Isolation, Characterization, and Genomic Elucidation of HRY1: An Unconventional but Highly Efficient Phosphate-Solubilizing Escherichia coli
Abstract
1. Introduction
2. Materials and Methods
2.1. Soil Sample Collection
2.2. Isolation and Screening of Bacterial Strains
2.3. Microbiological Identification of Bacterial Strains
2.3.1. Morphological Characteristics
2.3.2. 16S rRNA Gene Sequencing
2.4. Determination of Growth Curve
2.5. Optimization of Phosphate-Solubilizing Conditions
- (1)
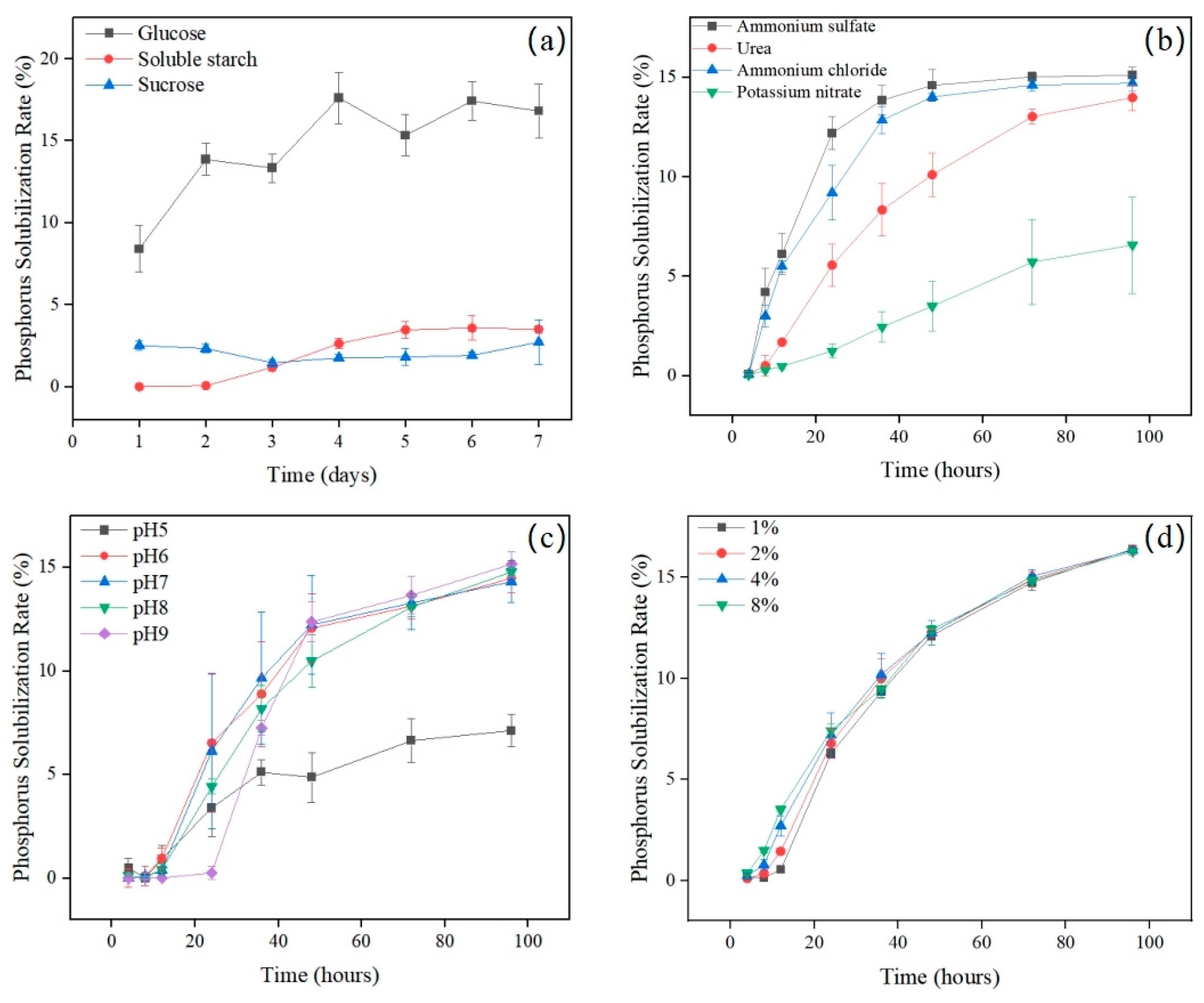
- Carbon source: glucose, soluble starch, and sucrose were tested individually at 10 g L−1;
- (2)
- Nitrogen source: (NH4)2SO4 (control), NH4Cl, urea, and KNO3 were supplied at 0.1 g L−1;
- (3)
- Initial pH: adjusted to 5.0, 6.0, 7.0, 8.0, or 9.0 using 1 M HCl or NaOH;
- (4)
- Inoculum size: 1%, 2%, 4%, or 8% (v/v) of a standardized suspension (OD600 = 1.0);
- (5)
- Ca3(PO4)2 concentration: Set at 2.5, 5.0, or 15.0 g L−1.
2.6. Whole-Genome Sequencing
2.7. Statistical Analysis
3. Results
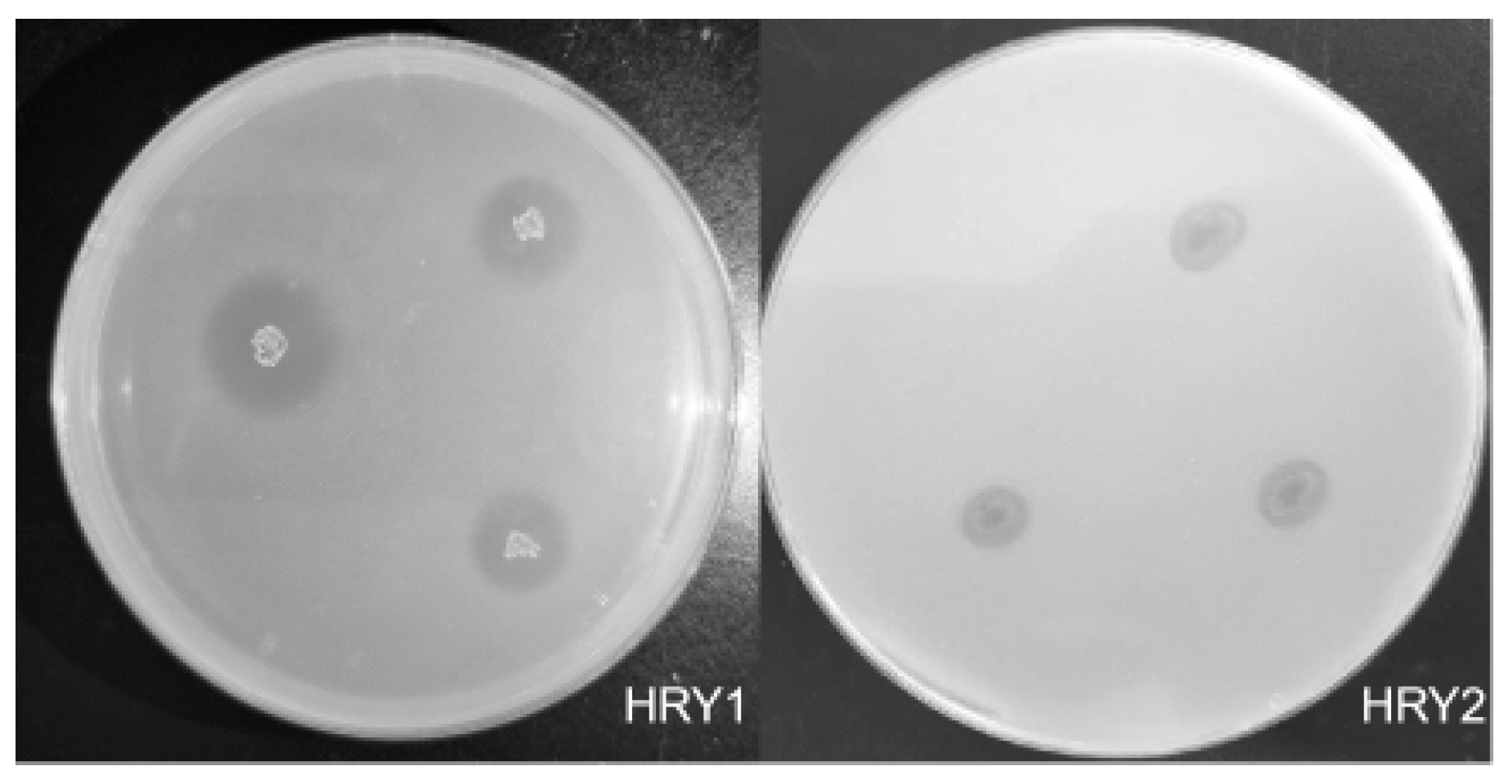
3.1. Screening and Identification of Phosphate-Solubilizing Bacteria
3.2. Bacterial Growth Curve
3.3. Optimization of Phosphate-Solubilizing Conditions
3.4. Whole-Genome Sequence Analysis of Strain HRY1
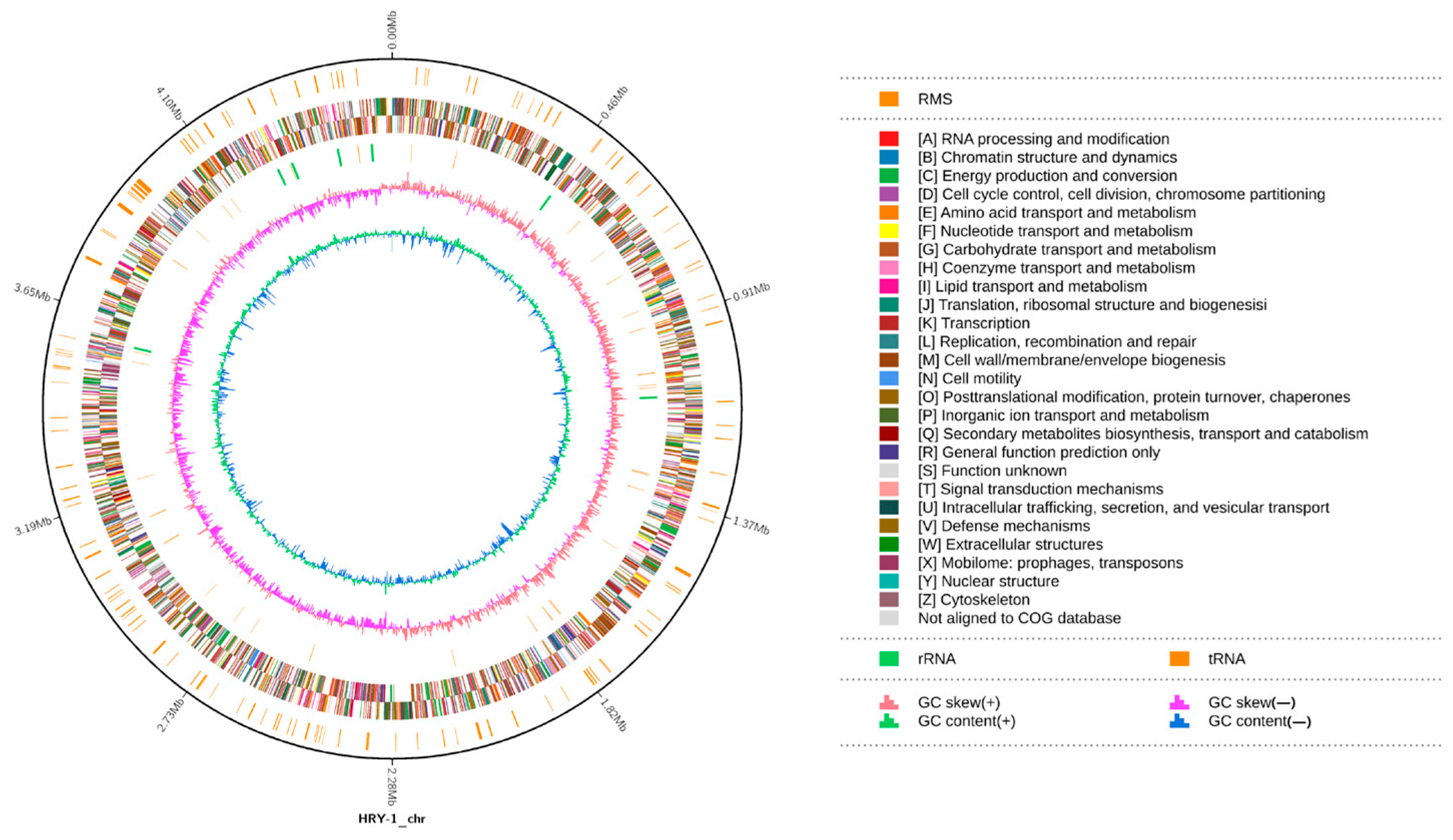
3.4.1. Basic Genomic Features
3.4.2. NR Database Annotation
3.4.3. KEGG Database Annotation and Analysis of Genes Related to Phosphorus Metabolism
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Wang, F.; Deng, M.; Xu, J.; Zhu, X.; Mao, C. Molecular mechanisms of phosphate transport and signaling in higher plants. Semin. Cell Dev. Biol. 2018, 74, 114–122. [Google Scholar] [CrossRef]
- McLaren, T.I.; Smernik, R.J.; McLaughlin, M.J.; Doolette, A.L.; Richardson, A.E.; Frossard, E. The chemical nature of soil organic phosphorus: A critical review and global compilation of quantitative data. Adv. Agron. 2020, 160, 51–124. [Google Scholar]
- Nassef, K.; Sahli, A.; Bouhdid, S.; Mezzoug, N.; Abrini, J.; Khay, E.O. Phosphate solubilizing microorganisms and their use in sustainable agriculture: A review. Geomicrobiol. J. 2025, 42, 224–243. [Google Scholar] [CrossRef]
- Yu, X.; Keitel, C.; Dijkstra, F.A. Global analysis of phosphorus fertilizer use efficiency in cereal crops. Glob. Food Secur. 2021, 29, 100545. [Google Scholar] [CrossRef]
- Joshi, S.; Gangola, S.; Jaggi, V.; Sahgal, M. Functional characterization and molecular fingerprinting of potential phosphate solubilizing bacterial candidates from Shisham rhizosphere. Sci. Rep. 2023, 13, 7003. [Google Scholar] [CrossRef] [PubMed]
- Kalayu, G. Phosphate Solubilizing Microorganisms: Promising Approach as Biofertilizers. Int. J. Agron. 2019, 2019, 4917256. [Google Scholar] [CrossRef]
- Silva, L.I.D.; Pereira, M.C.; Carvalho, A.M.X.D.; Buttrós, V.H.; Pasqual, M.; Dória, J. Phosphorus-solubilizing microorganisms: A key to sustainable agriculture. Agriculture 2023, 13, 462. [Google Scholar] [CrossRef]
- Kumar, S.; Diksha Sindhu, S.S.; Kumar, R. Harnessing phosphate-solubilizing microorganisms for mitigation of nutritional and environmental stresses, and sustainable crop production. Planta 2025, 261, 95. [Google Scholar] [CrossRef]
- Bakki, M.; Banane, B.; Marhane, O.; Esmaeel, Q.; Hatimi, A.; Barka, E.A.; Azim, K.; Bouizgarne, B. Phosphate solubilizing Pseudomonas and Bacillus combined with rock phosphates promoting tomato growth and reducing bacterial canker disease. Front. Microbiol. 2024, 15, 1289466. [Google Scholar] [CrossRef]
- Wang, Y.Z.; Jia, L.H.; Tian, G.; Dong, Y.H.; Zhang, X.; Zhou, Z.F.; Luo, X.; Li, Y.; Yao, W. shinyCircos-V2.0: Leveraging the creation of Circos plot with enhanced usability and advanced features. IMeta 2023, 2, e109. [Google Scholar] [CrossRef]
- Siles, J.A.; Starke, R.; Martinovic, T.; Fernandes, M.L.P.; Orgiazzi, A.; Bastida, F. Distribution of phosphorus cycling genes across land uses and microbial taxonomic groups based on metagenome and genome mining. Soil Biol. Biochem. 2022, 174, 108826. [Google Scholar] [CrossRef]
- Ughamba, K.T.; Ndukwe, J.K.; Lidbury, I.D.E.A.; Nnaji, N.D.; Eze, C.N.; Aduba, C.C.; Groenhof, S.; Chukwu, K.O.; Anyanwu, C.U.; Nwaiwu, O.; et al. Trends in the application of phosphate-solubilizing microbes as biofertilizers: Implications for soil improvement. Soil Syst. 2025, 9, 6. [Google Scholar] [CrossRef]
- Berza, B.; Sekar, J.; Vaiyapur, P.; Pagano, M.C.; Assefa, F. Evaluation of inorganic phosphate solubilizing efficiency and multiple plant growth promoting properties of endophytic bacteria isolated from root nodules of Erythrina brucei. BMC Microbiol. 2022, 22, 276. [Google Scholar] [CrossRef]
- Li, H.; Chen, S.; Wang, M.; Shi, S.; Zhao, W.; Xiong, G.; Zhou, J.; Qu, J. Phosphate solubilization and plant growth properties are promoted by a lactic acid bacterium in calcareous soil. Appl. Microbiol. Biotechnol. 2024, 108, 11–14. [Google Scholar] [CrossRef]
- Ma, Q.; Yi, J.; Tang, Y.; Geng, Z.; Zhang, C.; Sun, W.; Liu, Z.; Xiong, W.; Wu, H.; Xie, X. Co-utilization of carbon sources in microorganisms for the bioproduction of chemicals. Biotechnol. Adv. 2024, 73, 108380. [Google Scholar] [CrossRef]
- Tiwari, B.; Sindhu, V.; Mishra, A.K.; Singh, S.S. Carbon catabolite repression of methyl parathion degradation in a bacterial isolate characterized as a Cupriavidus sp. LMGR1. Water Air Soil Pollut. 2020, 231, 357. [Google Scholar] [CrossRef]
- Deb, S.; Lewicka-Szczebak, D.; Rohe, L. Microbial nitrogen transformations tracked by natural abundance isotope studies and microbiological methods: A review. Sci. Total Environ. 2024, 926, 172073. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Zhou, J.; Tang, S.; Wu, L.; Ma, Q.; Marsden, K.A.; Chadwick, D.R.; Jones, D.L. Utilisation and transformation of organic and inorganic nitrogen by soil microorganisms and its regulation by excessive carbon and nitrogen availability. Biol. Fertil. Soils 2023, 59, 379–389. [Google Scholar] [CrossRef]
- Scervino, J.M.; Papinutti, V.L.; Godoy, M.S.; Rodriguez, M.A.; Della Monica, I.; Recchi, M.; Pettinari, M.J. Medium pH, carbon and nitrogen concentrations modulate the phosphate solubilization efficiency of Penicillium purpurogenum through organic acid production. J. Appl. Microbiol. 2011, 110, 1215–1223. [Google Scholar] [CrossRef]
- Wang, J.; Yan, D.; Dixon, R.; Wang, Y.P. Deciphering the principles of bacterial nitrogen dietary preferences: A strategy for nutrient containment. mBio 2016, 7, e00792-16. [Google Scholar] [CrossRef]
- Sen, A.; Saha, N.; Sarkar, A.; Poddar, R.; Pramanik, K.; Samanta, A. Assessing the effectiveness of indigenous phosphate-solubilizing bacteria in mitigating phosphorus fixation in acid soils. 3 Biotech 2024, 14, 197. [Google Scholar] [CrossRef] [PubMed]
- Pereira, S.I.A.; Sousa, A.S.S.; Felpeto, A.B.; Miguel, A.; Pinto, J.C.; Castro, P.M.L.; Moreira, H. Lower inoculum sizes of phosphorus-solubilizing bacteria benefit maize growth and soil enzyme activity in P-fertilized soils. Symbiosis 2025, 96, 261–269. [Google Scholar] [CrossRef]
- Dai, Z.; Liu, G.; Chen, H.; Chen, C.; Wang, J.; Ai, S.; Wei, D.; Li, D.; Ma, B.; Tang, C.; et al. Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems. ISME J. 2020, 14, 757–770. [Google Scholar] [CrossRef]
- Ahash, S.; Manikandan, K.; Devi, T.S.; Elamathi, S.; Maragatham, S.; Subrahmaniyan, K. Phosphate-solubilizing microorganisms for sustainable phosphorus management in rice. Rhizosphere 2025, 34, 101096. [Google Scholar] [CrossRef]
- Adhikari, P.; Pandey, A. Phosphate solubilization potential of endophytic fungi isolated from Taxus wallichiana Zucc. roots. Rhizosphere 2019, 9, 2–9. [Google Scholar] [CrossRef]
- McKay Fletcher, D.M.; Ruiz, S.; Dias, T.; Petroselli, C.; Roose, T. Linking root structure to functionality: The impact of root system architecture on citrate-enhanced phosphate uptake. New Phytol. 2020, 227, 376–391. [Google Scholar] [CrossRef]
- Guo, D.J.; Yang, G.R.; Singh, P.; Wang, J.J.; Lan, X.M.; Singh, R.K.; Guo, J.; Dong, Y.D.; Li, D.P.; Yang, B. Comprehensive analysis of the physiological and molecular responses of phosphate-solubilizing bacterium Burkholderia gladioli DJB4-8 in promoting maize growth. Front. Plant Sci. 2025, 16, 1611674. [Google Scholar] [CrossRef]
- An, R.; Moe, L.A. Regulation of pyrroloquinoline quinone-dependent glucose dehydrogenase activity in the model rhizosphere-dwelling bacterium Pseudomonas putida KT2440. Appl. Environ. Microbiol. 2016, 82, 4955–4964. [Google Scholar] [CrossRef]
- Pang, F.; Li, Q.; Solanki, M.K.; Wang, Z.; Xing, Y.X.; Dong, D.F. Soil phosphorus transformation and plant uptake driven by phosphate-solubilizing microorganisms. Front. Microbiol. 2024, 15, 1383813. [Google Scholar] [CrossRef]
- Peng, Y.; Huang, Z.; Bian, F.; Wu, Z.; Zhong, Z.; Zhang, X. Characteristics of microbial phosphorus cycling in Phyllostachys praecox forest soils supplemented with the phosphate-solubilizing bacterium Pseudomonas sp. OP1 revealed by metagenomics. J. Environ. Manag. 2025, 394, 127363. [Google Scholar] [CrossRef]
- Li, X.L.; Lv, X.Y.; Ji, J.B.; Wang, W.D.; Wang, J.; Wang, C.; He, H.B.; Ben, A.L.; Liu, T.L. Complete genome sequence of Nguyenibacter sp. L1, a phosphate solubilizing bacterium isolated from Lespedeza bicolor rhizosphere. Front. Microbiol. 2023, 14, 1257442. [Google Scholar] [CrossRef] [PubMed]
- Cui, S.; Wang, Z.; Kong, L.; Zhang, M.; Xu, H.; Xiao, P.; Tao, Y. Dynamics of microbial communities and functional genes under the interactions between phosphorus inactivation agents and phosphorus-solubilizing bacteria in sediments. J. Water Process Eng. 2025, 77, 108395. [Google Scholar] [CrossRef]
- Oliverio, A.M.; Bissett, A.; McGuire, K.; Saltonstall, K.; Turner, B.L.; Fierer, N. The role of hosphorus limitation in shaping soil bacterial communities and their metabolic capabilities. mBio 2020, 11, e01718-20. [Google Scholar] [CrossRef] [PubMed]
- Khan, A.A.; Chen, Y.; Asghar, R.M.A.; Azeem, I.; Shah, T.; Cao, W.; Zhang, D.; Gao, Y. P-solubilising bacteria and P-cycling genes drive soil P transformation during green manure decomposition. Agric. Ecosyst. Environ. 2025, 394, 109875. [Google Scholar] [CrossRef]





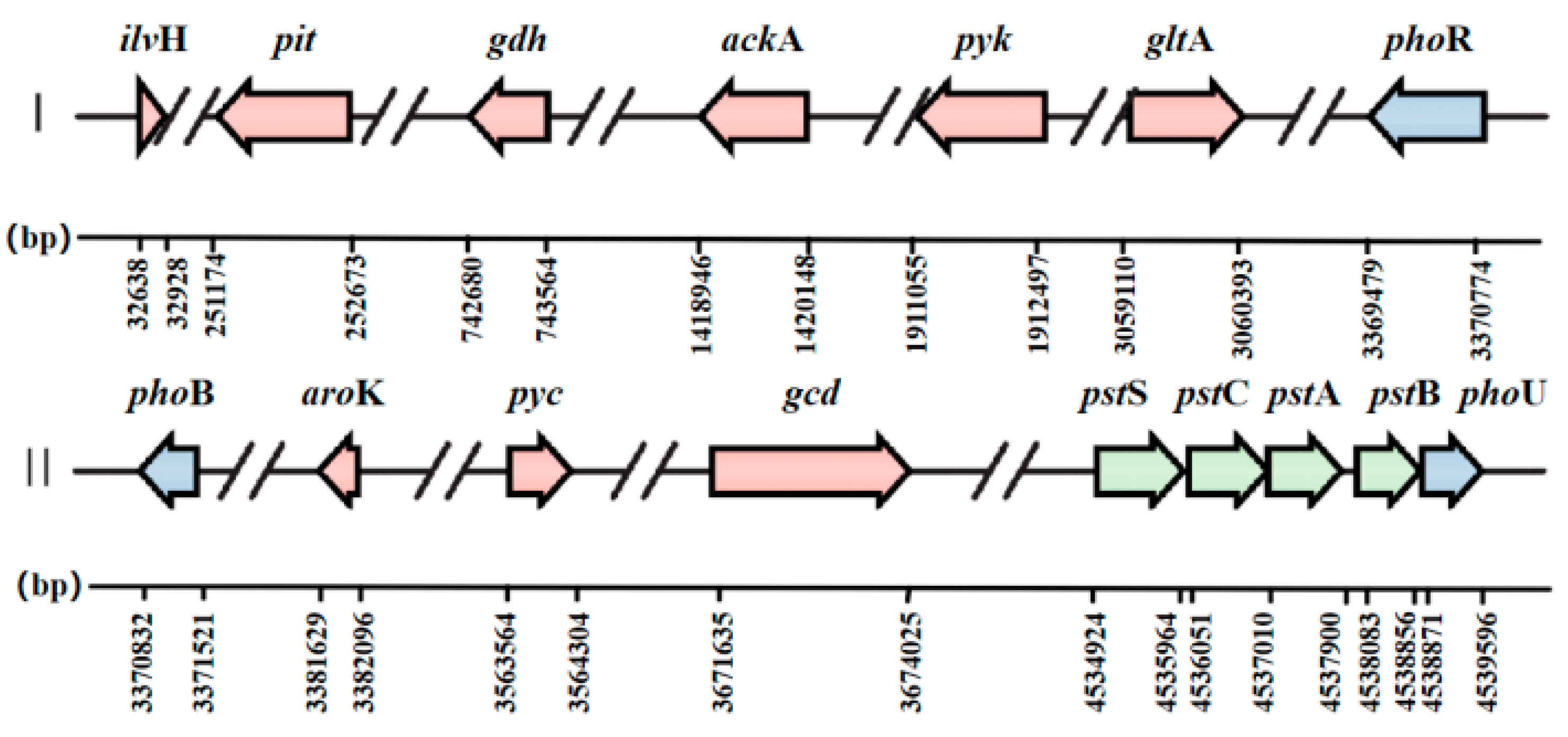
| Functional Group | Gene_ID | Gene | KEGG Annotation Function Description |
|---|---|---|---|
| Inorganic phosphorus decomposition | orf03540 | gcd | quinoprotein glucose dehydrogenase |
| orf01360 | ackA | acetate kinase | |
| orf00334 | aroK | shikimate kinase | |
| orf02944 | gltA | citrate synthase | |
| orf03587 | ilvH | acetolactate synthase I small subunit | |
| orf00717 | gdh | glucose 1-dehydrogenase | |
| orf03437 | pyc | pyruvate carboxylase | |
| orf02011 | pyk | pyruvate kinase | |
| Inorganic phosphate transport | orf00229 | pit | inorganic phosphate transporter |
| orf04331 | pstA | phosphate transport system permease protein | |
| orf04332 | pstB | phosphate transport system ATP-binding protein | |
| orf04330 | pstC | phosphate transport system permease protein | |
| orf04329 | pstS | phosphate transport system substrate-binding protein | |
| Phosphorus deficiency response regulation | orf03236 | phoB | phosphate regulon response regulator |
| orf03235 | phoR | phosphate regulon sensor histidine kinase | |
| orf04333 | phoU | Negative regulatory proteins of the PhoR/PhoB two-component regulators |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
He, R.; Zhang, S.; Liu, Y. Isolation, Characterization, and Genomic Elucidation of HRY1: An Unconventional but Highly Efficient Phosphate-Solubilizing Escherichia coli. Fermentation 2026, 12, 113. https://doi.org/10.3390/fermentation12020113
He R, Zhang S, Liu Y. Isolation, Characterization, and Genomic Elucidation of HRY1: An Unconventional but Highly Efficient Phosphate-Solubilizing Escherichia coli. Fermentation. 2026; 12(2):113. https://doi.org/10.3390/fermentation12020113
Chicago/Turabian StyleHe, Ruiyan, Sa Zhang, and Yuanwang Liu. 2026. "Isolation, Characterization, and Genomic Elucidation of HRY1: An Unconventional but Highly Efficient Phosphate-Solubilizing Escherichia coli" Fermentation 12, no. 2: 113. https://doi.org/10.3390/fermentation12020113
APA StyleHe, R., Zhang, S., & Liu, Y. (2026). Isolation, Characterization, and Genomic Elucidation of HRY1: An Unconventional but Highly Efficient Phosphate-Solubilizing Escherichia coli. Fermentation, 12(2), 113. https://doi.org/10.3390/fermentation12020113






