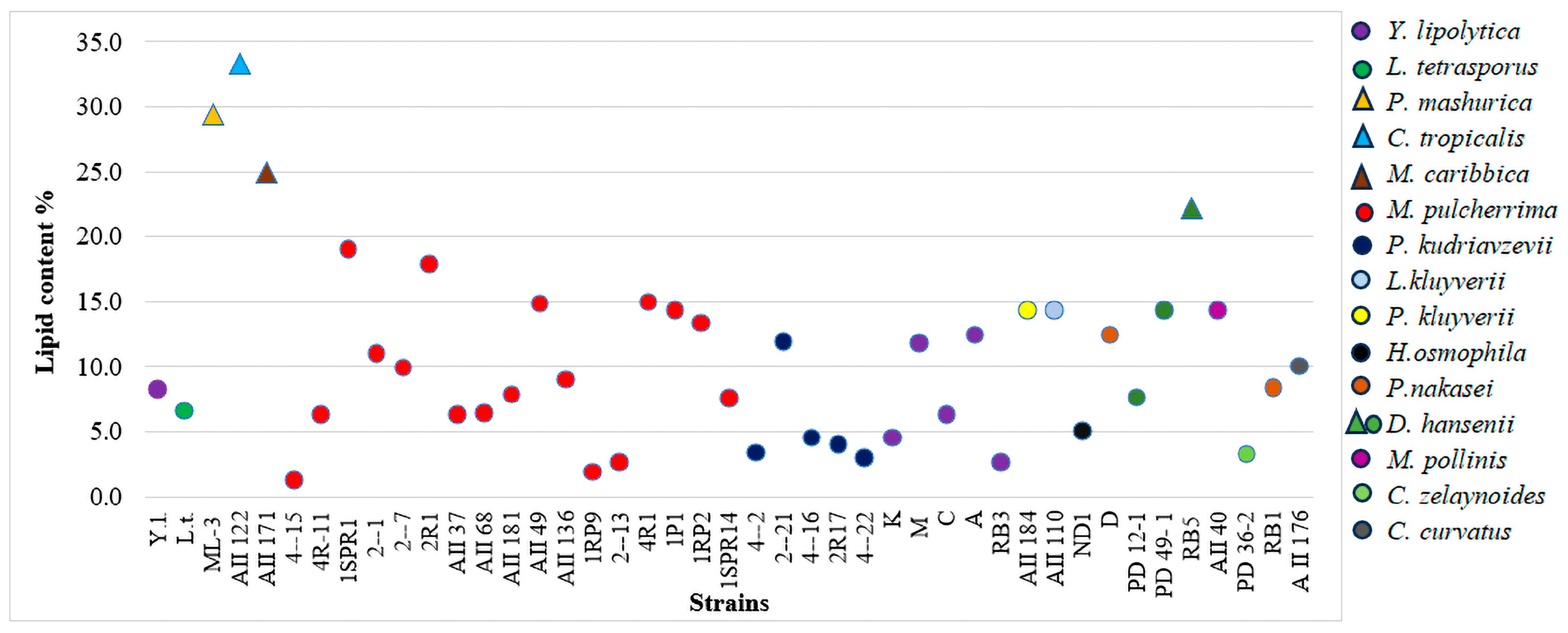
3.3. Quantitative Production of SCOs
3.4. Determination of the Lipase Activity
3.5. Lipid Production by Yeast Strains Grown in OMWW-Based Medium
3.6. Effect of the Potential Oleaginous Yeasts on OMWW-Based Medium Decolorization and Polyphenol Content Reduction
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Afonso, I.S.; Duarte, C.; Afonso, M.J.; Ribeiro, A.E.; Amaral, J.S.; Sousa, P.C.; Lima, R.A.; Ribeiro, J.E. Physicochemical characterisation of olive mill wastewaters based on extraction methods and filtration levels. Clean Technol. 2025, 7, 73. [Google Scholar] [CrossRef]
- Ginos, A.; Manios, T.; Mantzavinos, D. Treatment of olive mill effluents by coagulation–flocculation–hydrogen peroxide oxidation and effect on phytotoxicity. J. Hazard. Mater. 2006, 133, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Dermeche, S.; Nadour, M.; Larroche, C.; Moulti-Mati, F.; Michaud, P. Olive mill wastes: Biochemical characterizations and valorization strategies. Process Biochem. 2013, 48, 1532–1552. [Google Scholar] [CrossRef]
- Foti, P.; Romeo, F.V.; Occhipinti, P.S.; Timpanaro, N.; Musumeci, T.; Randazzo, C.L.; Caggia, C. Olive mill wastewater as renewable raw materials to generate high added-value ingredients for agro-food industries. Appl. Sci. 2021, 11, 7511. [Google Scholar] [CrossRef]
- Kavvadias, V.; Doula, M.; Theocharopoulos, S. Long-term effects on soil of the disposal of olive mill waste waters (OMW). Environ. Forensics 2014, 15, 37–51. [Google Scholar] [CrossRef]
- Elmansour, T.E.; Mandi, L.; Ahmali, A.; Hejjaj, A.; Bubba, M.D.; Ouazzani, N. Effect of polyphenols on activated sludge biomass during the treatment of highly diluted olive mill wastewaters. Water Sci. Technol. 2020, 82, 1416–1429. [Google Scholar] [CrossRef]
- Lindner, A.V.; Pleissner, D. Removal of phenolic compounds from olive mill wastewater by microalgae grown under dark and light conditions. Waste Biomass Valor. 2022, 13, 525–534. [Google Scholar] [CrossRef]
- Obied, H.K.; Allen, M.S.; Bedgood, D.R.; Prenzler, P.D.; Robards, K.; Stockmann, R. Bioactivity and analysis of biophenols recovered from olive mill waste. J. Agric. Food Chem. 2005, 53, 823–837. [Google Scholar] [CrossRef]
- Sayadi, S.; Ellouz, R. Roles of lignin peroxidase and manganese peroxidase from Phanerochaete chrysosporium in the decolorization of olive mill wastewater. Appl. Environ. Microbiol. 1995, 61, 1098–1103. [Google Scholar] [CrossRef]
- Ramires, F.A.; Durante, M.; Maiorano, G.; Migoni, D.; Rampino, P.; Fanizzi, F.P.; Perrotta, C.; Mita, G.; Grieco, F.; Bleve, G. Industrial scale bio-detoxification of raw olive mill wastewaters by the use of selected microbial yeast and bacterial strains to obtain a new source for fertigation. J. Environ. Manag. 2020, 265, 110574. [Google Scholar] [CrossRef]
- Mateescu, C.; Lungulescu, E.M.; Nicula, N.O. Effectiveness of biological approaches for removing persistent organic pollutants from wastewater: A mini review. Microorganisms 2024, 12, 1632. [Google Scholar] [CrossRef]
- Mishra, T.; Tiwari, P.B.; Kanchan, S.; Kesheri, M. Advances in microbial bioremediation for effective wastewater treatment. Water 2025, 17, 3196. [Google Scholar] [CrossRef]
- Bevilacqua, A.; Petruzzi, L.; Corbo, M.R.; Sinigaglia, M. Bioremediation of olive mill wastewater by yeasts—A review of the criteria for the selection of promising strains. In Applied Bioremediation—Active and Passive Approaches; Patil, Y.B., Rao, P., Eds.; IntechOpen: London, UK, 2013. [Google Scholar] [CrossRef]
- Gonçalves, C.; Lopes, M.; Ferreira, J.P.; Belo, I. Biological treatment of olive mill wastewater by non-conventional yeasts. Bioresour. Technol. 2009, 100, 3759–3763. [Google Scholar] [CrossRef]
- Caporusso, A.; Capece, A.; De Bari, I. Oleaginous yeasts as cell factories for the sustainable production of microbial lipids by the valorization of agri-food wastes. Fermentation 2021, 7, 50. [Google Scholar] [CrossRef]
- Ratledge, C.; Wynn, J.P. The biochemistry and molecular biology of lipid accumulation in oleaginous microorganisms. Adv. Appl. Microbiol. 2002, 51, 1–51. [Google Scholar] [CrossRef] [PubMed]
- Wierzchowska, K.; Szulc, K.; Zieniuk, B.; Fabiszewska, A. Bioconversion of liquid and solid lipid waste by Yarrowia lipolytica: Lipase biosynthesis and microbial lipid production. Molecules 2023, 28, 1187. [Google Scholar] [CrossRef]
- Ledesma-Amaro, R.; Nicaud, J.-M. Metabolic engineering for expanding the substrate range of Yarrowia lipolytica. Trends Biotechnol. 2016, 34, 798–809. [Google Scholar] [CrossRef]
- Donzella, S.; Serra, I.; Fumagalli, A.; Pellegrino, L.; Mosconi, G.; Scalzo, R.L.; Compagno, C. Recycling industrial food wastes for lipid production by oleaginous yeasts Rhodosporidiobolus azoricus and Cutaneotrichosporon oleaginosum. Biotechnol. Biofuels Bioprod. 2022, 15, 51. [Google Scholar] [CrossRef]
- Planonth, S.; Chantarasiri, A. The oleaginous yeast Pichia manshurica isolated from Lansium domesticum fruit in Thailand and its fatty acid composition of single cell oil. Biodiversitas 2022, 23, 801–809. [Google Scholar] [CrossRef]
- Thancharoen, K.; Malasri, A.; Leamsingkorn, W.; Boonyalit, P. Selection of oleaginous yeasts with lipid accumulation by the measurement of Sudan Black B for benefits of biodiesel. Int. J. Pharma Med. Biol. Sci. 2017, 6, 53–57. [Google Scholar] [CrossRef]
- Carrazco-Palafox, J.; Rivera-Chavira, B.E.; Ramírez-Baca, N.; Manzanares-Pérez, J.; Nevárez-Moorillón, G.V. Improved method for qualitative screening of lipolytic bacterial strains. MethodsX 2018, 5, 68–74. [Google Scholar] [CrossRef]
- Caporusso, A.; Casiello, M.; De Leo, V.; Perna, A.; Catucci, L.; D’Accolti, L.; Capece, A.; De Bari, I. Low impact methods for microbial lipids production for third generation biofuels and biobased polyurethanes. J. Environ. Chem. Eng. 2025, 13, 117375. [Google Scholar] [CrossRef]
- Dias, B.; Lopes, M.; Ramôa, R.; Pereira, A.S.; Belo, I. Candida tropicalis as a promising oleaginous yeast for olive mill wastewater bioconversion. Energies 2021, 14, 640. [Google Scholar] [CrossRef]
- Hammer, Ø.; Harper, D.A.T.; Ryan, P.D. PAST: Paleontological statistics software package for education and data analysis. Palaeontol. Electron. 2001, 4, 9. [Google Scholar]
- Russo, E.; Spallarossa, A.; Comite, A.; Pagliero, M.; Guida, P.; Belotti, V.; Caviglia, D.; Schito, A.M. Valorization and potential antimicrobial use of olive mill wastewater (OMW) from Italian olive oil production. Antioxidants 2022, 11, 903. [Google Scholar] [CrossRef]
- Issa, A.; El Riachy, M.; Bou-Mitri, C.; Doumit, J.; Skaff, W.; Karam, L. Influence of geographical origin, harvesting time and processing system on the characteristics of olive-mill wastewater: A step toward reducing the environmental impact of the olive oil sector. Environ. Technol. Innov. 2023, 32, 103365. [Google Scholar] [CrossRef]
- Ayadi, I.; Gargouri, A.; Guerfali, M. Sustainable Valorization of Olive Mill Wastewater Via Oleaginous Yeasts: Single-Cell Oil Production and Effluent Detoxification. Waste Biomass Valor. 2026, 17, 439–455. [Google Scholar] [CrossRef]
- Robles-Iglesias, R.; Naveira-Pazos, C.; Fernández-Blanco, C.; Veiga, M.C.; Kennes, C. Factors affecting the optimisation and scale-up of lipid accumulation in oleaginous yeasts for sustainable biofuels production. Renew. Sustain. Energy Rev. 2023, 171, 113043. [Google Scholar] [CrossRef]
- Poontawee, R.; Lorliam, W.; Polburee, P.; Limtong, S. Oleaginous yeasts: Biodiversity and cultivation. Fungal Biol. Rev. 2023, 44, 100295. [Google Scholar] [CrossRef]
- Lei, Y.; Wang, X.; Sun, S.; He, B.; Sun, W.; Wang, K.; Chen, Z.; Guo, Z.; Li, Z. A review of lipid accumulation by oleaginous yeasts: Culture mode. Sci. Total Environ. 2024, 919, 170385. [Google Scholar] [CrossRef]
- Pan, L.; Yang, D.; Shao, L.; Li, W.; Chen, G. Isolation of the Oleaginous Yeasts from the Soil and Studies of Their Lipid-Producing Capacities. Food Technol. Biotechnol. 2009, 47, 215–220. [Google Scholar]
- Dey, P.; Maiti, M.K. Molecular characterization of a novel isolate of Candida tropicalis for enhanced lipid production. J. Appl. Microbiol. 2013, 114, 1357–1368. [Google Scholar] [CrossRef] [PubMed]
- Legodi, L.M.; Moganedi, K.L.M. Selection of oleaginous yeasts isolated from sugarcane bagasse as the potential producers of single cell oils and other metabolites. Ann. Microbiol. 2023, 73, 21. [Google Scholar] [CrossRef]
- de Souza Miranda Muynarsk, E.; dos Santos, A.A.; Rüntzel, C.L.; Orozco Colonia, B.S.; de Oliveira Penha, R.; Vieira Marques, B.B.; Grunig Humberto da Silva, D.; Alves, S.L., Jr. From discovery to application: Twenty years of Meyerozyma caribbica. FEMS Yeast Res. 2026, 26, foag010. [Google Scholar] [CrossRef] [PubMed]
- Navarrete, C.; Estrada, M.; Martínez, J.L. Debaryomyces hansenii: An old acquaintance for a fresh start in the era of the green biotechnology. World J. Microbiol. Biotechnol. 2022, 38, 99. [Google Scholar] [CrossRef]
- Canonico, L.; Agarbati, A.; Galli, E.; Comitini, F.; Ciani, M. Metschnikowia pulcherrima as biocontrol agent and wine aroma enhancer in combination with a native Saccharomyces cerevisiae. LWT-Food Sci. Technol. 2023, 181, 114758. [Google Scholar] [CrossRef]
- Kántor, A.; Hutková, J.; Petrová, J.; Hleba, L.; Kačániová, M. Antimicrobial activity of pulcherrimin pigment produced by Metschnikowia pulcherrima against various yeast species. J. Microbiol. Biotechnol. Food Sci. 2015, 5, 282–285. [Google Scholar] [CrossRef]
- Morata, A.; Loira, I.; Escott, C.; del Fresno, J.M.; Bañuelos, M.A.; Suárez-Lepe, J.A. Applications of Metschnikowia pulcherrima in Wine Biotechnology. Fermentation 2019, 5, 63. [Google Scholar] [CrossRef]
- Karayannis, D.; Papanikolaou, S.; Vatistas, C.; Paris, C.; Chevalot, I. Yeast Lipid Produced through Glycerol Conversions and Its Use for Enzymatic Synthesis of Amino Acid-Based Biosurfactants. Int. J. Mol. Sci. 2023, 24, 714. [Google Scholar] [CrossRef]
- Di Fidio, N.; Minonne, F.; Antonetti, C.; Raspolli Galletti, A.M. Cutaneotrichosporon oleaginosus: A Versatile Whole-Cell Biocatalyst for the Production of Single-Cell Oil from Agro-Industrial Wastes. Catalysts 2021, 11, 1291. [Google Scholar] [CrossRef]
- Beopoulos, A.; Chardot, T.; Nicaud, J.-M. Yarrowia lipolytica: A model and a tool to understand the mechanisms implicated in lipid accumulation. Biochimie 2009, 91, 692–696. [Google Scholar] [CrossRef] [PubMed]
- Colacicco, M.; Ciliberti, C.; Agrimi, G.; Biundo, A.; Pisano, I. Towards the physiological understanding of Yarrowia lipolytica growth and lipase production using waste cooking oils. Energies 2022, 15, 5217. [Google Scholar] [CrossRef]
- da Silva, J.L.; Sales, M.B.; de Castro Bizerra, V.; Nobre, M.M.R.; de Sousa Braz, A.K.; da Silva Sousa, P.; Cavalcante, A.L.G.; Melo, R.L.F.; Gonçalves De Sousa Junior, P.; Neto, F.S.; et al. Lipase from Yarrowia lipolytica: Prospects as an Industrial Biocatalyst for Biotechnological Applications. Fermentation 2023, 9, 581. [Google Scholar] [CrossRef]
- Papagora, C.; Roukas, T.; Kotzekidou, P. Optimization of extracellular lipase production by Debaryomyces hansenii isolates from dry-salted olives using response surface methodology. Food Bioprod. Process. 2013, 91, 413–420. [Google Scholar] [CrossRef]
- Takaç, S.; Sengel, B.S. Extracellular lipolytic enzyme activity of a newly isolated Debaryomyces hansenii. Prep. Biochem. Biotechnol. 2010, 40, 28–37. [Google Scholar] [CrossRef]
- Fickers, P.; Marty, A.; Nicaud, J.-M. The lipases from Yarrowia lipolytica: Genetics, production, regulation, biochemical characterization and biotechnological applications. Biotechnol. Adv. 2011, 29, 632–644. [Google Scholar] [CrossRef]
- Arous, F.; Azabou, S.; Jaouani, A.; Zouari-Mechichi, H.; Nasri, M.; Mechichi, T. Biosynthesis of single-cell biomass from olive mill wastewater by newly isolated yeasts. Environ. Sci. Pollut. Res. 2016, 23, 6783–6792. [Google Scholar] [CrossRef]
- Laribi, A.; Zieniuk, B.; Bouchedja, D.; Elmechta, L.; Becila, S. Valorization of olive mill wastewater via Yarrowia lipolytica: Sustainable production of high-value metabolites and biocompounds—A review. Fermentation 2025, 11, 326. [Google Scholar] [CrossRef]
- Dourou, M.; Kancelista, A.; Juszczyk, P.; Sarris, D.; Bellou, S.; Triantaphyllidou, I.-E.; Rywinska, A.; Papanikolaou, S.; Aggelis, G. Bioconversion of Olive Mill Wastewater into High-Added Value Products. J. Clean. Prod. 2016, 139, 957–969. [Google Scholar] [CrossRef]
- Sayın, B.; Polat, Z.; Kaban, G. Enhanced Lipid Yield from Olive-Mill Wastewater by Yarrowia lipolytica NRRL YB-423. J. Agric. Prod. 2025, 6, 32–40. [Google Scholar] [CrossRef]
- Caporusso, A.; De Bari, I.; Liuzzi, F.; Albergo, R.; Valerio, V.; Viola, E.; Pietrafesa, R.; Siesto, G.; Capece, A. Optimized Conversion of Wheat Straw into Single Cell Oils by Yarrowia lipolytica and Lipomyces tetrasporus and Synthesis of Advanced Biofuels. Renew. Energy 2023, 202, 184–195. [Google Scholar] [CrossRef]
- Abeln, F.; Chuck, C.J. The history, state of the art and future prospects for oleaginous yeast research. Microb. Cell Factories 2021, 20, 221. [Google Scholar] [CrossRef]
- Rharrabti, Y.; El Yamani, M. Olive Mill Wastewater: Treatment and Valorization Technologies. In Handbook of Environmental Materials Management; Hussain, C., Ed.; Springer: Cham, Switzerland, 2019; pp. 1659–1686. [Google Scholar] [CrossRef]
- El Hassani, F.Z.; El Karkouri, A.; Errachidi, F.; Merzouki, M.; Benlemlih, M. The impact of olive mill wastewater spreading on soil and plant in arid and semi-arid areas. Environ. Nanotechnol. Monit. Manag. 2023, 20, 100798. [Google Scholar] [CrossRef]
- Ghilardi, C.; Sanmartin Negrete, P.T.; Rodríguez Gutiérrez, G.; Monetta, P.M.; Arroyo López, F.N.; Hornero-Méndez, D.; Carelli, A.A.; Borroni, V. Influence of olive mill waste phenolic compounds levels on carotenoid production by Rhodotorula spp. Process Biochem. 2022, 120, 275–286. [Google Scholar] [CrossRef]
- Crognale, S.; D’Annibale, A.; Federici, F.; Fenice, M.; Quaratino, D.; Petruccioli, M. Olive oil mill wastewater valorization by fungi. J. Chem. Technol. Biotechnol. 2006, 81, 1547–1555. [Google Scholar] [CrossRef]
- Sayadi, S.; Allouche, N.; Jaoua, M.; Aloui, F. Detrimental effects of high molecular-mass polyphenols on olive mill wastewater biotreatment. Process Biochem. 2000, 35, 725–735. [Google Scholar] [CrossRef]



| Species | Strain Code | Source |
|---|---|---|
| Pichia kudriavzevii | 2-21, 4-16, 2R17, 4-22 | Grape must |
| Hanseniaspora osmophila | ND1 | |
| Metschnikowia pulcherrima | 1P1, 1RP2, 1RP9,1SPR14, 1SPR1, 2-1, 2-7, 2-13, 2R1, 4-2, 4-15, 4R1, 4R-11 | |
| Metschnikowia pulcherrima | AII 37, AII 68, AII 136, AII 181, AII 49 | Honey bees |
| Candida tropicalis | AII 122 | |
| Meyerozyma caribbica | AII 171 | |
| Pichia kluyveri | AII 110 | |
| Lachancea kluyveri | AII 184 | |
| Moniliella pollinis | AII 40 | |
| Cutaneotrichosporon curvatus | AII 176 | |
| Yarrowia lipolytica | RB3, K, M, C, A | Cheese |
| Pichia nakasei | D, RB1 | |
| Debaryomyceshansenii | RB5, PD 12-1, PD 49- 1 | |
| Candida zeylanoides | PD 36-2 | |
| Pichia manshurica | ML-3 | Olive mill wastewater |
| Yarrowia lipolytica ATCC 46483 (DSM 8218) | Y.l. | Control |
| Lipomyces tetrasporus Li-0407 (DSM 70314) | L.t. |
| Parameters | Value (In Present Work) | Value from Reference [28] | Value from Reference [24] |
|---|---|---|---|
| pH | 4.7 | 5.4 | 5.1 |
| Total reducing sugars (g/L) | 9.9 | 9.4 | 10.1 |
| Phenolic compounds (g/L) | 5.1 | 3.4 | 4.8 |
| Lipids | - | 0.76% | - |
| Strain | Lipid Production (%) | DCW (g/L) |
|---|---|---|
| RB5 (Debaryomyces hansenii) | 2.88 ± 0.06 | 9.05 ± 0.04 |
| ML-3 (Pichia manshurica) | 1.84 ± 0.05 | 6.73 ± 0.13 (#) |
| M (Yarrowia lipolytica) | 2.78 ± 1.47 | 8.31 ± 0.14 |
| D (P. nakasei) | 2.71 ± 0.52 | 5.43 ± 0.73 (#) |
| 2R1 (Metschnikowia pulcherrima) | 6.48 ± 0.27 (#;*) | 8.30 ± 0.36 |
| AII 171 (Meyerozyma caribbica) | 4.47 ± 0.72 (#;*) | 8.13 ± 0.20 |
| AII 110 (P. kluyverii) | 4.60 ± 0.07 (#;*) | 6.11 ± 0.06 (#) |
| AII 122 (Candida tropicalis) | 2.68 ± 0.15 | 9.86 ± 0.24 |
| Y.l. (Y. lipolytica) | 1.80 ± 0.36 | 7.54 ± 1.93 |
| L.t. (Lipomyces tetrasporus) | 1.67 ± 0.02 | 8.69 ± 0.28 |
| Strains | Phenolic Compounds Reduction (%) | Decolorization (%) |
|---|---|---|
| RB5 (Debaryomyces hansenii) | 20.42 ± 2.02 (#;*) | 26.39 ± 1.94 |
| ML-3 (Pichia manshurica) | 22.95 ± 0.46 (#;*) | 17.25 ± 0.52 (#;*) |
| M (Yarrowia lipolytica) | 34.17 ± 1.43 | 22.22 ± 1.22 (*) |
| D (P. nakasei) | 23.20 ± 1.64 (#;*) | 18.98 ± 2.76 (*) |
| 2R1 (Metschnikowia pulcherrima) | 15.51 ± 0.71 (#;*) | 3.47 ± 0.02 (#;*) |
| AII 171 (Meyerozyma caribbica) | 37.07 ± 3.11 | 1.04 ± 0.16 (#;*) |
| AII 110 (P. kluyverii) | 11.6 ± 0.14 (#;*) | 25.00 ± 0.33 |
| AII 122 (Candida tropicalis) | 40.35 ± 3.05 (*) | 41.78 ± 1.80 (#;*) |
| Y.l. (Y. lipolytica) | 31.90 ± 2.85 | 29.97 ± 0.49 (#) |
| L.t. (Lipomyces tetrasporus) | 32.53 ± 2.32 | 24.07 ± 0.89 (*) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Siesto, G.; Pietrafesa, R.; Caporusso, A.; La Rocca, G.; Alberico, G.; Valerio, V.; Capece, A. Preliminary Screening of Non-Conventional Yeasts for Olive Mill Wastewater Valorization. Fermentation 2026, 12, 188. https://doi.org/10.3390/fermentation12040188
Siesto G, Pietrafesa R, Caporusso A, La Rocca G, Alberico G, Valerio V, Capece A. Preliminary Screening of Non-Conventional Yeasts for Olive Mill Wastewater Valorization. Fermentation. 2026; 12(4):188. https://doi.org/10.3390/fermentation12040188
Chicago/Turabian StyleSiesto, Gabriella, Rocchina Pietrafesa, Antonio Caporusso, Giorgia La Rocca, Grazia Alberico, Vito Valerio, and Angela Capece. 2026. "Preliminary Screening of Non-Conventional Yeasts for Olive Mill Wastewater Valorization" Fermentation 12, no. 4: 188. https://doi.org/10.3390/fermentation12040188
APA StyleSiesto, G., Pietrafesa, R., Caporusso, A., La Rocca, G., Alberico, G., Valerio, V., & Capece, A. (2026). Preliminary Screening of Non-Conventional Yeasts for Olive Mill Wastewater Valorization. Fermentation, 12(4), 188. https://doi.org/10.3390/fermentation12040188








