Ethnobiotechnological Analysis of the Lactic Bacterial Diversity in the Mezcal Fermentation of Four Palenques in Oaxaca, Mexico
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling, Isolation, and Propagation of Bacteria
2.2. DNA Extraction, PCR Amplification, and Phylogenetic Analysis
2.3. Phenotypic Characterization of Bacterial Isolates
2.4. Tolerance Resistance Experiment
Strategy for Studying Tolerance to Ethanol
2.5. Ethnobiotechnological Data Acquisition and Analysis
3. Results
3.1. Molecular and Phenotypic Identification of Bacteria in the Must of Mezcal Fermentation
3.2. Phylogenetic Analysis of Isolates of Must
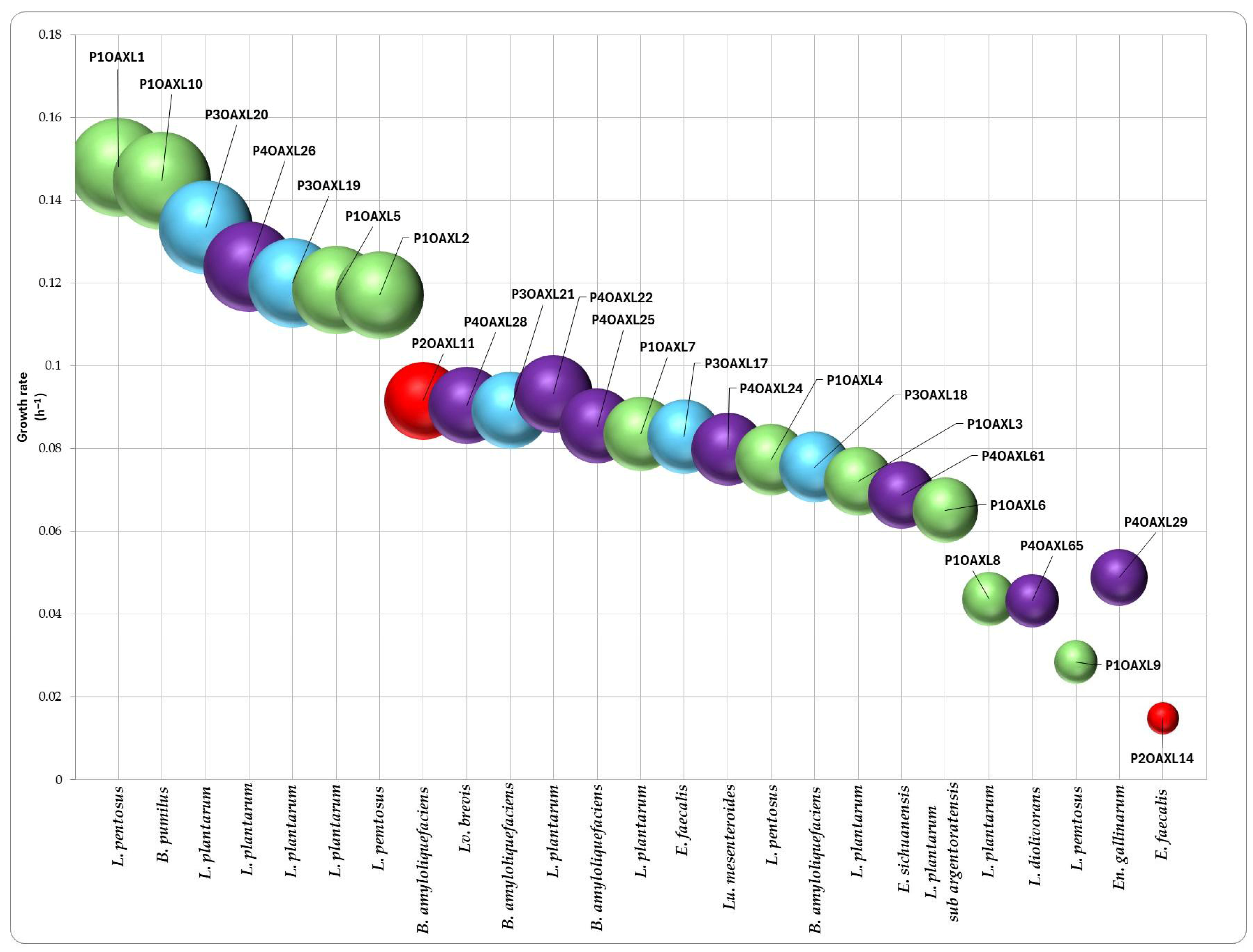
3.3. Ethanol Tolerance
3.4. Bacterial Diversity and Ethnobiotechnological Context
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- De León Rodríguez, A.; González Hernández, L.; Barba de la Rosa, A.P.; Escalante Minakata, M.d.P.; López, M.G. Characterization of Volatile Compounds of Mezcal, an Ethnic Alcoholic Beverage Obtained from Agave salmiana. J. Agric. Food Chem. 2006, 54, 1337–1341. [Google Scholar] [CrossRef]
- Páez-Lerma, J.B.; Arias-García, A.; Rutiaga-Quiñones, O.M.; Barrio, E.; Soto-Cruz, N.O. Yeasts Isolated from the Alcoholic Fermentation of Agave duranguensis During Mezcal Production. Food Biotechnol. 2013, 27, 342–356. [Google Scholar] [CrossRef]
- Becerra-Lucio, P.A.; Diego-García, E.; Guillén-Navarro, K.; Peña-Ramírez, Y.J. Unveiling the Microbial Ecology behind Mezcal: A Spirit Drink with a Growing Global Demand. Fermentation 2022, 8, 662. [Google Scholar] [CrossRef]
- Contreras-Negrete, G.; Valiente-Banuet, A.; Molina-Freaner, F.; Partida-Martínez, L.P.; Hernández-López, A. Agricultural Practices and Environmental Factors Drive Microbial Communities in the Mezcal-Producing Agave angustifolia Haw. Microb. Ecol. 2025, 87, 181. [Google Scholar] [CrossRef] [PubMed]
- Axelsson, L. Lactic Acid Bacteria. 2004. Available online: https://books.google.rs/books?hl=sr&lr=&id=P0p5_uXL9uQC&oi=fnd&pg=PA1&dq=Axelsson,+L.+Lactic+Acid+Bacteria.+2004&ots=gQa2VZXHGf&sig=dL45XYC0oVUILoNKZd5En9-n4J4&redir_esc=y#v=onepage&q=Axelsson%2C%20L.%20Lactic%20Acid%20Bacteria.%202004&f=false (accessed on 18 December 2025).
- Echegaray, N.; Yilmaz, B.; Sharma, H.; Kumar, M.; Pateiro, M.; Ozogul, F.; Lorenzo, J.M. A novel approach to Lactiplantibacillus plantarum: From probiotic properties to the omics insights. Microbiol. Res. 2023, 268, 127289. [Google Scholar] [CrossRef]
- Jara-Servin, A.; Alcaraz, L.D.; Juarez-Serrano, S.I.; Espinosa-Jaime, A.; Barajas, I.; Morales, L.; DeLuna, A.; Hernández-López, A.; Mancera, E. Microbial Communities in Agave Fermentations Vary by Local Biogeographic Regions. Environ. Microbiol. Rep. 2025, 17, e70057. [Google Scholar] [CrossRef]
- Ali, M.A.; Ahmed, T.; Ibrahim, E.; Rizwan, M.; Chong, K.P.; Yong, J.W.H. A review on mechanisms and prospects of endophytic bacteria in biocontrol of plant pathogenic fungi and their plant growth-promoting activities. Heliyon 2024, 10, e31573. [Google Scholar] [CrossRef]
- Shahid, M.; Singh, U.B.; Khan, M.S.; Singh, P.; Kumar, R.; Singh, R.N.; Kumar, A.; Singh, H.V. Bacterial ACC deaminase: Insights into enzymology, biochemistry, genetics, and potential role in amelioration of environmental stress in crop plants. Front. Microbiol. 2023, 14, 1132770. [Google Scholar] [CrossRef]
- Hamane, S.; El Yemlahi, A.; Hassani Zerrouk, M.; El Galiou, O.; Laglaoui, A.; Bakkali, M.; Arakrak, A. Plant Growth Promotion and Biocontrol Potentiality of Endophytes Isolated from Root Nodules of Sulla flexuosa L. Plants. Int. J. Agron. 2023, 2023, 2451806. [Google Scholar] [CrossRef]
- Lu, P.; Gao, T.; Bai, R.; Yang, J.; Xu, Y.; Chu, W.; Jiang, K.; Zhang, J.; Xu, F.; Zhao, H. Regulation of carbon flux and NADH/NAD+ supply to enhance 2,3-butanediol production in Enterobacter aerogenes. J. Biotechnol. 2022, 358, 67–75. [Google Scholar] [CrossRef]
- Liu, S.; Chen, L.; Wang, L.; Zhou, B.; Ye, D.; Zheng, X.; Lin, Y.; Zeng, W.; Zhou, T.; Ye, J. Cluster Differences in Antibiotic Resistance, Biofilm Formation, Mobility, and Virulence of Clinical Enterobacter cloacae Complex. Front. Microbiol. 2022, 13, 814831. [Google Scholar] [CrossRef] [PubMed]
- Singh, S.; Singh, S.K.; Chowdhury, I.; Singh, R. Understanding the mechanism of bacterial biofilms resistance to antimicrobial agents. Open Microbiol. J. 2017, 11, 53–62. [Google Scholar] [CrossRef]
- Sachman-Ruíz, B.; Wong-Villarreal, A.; Aguilar-Marcelino, L.; Lozano-Aguirre, L.F.; Espinosa-Zaragoza, S.; Reyes-Reyes, A.L.; Sanzón-Gómez, D.; Mireles-Arriaga, A.I.; Romero-Tirado, R.; Rocha-Martínez, M.K.; et al. Nematicidal, Acaricidal and Plant Growth-Promoting Activity of Enterobacter Endophytic Strains and Identification of Genes Associated with These Biological Activities in the Genomes. Plants 2022, 11, 3136. [Google Scholar] [CrossRef] [PubMed]
- Escalante, A.; Giles-Gómez, M.; Hernández, G.; Córdova-Aguilar, M.S.; López-Munguía, A.; Gosset, G.; Bolívar, F. Analysis of bacterial community during the fermentation of pulque, a traditional Mexican alcoholic beverage, using a polyphasic approach. Int. J. Food Microbiol. 2008, 124, 126–134. [Google Scholar] [CrossRef] [PubMed]
- Escalante-Minakata, P.; Blaschek, H.P.; Barba De La Rosa, A.P.; Santos, L.; De León-Rodríguez, A. Identification of yeast and bacteria involved in the mezcal fermentation of Agave salmiana. Lett. Appl. Microbiol. 2008, 46, 626–630. [Google Scholar] [CrossRef]
- De León Rodríguez, A.; Escalante Minakata, M.d.P.; Jiménez García, M.I.; Ordoñez Acevedo, L.G.; Flores Flores, J.L.; Barba de la Rosa, A.P. Characterization of volatile compounds from ethnic Agave alcoholic beverages by gas chromatography-mass spectrometry. Food Technol. Biotechnol. 2008, 46, 448–455. [Google Scholar]
- Kirchmayr, M.R.; Segura-García, L.E.; Lappe-Oliveras, P.; Moreno-Terrazas, R.; de la Rosa, M.; Gschaedler Mathis, A. Impact of environmental conditions and process modifications on microbial diversity, fermentation efficiency and chemical profile during the fermentation of Mezcal in Oaxaca. LWT-Food Sci. Technol. 2017, 79, 160–169. [Google Scholar] [CrossRef]
- Espinoza-Martínez, V.A.; Alvarez-Gutierrez, P.E.; Palma-Cruz, F.d.J.; Enriquez-Valencia, R.; Ramírez-Lopez, M.P.; Lopez-Sanchez, C.; Vazquez-Lopez, H.G. Influence of the Biotechnological Process of Mezcal Fermentation on Yeast Diversity in Four Palenques of Oaxaca, Mexico. Beverages 2023, 9, 99. [Google Scholar] [CrossRef]
- De Man, J.C.; Rogosa, M.; Sharpe, M.E. A Medium for the cultivation of Lactobacilli. J. Appl. Bacteriol. 1960, 23, 130–135. [Google Scholar] [CrossRef]
- Rogosa, M.; Mitchelli, J.A.; Wiseman, R.F. A selective medium for the isolation and enumeration of oral and fecal lactobacilli. J. Bacteriol. 1951, 62, 132–133. [Google Scholar] [CrossRef]
- Terzaghi, B.E.; Sandine, W.E. Improved medium for lactic streptococci and their bacteriophages. Appl. Microbiol. 1975, 29, 807–813. [Google Scholar] [CrossRef]
- Benkerroum, N.; Misbah, M.; Sandine, W.E.; Elaraki, A.T. Development and use of a selective medium for isolation of Leuconostoc spp. from vegetables and dairy products. Appl. Environ. Microbiol. 1993, 59, 607. [Google Scholar] [CrossRef] [PubMed]
- Amanullah, S.; Kabir, M.; Rahman, M.; Halder, P.; Hossain, S.M.J.; Samad, M.A. Isolation, identification and biochemical characterization of lactic acid bacteria from selected yogurt samples. Bangladesh J. Livest. Res. 2021, 27, 64–72. [Google Scholar] [CrossRef]
- Ashraf, R.; Smith, S.C. Selective enumeration of dairy based strains of probiotic and lactic acid bacteria. Int. Food Res. J. 2015, 22, 2576–2586. [Google Scholar]
- Weisburg, W.G.; Barns, S.M.; Pelletier, D.A.; Lane, D.J. 16S ribosomal DNA amplification for phylogenetic study. J. Bacteriol. 1991, 173, 697–703. [Google Scholar] [CrossRef]
- Altschul, S.F.; Madden, T.L.; Schäffer, A.A.; Zhang, J.; Zhang, Z.; Miller, W.; Lipman, D.J. Gapped BLAST and PSI-BLAST: A new generation of protein database search programs. Nucleic Acids Res. 1997, 25, 3389–3402. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Higgins, D.G. Multiple Sequence Alignment Using ClustalW and ClustalX. Curr. Protoc. Bioinform. 2003, 1, 2.3.1–2.3.22. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across Computing Platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Biswas, B.B.; Basu, P.S.; Pal, M.K. Gram Staining and Its Molecular Mechanism. Int. Rev. Cytol. 1970, 29, 1–27. [Google Scholar]
- Settanni, L.; Moschetti, G. Non-starter lactic acid bacteria used to improve cheese quality and provide health benefits. Food Microbiol. 2010, 27, 691–697. [Google Scholar] [CrossRef]
- Zúñiga, M.; Pardo, I.; Ferrer, S. An improved medium for distinguishing between homofermentative and heterofermentative lactic acid bacteria. Int. J. Food Microbiol. 1993, 18, 37–42. [Google Scholar] [CrossRef] [PubMed]
- McDonald, L.C.; McFeeters, R.F.; Daeschel, M.A.; Fleming, H.P. A differential medium for the enumeration of homofermentative and heterofermentative lactic Acid bacteria. Appl. Environ. Microbiol. 1987, 53, 1382–1384. [Google Scholar] [CrossRef]
- Kandler, O.; Schillinger, U.; Weiss, N. Lactobacillus bifermentans sp. nov., nom. rev., an organism forming CO2 and H2 from lactic acid. Syst. Appl. Microbiol. 1983, 4, 408–412. [Google Scholar] [CrossRef] [PubMed]
- Rogosa, M.; Franklin, J.G.; Perry, K.D. Correlation of the vitamin requirements with cultural and biochemical characters of Lactobacillus spp. J. Gen. Microbiol. 1961, 25, 473–482. [Google Scholar] [CrossRef] [PubMed]
- Tiso, M.; Schechter, A.N. Nitrate Reduction to Nitrite, Nitric Oxide and Ammonia by Gut Bacteria under Physiological Conditions. PLoS ONE 2015, 10, e0119712. [Google Scholar]
- Nandha, M.C.; Shukla, R.M. Exploration of probiotic attributes in lactic acid bacteria isolated from fermented Theobroma cacao L. fruit using in vitro techniques. Front. Microbiol. 2023, 14, 1274636. [Google Scholar] [CrossRef]
- Jin, G.; Jiranek, V.; Hayes, A.M.; Grabin, P.R. Isolation and Characterization of High-Ethanol-Tolerance Lactic Acid Bacteria from Australian Wine. Foods 2022, 11, 1231. [Google Scholar] [CrossRef]
- Costa, M.A.S.; Cerri, B.C.; Ceccato-Antonini, S.R. Ethanol addition enhances acid treatment to eliminate Lactobacillus fermentum from the fermentation process for fuel ethanol production. Lett. Appl. Microbiol. 2018, 66, 77–85. [Google Scholar] [CrossRef]
- Yun, M.; Jo, H.E.; Kim, N.; Park, H.K.; Jang, Y.S.; Choi, G.H.; Jo, H.E.; Seo, J.H.; Mok, J.Y.; Park, S.M.; et al. Oral Administration of Alcohol-Tolerant Lactic Acid Bacteria Alleviates Blood Alcohol Concentration and Ethanol-Induced Liver Damage in Rodents. J. Microbiol. Biotechnol. 2024, 34, 838. [Google Scholar] [CrossRef]
- Instituto Nacional de Estadística y Geografía (INEGI), México. Anuario Estadístico y Geográfico por Entidad Federativa. 2023. Available online: https://www.inegi.org.mx/contenidos/productos/prod_serv/contenidos/espanol/bvinegi/productos/nueva_estruc/889463916376.pdf (accessed on 15 June 2025).
- Instituto Nacional de Estadística y Geografía (INEGI), México. Censo de Población y Vivienda 2020, Principales Resultados por Localidad. 2020. Available online: https://www.inegi.org.mx/app/areasgeograficas/#collapse-Resumen (accessed on 16 June 2025).
- Ontoy, D.S.; Ontoy, D.S.; Padua, R.N. Measuring species diversity for conservation biology: Incorporating social and ecological importance of species. Biodivers. J. 2014, 5, 387–390. [Google Scholar]
- Chao, A. Nonparametric estimation of the number of classes in a population. Scand. J. Stat. 1984, 11, 265–270. [Google Scholar]
- Pearson, K. Note on Regression and Inheritance in the Case of Two Parents. Proc. R. Soc. Lond. 1895, 58, 240–242. [Google Scholar] [CrossRef]
- Zhang, T.; Li, L.; Wang, X.F.; Zeng, Z.H.; Hu, Y.G.; Cui, Z.J. Effects of Lactobacillus buchneri and Lactobacillus plantarum on fermentation, aerobic stability, bacteria diversity and ruminal degradability of alfalfa silage. World J. Microbiol. Biotechnol. 2009, 25, 965–971. [Google Scholar] [CrossRef]
- Doijad, S.; Imirzalioglu, C.; Yao, Y.; Pati, N.B.; Falgenhauer, L.; Hain, T.; Foesel, B.U.; Abt, B.; Overmann, J.; Mirambo, M.M.; et al. Enterobacter bugandensis sp. nov., isolated from neonatal blood. Int. J. Syst. Evol. Microbiol. 2016, 66, 968–974. [Google Scholar] [CrossRef]
- Wu, W.; Feng, Y.; Zong, Z. Enterobacter sichuanensis sp. nov., recovered from human urine. Int. J. Syst. Evol. Microbiol. 2018, 68, 3922–3927. [Google Scholar] [CrossRef]
- Leroy, F.; De Vuyst, L. Lactic acid bacteria as functional starter cultures for the food fermentation industry. Trends Food Sci. Technol. 2004, 15, 67–78. [Google Scholar] [CrossRef]
- Verdugo Valdez, A.; Segura García, L.; Kirchmayr, M.; Ramírez Rodríguez, P.; González Esquinca, A.; Coria, R.; Gschaedler Mathis, A. Yeast communities associated with artisanal mezcal fermentations from Agave salmiana. Antonie Van Leeuwenhoek 2011, 100, 497–506. [Google Scholar] [CrossRef]
- Ramírez-Pérez, J.I.; Álvarez-Gutiérrez, P.E.; Luján-Hidalgo, M.C.; Ovando-Chacón, S.L.; Soria-Guerra, R.E.; Ruiz-Cabrera, M.Á.; Grajales-Lagunes, A.; Abud-Archila, M. Effect of linear and branched fructans on growth and probiotic characteristics of seven Lactobacillus spp. isolated from an autochthonous beverage from Chiapas, Mexico. Arch. Microbiol. 2022, 204, 364. [Google Scholar] [CrossRef]
- Beaumont, M. Flavouring composition prepared by fermentation with Bacillus spp. Int. J. Food Microbiol. 2002, 75, 189–196. [Google Scholar] [CrossRef]
- Sawers, R.G.; Clark, D.P. Fermentative Pyruvate and Acetyl-Coenzyme A Metabolism. EcoSal Plus 2004, 1, 10–1128. [Google Scholar] [CrossRef]
- Muñoz, R.; Rivas, B.d.L.; Rodríguez, H.; Esteban-Torres, M.; Reverón, I.; Santamaría, L.; Landete, J.M.; Plaza-Vinuesa, L.; Sánchez-Arroyo, A.; Jiménez, N.; et al. Food phenolics and Lactiplantibacillus plantarum. Int. J. Food Microbiol. 2024, 412, 110555. [Google Scholar] [CrossRef]
- Fidanza, M.; Panigrahi, P.; Kollmann, T.R. Lactiplantibacillus plantarum–Nomad and Ideal Probiotic. Front. Microbiol. 2021, 12, 712236. [Google Scholar] [CrossRef]

| Strain | Query Cover | % Ident | E-Value | Gene Bank Accession Number |
|---|---|---|---|---|
| NBRC 15535 Bacillus amyloliquefaciens T | MK182997.1 | |||
| P2OAXL11 Bacillus amyloliquefaciens | 97% | 87.91% | 0.0 | PP694281 |
| P3OAXL18 Bacillus amyloliquefaciens | 100% | 98.51% | 0.0 | PP694284 |
| P3OAXL21 Bacillus amyloliquefaciens | 99% | 93.78% | 0.0 | PP694287 |
| P4OAXL25 Bacillus amyloliquefaciens | 99% | 98.47% | 0.0 | PP694290 |
| ATCC 7061 Bacillus pumilus T | KY034389.1 | |||
| P1OAXL10 Bacillus pumilus | 93% | 78.02% | 3.00 × 10−63 | |
| P4OAXL11 Bacillus sp. * | * | * | * | PP694281 |
| P4OAXL27 Bacillus sp. * | * | * | * | |
| WCHECL1597 Enterobacter sichuanensis T | MG832788.1 | |||
| P4OAXL61 Enterobacter sichuanensis | 100% | 97.41% | 0.0 | PP694294 |
| P4OAXL62 Enterobacter sp. * | * | * | * | PP694295 |
| P4OAXL66 Enterobacter sp. * | * | * | * | PP694297 |
| ATCC19433 Enterococcus faecalis T | NR_115765.1 | |||
| P2OAXL14 Enterococcus faecalis | 100% | 99.93% | 0.0 | PP694282 |
| P3OAXL17 Enterococcus faecalis | 99% | 99.35% | 0.0 | PP694283 |
| ATCC 49573 Enterococcus gallinarum T | LC097066.1 | |||
| P4OAXL29 Enterococcus gallinarum | 100% | 99.62% | 0.0 | PP694293 |
| JCM 1149 Lactiplantibacillus pentosus T | LC064896.1 | |||
| P1OAXL1 Lactiplantibacillus pentosus | 99% | 99.42% | 0.0 | PP694274 |
| P1OAXL2 Lactiplantibacillus pentosus | 100% | 90.65% | 0.0 | PP694275 |
| P1OAXL4 Lactiplantibacillus pentosus | 99% | 99.78% | 0.0 | PP694276 |
| P1OAXL9 Lactiplantibacillus pentosus | 100% | 99.86% | 0.0 | PP694280 |
| CIP 103151 Lactiplantibacillus plantarum | MN326667.1 | |||
| P1OAXL3 Lactiplantibacillus plantarum | 90% | 91.67% | 6 × 10−53 | |
| P1OAXL5 Lactiplantibacillus plantarum * | * | * | * | |
| P1OAXL6 Lactiplantibacillus plantarum subargentoratensis | 100% | 99.50% | 0.0 | PP694278 |
| P1OAXL7 Lactiplantibacillus plantarum * | * | * | * | PP694279 |
| P1OAXL8 Lactiplantibacillus plantarum | 79% | 97.40% | 0.0 | PP694280 |
| P3OAXL19 Lactiplantibacillus plantarum * | * | * | * | PP694285 |
| P3OAXL20 Lactiplantibacillus plantarum * | * | * | * | PP694286 |
| P4OAXL22 Lactiplantibacillus plantarum * | * | * | * | PP694288 |
| P4OAXL26 Lactiplantibacillus plantarum * | * | * | * | PP694291 |
| JCM 12183 Lactiplantibacillus diolivorans T | LC311745.1 | |||
| P4OAXL65 Lactiplantibacillus diolivorans | 98% | 97.75% | 0.0 | PP694296 |
| P3OAXL15 Lactiplantibacillus sp. * | * | * | * | |
| P4OAXL64 Lacticaseibacillus sp. * | * | * | * | |
| ATCC 8293 Leuconostoc mesenteroides T | NR_074957.1 | |||
| P4OAXL24 Leuconostoc mesenteroides | 99% | 99.19% | 0.0 | PP694289 |
| ATCC 14869 Levilactobacillus brevis T | NR_116238.1 | |||
| P4OAXL28 Levilactobacillus brevis | 99% | 96.48% | 0.0 | PP694292 |
| Group | Specie | Strain | GAL a | GLU a | FRU a | MAN a | LAC a | SUC a | GAS b | NO3 c | LM d | LBS e | M17 f | ELLI f | MSE f | CAT g | GRAM h |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | Lactiplantibacillus pentosus | P1OAXL1 | 0 | 2 | 2 | 0 | 0 | 2 | 0 | 2 | 1 | 2 | 2 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus pentosus | P1OAXL2 | 2 | 2 | 2 | 0 | 0 | 2 | 0 | 2 | 1 | 2 | 0 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus plantarum | P1OAXL3 | 2 | 2 | 2 | 0 | 2 | 2 | 2 | 0 | 2 | 1 | 0 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus pentosus | P1OAXL4 | 2 | 2 | 2 | 0 | 0 | 2 | 0 | 2 | 1 | 0 | 2 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus plantarum | P1OAXL5 | 2 | 2 | 2 | 0 | 0 | 0 | 0 | 0 | 3 | 2 | 2 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus subsp. argentoratensis | P1OAXL6 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 3 | 2 | 2 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus plantarum | P1OAXL7 | 2 | 2 | 2 | 2 | 2 | 0 | 0 | 2 | 3 | 0 | 0 | 2 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus plantarum | P1OAXL8 | 2 | 2 | 2 | 0 | 2 | 0 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 1 |
| 1 | Lactiplantibacillus pentosus | P1OAXL9 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 1 | 1 | 1 | 0 | 1 | 0 | 0 | 1 |
| 1 | Bacillus pumilus | P1OAXL10 | 2 | 2 | 2 | 2 | 2 | 0 | 0 | 1 | 1 | 0 | 0 | 1 | 0 | 0 | 1 |
| 2 | Bacillus amyloliquefaciens | P2OAXL11 | 1 | 1 | 2 | 0 | 1 | 1 | 0 | 2 | 4 | 0 | 0 | 2 | 0 | 1 | 1 |
| 2 | Enterococcus faecalis | P2OAXL14 | 0 | 1 | 2 | 1 | 0 | 1 | 2 | 2 | 1 | 0 | 0 | 2 | 0 | 0 | 1 |
| 3 | Lactiplantibacillus sp. | P3OAXL15 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 2 | 1 | 0 | 2 | 2 | 0 | 0 | 1 |
| 3 | Staphylococcus sp. | P3OAXL16 | 2 | 2 | 2 | 0 | 0 | 0 | 0 | 2 | 1 | 1 | 0 | 2 | 0 | 0 | 1 |
| 3 | Enterococcus faecalis | P3OAXL17 | 1 | 1 | 2 | 2 | 0 | 2 | 2 | 0 | 2 | 1 | 0 | 2 | 0 | 0 | 1 |
| 3 | Bacillus amyloliquefaciens | P3OAXL18 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 2 | 1 | 0 | 2 | 2 | 0 | 0 | 1 |
| 3 | Lactiplantibacillus plantarum | P3OAXL19 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 0 | 3 | 2 | 2 | 2 | 0 | 0 | 1 |
| 3 | Lactiplantibacillus plantarum | P3OAXL20 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 3 | 2 | 2 | 2 | 0 | 0 | 1 |
| 3 | Bacillus amyloliquefaciens | P3OAXL21 | 0 | 1 | 2 | 0 | 0 | 2 | 0 | 2 | 3 | 0 | 0 | 2 | 0 | 0 | 1 |
| 4 | Enterobacter sichuanensis | P4OAXL61 | 1 | 1 | 2 | 0 | 2 | 1 | 0 | 2 | 1 | 2 | 2 | 2 | 0 | 0 | 1 |
| 4 | Enterobacter sp. | P4OAXL62 | 1 | 2 | 2 | 0 | 2 | 1 | 0 | 0 | 3 | 2 | 2 | 2 | 0 | 0 | 0 |
| 4 | Lacticaseibacillus sp. | P4OAXL64 | 1 | 1 | 2 | 0 | 2 | 2 | 0 | 2 | 3 | 2 | 2 | 2 | 0 | 0 | 1 |
| 4 | Lactiplantibacillus diolivorans | P4OAXL65 | 2 | 1 | 1 | 0 | 2 | 1 | 0 | 2 | 1 | 2 | 2 | 2 | 0 | 0 | 1 |
| 4 | Enterobacter sp. | P4OAXL66 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 2 | 3 | 2 | 2 | 2 | 0 | 0 | 0 |
| 4 | Bacillus sp. | P4OXL11 | 2 | 2 | 2 | 2 | 0 | 2 | 0 | 2 | 1 | 0 | 0 | 1 | 1 | 1 | 1 |
| 4 | Lactiplantibacillus plantarum | P4OAXL22 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 0 | 0 | 1 |
| 4 | Leuconostoc mesenteroides | P0AXL24 | 2 | 2 | 2 | 1 | 2 | 2 | 2 | 0 | 0 | 1 | 1 | 2 | 2 | 0 | 1 |
| 4 | Bacillus amyloliquefaciens | P4OAXL25 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 1 | 2 | 2 | 2 | 2 | 0 | 1 |
| 4 | Lactiplantibacillus plantarum | P4OAXL26 | 2 | 2 | 2 | 1 | 2 | 2 | 0 | 0 | 1 | 2 | 2 | 2 | 2 | 0 | 1 |
| 4 | Bacillus sp. | P4OAXL27 | 2 | 2 | 2 | 0 | 2 | 2 | 0 | 0 | 1 | 2 | 2 | 2 | 2 | 0 | 1 |
| 4 | Levillactobacillus brevis | P4OAXL28 | 2 | 2 | 1 | 1 | 1 | 1 | 2 | 2 | 1 | 2 | 0 | 2 | 2 | 0 | 1 |
| 4 | Enterococcus gallinarum | P4OAXL29 | 2 | 2 | 2 | 1 | 2 | 0 | 0 | 0 | 2 | 0 | 2 | 2 | 2 | 0 | 1 |
| Strain | Biomass Growth in MRS Media with Ethanol (×106 UFC/mL) | Maximum Population Relative | ||||
|---|---|---|---|---|---|---|
| 0% | 6% | 8% | 6% | 8% | ||
| P1OAXL1 | Lactiplantibacillus pentosus | 40.20 | 14.00 | 4.60 | 35 | 33 |
| P1OAXL2 | Lactiplantibacillus pentosus | 40.10 | 38.00 | 26.80 | 95 | 71 |
| P1OAXL3 | Lactiplantibacillus plantarum | 40.10 | 12.00 | 2.36 | 30 | 20 |
| P1OAXL4 | Lactiplantibacillus pentosus | 40.40 | 36.00 | 14.02 | 89 | 39 |
| P1OAXL5 | Lactiplantibacillus plantarum | 38.80 | 34.50 | 26.60 | 89 | 77 |
| P1OAXL6 | Lactiplantibacillus plantarum subsp. argentoratensis | 40.50 | 14.20 | 2.61 | 35 | 18 |
| P1OAXL7 | Lactiplantibacillus plantarum | 18.20 | 6.70 | 0.84 | 37 | 13 |
| P1OAXL8 | Lactiplantibacillus plantarum | 40.50 | 9.94 | 4.69 | 25 | 47 |
| P1OAXL9 | Lactiplantibacillus pentosus | 28.40 | 9.67 | 6.42 | 34 | 66 |
| P1OAXL10 | Bacillus pumilus | 27.70 | 11.90 | 4.26 | 43 | 36 |
| P2OAXL11 | Bacillus amyloliquefaciens | 16.10 | 7.34 | 4.26 | 46 | 58 |
| P3OAXL15 | Lactiplantibacillus sp. | 28.40 | 8.94 | 4.66 | 31 | 52 |
| P3OAXL18 | Bacillus amyloliquefaciens | 16.50 | 5.66 | 3.16 | 34 | 56 |
| P3OAXL19 | Lactiplantibacillus plantarum | 40.40 | 39.10 | 32.10 | 97 | 82 |
| P3OAXL20 | Lactiplantibacillus plantarum | 37.50 | 35.80 | 29.40 | 95 | 82 |
| P3OAXL21 | Bacillus amyloliquefaciens | 15.20 | 4.11 | 2.16 | 27 | 53 |
| P4OAXL11 | Bacillus sp. | 51.10 | 33.30 | 20.30 | 65 | 61 |
| P4OAXL64 | Lacticaseibacillus sp. | 12.70 | 3.86 | 2.15 | 30 | 56 |
| P4OAXL65 | Lactiplantibacillus diolivorans | 19.90 | 3.98 | 1.48 | 20 | 37 |
| Palenque 1 | Palenque 2 | Palenque 3 | Palenque 4 | ||
|---|---|---|---|---|---|
| Localization | Municipality | San Dionisio Ocotepec | San Pablo Etla | Concepción Buenavista | San Pedro Teozacoalco |
| District | Tlacolula | Etla | Coixtlahuaca | Nochixtlán | |
| Region | Valles Centrales | Valles Centrales | Cañada | Mixteca | |
| Location geographic | North latitude (parallels) | 16°41′39.84″–16°52′24.24″ | 17°07′21.00″–17°11′15.72″ | 17°51’35.28″–18°06’27.00″ | 16°55’49.44″–17°04’31.80″ |
| Western longitude (meridians) | 96°25′26.76″–96°11′54.24″ | 96°48′11.52″–96°39′11.88″ | 97°34’58.80″–97°21’46.44″ | 97°19’26.76″–97°13’36.48″ | |
| Altitude (m asl) | 1700–2200 | 1500–3300 | 1600–2300 | 1300–2400 | |
| Climate | Type | Semi-dry, semi-warm | Semi-warm, sub-humid, and temperate sub-humid | Semi-dry, semi-warm | Semi-warm, Asub-humid |
| Average annual temperature (°C) | 20 | 18 | 20 | 22 | |
| Average annual precipitation (mm) | 600–700 | 700–800 | 500–600 | 800–1000 | |
| Hydrography | Basin | Tehuantepec river | Atoyac river | Hondo river | Atoyac river |
| Land use and vegetation (% of territory) [41,42] | Agricultural area | 9.92 | 23.03 | 0.97 | 23.19 |
| Urban area | 1.74 | 16.87 | 0.7 | 1.03 | |
| Pine-Oak Forest | 47.9 | 54.4 | 42.16 | 59.98 | |
| Tropical deciduous forest | 32.06 | NF | 23.79 | 1.72 | |
| Matorral hierophile | NF | NF | 2.11 | NF | |
| Induced grassland [Reference] | 8.38 | 5.7 | 30.27 | 14.08 | |
| Population [41,42,43] | Total number of inhabitants | 11,411 | 17,116 | 752 | 1153 |
| Total of women | 5998 | 9083 | 377 | 578 | |
| Total of men | 5413 | 8033 | 375 | 575 | |
| Population growth % (2010–2020) | 8.68 | 10.20 | −12.70 | −9.83 | |
| Native language | Zapoteco | Zapoteco | Chocholteco | Mixteco | |
| The population in a situation of poverty (%) | 99 | 22 | 75.93 | 91.67 | |
| Marginalization index | High | Extremely low | High | High | |
| Social backwardness index | High | Extremely low | Medium | High |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
López-Sánchez, C.; Espinoza-Martínez, V.A.; Palma-Cruz, F.d.J.; Enríquez-Valencia, R.; Ramírez-López, M.P.; Ortiz-Cortés, L.Y.; Álvarez-Gutiérrez, P.E. Ethnobiotechnological Analysis of the Lactic Bacterial Diversity in the Mezcal Fermentation of Four Palenques in Oaxaca, Mexico. Fermentation 2026, 12, 89. https://doi.org/10.3390/fermentation12020089
López-Sánchez C, Espinoza-Martínez VA, Palma-Cruz FdJ, Enríquez-Valencia R, Ramírez-López MP, Ortiz-Cortés LY, Álvarez-Gutiérrez PE. Ethnobiotechnological Analysis of the Lactic Bacterial Diversity in the Mezcal Fermentation of Four Palenques in Oaxaca, Mexico. Fermentation. 2026; 12(2):89. https://doi.org/10.3390/fermentation12020089
Chicago/Turabian StyleLópez-Sánchez, Claudia, Víctor Adrián Espinoza-Martínez, Felipe de Jesús Palma-Cruz, Raúl Enríquez-Valencia, Marcos Pedro Ramírez-López, Lourdes Yaret Ortiz-Cortés, and Peggy Elizabeth Álvarez-Gutiérrez. 2026. "Ethnobiotechnological Analysis of the Lactic Bacterial Diversity in the Mezcal Fermentation of Four Palenques in Oaxaca, Mexico" Fermentation 12, no. 2: 89. https://doi.org/10.3390/fermentation12020089
APA StyleLópez-Sánchez, C., Espinoza-Martínez, V. A., Palma-Cruz, F. d. J., Enríquez-Valencia, R., Ramírez-López, M. P., Ortiz-Cortés, L. Y., & Álvarez-Gutiérrez, P. E. (2026). Ethnobiotechnological Analysis of the Lactic Bacterial Diversity in the Mezcal Fermentation of Four Palenques in Oaxaca, Mexico. Fermentation, 12(2), 89. https://doi.org/10.3390/fermentation12020089









