Recent Advances in Natural Product Biosynthesis and Yield Improvement Strategies Using Yarrowia lipolytica
Abstract
1. Introduction
2. Progress and Current Strategies in Natural Product Synthesis in Y. lipolytica
2.1. Lipids and Polymers
2.2. Sugar Alcohols
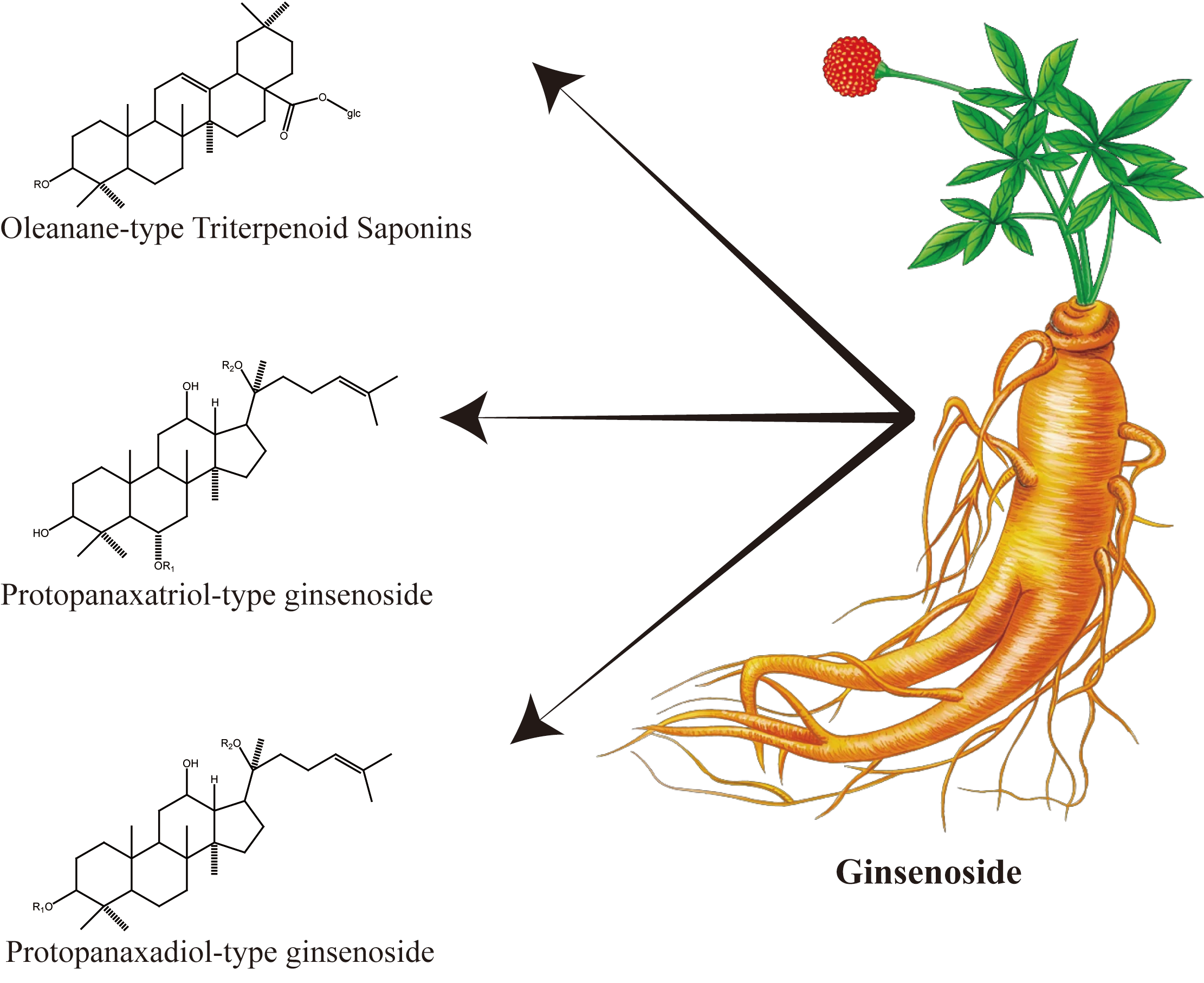
2.3. Terpenoids
2.4. Flavonoids
2.5. Organic Acids
2.6. Others
3. Upcoming Yield Improvement Strategies
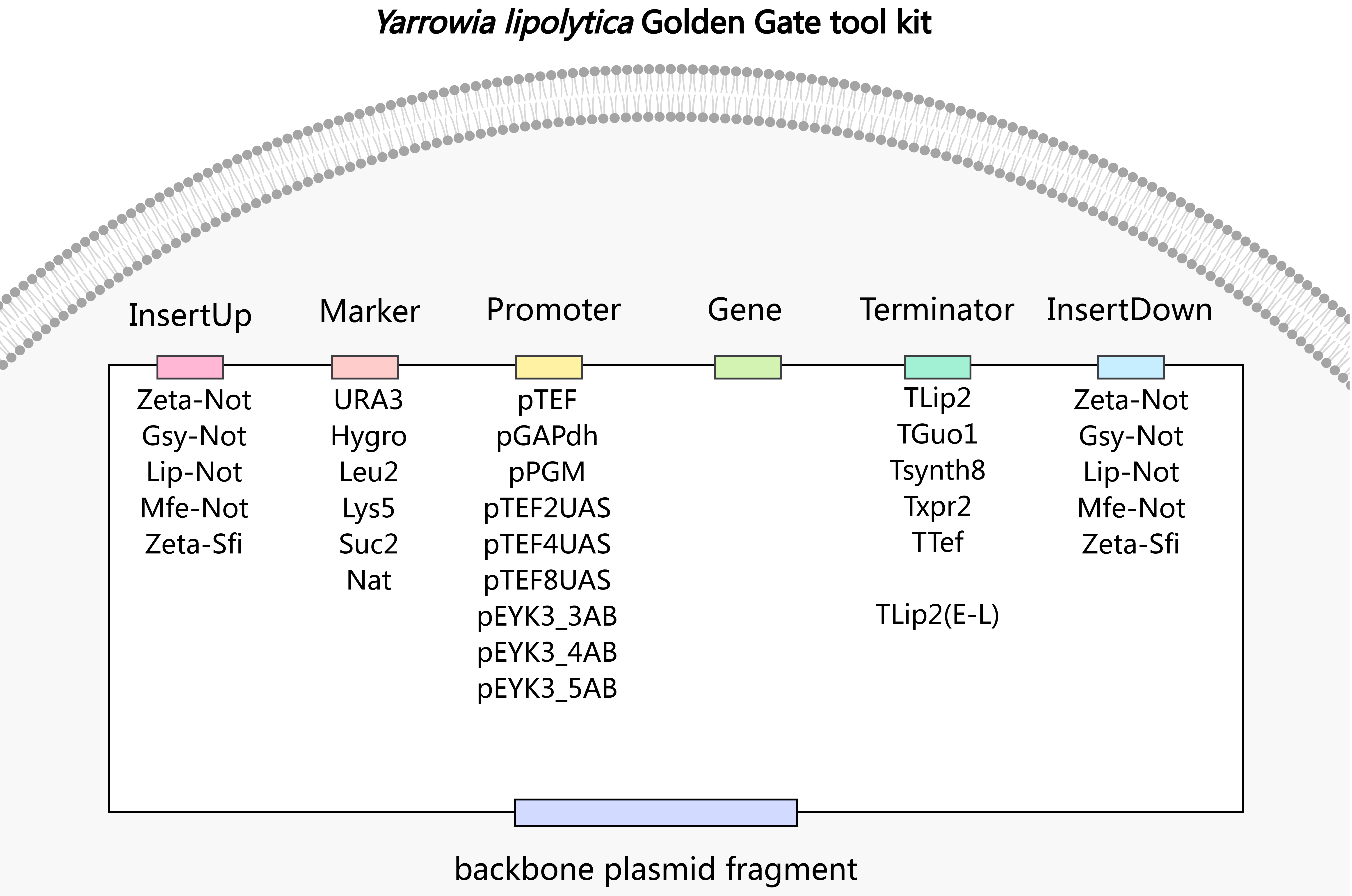
3.1. Establishment of DNA Modular Assembly Platforms
3.2. Acetyl-CoA and NADPH Supply Optimization
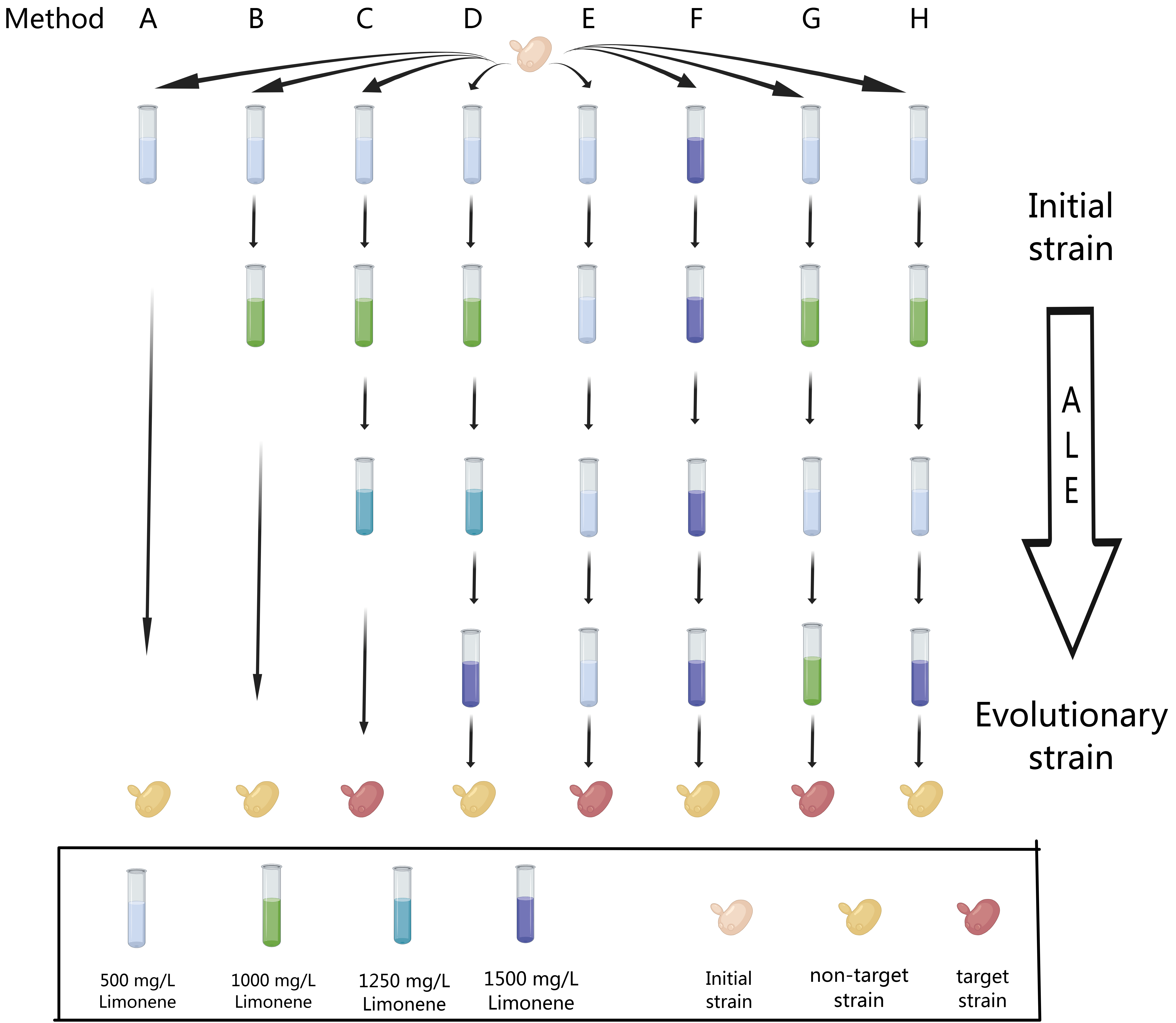
3.3. Adaptive Laboratory Evolution (ALE)
3.4. Regulation and Design of Metabolic Pathways
3.5. Substrate Selection and Optimization
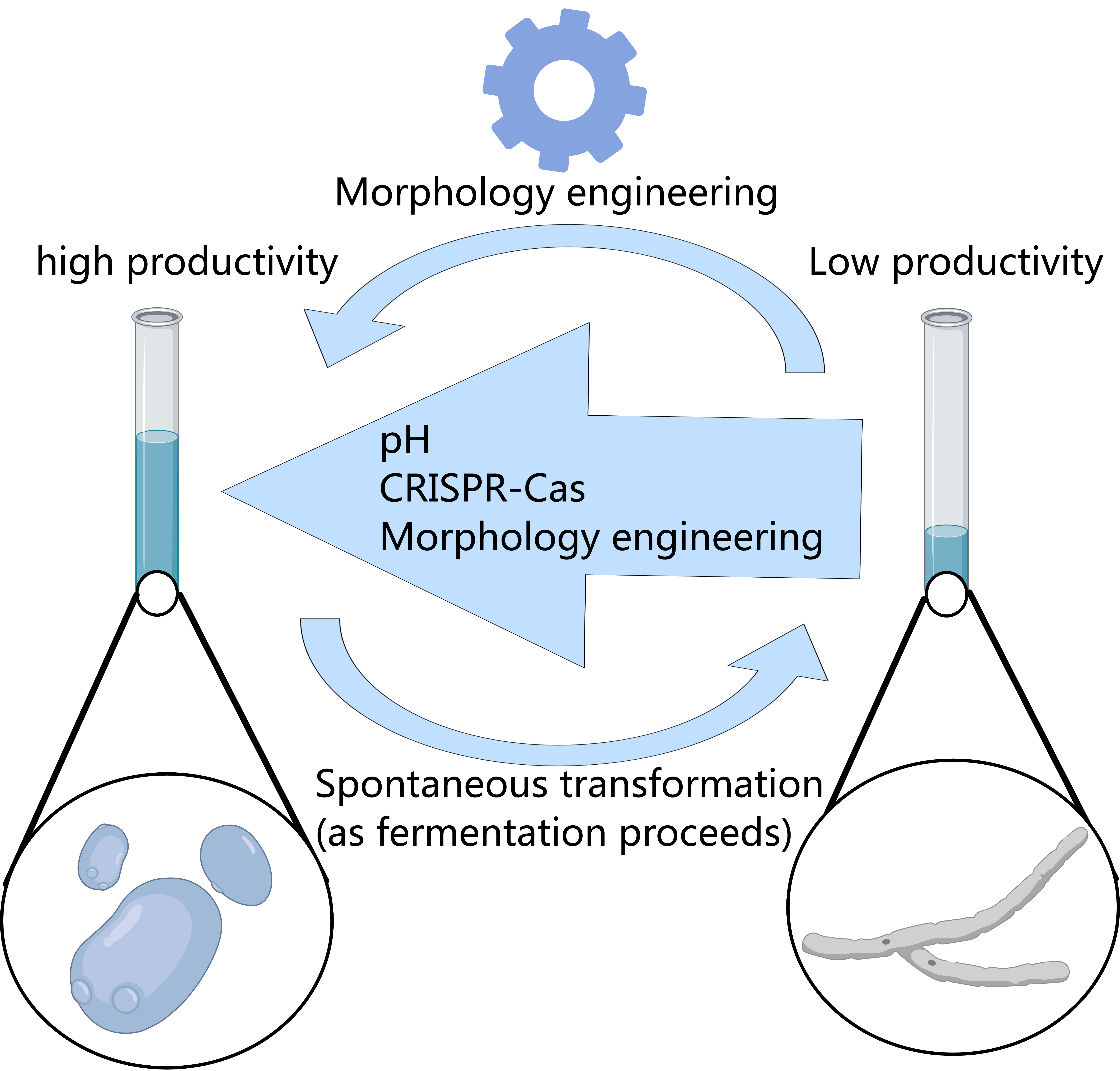
3.6. Morphological Engineering
4. Conclusions and Prospects
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ACL | ATP-citrate lyase |
| α-KG | α-ketoglutarate |
| ALE | Adaptive laboratory evolution |
| C/N | Carbon to nitrogen |
| DCW | Dry cell weight |
| DGK1 | Diacylglycerol kinase 1 |
| DMAPP | Dimethylallyl pyrophosphate |
| EPA | Eicosapentaenoic acid |
| FAP | Fatty acid photodecarboxylase |
| FPP | Farnesyl pyrophosphate |
| FPPS | Farnesyl pyrophosphate synthase |
| GGPP | Geranylgeranyl pyrophosphate |
| GGPPS | Geranylgeranyl pyrophosphate synthase |
| GPP | Geranyl pyrophosphate |
| GRAS | Generally regarded as safe |
| HAT | Histone acetyl transferase |
| HDAC | Histone deacetylase |
| IA | Itaconic acid |
| IDI | Isopentenyl pyrophosphate isomerase |
| IPP | Isopentenyl pyrophosphate |
| LB | Lipid bodies |
| LS | Limonene synthase |
| MAN | Mannitol |
| MAPK | Mitogen-activated protein kinase |
| MVA | Mevalonate |
| NDPS1 | Neryl diphosphate synthase 1 |
| PPP | Pentose phosphate pathway |
| PUFAs | Polyunsaturated fatty acids |
| SA/V | Surface-area-to-volume |
| SQE | Squalene epoxidase |
| SQS | Squalene synthase |
| TOL | Threitol |
| XDH | Xylitol dehydrogenase |
References
- Paul, P.E.V.; Sangeetha, V.; Deepika, R.G. Emerging trends in the industrial production of chemical products by microorganisms. In Recent Developments in Applied Microbiology and Biochemistry; Academic Press: Cambridge, MA, USA, 2019; pp. 107–125. [Google Scholar] [CrossRef]
- Cutlan, R.; De Rose, S.; Isupov, M.N.; Littlechild, J.A.; Harmer, N.J. Using enzyme cascades in biocatalysis: Highlight on transaminases and carboxylic acid reductases. Biochim. Biophys. Acta (BBA)-Proteins Proteom. 2020, 1868, 140322. [Google Scholar]
- Shaw, A.J.; Lam, F.H.; Hamilton, M.; Consiglio, A.; MacEwen, K.; Brevnova, E.E.; Greenhagen, E.; LaTouf, W.G.; South, C.R.; Van Dijken, H. Metabolic engineering of microbial competitive advantage for industrial fermentation processes. Science 2016, 353, 583–586. [Google Scholar] [CrossRef]
- Zhou, S.; Causey, T.; Hasona, A.; Shanmugam, K.; Ingram, L. Production of optically pure D-lactic acid in mineral salts medium by metabolically engineered Escherichia coli W3110. Appl. Environ. Microbiol. 2003, 69, 399–407. [Google Scholar] [PubMed]
- Becker, J.; Zelder, O.; Häfner, S.; Schröder, H.; Wittmann, C. From zero to hero—Design-based systems metabolic engineering of Corynebacterium glutamicum for L-lysine production. Metab. Eng. 2011, 13, 159–168. [Google Scholar] [CrossRef]
- Lee, K.H.; Park, J.H.; Kim, T.Y.; Kim, H.U.; Lee, S.Y. Systems metabolic engineering of Escherichia coli for L-threonine production. Mol. Syst. Biol. 2007, 3, 149. [Google Scholar] [PubMed]
- Widner, B.; Behr, R.; Von Dollen, S.; Tang, M.; Heu, T.; Sloma, A.; Sternberg, D.; DeAngelis, P.L.; Weigel, P.H.; Brown, S. Hyaluronic acid production in Bacillus subtilis. Appl. Environ. Microbiol. 2005, 71, 3747–3752. [Google Scholar] [CrossRef]
- Jin, P.; Zhang, L.; Yuan, P.; Kang, Z.; Du, G.; Chen, J. Efficient biosynthesis of polysaccharides chondroitin and heparosan by metabolically engineered Bacillus subtilis. Carbohydr. Polym. 2016, 140, 424–432. [Google Scholar] [CrossRef]
- Liu, L.; Li, G.; Cao, H. The Multiomics Response of Bacillus subtilis to Simultaneous Genetic and Environmental Perturbations. Microorganisms 2023, 11, 1949. [Google Scholar] [CrossRef]
- Mahmood, Z.A. Microbial amino acids production. In Microbial Biotechnology; CRC Press: Boca Raton, FL, USA, 2018; pp. 202–227. [Google Scholar]
- Pérez-García, F.; Wendisch, V.F. Transport and metabolic engineering of the cell factory Corynebacterium glutamicum. FEMS Microbiol. Lett. 2018, 365, fny166. [Google Scholar] [CrossRef]
- Park, Y.-K.; Ledesma-Amaro, R. What makes Yarrowia lipolytica well suited for industry? Trends Biotechnol. 2023, 41, 242–254. [Google Scholar] [PubMed]
- Barth, G.; Gaillardin, C. Physiology and genetics of the dimorphic fungus Yarrowia lipolytica. FEMS Microbiol. Rev. 1997, 19, 219–237. [Google Scholar] [CrossRef]
- Groenewald, M.; Boekhout, T.; Neuvéglise, C.; Gaillardin, C.; van Dijck, P.W.; Wyss, M. Yarrowia lipolytica: Safety assessment of an oleaginous yeast with a great industrial potential. Crit. Rev. Microbiol. 2014, 40, 187–206. [Google Scholar]
- Liu, L.; Alper, H.S. Draft genome sequence of the oleaginous yeast Yarrowia lipolytica PO1f, a commonly used metabolic engineering host. Genome Announc. 2014, 2, e00652-14. [Google Scholar] [CrossRef]
- Bigey, F.; Pasteur, E.; Połomska, X.; Thomas, S.; Crutz-Le Coq, A.-M.; Devillers, H.; Neuvéglise, C. Insights into the Genomic and Phenotypic Landscape of the Oleaginous Yeast Yarrowia lipolytica. J. Fungi 2023, 9, 76. [Google Scholar] [CrossRef]
- Müller, S.; Sandal, T.; Kamp-Hansen, P.; Dalbøge, H. Comparison of expression systems in the yeasts Saccharomyces cerevisiae, Hansenula polymorpha, Klyveromyces lactis, Schizosaccharomyces pombe and Yarrowia lipolytica. Cloning of two novel promoters from Yarrowia lipolytica. Yeast 1998, 14, 1267–1283. [Google Scholar] [CrossRef]
- Ji, Q.; Mai, J.; Ding, Y.; Wei, Y.; Ledesma-Amaro, R.; Ji, X.-J. Improving the homologous recombination efficiency of Yarrowia lipolytica by grafting heterologous component from Saccharomyces cerevisiae. Metab. Eng. Commun. 2020, 11, e00152. [Google Scholar] [CrossRef]
- Domınguez, A.; Ferminan, E.; Gaillardin, C. Yarrowia lipolytica: An organism amenable to genetic manipulation as a model for analyzing dimorphism in fungi. Dimorphism Hum. Pathog. Apathog. Yeasts 2000, 5, 151–172. [Google Scholar]
- Barth, G.; Gaillardin, C. Yarrowia lipolytica. In Nonconventional Yeasts in Biotechnology: A Handbook; Springer: Berlin/Heidelberg, Germany, 2013; pp. 313–388. [Google Scholar]
- Ran, M.; Zhao, G.; Jiao, L.; Gu, Z.; Yang, K.; Wang, L.; Cao, X.; Xu, L.; Yan, J.; Yan, Y. Copper ion mediates yeast-to-hypha transition in Yarrowia lipolytica. J. Fungi 2023, 9, 249. [Google Scholar]
- Timoumi, A.; Bideaux, C.; Guillouet, S.E.; Allouche, Y.; Molina-Jouve, C.; Fillaudeau, L.; Gorret, N. Influence of oxygen availability on the metabolism and morphology of Yarrowia lipolytica: Insights into the impact of glucose levels on dimorphism. Appl. Microbiol. Biotechnol. 2017, 101, 7317–7333. [Google Scholar] [CrossRef]
- Li, M.; Li, Y.-Q.; Zhao, X.-F.; Gao, X.-D. Roles of the three Ras proteins in the regulation of dimorphic transition in the yeast Yarrowia lipolytica. FEMS Yeast Res. 2014, 14, 451–463. [Google Scholar] [CrossRef]
- González-López, C.I.; Ortiz-Castellanos, L.; Ruiz-Herrera, J. The ambient pH response Rim pathway in Yarrowia lipolytica: Identification of YlRIM9 and characterization of its role in dimorphism. Curr. Microbiol. 2006, 53, 8–12. [Google Scholar] [CrossRef]
- Bellou, S.; Makri, A.; Triantaphyllidou, I.-E.; Papanikolaou, S.; Aggelis, G. Morphological and metabolic shifts of Yarrowia lipolytica induced by alteration of the dissolved oxygen concentration in the growth environment. Microbiology 2014, 160, 807–817. [Google Scholar] [CrossRef] [PubMed]
- Szabo, R. Dimorphism in Yarrowia lipolytica: Filament formation is suppressed by nitrogen starvation and inhibition of respiration. Folia Microbiol. 1999, 44, 19–24. [Google Scholar] [CrossRef]
- Hackenschmidt, S.; Bracharz, F.; Daniel, R.; Thürmer, A.; Bruder, S.; Kabisch, J. Effects of a high-cultivation temperature on the physiology of three different Yarrowia lipolytica strains. FEMS Yeast Res. 2019, 19, foz068. [Google Scholar] [CrossRef]
- Pérez-Campo, F.M.; Domínguez, A. Factors affecting the morphogenetic switch in Yarrowia lipolytica. Curr. Microbiol. 2001, 43, 429–433. [Google Scholar] [CrossRef]
- Alamillo, E.; Reyes-Becerril, M.; Cuesta, A.; Angulo, C. Marine yeast Yarrowia lipolytica improves the immune responses in Pacific red snapper (Lutjanus peru) leukocytes. Fish Shellfish Immunol. 2017, 70, 48–56. [Google Scholar]
- Bankar, A.; Zinjarde, S.; Telmore, A.; Walke, A.; Ravikumar, A. Morphological response of Yarrowia lipolytica under stress of heavy metals. Can. J. Microbiol. 2018, 64, 559–566. [Google Scholar] [CrossRef]
- Fickers, P.; Benetti, P.-H.; Waché, Y.; Marty, A.; Mauersberger, S.; Smit, M.; Nicaud, J.-M. Hydrophobic substrate utilisation by the yeast Yarrowia lipolytica, and its potential applications. FEMS Yeast Res. 2005, 5, 527–543. [Google Scholar] [CrossRef] [PubMed]
- Abghari, A.; Chen, S. Yarrowia lipolytica as an oleaginous cell factory platform for production of fatty acid-based biofuel and bioproducts. Front. Energy Res. 2014, 2, 21. [Google Scholar] [CrossRef]
- Fukuda, R.; Ohta, A. Utilization of hydrophobic substrate by Yarrowia lipolytica. In Yarrowia lipolytica: Genetics, Genomics, and Physiology; Springer: Berlin/Heidelberg, Germany, 2013; pp. 111–119. [Google Scholar]
- Koch, B.; Schmidt, C.; Daum, G. Storage lipids of yeasts: A survey of nonpolar lipid metabolism in Saccharomyces cerevisiae, Pichia pastoris, and Yarrowia lipolytica. FEMS Microbiol. Rev. 2014, 38, 892–915. [Google Scholar] [CrossRef] [PubMed]
- Mlícková, K.; Roux, E.; Athenstaedt, K.; d’Andrea, S.; Daum, G.n.; Chardot, T.; Nicaud, J.-M. Lipid accumulation, lipid body formation, and acyl coenzyme A oxidases of the yeast Yarrowia lipolytica. Appl. Environ. Microbiol. 2004, 70, 3918–3924. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Marsafari, M.; Wang, F.; Deng, L.; Xu, P. Engineering acetyl-CoA metabolic shortcut for eco-friendly production of polyketides triacetic acid lactone in Yarrowia lipolytica. Metab. Eng. 2019, 56, 60–68. [Google Scholar] [CrossRef]
- Arnesen, J.A.; Kildegaard, K.R.; Cernuda Pastor, M.; Jayachandran, S.; Kristensen, M.; Borodina, I. Yarrowia lipolytica strains engineered for the production of terpenoids. Front. Bioeng. Biotechnol. 2020, 8, 945. [Google Scholar] [CrossRef]
- Qiao, K.; Wasylenko, T.M.; Zhou, K.; Xu, P.; Stephanopoulos, G. Lipid production in Yarrowia lipolytica is maximized by engineering cytosolic redox metabolism. Nat. Biotechnol. 2017, 35, 173–177. [Google Scholar] [CrossRef]
- Wasylenko, T.M.; Ahn, W.S.; Stephanopoulos, G. The oxidative pentose phosphate pathway is the primary source of NADPH for lipid overproduction from glucose in Yarrowia lipolytica. Metab. Eng. 2015, 30, 27–39. [Google Scholar] [CrossRef]
- Liu, M.; Zhang, J.; Ye, J.; Qi, Q.; Hou, J. Morphological and metabolic engineering of Yarrowia lipolytica to increase β-carotene production. ACS Synth. Biol. 2021, 10, 3551–3560. [Google Scholar] [CrossRef]
- Biryukova, E.; Medentsev, A.; Arinbasarova, A.Y.; Akimenko, V. Tolerance of the yeast Yarrowia lipolytica to oxidative stress. Microbiology 2006, 75, 243–247. [Google Scholar] [CrossRef]
- Cordova, L.T.; Alper, H.S. Production of α-linolenic acid in Yarrowia lipolytica using low-temperature fermentation. Appl. Microbiol. Biotechnol. 2018, 102, 8809–8816. [Google Scholar]
- Andreishcheva, E.; Isakova, E.; Sidorov, N.; Abramova, N.; Ushakova, N.; Shaposhnikov, G.; Soares, M.; Zvyagilskaya, R. Adaptation to salt stress in a salt-tolerant strain of the yeast Yarrowia lipolytica. Biochem. C/C Biokhimiia 1999, 64, 1061–1067. [Google Scholar]
- Kerkhoven, E.J.; Pomraning, K.R.; Baker, S.E.; Nielsen, J. Regulation of amino-acid metabolism controls flux to lipid accumulation in Yarrowia lipolytica. NPJ Syst. Biol. Appl. 2016, 2, 16005. [Google Scholar]
- Gálvez-López, D.; Chávez-Meléndez, B.; Vázquez-Ovando, A.; Rosas-Quijano, R. The metabolism and genetic regulation of lipids in the oleaginous yeast Yarrowia lipolytica. Braz. J. Microbiol. 2019, 50, 23–31. [Google Scholar] [PubMed]
- Motter, A.E.; Gulbahce, N.; Almaas, E.; Barabási, A.L. Predicting synthetic rescues in metabolic networks. Mol. Syst. Biol. 2008, 4, 168. [Google Scholar] [CrossRef]
- Jiang, X.-R.; Chen, G.-Q. Morphology engineering of bacteria for bio-production. Biotechnol. Adv. 2016, 34, 435–440. [Google Scholar] [CrossRef] [PubMed]
- Beopoulos, A.; Cescut, J.; Haddouche, R.; Uribelarrea, J.-L.; Molina-Jouve, C.; Nicaud, J.-M. Yarrowia lipolytica as a model for bio-oil production. Prog. Lipid Res. 2009, 48, 375–387. [Google Scholar] [CrossRef]
- Papanikolaou, S.; Chevalot, I.; Komaitis, M.; Marc, I.; Aggelis, G. Single cell oil production by Yarrowia lipolytica growing on an industrial derivative of animal fat in batch cultures. Appl. Microbiol. Biotechnol. 2002, 58, 308–312. [Google Scholar] [CrossRef]
- Zárate, R.; el Jaber-Vazdekis, N.; Tejera, N.; Pérez, J.A.; Rodríguez, C. Significance of long chain polyunsaturated fatty acids in human health. Clin. Transl. Med. 2017, 6, 25. [Google Scholar] [CrossRef]
- Liu, H.-H.; Wang, C.; Lu, X.-Y.; Huang, H.; Tian, Y.; Ji, X.-J. Improved production of arachidonic acid by combined pathway engineering and synthetic enzyme fusion in Yarrowia lipolytica. J. Agric. Food Chem. 2019, 67, 9851–9857. [Google Scholar] [CrossRef]
- Li, Z.-J.; Qiao, K.; Liu, N.; Stephanopoulos, G. Engineering Yarrowia lipolytica for poly-3-hydroxybutyrate production. J. Ind. Microbiol. Biotechnol. 2017, 44, 605–612. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Liu, N.; Qiao, K.; Vogg, S.; Stephanopoulos, G. Application of metabolic controls for the maximization of lipid production in semicontinuous fermentation. Proc. Natl. Acad. Sci. USA 2017, 114, E5308–E5316. [Google Scholar]
- Chuang, L.-T.; Chen, D.-C.; Nicaud, J.-M.; Madzak, C.; Chen, Y.-H.; Huang, Y.-S. Co-expression of heterologous desaturase genes in Yarrowia lipolytica. New Biotechnol. 2010, 27, 277–282. [Google Scholar] [CrossRef]
- Tai, M.; Stephanopoulos, G. Engineering the push and pull of lipid biosynthesis in oleaginous yeast Yarrowia lipolytica for biofuel production. Metab. Eng. 2013, 15, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.-H.; Ji, X.-J.; Huang, H. Biotechnological applications of Yarrowia lipolytica: Past, present and future. Biotechnol. Adv. 2015, 33, 1522–1546. [Google Scholar] [CrossRef]
- Dulermo, T.; Nicaud, J.-M. Involvement of the G3P shuttle and β-oxidation pathway in the control of TAG synthesis and lipid accumulation in Yarrowia lipolytica. Metab. Eng. 2011, 13, 482–491. [Google Scholar] [CrossRef] [PubMed]
- Farré, J.C.; Mahalingam, S.S.; Proietto, M.; Subramani, S. Peroxisome biogenesis, membrane contact sites, and quality control. EMBO Rep. 2019, 20, e46864. [Google Scholar] [CrossRef]
- Xue, Z.; Sharpe, P.L.; Hong, S.-P.; Yadav, N.S.; Xie, D.; Short, D.R.; Damude, H.G.; Rupert, R.A.; Seip, J.E.; Wang, J. Production of omega-3 eicosapentaenoic acid by metabolic engineering of Yarrowia lipolytica. Nat. Biotechnol. 2013, 31, 734–740. [Google Scholar] [CrossRef]
- Xie, D.; Miller, E.; Tyreus, B.; Jackson, E.N.; Zhu, Q. Sustainable production of omega-3 eicosapentaenoic acid by fermentation of metabolically engineered Yarrowia lipolytica. In Quality Living Through Chemurgy and Green Chemistry; Springer: Berlin/Heidelberg, Germany, 2016; pp. 17–33. [Google Scholar]
- Cotton, C.A.; Claassens, N.J.; Benito-Vaquerizo, S.; Bar-Even, A. Renewable methanol and formate as microbial feedstocks. Curr. Opin. Biotechnol. 2020, 62, 168–180. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.; Liang, H.; Panda, S.; Fung, V.K.Y.; Zhou, J.F.J.; Zhou, K. C2 feedstock-based biomanufacturing of value-added chemicals. Curr. Opin. Biotechnol. 2022, 73, 240–245. [Google Scholar] [CrossRef]
- Christen, S.; Sauer, U. Intracellular characterization of aerobic glucose metabolism in seven yeast species by 13C flux analysis and metabolomics. FEMS Yeast Res. 2011, 11, 263–272. [Google Scholar] [CrossRef]
- Carly, F.; Fickers, P. Erythritol production by yeasts: A snapshot of current knowledge. Yeast 2018, 35, 455–463. [Google Scholar] [CrossRef]
- Rzechonek, D.A.; Day, A.M.; Quinn, J.; Mirończuk, A.M. Influence of ylHog1 MAPK kinase on Yarrowia lipolytica stress response and erythritol production. Sci. Rep. 2018, 8, 14735. [Google Scholar] [CrossRef]
- Janek, T.; Dobrowolski, A.; Biegalska, A.; Mirończuk, A.M. Characterization of erythrose reductase from Yarrowia lipolytica and its influence on erythritol synthesis. Microb. Cell Factories 2017, 16, 118. [Google Scholar] [CrossRef]
- Rakicka, M.; Rywińska, A.; Cybulski, K.; Rymowicz, W. Enhanced production of erythritol and mannitol by Yarrowia lipolytica in media containing surfactants. Braz. J. Microbiol. 2016, 47, 417–423. [Google Scholar] [CrossRef]
- Chi, P.; Wang, S.; Ge, X.; Bilal, M.; Fickers, P.; Cheng, H. Efficient D-threitol production by an engineered strain of Yarrowia lipolytica overexpressing xylitol dehydrogenase gene from Scheffersomyces stipitis. Biochem. Eng. J. 2019, 149, 107259. [Google Scholar] [CrossRef]
- Meena, M.; Prasad, V.; Zehra, A.; Gupta, V.K.; Upadhyay, R.S. Mannitol metabolism during pathogenic fungal–host interactions under stressed conditions. Front. Microbiol. 2015, 6, 1019. [Google Scholar] [CrossRef]
- Tomaszewska, L.; Rywińska, A.; Gładkowski, W. Production of erythritol and mannitol by Yarrowia lipolytica yeast in media containing glycerol. J. Ind. Microbiol. Biotechnol. 2012, 39, 1333–1343. [Google Scholar] [CrossRef]
- Zhang, Y.; Zhang, X.; Xu, Y.; Xu, S.; Bilal, M.; Cheng, H. Engineering thermotolerant Yarrowia lipolytica for sustainable biosynthesis of mannitol and fructooligosaccharides. Biochem. Eng. J. 2022, 187, 108604. [Google Scholar] [CrossRef]
- Sarris, D.; Rapti, A.; Papafotis, N.; Koutinas, A.A.; Papanikolaou, S. Production of added-value chemical compounds through bioconversions of olive-mill wastewaters blended with crude glycerol by a Yarrowia lipolytica strain. Molecules 2019, 24, 222. [Google Scholar] [CrossRef]
- Yang, L.; Kong, W.; Yang, W.; Li, D.; Zhao, S.; Wu, Y.; Zheng, S. High D-arabitol production with osmotic pressure control fed-batch fermentation by Yarrowia lipolytica and proteomic analysis under nitrogen source perturbation. Enzym. Microb. Technol. 2021, 152, 109936. [Google Scholar] [CrossRef] [PubMed]
- Kordowska-Wiater, M. Production of arabitol by yeasts: Current status and future prospects. J. Appl. Microbiol. 2015, 119, 303–314. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Wang, X. Biosynthesis of monoterpenoid and sesquiterpenoid as natural flavors and fragrances. Biotechnol. Adv. 2023, 65, 108151. [Google Scholar] [CrossRef]
- Zhang, H.; Wu, J.; Guo, J.; Hu, J. Natural terpenoid-based sustainable thermoplastics, cross-linked polymers, and supramolecular materials. Polym. Rev. 2024, 64, 119–161. [Google Scholar] [CrossRef]
- Shaikh, K.M.; Odaneth, A.A. Metabolic engineering of Yarrowia lipolytica for the production of isoprene. Biotechnol. Prog. 2021, 37, e3201. [Google Scholar] [CrossRef] [PubMed]
- Sun, P.; Schuurink, R.C.; Caissard, J.-C.; Hugueney, P.; Baudino, S. My way: Noncanonical biosynthesis pathways for plant volatiles. Trends Plant Sci. 2016, 21, 884–894. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.-R.; Wang, K.-F.; Wang, W.-J.; Ding, Y.; Shi, T.-Q.; Huang, H.; Ji, X.-J. Advances in the metabolic engineering of Yarrowia lipolytica for the production of terpenoids. Bioresour. Technol. 2019, 281, 449–456. [Google Scholar] [CrossRef] [PubMed]
- Kang, A.; George, K.W.; Wang, G.; Baidoo, E.; Keasling, J.D.; Lee, T.S. Isopentenyl diphosphate (IPP)-bypass mevalonate pathways for isopentenol production. Metab. Eng. 2016, 34, 25–35. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Rao, S.; Zhou, X.; Li, L. Plant carotenoids: Recent advances and future perspectives. Mol. Hortic. 2022, 2, 3. [Google Scholar] [CrossRef]
- Matthäus, F.; Ketelhot, M.; Gatter, M.; Barth, G. Production of lycopene in the non-carotenoid-producing yeast Yarrowia lipolytica. Appl. Environ. Microbiol. 2014, 80, 1660–1669. [Google Scholar] [CrossRef]
- Schwartz, C.; Frogue, K.; Misa, J.; Wheeldon, I. Host and pathway engineering for enhanced lycopene biosynthesis in Yarrowia lipolytica. Front. Microbiol. 2017, 8, 2233. [Google Scholar] [CrossRef]
- Zhang, X.-K.; Nie, M.-Y.; Chen, J.; Wei, L.-J.; Hua, Q. Multicopy integrants of crt genes and co-expression of AMP deaminase improve lycopene production in Yarrowia lipolytica. J. Biotechnol. 2019, 289, 46–54. [Google Scholar] [CrossRef]
- Gao, S.; Tong, Y.; Zhu, L.; Ge, M.; Zhang, Y.; Chen, D.; Jiang, Y.; Yang, S. Iterative integration of multiple-copy pathway genes in Yarrowia lipolytica for heterologous β-carotene production. Metab. Eng. 2017, 41, 192–201. [Google Scholar] [CrossRef]
- Celińska, E.; Ledesma-Amaro, R.; Larroude, M.; Rossignol, T.; Pauthenier, C.; Nicaud, J.M. Golden Gate Assembly system dedicated to complex pathway manipulation in Yarrowia lipolytica. Microb. Biotechnol. 2017, 10, 450–455. [Google Scholar] [CrossRef]
- Larroude, M.; Celinska, E.; Back, A.; Thomas, S.; Nicaud, J.M.; Ledesma-Amaro, R. A synthetic biology approach to transform Yarrowia lipolytica into a competitive biotechnological producer of β-carotene. Biotechnol. Bioeng. 2018, 115, 464–472. [Google Scholar] [CrossRef] [PubMed]
- Ma, Y.; Liu, N.; Greisen, P.; Li, J.; Qiao, K.; Huang, S.; Stephanopoulos, G. Removal of lycopene substrate inhibition enables high carotenoid productivity in Yarrowia lipolytica. Nat. Commun. 2022, 13, 572. [Google Scholar] [CrossRef]
- Abdullah, C.N.; Liu, M.; Chen, Q.; Gao, S.; Zhang, C.; Liu, S.; Zhou, J. Efficient production of astaxanthin in Yarrowia lipolytica through metabolic and enzyme engineering. Synth. Syst. Biotechnol. 2025, 10, 737–750. [Google Scholar] [CrossRef]
- Santana, H.S.; de Carvalho, F.O.; Silva, E.R.; Santos, N.G.; Shanmugam, S.; Santos, D.N.; Wisniewski, J.O.; Junior, J.S.C.; Nunes, P.S.; Araujo, A.A. Anti-inflammatory activity of limonene in the prevention and control of injuries in the respiratory system: A systematic review. Curr. Pharm. Des. 2020, 26, 2182–2191. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Sun, Z.; Chen, W. Antimicrobial susceptibility and antibacterial mechanism of limonene against Listeria monocytogenes. Molecules 2019, 25, 33. [Google Scholar] [CrossRef]
- Cao, X.; Lv, Y.-B.; Chen, J.; Imanaka, T.; Wei, L.-J.; Hua, Q. Metabolic engineering of oleaginous yeast Yarrowia lipolytica for limonene overproduction. Biotechnol. Biofuels 2016, 9, 214. [Google Scholar] [CrossRef] [PubMed]
- Cheng, B.-Q.; Wei, L.-J.; Lv, Y.-B.; Chen, J.; Hua, Q. Elevating limonene production in oleaginous yeast Yarrowia lipolytica via genetic engineering of limonene biosynthesis pathway and optimization of medium composition. Biotechnol. Bioprocess Eng. 2019, 24, 500–506. [Google Scholar] [CrossRef]
- Wu, J.; Cheng, S.; Cao, J.; Qiao, J.; Zhao, G.-R. Systematic optimization of limonene production in engineered Escherichia coli. J. Agric. Food Chem. 2019, 67, 7087–7097. [Google Scholar] [CrossRef]
- Odani, T.; Tanizawa, H.; Takino, Y. Studies on the absorption, distribution, excretion and metabolism of ginseng saponins. II. The absorption, distribution and excretion of ginsenoside Rg1 in the rat. Chem. Pharm. Bull. 1983, 31, 292–298. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Luan, X.; Yuan, Y.; Dou, D.; Huang, L. The characteristics of ginsenosides and oligosaccharides in mountain-and garden-cultivated ginseng. J. Sci. Food Agric. 2021, 101, 1491–1498. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Jeong, E.-J.; Hwang, B.; Lee, H.-D.; Lee, S.; Jang, M.; Yeo, K.; Shin, Y.; Park, S.; Lim, W.T. Pharmacological effects of biologically synthesized ginsenoside CK-rich preparation (AceCK40) on the colitis symptoms in DSS-induced Caco-2 cells and C57BL mice. Phytomedicine 2023, 124, 155301. [Google Scholar] [CrossRef]
- Shahrajabian, M.H.; Sun, W.; Cheng, Q. A review of ginseng species in different regions as a multipurpose herb in traditional Chinese medicine, modern herbology and pharmacological science. J. Med. Plants Res. 2019, 13, 213–226. [Google Scholar]
- Xu, R.; Zhang, J.; You, J.; Gao, L.; Li, Y.; Zhang, S.; Zhu, W.; Shu, S.; Xiong, C.; Xiong, H. Full-length transcriptome sequencing and modular organization analysis of oleanolic acid-and dammarane-type saponins related gene expression patterns in Panax japonicus. Genomics 2020, 112, 4137–4147. [Google Scholar]
- Cheng, Y.; Shen, L.-H.; Zhang, J.-T. Anti-amnestic and anti-aging effects of ginsenoside Rg1 and Rb1 and its mechanism of action. ACTA Pharmacol. Sin. 2005, 26, 143–149. [Google Scholar]
- Matsuda, H.; Samukawa, K.-I.; Kubo, M. Anti-inflammatory activity of ginsenoside Ro1. Planta Medica 1990, 56, 19–23. [Google Scholar] [CrossRef]
- Zhou, L.; Li, Z.K.; Li, C.Y.; Liang, Y.Q.; Yang, F. Anticancer properties and pharmaceutical applications of ginsenoside compound K: A review. Chem. Biol. Drug Des. 2022, 99, 286–300. [Google Scholar]
- Carabin, I.G.; Burdock, G.A.; Chatzidakis, C. Safety assessment of Panax ginseng. Int. J. Toxicol. 2000, 19, 293–301. [Google Scholar] [CrossRef]
- Li, X.-Y.; Sun, L.-W.; Zhao, D.-Q. Current status and problem-solving strategies for ginseng industry. Chin. J. Integr. Med. 2019, 25, 883–886. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-J.; Silva, J.; Zhang, D.; Shi, J.; Joo, S.C.; Jang, M.-G.; Kwon, W.-S.; Yang, D.-C. Development of interspecies hybrids to increase ginseng biomass and ginsenoside yield. Plant Cell Rep. 2016, 35, 779–790. [Google Scholar] [CrossRef]
- Li, D.; Wu, Y.; Wei, P.; Gao, X.; Li, M.; Zhang, C.; Zhou, Z.; Lu, W. Metabolic engineering of Yarrowia lipolytica for heterologous oleanolic acid production. Chem. Eng. Sci. 2020, 218, 115529. [Google Scholar] [CrossRef]
- Wu, Y.; Xu, S.; Gao, X.; Li, M.; Li, D.; Lu, W. Enhanced protopanaxadiol production from xylose by engineered Yarrowia lipolytica. Microb. Cell Factories 2019, 18, 83. [Google Scholar] [CrossRef]
- Rautela, A.; Chatterjee, R.; Yadav, I.; Kumar, S. A comprehensive review on engineered microbial production of farnesene for versatile applications. J. Environ. Chem. Eng. 2024, 12, 112398. [Google Scholar] [CrossRef]
- Cao, X.; Wei, L.-J.; Lin, J.-Y.; Hua, Q. Enhancing linalool production by engineering oleaginous yeast Yarrowia lipolytica. Bioresour. Technol. 2017, 245, 1641–1644. [Google Scholar] [CrossRef]
- Yang, X.; Nambou, K.; Wei, L.; Hua, Q. Heterologous production of α-farnesene in metabolically engineered strains of Yarrowia lipolytica. Bioresour. Technol. 2016, 216, 1040–1048. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Jiang, X.; Cui, Z.; Wang, Z.; Qi, Q.; Hou, J. Engineering the oleaginous yeast Yarrowia lipolytica for production of α-farnesene. Biotechnol. Biofuels 2019, 12, 296. [Google Scholar] [CrossRef]
- Taratynova, M.O.; Tikhonova, E.E.; Fedyaeva, I.M.; Dementev, D.A.; Yuzbashev, T.V.; Solovyev, A.I.; Sineoky, S.P.; Yuzbasheva, E.Y. Boosting Geranyl Diphosphate Synthesis for Linalool Production in Engineered Yarrowia lipolytica. Appl. Biochem. Biotechnol. 2023, 196, 1304–1315. [Google Scholar] [CrossRef] [PubMed]
- Bao, Y.; Feng, P.; Xu, C.; Bi, H.; Wang, M.; Fang, Y.; Tan, T. Enhancing β-farnesene production in engineered Yarrowia lipolytica: A new process control strategy. Ind. Crops Prod. 2024, 209, 117977. [Google Scholar]
- Bi, H.; Xu, C.; Bao, Y.; Zhang, C.; Wang, K.; Zhang, Y.; Wang, M.; Chen, B.; Fang, Y.; Tan, T. Enhancing precursor supply and modulating metabolism to achieve high-level production of β-farnesene in Yarrowia lipolytica. Bioresour. Technol. 2023, 382, 129171. [Google Scholar] [CrossRef]
- Ning, Y.; Liu, M.; Ru, Z.; Zeng, W.; Liu, S.; Zhou, J. Efficient synthesis of squalene by cytoplasmic-peroxisomal engineering and regulating lipid metabolism in Yarrowia lipolytica. Bioresour. Technol. 2024, 395, 130379. [Google Scholar] [CrossRef]
- Liu, Y.; Wang, Z.; Cui, Z.; Qi, Q.; Hou, J. α-Farnesene production from lipid by engineered Yarrowia lipolytica. Bioresour. Bioprocess. 2021, 8, 78. [Google Scholar] [CrossRef]
- Boo, Y.C. p-Coumaric acid as an active ingredient in cosmetics: A review focusing on its antimelanogenic effects. Antioxidants 2019, 8, 275. [Google Scholar] [CrossRef]
- Panche, A.N.; Diwan, A.D.; Chandra, S.R. Flavonoids: An overview. J. Nutr. Sci. 2016, 5, e47. [Google Scholar] [CrossRef]
- Vesaghhamedani, S.; Kiapey, S.S.M.; Shabgah, A.G.; Amiresmaili, S.; Jahanara, A.; Oveisee, M.; Shekarchi, A.; Gheibihayat, S.M.; Jadidi-Niaragh, F.; Navashenaq, J.G. From traditional medicine to modern oncology: Scutellarin, a promising natural compound in cancer treatment. Prog. Biophys. Mol. Biol. 2023, 180, 19–27. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Ma, X.; Han, J.; Zhou, M.; Ren, H.; Pan, Q.; Zheng, C.; Zheng, Q. Neuroprotective effect of scutellarin on ischemic cerebral injury by down-regulating the expression of angiotensin-converting enzyme and AT1 receptor. PLoS ONE 2016, 11, e0146197. [Google Scholar]
- Liu, Q.; Liu, Y.; Li, G.; Savolainen, O.; Chen, Y.; Nielsen, J. De novo biosynthesis of bioactive isoflavonoids by engineered yeast cell factories. Nat. Commun. 2021, 12, 6085. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, W.-B.; Li, Y.-H.; Shu, X.-C.; Pu, Y.-T.; Wang, X.-J.; Wang, T.; Wang, Z. The classification, molecular structure and biological biosynthesis of flavonoids, and their roles in biotic and abiotic stresses. Molecules 2023, 28, 3599. [Google Scholar] [CrossRef]
- Konzock, O.; Tous-Mohedano, M.; Cibin, I.; Chen, Y.; Norbeck, J. Cinnamic acid and p-coumaric acid are metabolized to 4-hydroxybenzoic acid by Yarrowia lipolytica. AMB Express 2023, 13, 84. [Google Scholar] [CrossRef]
- Gu, Y.; Ma, J.; Zhu, Y.; Ding, X.; Xu, P. Engineering Yarrowia lipolytica as a chassis for de novo synthesis of five aromatic-derived natural products and chemicals. ACS Synth. Biol. 2020, 9, 2096–2106. [Google Scholar] [CrossRef]
- Lv, Y.; Marsafari, M.; Koffas, M.; Zhou, J.; Xu, P. Optimizing oleaginous yeast cell factories for flavonoids and hydroxylated flavonoids biosynthesis. ACS Synth. Biol. 2019, 8, 2514–2523. [Google Scholar] [CrossRef]
- Wei, W.; Zhang, P.; Shang, Y.; Zhou, Y.; Ye, B.-C. Metabolically engineering of Yarrowia lipolytica for the biosynthesis of naringenin from a mixture of glucose and xylose. Bioresour. Technol. 2020, 314, 123726. [Google Scholar] [CrossRef]
- Hernández-Guzmán, C.; Prado-Barragán, A.; Gimeno, M.; Román-Guerrero, A.; Rutiaga-Quiñones, O.M.; Rocha Guzmán, N.E.; Huerta-Ochoa, S. Whole-cell bioconversion of naringenin to high added value hydroxylated compounds using Yarrowia lipolytica 2.2 ab in surface and liquid cultures. Bioprocess Biosyst. Eng. 2020, 43, 1219–1230. [Google Scholar] [CrossRef]
- Ru, Z.; Liu, M.; Chen, Q.; Li, H.; Ning, Y.; Zeng, W.; Zhou, J. High-Level De Novo Production of (2 S)-Naringenin in Yarrowia lipolytica Using Metabolic and Enzyme Engineering. ACS Agric. Sci. Technol. 2025, 5, 784–793. [Google Scholar] [CrossRef]
- Wang, Y.; Liu, X.; Chen, B.; Liu, W.; Guo, Z.; Liu, X.; Zhu, X.; Liu, J.; Zhang, J.; Li, J. Metabolic engineering of Yarrowia lipolytica for scutellarin production. Synth. Syst. Biotechnol. 2022, 7, 958–964. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zong, Z.; Liu, Y.; Zheng, M.; Li, D.; Wang, C.; Zheng, F.; Madzak, C.; Liu, Z. Metabolic engineering of Yarrowia lipolytica for the biosynthesis of crotonic acid. Bioresour. Technol. 2019, 287, 121484. [Google Scholar] [CrossRef]
- Wu, L.; Zhang, C.; Long, Y.; Chen, Q.; Zhang, W.; Liu, G. Food additives: From functions to analytical methods. Crit. Rev. Food Sci. Nutr. 2022, 62, 8497–8517. [Google Scholar] [CrossRef]
- Otten, A.; Brocker, M.; Bott, M. Metabolic engineering of Corynebacterium glutamicum for the production of itaconate. Metab. Eng. 2015, 30, 156–165. [Google Scholar] [CrossRef] [PubMed]
- Tsugawa, R.; Nakase, T.; Kobayashi, T.; Yamashita, K.; Okumura, S. Fermentation of n-Paraffins by Yeast: Part I. Fermentative Production of α-Ketoglutaric Acid by Candida lipolytica. Agric. Biol. Chem. 1969, 33, 158–167. [Google Scholar]
- Ryu, S.; Labbé, N.; Trinh, C.T. Simultaneous saccharification and fermentation of cellulose in ionic liquid for efficient production of α-ketoglutaric acid by Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2015, 99, 4237–4244. [Google Scholar] [CrossRef]
- Kamzolova, S.V.; Morgunov, I.G. α-Ketoglutaric acid production from rapeseed oil by Yarrowia lipolytica yeast. Appl. Microbiol. Biotechnol. 2013, 97, 5517–5525. [Google Scholar] [CrossRef]
- Yin, X.; Madzak, C.; Du, G.; Zhou, J.; Chen, J. Enhanced alpha-ketoglutaric acid production in Yarrowia lipolytica WSH-Z06 by regulation of the pyruvate carboxylation pathway. Appl. Microbiol. Biotechnol. 2012, 96, 1527–1537. [Google Scholar] [CrossRef]
- Yovkova, V.; Otto, C.; Aurich, A.; Mauersberger, S.; Barth, G. Engineering the α-ketoglutarate overproduction from raw glycerol by overexpression of the genes encoding NADP+-dependent isocitrate dehydrogenase and pyruvate carboxylase in Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2014, 98, 2003–2013. [Google Scholar] [CrossRef]
- Kamzolova, S.V.; Vinokurova, N.G.; Dedyukhina, E.G.; Samoilenko, V.A.; Lunina, J.N.; Mironov, A.A.; Allayarov, R.K.; Morgunov, I.G. The peculiarities of succinic acid production from rapeseed oil by Yarrowia lipolytica yeast. Appl. Microbiol. Biotechnol. 2014, 98, 4149–4157. [Google Scholar] [CrossRef]
- Bondarenko, P.Y.; Fedorov, A.; Sineoky, S. Optimization of repeated-batch fermentation of a recombinant strain of the yeast Yarrowia lipolytica for succinic acid production at low pH. Appl. Biochem. Microbiol. 2017, 53, 882–887. [Google Scholar] [CrossRef]
- Yang, X.; Wang, H.; Li, C.; Lin, C.S.K. Restoring of glucose metabolism of engineered Yarrowia lipolytica for succinic acid production via a simple and efficient adaptive evolution strategy. J. Agric. Food Chem. 2017, 65, 4133–4139. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-L.; Morcelle, C.; Cheng, Z.-L.; Chen, X.; Xu, Y.; Gao, Y.; Song, J.; Li, Z.; Smith, M.D.; Shi, M. Itaconate inhibits TET DNA dioxygenases to dampen inflammatory responses. Nat. Cell Biol. 2022, 24, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Doll, K.M.; Moser, B.R. Industrial Chemicals via Decarboxylation of Natural Carboxylic Acids. In Green Chemistry and Green Materials from Plant Oils and Natural Acids; Liu, Z., Kraus, G., Eds.; Royal Society of Chemistry: Cambridge, UK, 2023; pp. 144–158. [Google Scholar]
- Blazeck, J.; Hill, A.; Jamoussi, M.; Pan, A.; Miller, J.; Alper, H.S. Metabolic engineering of Yarrowia lipolytica for itaconic acid production. Metab. Eng. 2015, 32, 66–73. [Google Scholar] [CrossRef]
- Zhao, C.; Cui, Z.; Zhao, X.; Zhang, J.; Zhang, L.; Tian, Y.; Qi, Q.; Liu, J. Enhanced itaconic acid production in Yarrowia lipolytica via heterologous expression of a mitochondrial transporter MTT. Appl. Microbiol. Biotechnol. 2019, 103, 2181–2192. [Google Scholar] [CrossRef] [PubMed]
- Parodi, A.; Jorea, A.; Fagnoni, M.; Ravelli, D.; Samorì, C.; Torri, C.; Galletti, P. Bio-based crotonic acid from polyhydroxybutyrate: Synthesis and photocatalyzed hydroacylation. Green Chem. 2021, 23, 3420–3427. [Google Scholar] [CrossRef]
- Blazeck, J.; Liu, L.; Knight, R.; Alper, H.S. Heterologous production of pentane in the oleaginous yeast Yarrowia lipolytica. J. Biotechnol. 2013, 165, 184–194. [Google Scholar] [CrossRef]
- Xu, P.; Qiao, K.; Ahn, W.S.; Stephanopoulos, G. Engineering Yarrowia lipolytica as a platform for synthesis of drop-in transportation fuels and oleochemicals. Proc. Natl. Acad. Sci. USA 2016, 113, 10848–10853. [Google Scholar] [CrossRef]
- Li, J.; Ma, Y.; Liu, N.; Eser, B.E.; Guo, Z.; Jensen, P.R.; Stephanopoulos, G. Synthesis of high-titer alka (e) nes in Yarrowia lipolytica is enabled by a discovered mechanism. Nat. Commun. 2020, 11, 6198. [Google Scholar] [CrossRef]
- Tan, L.; Song, X.; Ren, Y.; Wang, M.; Guo, C.; Guo, D.; Gu, Y.; Li, Y.; Cao, Z.; Deng, Y. Anti-inflammatory effects of cordycepin: A review. Phytother. Res. 2021, 35, 1284–1297. [Google Scholar] [CrossRef]
- Radhi, M.; Ashraf, S.; Lawrence, S.; Tranholm, A.A.; Wellham, P.A.D.; Hafeez, A.; Khamis, A.S.; Thomas, R.; McWilliams, D.; De Moor, C.H. A systematic review of the biological effects of cordycepin. Molecules 2021, 26, 5886. [Google Scholar] [CrossRef]
- Yang, L.; Li, G.; Chai, Z.; Gong, Q.; Guo, J. Synthesis of cordycepin: Current scenario and future perspectives. Fungal Genet. Biol. 2020, 143, 103431. [Google Scholar] [CrossRef]
- Duan, X.-Y.; Tian, Y.; Song, Z.-Q.; Song, L.-P.; Lin, W.-B.; Wang, C.; Yang, H.; Lu, X.-Y.; Ji, X.-J.; Liu, H.-H. High-level de novo biosynthesis of cordycepin by systems metabolic engineering in Yarrowia lipolytica. Bioresour. Technol. 2022, 363, 127862. [Google Scholar] [CrossRef] [PubMed]
- Song, Z.; Lin, W.; Duan, X.; Song, L.; Wang, C.; Yang, H.; Lu, X.; Ji, X.; Tian, Y.; Liu, H. Increased Cordycepin Production in Yarrowia lipolytica Using Combinatorial Metabolic Engineering Strategies. ACS Synth. Biol. 2023, 12, 780–787. [Google Scholar] [CrossRef] [PubMed]
- Jeennor, S.; Anantayanon, J.; Panchanawaporn, S.; Chutrakul, C.; Vongsangnak, W.; Laoteng, K. Efficient de novo production of bioactive cordycepin by Aspergillus oryzae using a food-grade expression platform. Microb. Cell Factories 2023, 22, 253. [Google Scholar] [CrossRef] [PubMed]
- Shang, Y.; Wei, W.; Zhang, P.; Ye, B.-C. Engineering Yarrowia lipolytica for enhanced production of arbutin. J. Agric. Food Chem. 2020, 68, 1364–1372. [Google Scholar] [CrossRef]
- Saeedi, M.; Khezri, K.; Seyed Zakaryaei, A.; Mohammadamini, H. A comprehensive review of the therapeutic potential of α-arbutin. Phytother. Res. 2021, 35, 4136–4154. [Google Scholar]
- Tong, Y.; Zhou, J.; Zhang, L.; Xu, P. A golden-gate based cloning toolkit to build violacein pathway libraries in Yarrowia lipolytica. ACS Synth. Biol. 2021, 10, 115–124. [Google Scholar] [CrossRef]
- Durán, N.; Justo, G.Z.; Ferreira, C.V.; Melo, P.S.; Cordi, L.; Martins, D. Violacein: Properties and biological activities. Biotechnol. Appl. Biochem. 2007, 48, 127–133. [Google Scholar] [CrossRef] [PubMed]
- Obydennov, D.L.; El-Tantawy, A.I.; Sosnovskikh, V.Y. Triacetic acid lactone as a bioprivileged molecule in organic synthesis. Mendeleev Commun. 2019, 29, 1–10. [Google Scholar] [CrossRef]
- Yu, J.; Landberg, J.; Shavarebi, F.; Bilanchone, V.; Okerlund, A.; Wanninayake, U.; Zhao, L.; Kraus, G.; Sandmeyer, S. Bioengineering triacetic acid lactone production in Yarrowia lipolytica for pogostone synthesis. Biotechnol. Bioeng. 2018, 115, 2383–2388. [Google Scholar] [CrossRef] [PubMed]
- Anderson, L.A.; Islam, M.A.; Prather, K.L. Synthetic biology strategies for improving microbial synthesis of “green” biopolymers. J. Biol. Chem. 2018, 293, 5053–5061. [Google Scholar] [CrossRef]
- Zhang, Y.; Yu, J.; Wu, Y.; Li, M.; Zhao, Y.; Zhu, H.; Chen, C.; Wang, M.; Chen, B.; Tan, T. Efficient production of chemicals from microorganism by metabolic engineering and synthetic biology. Chin. J. Chem. Eng. 2021, 30, 14–28. [Google Scholar] [CrossRef]
- Celińska, E.; Grajek, W. A novel multigene expression construct for modification of glycerol metabolism in Yarrowia lipolytica. Microb. Cell Factories 2013, 12, 102. [Google Scholar] [CrossRef]
- Wong, L.; Engel, J.; Jin, E.; Holdridge, B.; Xu, P. YaliBricks, a versatile genetic toolkit for streamlined and rapid pathway engineering in Yarrowia lipolytica. Metab. Eng. Commun. 2017, 5, 68–77. [Google Scholar] [CrossRef]
- Larroude, M.; Park, Y.; Soudier, P.; Kubiak, M.; Nicaud, J.; Rossignol, T. A modular Golden Gate toolkit for Yarrowia lipolytica synthetic biology. Microb. Biotechnol. 2019, 12, 1249–1259. [Google Scholar] [CrossRef]
- Larroude, M.; Trabelsi, H.; Nicaud, J.-M.; Rossignol, T. A set of Yarrowia lipolytica CRISPR/Cas9 vectors for exploiting wild-type strain diversity. Biotechnol. Lett. 2020, 42, 773–785. [Google Scholar] [CrossRef] [PubMed]
- Holkenbrink, C.; Dam, M.I.; Kildegaard, K.R.; Beder, J.; Dahlin, J.; Doménech Belda, D.; Borodina, I. EasyCloneYALI: CRISPR/Cas9-based synthetic toolbox for engineering of the yeast Yarrowia lipolytica. Biotechnol. J. 2018, 13, 1700543. [Google Scholar] [CrossRef] [PubMed]
- Dulermo, T.; Lazar, Z.; Dulermo, R.; Rakicka, M.; Haddouche, R.; Nicaud, J.-M. Analysis of ATP-citrate lyase and malic enzyme mutants of Yarrowia lipolytica points out the importance of mannitol metabolism in fatty acid synthesis. Biochim. Biophys. Acta (BBA)-Mol. Cell Biol. Lipids 2015, 1851, 1107–1117. [Google Scholar] [CrossRef] [PubMed]
- Markham, K.A.; Palmer, C.M.; Chwatko, M.; Wagner, J.M.; Murray, C.; Vazquez, S.; Swaminathan, A.; Chakravarty, I.; Lynd, N.A.; Alper, H.S. Rewiring Yarrowia lipolytica toward triacetic acid lactone for materials generation. Proc. Natl. Acad. Sci. USA 2018, 115, 2096–2101. [Google Scholar] [CrossRef]
- Huang, Y.-Y.; Jian, X.-X.; Lv, Y.-B.; Nian, K.-Q.; Gao, Q.; Chen, J.; Wei, L.-J.; Hua, Q. Enhanced squalene biosynthesis in Yarrowia lipolytica based on metabolically engineered acetyl-CoA metabolism. J. Biotechnol. 2018, 281, 106–114. [Google Scholar]
- Wang, Y.; San, K.-Y.; Bennett, G.N. Cofactor engineering for advancing chemical biotechnology. Curr. Opin. Biotechnol. 2013, 24, 994–999. [Google Scholar] [CrossRef]
- Li, W.; Wu, H.; Li, M.; San, K.Y. Effect of NADPH availability on free fatty acid production in Escherichia coli. Biotechnol. Bioeng. 2018, 115, 444–452. [Google Scholar] [CrossRef]
- Liu, H.; Marsafari, M.; Deng, L.; Xu, P. Understanding lipogenesis by dynamically profiling transcriptional activity of lipogenic promoters in Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2019, 103, 3167–3179. [Google Scholar] [CrossRef]
- Liu, L.; Pan, A.; Spofford, C.; Zhou, N.; Alper, H.S. An evolutionary metabolic engineering approach for enhancing lipogenesis in Yarrowia lipolytica. Metab. Eng. 2015, 29, 36–45. [Google Scholar] [CrossRef]
- Sandberg, T.E.; Salazar, M.J.; Weng, L.L.; Palsson, B.O.; Feist, A.M. The emergence of adaptive laboratory evolution as an efficient tool for biological discovery and industrial biotechnology. Metab. Eng. 2019, 56, 1–16. [Google Scholar] [CrossRef]
- Mavrommati, M.; Daskalaki, A.; Papanikolaou, S.; Aggelis, G. Adaptive laboratory evolution principles and applications in industrial biotechnology. Biotechnol. Adv. 2022, 54, 107795. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.; Kim, P. Current Status and Applications of Adaptive Laboratory Evolution in Industrial Microorganisms. J. Microbiol. Biotechnol. 2020, 30, 793–803. [Google Scholar] [CrossRef] [PubMed]
- Mohamed, E.T.; Mundhada, H.; Landberg, J.; Cann, I.; Mackie, R.I.; Nielsen, A.T.; Herrgård, M.J.; Feist, A.M. Generation of an E. coli platform strain for improved sucrose utilization using adaptive laboratory evolution. Microb. Cell Factories 2019, 18, 116. [Google Scholar] [CrossRef]
- Portnoy, V.A.; Bezdan, D.; Zengler, K. Adaptive laboratory evolution—Harnessing the power of biology for metabolic engineering. Curr. Opin. Biotechnol. 2011, 22, 590–594. [Google Scholar] [CrossRef]
- Wang, J.; Ledesma-Amaro, R.; Wei, Y.; Ji, B.; Ji, X.-J. Metabolic engineering for increased lipid accumulation in Yarrowia lipolytica—A review. Bioresour. Technol. 2020, 313, 123707. [Google Scholar]
- Daskalaki, A.; Perdikouli, N.; Aggeli, D.; Aggelis, G. Laboratory evolution strategies for improving lipid accumulation in Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2019, 103, 8585–8596. [Google Scholar] [CrossRef]
- Li, J.; Zhu, K.; Miao, L.; Rong, L.; Zhao, Y.; Li, S.; Ma, L.; Li, J.; Zhang, C.; Xiao, D. Simultaneous improvement of limonene production and tolerance in Yarrowia lipolytica through tolerance engineering and evolutionary engineering. ACS Synth. Biol. 2021, 10, 884–896. [Google Scholar] [CrossRef]
- Liu, D.; Mannan, A.A.; Han, Y.; Oyarzún, D.A.; Zhang, F. Dynamic metabolic control: Towards precision engineering of metabolism. J. Ind. Microbiol. Biotechnol. 2018, 45, 535–543. [Google Scholar] [CrossRef]
- Zhang, S.; Ito, M.; Skerker, J.M.; Arkin, A.P.; Rao, C.V. Metabolic engineering of the oleaginous yeast Rhodosporidium toruloides IFO0880 for lipid overproduction during high-density fermentation. Appl. Microbiol. Biotechnol. 2016, 100, 9393–9405. [Google Scholar] [CrossRef]
- Yan, F.X.; Dong, G.R.; Qiang, S.; Niu, Y.J.; Hu, C.Y.; Meng, Y.H. Overexpression of △12, △15-desaturases for enhanced lipids synthesis in Yarrowia lipolytica. Front. Microbiol. 2020, 11, 289. [Google Scholar] [CrossRef] [PubMed]
- Sun, T.; Yu, Y.; Wang, L.; Qi, Y.; Xu, T.; Wang, Z.; Lin, L.; Ledesma-Amaro, R.; Ji, X.-J. Combination of a Push–Pull–Block Strategy with a Heterologous Xylose Assimilation Pathway toward Lipid Overproduction from Lignocellulose in Yarrowia lipolytica. ACS Synth. Biol. 2023, 12, 761–767. [Google Scholar] [CrossRef]
- Sagnak, R.; Cochot, S.; Molina-Jouve, C.; Nicaud, J.-M.; Guillouet, S.E. Modulation of the glycerol phosphate availability led to concomitant reduction in the citric acid excretion and increase in lipid content and yield in Yarrowia lipolytica. J. Biotechnol. 2018, 265, 40–45. [Google Scholar] [CrossRef]
- Shi, T.; Li, Y.; Zhu, L.; Tong, Y.; Yang, J.; Fang, Y.; Wang, M.; Zhang, J.; Jiang, Y.; Yang, S. Engineering the oleaginous yeast Yarrowia lipolytica for β-farnesene overproduction. Biotechnol. J. 2021, 16, 2100097. [Google Scholar]
- Kildegaard, K.R.; Adiego-Pérez, B.; Belda, D.D.; Khangura, J.K.; Holkenbrink, C.; Borodina, I. Engineering of Yarrowia lipolytica for production of astaxanthin. Synth. Syst. Biotechnol. 2017, 2, 287–294. [Google Scholar] [CrossRef]
- Yocum, H.C.; Pham, A.; Da Silva, N.A. Successful enzyme colocalization strategies in yeast for increased synthesis of non-native products. Front. Bioeng. Biotechnol. 2021, 9, 606795. [Google Scholar] [CrossRef]
- Zhao, M.; Zhao, Y.; Yao, M.; Iqbal, H.; Hu, Q.; Liu, H.; Qiao, B.; Li, C.; Skovbjerg, C.A.; Nielsen, J.C. Pathway engineering in yeast for synthesizing the complex polyketide bikaverin. Nat. Commun. 2020, 11, 6197. [Google Scholar] [CrossRef]
- Liu, M.; Wang, C.; Ren, X.; Gao, S.; Yu, S.; Zhou, J. Remodelling metabolism for high-level resveratrol production in Yarrowia lipolytica. Bioresour. Technol. 2022, 365, 128178. [Google Scholar] [CrossRef]
- Liu, H.; Chen, S.-L.; Xu, J.-Z.; Zhang, W.-G. Dual regulation of cytoplasm and peroxisomes for improved α-farnesene production in recombinant Pichia pastoris. ACS Synth. Biol. 2021, 10, 1563–1573. [Google Scholar] [CrossRef] [PubMed]
- Bhataya, A.; Schmidt-Dannert, C.; Lee, P.C. Metabolic engineering of Pichia pastoris X-33 for lycopene production. Process Biochem. 2009, 44, 1095–1102. [Google Scholar]
- Ma, Y.; Li, J.; Huang, S.; Stephanopoulos, G. Targeting pathway expression to subcellular organelles improves astaxanthin synthesis in Yarrowia lipolytica. Metab. Eng. 2021, 68, 152–161. [Google Scholar] [CrossRef]
- Yang, K.; Qiao, Y.; Li, F.; Xu, Y.; Yan, Y.; Madzak, C.; Yan, J. Subcellular engineering of lipase dependent pathways directed towards lipid related organelles for highly effectively compartmentalized biosynthesis of triacylglycerol derived products in Yarrowia lipolytica. Metab. Eng. 2019, 55, 231–238. [Google Scholar] [CrossRef]
- Ayer, A.; Sanwald, J.; Pillay, B.A.; Meyer, A.J.; Perrone, G.G.; Dawes, I.W. Distinct redox regulation in sub-cellular compartments in response to various stress conditions in Saccharomyces cerevisiae. PLoS ONE 2013, 8, e65240. [Google Scholar] [CrossRef]
- Hammer, S.K.; Avalos, J.L. Harnessing yeast organelles for metabolic engineering. Nat. Chem. Biol. 2017, 13, 823–832. [Google Scholar] [CrossRef]
- Sheng, J.; Stevens, J.; Feng, X. Pathway compartmentalization in peroxisome of Saccharomyces cerevisiae to produce versatile medium chain fatty alcohols. Sci. Rep. 2016, 6, 26884. [Google Scholar] [CrossRef]
- Yuan, J.; Ching, C.-B. Mitochondrial acetyl-CoA utilization pathway for terpenoid productions. Metab. Eng. 2016, 38, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Szczebara, F.M.; Chandelier, C.; Villeret, C.; Masurel, A.; Bourot, S.; Duport, C.; Blanchard, S.; Groisillier, A.; Testet, E.; Costaglioli, P. Total biosynthesis of hydrocortisone from a simple carbon source in yeast. Nat. Biotechnol. 2003, 21, 143–149. [Google Scholar] [CrossRef] [PubMed]
- Fuhr, A.C.F.P.; da Mata Gonçalves, I.; Santos, L.O.; Salau, N.P.G. Machine learning modeling and additive explanation techniques for glutathione production from multiple experimental growth conditions of Saccharomyces cerevisiae. Int. J. Biol. Macromol. 2024, 262, 130035. [Google Scholar] [CrossRef]
- Pensupa, N.; Treebuppachartsakul, T.; Pechprasarn, S. Machine learning models using data mining for biomass production from Yarrowia lipolytica fermentation. Fermentation 2023, 9, 239. [Google Scholar] [CrossRef]
- Antony, J. Design of Experiments for Engineers and Scientists; Elsevier: Amsterdam, The Netherlands, 2023. [Google Scholar]
- Fabiszewska, A.; Zieniuk, B.; Jasińska, K.; Nowak, D.; Sasal, K.; Kobus, J.; Jankiewicz, U. Extracellular Lipases of Yarrowia lipolytica Yeast in Media Containing Plant Oils—Studies Supported by the Design of Experiment Methodology. Appl. Sci. 2024, 14, 11449. [Google Scholar] [CrossRef]
- Małajowicz, J.; Fabiszewska, A.; Nowak, D.; Kuśmirek, S. Improved gamma-decalactone synthesis by Yarrowia lipolytica yeast using Taguchi robust design method. Appl. Sci. 2022, 12, 10231. [Google Scholar] [CrossRef]
- Dietrich, D.; Qi, H.; Jovanovic Gasovic, S.; Kohlstedt, M.; Wittmann, C. Defined YNB-free mineral medium improves reproducibility and enables high-titer production in Yarrowia lipolytica. Microb. Cell Factories 2026, 25, 59. [Google Scholar] [CrossRef]
- Meng, Q.; Kerley, M.; Ludden, P.; Belyea, R. Fermentation substrate and dilution rate interact to affect microbial growth and efficiency. J. Anim. Sci. 1999, 77, 206–214. [Google Scholar] [CrossRef]
- Li, T.; Chen, X.b.; Chen, J.c.; Wu, Q.; Chen, G.Q. Open and continuous fermentation: Products, conditions and bioprocess economy. Biotechnol. J. 2014, 9, 1503–1511. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, G.P.; Mack, M.; Contiero, J. Glycerol: A promising and abundant carbon source for industrial microbiology. Biotechnol. Adv. 2009, 27, 30–39. [Google Scholar] [CrossRef]
- Qian, Z.; Yu, J.; Chen, X.; Kang, Y.; Ren, Y.; Liu, Q.; Lu, J.; Zhao, Q.; Cai, M. De novo production of plant 4′-deoxyflavones baicalein and oroxylin A from ethanol in crabtree-negative yeast. ACS Synth. Biol. 2022, 11, 1600–1612. [Google Scholar] [CrossRef]
- Guo, J.; Zhang, Y.; Fang, J.; Ma, Z.; Li, C.; Yan, M.; Qiao, N.; Liu, Y.; Bian, M. Reduction and reuse of forestry and agricultural bio-waste through innovative green utilization approaches: A review. Forests 2024, 15, 1372. [Google Scholar] [CrossRef]
- Blasi, A.; Verardi, A.; Lopresto, C.G.; Siciliano, S.; Sangiorgio, P. Lignocellulosic agricultural waste valorization to obtain valuable products: An overview. Recycling 2023, 8, 61. [Google Scholar] [CrossRef]
- Liu, X.; Lv, J.; Xu, J.; Xia, J.; He, A.; Zhang, T.; Li, X.; Xu, J. Effects of osmotic pressure and pH on citric acid and erythritol production from waste cooking oil by Yarrowia lipolytica. Eng. Life Sci. 2018, 18, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Sarris, D.; Stoforos, N.G.; Mallouchos, A.; Kookos, I.K.; Koutinas, A.A.; Aggelis, G.; Papanikolaou, S. Production of added-value metabolites by Yarrowia lipolytica growing in olive mill wastewater-based media under aseptic and non-aseptic conditions. Eng. Life Sci. 2017, 17, 695–709. [Google Scholar] [CrossRef]
- Hassanshahian, M.; Tebyanian, H.; Cappello, S. Isolation and characterization of two crude oil-degrading yeast strains, Yarrowia lipolytica PG-20 and PG-32, from the Persian Gulf. Mar. Pollut. Bull. 2012, 64, 1386–1391. [Google Scholar] [CrossRef]
- Tan, E.C.; Lamers, P. Circular bioeconomy concepts—A perspective. Front. Sustain. 2021, 2, 701509. [Google Scholar] [CrossRef]
- Muscat, A.; de Olde, E.M.; Ripoll-Bosch, R.; Van Zanten, H.H.; Metze, T.A.; Termeer, C.J.; van Ittersum, M.K.; de Boer, I.J. Principles, drivers and opportunities of a circular bioeconomy. Nat. Food 2021, 2, 561–566. [Google Scholar] [CrossRef]
- Bijon, N.; Wassenaar, T.; Junqua, G.; Dechesne, M. Towards a sustainable bioeconomy through industrial symbiosis: Current situation and perspectives. Sustainability 2022, 14, 1605. [Google Scholar] [CrossRef]
- Dobrowolski, A.; Mituła, P.; Rymowicz, W.; Mirończuk, A.M. Efficient conversion of crude glycerol from various industrial wastes into single cell oil by yeast Yarrowia lipolytica. Bioresour. Technol. 2016, 207, 237–243. [Google Scholar] [CrossRef] [PubMed]
- Ledesma-Amaro, R.; Nicaud, J.-M. Metabolic engineering for expanding the substrate range of Yarrowia lipolytica. Trends Biotechnol. 2016, 34, 798–809. [Google Scholar] [CrossRef] [PubMed]
- Dobrowolski, A.; Drzymała, K.; Mituła, P.; Mirończuk, A.M. Production of tailor-made fatty acids from crude glycerol at low pH by Yarrowia lipolytica. Bioresour. Technol. 2020, 314, 123746. [Google Scholar] [CrossRef]
- Zhu, J.; Yang, S.; Cao, Q.; Li, X.; Jiao, L.; Shi, Y.; Yan, Y.; Xu, L.; Yang, M.; Xie, X.; et al. Engineering Yarrowia lipolytica as a Cellulolytic Cell Factory for Production of p-Coumaric Acid from Cellulose and Hemicellulose. J. Agric. Food Chem. 2024, 72, 5867–5877. [Google Scholar] [CrossRef]
- Bhukya, G.; Pilli, S.; Chinthala, S.; Tyagi, R. Pre-treated crude glycerol a valuable green energy source in the era of circular bioeconomy—A review. Circ. Econ. Sustain. 2024, 4, 877–904. [Google Scholar] [CrossRef]
- Elsayed, M.; Eraky, M.; Osman, A.I.; Wang, J.; Farghali, M.; Rashwan, A.K.; Yacoub, I.H.; Hanelt, D.; Abomohra, A. Sustainable valorization of waste glycerol into bioethanol and biodiesel through biocircular approaches: A review. Environ. Chem. Lett. 2024, 22, 609–634. [Google Scholar] [CrossRef]
- Kumar, L.R.; Yellapu, S.K.; Tyagi, R.D.; Drogui, P. Microbial lipid and biodiesel production from municipal sludge fortified with crude glycerol medium using pH-based fed-batch strategy. J. Environ. Chem. Eng. 2021, 9, 105032. [Google Scholar] [CrossRef]
- Devi, A.; Bajar, S.; Kour, H.; Kothari, R.; Pant, D.; Singh, A. Lignocellulosic biomass valorization for bioethanol production: A circular bioeconomy approach. Bioenergy Res. 2022, 15, 1820–1841. [Google Scholar] [CrossRef]
- Dale, B.E.; Linden, J.C. Fermentation substrates and economics. Annu. Rep. Ferment. Process. 1984, 7, 107–134. [Google Scholar]
- Ioannidou, S.M.; Stylianou, E.; Pateraki, C.; Kookos, I.; Rabaey, K.; Koutinas, A.; Ladakis, D. Techno-economic and environmental sustainability assessment of succinic acid production from municipal biowaste using an electrochemical membrane bioreactor. Chem. Eng. J. 2023, 473, 145070. [Google Scholar] [CrossRef]
- Vickram, S.; Infant, S.S.; Balamurugan, B.S.; Jayanthi, P.; Sivasubramanian, M. Techno-economic and life cycle analysis of biorefineries: Assessing sustainability and scalability in the bioeconomy. Environ. Qual. Manag. 2025, 34, e70077. [Google Scholar] [CrossRef]
- Scown, C.D.; Baral, N.R.; Yang, M.; Vora, N.; Huntington, T. Technoeconomic analysis for biofuels and bioproducts. Curr. Opin. Biotechnol. 2021, 67, 58–64. [Google Scholar] [CrossRef]
- Huo, K.; Zhao, F.; Zhang, F.; Liu, R.; Yang, C. Morphology engineering: A new strategy to construct microbial cell factories. World J. Microbiol. Biotechnol. 2020, 36, 127. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-C.; Guo, Y.; Liu, X.; Chen, X.-G.; Wu, Q.; Chen, G.-Q. Engineering cell wall synthesis mechanism for enhanced PHB accumulation in E. coli. Metab. Eng. 2018, 45, 32–42. [Google Scholar] [CrossRef]
- Ward, J.E., Jr.; Lutkenhaus, J. Overproduction of FtsZ induces minicell formation in E. coli. Cell 1985, 42, 941–949. [Google Scholar] [CrossRef]
- Timoumi, A.; Guillouet, S.E.; Molina-Jouve, C.; Fillaudeau, L.; Gorret, N. Impacts of environmental conditions on product formation and morphology of Yarrowia lipolytica. Appl. Microbiol. Biotechnol. 2018, 102, 3831–3848. [Google Scholar] [CrossRef]
- Ferreira, P.; Lopes, M.; Mota, M.; Belo, I. Oxygen transfer rate and pH as major operating parameters of citric acid production from glycerol by Yarrowia lipolytica W29 and CBS 2073. Chem. Pap. 2016, 70, 869–876. [Google Scholar] [CrossRef]
- Bergs, A.; Ishitsuka, Y.; Evangelinos, M.; Nienhaus, G.U.; Takeshita, N. Dynamics of actin cables in polarized growth of the filamentous fungus Aspergillus nidulans. Front. Microbiol. 2016, 7, 682. [Google Scholar] [CrossRef]
- Rollenhagen, C.; Mamtani, S.; Ma, D.; Dixit, R.; Eszterhas, S.; Lee, S.A. The role of secretory pathways in Candida albicans pathogenesis. J. Fungi 2020, 6, 26. [Google Scholar] [CrossRef]
- Celińska, E.; Nicaud, J.-M. Filamentous fungi-like secretory pathway strayed in a yeast system: Peculiarities of Yarrowia lipolytica secretory pathway underlying its extraordinary performance. Appl. Microbiol. Biotechnol. 2019, 103, 39–52. [Google Scholar] [CrossRef]
- Titorenko, V.I.; Ogrydziak, D.M.; Rachubinski, R.A. Four distinct secretory pathways serve protein secretion, cell surface growth, and peroxisome biogenesis in the yeast Yarrowia lipolytica. Mol. Cell. Biol. 1997, 17, 5210–5226. [Google Scholar] [CrossRef] [PubMed]
- Lopez, M.C.; Nicaud, J.-M.; Skinner, H.B.; Vergnolle, C.; Kader, J.C.; Bankaitis, V.A.; Gaillardin, C. A phosphatidylinositol/phosphatidylcholine transfer protein is required for differentiation of the dimorphic yeast Yarrowia lipolytica from the yeast to the mycelial form. J. Cell Biol. 1994, 125, 113–127. [Google Scholar] [CrossRef]
- Käppeli, O.; Walther, P.; Mueller, M.; Fiechter, A. Structure of the cell surface of the yeast Candida tropicalis and its relation to hydrocarbon transport. Arch. Microbiol. 1984, 138, 279–282. [Google Scholar] [CrossRef] [PubMed]
- Harris, L.K.; Theriot, J.A. Surface area to volume ratio: A natural variable for bacterial morphogenesis. Trends Microbiol. 2018, 26, 815–832. [Google Scholar] [CrossRef]
- Taheri-Araghi, S.; Bradde, S.; Sauls, J.T.; Hill, N.S.; Levin, P.A.; Paulsson, J.; Vergassola, M.; Jun, S. Cell-size control and homeostasis in bacteria. Curr. Biol. 2015, 25, 385–391. [Google Scholar] [CrossRef]
- Olsvik, E.; Kristiansen, B. Rheology of filamentous fermentations. Biotechnol. Adv. 1994, 12, 1–39. [Google Scholar] [CrossRef]
- Caşcaval, D.; Galaction, A.-I.; Turnea, M. Comparative analysis of oxygen transfer rate distribution in stirred bioreactor for simulated and real fermentation broths. J. Ind. Microbiol. Biotechnol. 2011, 38, 1449–1466. [Google Scholar] [CrossRef] [PubMed]
- Bliatsiou, C.; Schrinner, K.; Waldherr, P.; Tesche, S.; Böhm, L.; Kraume, M.; Krull, R. Rheological characteristics of filamentous cultivation broths and suitable model fluids. Biochem. Eng. J. 2020, 163, 107746. [Google Scholar] [CrossRef]
- Albaek, M.O.; Gernaey, K.V.; Hansen, M.S.; Stocks, S.M. Modeling enzyme production with Aspergillus oryzae in pilot scale vessels with different agitation, aeration, and agitator types. Biotechnol. Bioeng. 2011, 108, 1828–1840. [Google Scholar] [CrossRef]
- Magdouli, S.; Brar, S.K.; Blais, J.F. Morphology and rheological behaviour of Yarrowia lipolytica: Impact of dissolved oxygen level on cell growth and lipid composition. Process Biochem. 2018, 65, 1–10. [Google Scholar] [CrossRef]
- Rupeš, I.; Webb, B.A.; Mak, A.; Young, P.G. G2/M arrest caused by actin disruption is a manifestation of the cell size checkpoint in fission yeast. Mol. Biol. Cell 2001, 12, 3892–3903. [Google Scholar] [CrossRef]
- Pomraning, K.R.; Bredeweg, E.L.; Kerkhoven, E.J.; Barry, K.; Haridas, S.; Hundley, H.; LaButti, K.; Lipzen, A.; Yan, M.; Magnuson, J.K. Regulation of yeast-to-hyphae transition in Yarrowia lipolytica. mSphere 2018, 3, e00541-18. [Google Scholar] [CrossRef]
- Hurtado, C.A.; Rachubinski, R.A. Isolation and characterization of YlBEM1, a gene required for cell polarization and differentiation in the dimorphic yeast Yarrowia lipolytica. Eukaryot. Cell 2002, 1, 526–537. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.-F.; Li, M.; Li, Y.-Q.; Chen, X.-D.; Gao, X.-D. The TEA/ATTS transcription factor YlTec1p represses the yeast-to-hypha transition in the dimorphic yeast Yarrowia lipolytica. FEMS Yeast Res. 2013, 13, 50–61. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Vazquez, A.; Gonzalez-Hernandez, A.; Domínguez, Á.; Rachubinski, R.; Riquelme, M.; Cuellar-Mata, P.; Guzman, J.C.T. Identification of the transcription factor Znc1p, which regulates the yeast-to-hypha transition in the dimorphic yeast Yarrowia lipolytica. PLoS ONE 2013, 8, e66790. [Google Scholar] [CrossRef]
- Toya, Y.; Shimizu, H. Flux controlling technology for central carbon metabolism for efficient microbial bio-production. Curr. Opin. Biotechnol. 2020, 64, 169–174. [Google Scholar] [CrossRef] [PubMed]
- Stephanopoulos, G.; Aristidou, A.A.; Nielsen, J. Metabolic Engineering: Principles and Methodologies; Academic Press: Orlando, FL, USA, 1998. [Google Scholar]
- Liang, B.; Yang, Q.; Zhang, X.; Zhao, Y.; Liu, Y.; Yang, J.; Wang, Z. Switching carbon metabolic flux for enhancing the production of sesquiterpene-based high-density biofuel precursor in Saccharomyces cerevisiae. Biotechnol. Biofuels Bioprod. 2023, 16, 124. [Google Scholar] [CrossRef]
- Steinberg, G. Hyphal growth: A tale of motors, lipids, and the Spitzenkorper. Eukaryot. Cell 2007, 6, 351–360. [Google Scholar] [CrossRef] [PubMed]
- Lew, R.R. How does a hypha grow? The biophysics of pressurized growth in fungi. Nat. Rev. Microbiol. 2011, 9, 509–518. [Google Scholar] [CrossRef] [PubMed]
- Steinberg, G.; Peñalva, M.A.; Riquelme, M.; Wösten, H.A.; Harris, S.D. Cell biology of hyphal growth. Microbiol. Spectr. 2017, 5. [Google Scholar] [CrossRef] [PubMed]
- Bedsole, C.O.; Vasselli, J.G.; Shaw, B.D. Endocytosis in filamentous Fungi: Coordinating polarized hyphal growth and membrane recycling. Fungal Genet. Biol. 2025, 179, 104000. [Google Scholar] [CrossRef]
- Cervantes-Chávez, J.A.; Ruiz-Herrera, J. STE11 disruption reveals the central role of a MAPK pathway in dimorphism and mating in Yarrowia lipolytica. FEMS Yeast Res. 2006, 6, 801–815. [Google Scholar]
- Shu, T.; He, X.-Y.; Chen, J.-W.; Mao, Y.-S.; Gao, X.-D. The pH-responsive transcription factors YlRim101 and Mhy1 regulate alkaline pH-induced filamentation in the dimorphic yeast Yarrowia lipolytica. mSphere 2021, 6, e00179-21. [Google Scholar] [CrossRef]
- Torres-Guzmán, J.C.; Domínguez, A. HOY1, a homeo gene required for hyphal formation in Yarrowia lipolytica. Mol. Cell. Biol. 1997, 17, 6283–6293. [Google Scholar]
- Chen, J.-W.; Mao, Y.-S.; Yan, L.-Q.; Gao, X.-D. The zinc finger transcription factor Fts2 represses the yeast-to-filament transition in the dimorphic yeast Yarrowia lipolytica. mSphere 2022, 7, e00450-22. [Google Scholar]
- Csank, C.; Schröppel, K.; Leberer, E.; Harcus, D.; Mohamed, O.; Meloche, S.; Thomas, D.Y.; Whiteway, M. Roles of the Candida albicans mitogen-activated protein kinase homolog, Cek1p, in hyphal development and systemic candidiasis. Infect. Immun. 1998, 66, 2713–2721. [Google Scholar]
- Mösch, H.-U.; Roberts, R.L.; Fink, G.R. Ras2 signals via the Cdc42/Ste20/mitogen-activated protein kinase module to induce filamentous growth in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 1996, 93, 5352–5356. [Google Scholar]
- Leberer, E.; Harcus, D.; Dignard, D.; Johnson, L.; Ushinsky, S.; Thomas, D.Y.; Schröppel, K. Ras links cellular morphogenesis to virulence by regulation of the MAP kinase and cAMP signalling pathways in the pathogenic fungus Candida albicans. Mol. Microbiol. 2001, 42, 673–687. [Google Scholar] [CrossRef] [PubMed]
- Robertson, L.S.; Fink, G.R. The three yeast A kinases have specific signaling functions in pseudohyphal growth. Proc. Natl. Acad. Sci. USA 1998, 95, 13783–13787. [Google Scholar] [CrossRef]
- Zhang, C.; Tian, J.; Zhang, J.; Liu, R.; Zhao, X.; Lu, W. Engineering and transcriptome study of Saccharomyces cerevisiae to produce ginsenoside compound K by glycerol. Biotechnol. J. 2024, 19, e2300383. [Google Scholar] [CrossRef]
- Cervantes-Chávez, J.A.; Kronberg, F.; Passeron, S.; Ruiz-Herrera, J. Regulatory role of the PKA pathway in dimorphism and mating in Yarrowia lipolytica. Fungal Genet. Biol. 2009, 46, 390–399. [Google Scholar] [CrossRef]
- Wu, H.; Shu, T.; Mao, Y.-S.; Gao, X.-D. Characterization of the promoter, downstream target genes and recognition DNA sequence of Mhy1, a key filamentation-promoting transcription factor in the dimorphic yeast Yarrowia lipolytica. Curr. Genet. 2020, 66, 245–261. [Google Scholar]
- Kouzarides, T. Histone acetylases and deacetylases in cell proliferation. Curr. Opin. Genet. Dev. 1999, 9, 40–48. [Google Scholar] [CrossRef]
- Ververis, K.; Karagiannis, T.C. Overview of the classical histone deacetylase enzymes and histone deacetylase inhibitors. Int. Sch. Res. Not. 2012, 2012, 130360. [Google Scholar] [CrossRef]
- Baidyaroy, D.; Brosch, G.; Ahn, J.-h.; Graessle, S.; Wegener, S.; Tonukari, N.J.; Caballero, O.; Loidl, P.; Walton, J.D. A gene related to yeast HOS2 histone deacetylase affects extracellular depolymerase expression and virulence in a plant pathogenic fungus. Plant Cell 2001, 13, 1609–1624. [Google Scholar][Green Version]
- Brandão, F.; Esher, S.K.; Ost, K.S.; Pianalto, K.; Nichols, C.B.; Fernandes, L.; Bocca, A.L.; Poças-Fonseca, M.J.; Alspaugh, J.A. HDAC genes play distinct and redundant roles in Cryptococcus neoformans virulence. Sci. Rep. 2018, 8, 5209. [Google Scholar] [CrossRef] [PubMed]
- Dang, V.D.; Benedik, M.J.; Ekwall, K.; Choi, J.; Allshire, R.C.; Levin, H.L. A new member of the Sin3 family of corepressors is essential for cell viability and required for retroelement propagation in fission yeast. Mol. Cell. Biol. 1999, 19, 2351–2365. [Google Scholar] [CrossRef] [PubMed]
- Ding, S.-L.; Liu, W.; Iliuk, A.; Ribot, C.; Vallet, J.; Tao, A.; Wang, Y.; Lebrun, M.-H.; Xu, J.-R. The tig1 histone deacetylase complex regulates infectious growth in the rice blast fungus Magnaporthe oryzae. Plant Cell 2010, 22, 2495–2508. [Google Scholar] [CrossRef]
- Li, Y.; Wang, C.; Liu, W.; Wang, G.; Kang, Z.; Kistler, H.C.; Xu, J.-R. The HDF1 histone deacetylase gene is important for conidiation, sexual reproduction, and pathogenesis in Fusarium graminearum. Mol. Plant-Microbe Interact. 2011, 24, 487–496. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.; Oh, J.-H.; Shwab, E.K.; Dagenais, T.R.; Andes, D.; Keller, N.P. HdaA, a class 2 histone deacetylase of Aspergillus fumigatus, affects germination and secondary metabolite production. Fungal Genet. Biol. 2009, 46, 782–790. [Google Scholar] [CrossRef] [PubMed]
- Roa-Cordero, M.V.; Arenas-Sepúlveda, C.A.; Herrera-Plata, M.C.; Leal-Pinto, S.M.; Villota-Salazar, N.A.; González-Prieto, J.M. Switching off the yeast-to-hyphae transition in Yarrowia lipolytica through histone deacetylase inhibitors. Res. Microbiol. 2025, 176, 104299. [Google Scholar] [CrossRef] [PubMed]




| Product (by Class) | Maximum Yields Achieved | Metabolic Engineering Strategy | Reference(s) |
|---|---|---|---|
| Lipids | |||
| Arachidonic acid | 118.1 mg/L | Fuse Δ-9 elongase and Δ-8 desaturase | [51] |
| Eicosapentaenoic acid | EPA at 15% of dry cell weight | Inactivate PEX10 to reduce peroxisome biogenesis | [59] |
| poly-3-hydroxybutyrate | 7.35 g/L | Utilize acetate as substrate; use fed-batch fermentation | [52] |
| Sugar alcohols | |||
| Erythritol #1 | 44.5 g/L; 20% yield increase compared to control | Overexpress erythrose reductase | [66] |
| Erythritol #2 | 142 g/L; 15% yield increase compared to control | Add sorbitan monolaurate to fed-batch cultures with glycerol as a carbon source | [67] |
| Threitol | 112 g/L | Overexpress xylitol dehydrogenase | [68] |
| Mannitol | 5.3 g/L ~ 13.1 g/L | Overexpress Hsp90; add olive mill wastewater to media | [72] |
| Arabitol | 118.5 g/L | Control osmolarity and optimize C/N ratio | [72,73] |
| Terpenoids | |||
| β-carotene #1 | 4 g/L | Overexpress 11 biosynthetic genes; use fed-batch fermentation | [85] |
| β-carotene #2 | 6.5 g/L | Optimize promoter–gene pairs for expression | [87] |
| β-carotene #3 | 39.5 g/L | Regulate metabolic flux | [88] |
| Lycopene | 46–60 mg/g DCW (dry cell weight) | Optimize and overexpress related genes | [84] |
| Astaxanthin | 2820 mg/L | Employ metabolic pathway engineering and enzyme complex construction | [89] |
| Limonene #1 | 23.56 mg/L | Optimize the substrate pyruvic acid and dodecane concentrations in flask culture | [92] |
| Limonene #2 | 165.3 mg/L | Introduce an additional copy of limonene synthesis gene | [93] |
| Oleanolic acid | 540.7 mg/L | Fuse cytochrome P450 (CYP716A12) to NADPH-P450 reductase | [106] |
| Protopanaxadiol | 300 mg/L | Introduce xylose reductase (XR) and xylitol dehydrogenase (XDH); utilize xylose as sole carbon source | [107] |
| Linalool #1 | 7.0 ± 0.3 mg/L | Overexpress genes in the MVA pathway | [109] |
| Linalool #2 | 110 mg/L | Disrupt diacylglycerol kinase (DGK1) | [112] |
| β-Farnesene #1 | 22 g/L | Apply nitrogen-limited conditions to maximize carbon flux | [113] |
| β-Farnesene #2 | 28.9 g/L | Regulate carbon flux and optimize metabolic Pathways | [114] |
| α-Farnesene #1 | 57 ± 1 mg/L | Overexpress genes in the MVA pathway | [110] |
| α-Farnesene #2 | 25.6 g/L | Overexpress genes in the MVA pathway highlighted as bottlenecks; employ strain selection | [111] |
| Squalene | 51.2 g/L | Increase the supply of acetyl-CoA in peroxisomes and cytoplasm | [115] |
| Flavonoids | |||
| p-Coumaric acid | 593.53 ± 28.75 mg/L | Overexpress bottleneck genes; remove competing pathways | [124] |
| Naringenin #1 | 252.4 mg/L | Overexpress bottleneck genes; control pH and C/N ratio | [125] |
| Naringenin #2 | 715.3 ± 12.8 mg/L | Introduce xylose reductase (XR) and xylitol dehydrogenase (XDH); utilize xylose as a carbon source | [126] |
| (2S)-Naringenin | 8.65 g/L | Employ enzyme engineering, precursor supply enhancement, and multicopy pathway integration | [128] |
| Scutellarin | 346 mg/L | Express multiple copies of biosynthetic genes and use a fed-batch bioreactor | [129] |
| Organic acids | |||
| α-Ketoglutarate | 186 g/L | Co-express orthologous biosynthetic genes | [137] |
| Succinic acid | 87.9 g/L | Evolve a natural mutant in vitro; employ batch fermentation with food-waste hydrolysate | [140] |
| Itaconic acid #1 | 4.6 g/L | Overexpress precursor pathway; inhibit competing pathway; optimize C/N ratio | [143] |
| Itaconic acid #2 | 22 g/L | Overexpress the mitochondrial cis-aconitate transporter MTT | [144] |
| Crotonic acid | 220 ± 8 mg/L | Overexpress orthologous genes to produce precursors | [130] |
| Others | |||
| Pentane | 4.98 mg/L | Overexpress lipoxygenase | [146] |
| Diverse alkanes and alcohols | 142.5 mg/L of FAEEs, 23.3 mg/L of fatty alkanes, 2.15 g/L of fatty alcohols, 9.67 g/L of free fatty acids, and 66.4 g/L of triacylglyceride | Activate free fatty acids by conjugation to CoA; overexpress processing enzymes; decouple nitrogen starvation from lipogenesis | [147] |
| Alkanes | 1.47 g/L | Overexpress fatty acid photodecarboxylase; optimize C/N ratio | [148] |
| Cordycepin | 4.36 g/L | Optimize promoters; overexpress genes supporting metabolic precursors; optimize feedstock | [153] |
| Arbutin | 8.6 ± 0.7 g/L | Optimize promoters | [155] |
| Violacein | 366.30 ± 28.99 mg/L | Optimize promoters; overexpress genes in the shikimate, pentose phosphate, and glycolytic pathways | [124] |
| Triacetic acid lactone | 4.76 g/L | Overexpress genes in precursor pathways; utilize glacial acetic acid as a substrate; optimize C/N ratio; inhibit fatty acid synthesis | [36] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Gu, Z.; Li, X.; Moore, F.; Jamithireddy, A.K.; Bates, S.; Harmer, N.J. Recent Advances in Natural Product Biosynthesis and Yield Improvement Strategies Using Yarrowia lipolytica. Fermentation 2026, 12, 182. https://doi.org/10.3390/fermentation12040182
Gu Z, Li X, Moore F, Jamithireddy AK, Bates S, Harmer NJ. Recent Advances in Natural Product Biosynthesis and Yield Improvement Strategies Using Yarrowia lipolytica. Fermentation. 2026; 12(4):182. https://doi.org/10.3390/fermentation12040182
Chicago/Turabian StyleGu, Zhaorui, Xiaojing Li, Freddie Moore, Anil Kumar Jamithireddy, Steven Bates, and Nicholas J. Harmer. 2026. "Recent Advances in Natural Product Biosynthesis and Yield Improvement Strategies Using Yarrowia lipolytica" Fermentation 12, no. 4: 182. https://doi.org/10.3390/fermentation12040182
APA StyleGu, Z., Li, X., Moore, F., Jamithireddy, A. K., Bates, S., & Harmer, N. J. (2026). Recent Advances in Natural Product Biosynthesis and Yield Improvement Strategies Using Yarrowia lipolytica. Fermentation, 12(4), 182. https://doi.org/10.3390/fermentation12040182







