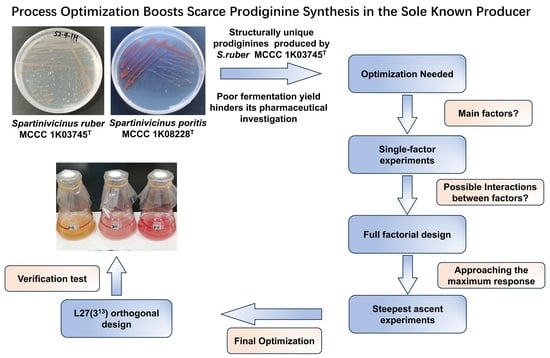
Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T
Abstract
1. Introduction
2. Materials and Methods
2.1. Reagents
2.2. Single-Factor Experiments
2.3. Factorial Design and Steepest Ascent Experiments
2.4. Orthogonal Design
2.5. Quantification of Prodiginines
2.6. Statistical Analysis
3. Results
3.1. Effects of Single Factors on Fermentation Titer
3.2. Analysis of Main Effects and Interactions by Full Factorial Design
3.3. Process Optimization via Sequential Steepest Ascent Experiments
3.4. Optimization of Prodiginine Production Medium by Orthogonal Design
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
Appendix A.1
| Batch 1 | Batch 2 | Batch 3 | |
|---|---|---|---|
| TN (mg/L) | 1.37 ± 0.11 | 0.64 ± 0.04 | 1.10 ± 0.01 |
| TP (mg/L) | 0.040 ± 0.005 | 0.023 ± 0.003 | 0.044 ± 0.003 |
| TOC (mg/L) | 1.8 ± 0.1 | 1.3 ± 0.1 | 1.5 ± 0.1 |
| Salinity (‰) | 20.4 ± 0.2 | 28.4 ± 0.3 | 23.5 ± 0.3 |
| pH | 6.56 ± 0.04 | 6.37 ± 0.03 | 7.01 ± 0.04 |
Appendix A.2
| Component | Amount (/L) | Grade | Supplier | Unit Price (CNY/g or mL) | Quotation Date | Cost (CNY/L) |
|---|---|---|---|---|---|---|
| Peptone | 11 g | - | Solarbio | 0.400 | October 2025 | 4.40 |
| Yeast extract | 1 g | - | Oxoid | 0.270 | October 2025 | 0.27 |
| Soybean oil | 5 mL | Food | Yihai Kerry | 0.015 | October 2025 | 0.08 |
| MgCl2·6H2O | 3 g | Analytical | Sinopharm | 0.068 | October 2025 | 0.20 |
| total | 4.95 |
References
- Williamson, N.R.; Fineran, P.C.; Leeper, F.J.; Salmond, G.P. The biosynthesis and regulation of bacterial prodiginines. Nat. Rev. Microbiol. 2006, 4, 887–899. [Google Scholar] [CrossRef]
- You, Z.; Zhang, S.; Liu, X.; Zhang, J.; Wang, Y.; Peng, Y.; Wu, W. Insights into the anti-infective properties of prodiginines. Appl. Microbiol. Biotechnol. 2019, 103, 2873–2887. [Google Scholar] [CrossRef]
- Llagostera, E.; Soto-Cerrato, V.; Joshi, R.; Montaner, B.; Gimenez-Bonafe, P.; Perez-Tomas, R. High cytotoxic sensitivity of the human small cell lung doxorubicin-resistant carcinoma (GLC4/ADR) cell line to prodigiosin through apoptosis activation. Anticancer. Drugs 2005, 16, 393–399. [Google Scholar] [CrossRef]
- Soto-Cerrato, V.; Llagostera, E.; Montaner, B.; Scheffer, G.L.; Perez-Tomas, R. Mitochondria-mediated apoptosis operating irrespective of multidrug resistance in breast cancer cells by the anticancer agent prodigiosin. Biochem. Pharmacol. 2004, 68, 1345–1352. [Google Scholar] [CrossRef]
- Elahian, F.; Moghimi, B.; Dinmohammadi, F.; Ghamghami, M.; Hamidi, M.; Mirzaei, S.A. The anticancer agent prodigiosin is not a multidrug resistance protein substrate. DNA Cell Biol. 2013, 32, 90–97. [Google Scholar] [CrossRef] [PubMed]
- Huang, Z.; Dong, L.; Lai, Q.; Liu, J. Spartinivicinus ruber gen. nov., sp. nov., a Novel Marine Gammaproteobacterium Producing Heptylprodigiosin and Cycloheptylprodigiosin as Major Red Pigments. Front. Microbiol. 2020, 11, 2056. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.Y.; Chen, N.F.; Kuo, H.M.; Yang, S.N.; Sung, C.S.; Sung, P.J.; Wen, Z.H.; Chen, W.F. Prodigiosin stimulates endoplasmic reticulum stress and induces autophagic cell death in glioblastoma cells. Apoptosis 2018, 23, 314–328. [Google Scholar] [CrossRef]
- Wang, J.; Liu, H.; Zhu, L.; Wang, J.; Luo, X.; Liu, W.; Ma, Y. Prodigiosin from Serratia Marcescens in Cockroach Inhibits the Proliferation of Hepatocellular Carcinoma Cells through Endoplasmic Reticulum Stress-Induced Apoptosis. Molecules 2022, 27, 7281. [Google Scholar] [CrossRef]
- Lin, X.; Dong, L.; Miao, Q.; Huang, Z.; Wang, F. Cycloheptylprodigiosin from marine bacterium Spartinivicinus ruber MCCC 1K03745T induces a novel form of cell death characterized by Golgi disruption and enhanced secretion of cathepsin D in non-small cell lung cancer cell lines. Eur. J. Pharmacol. 2024, 974, 176608. [Google Scholar] [CrossRef] [PubMed]
- Furstner, A. Chemistry and biology of roseophilin and the prodigiosin alkaloids: A survey of the last 2500 years. Angew. Chem. Int. Ed. Engl. 2003, 42, 3582–3603. [Google Scholar] [CrossRef]
- Lapenda, J.C.; Silva, P.A.; Vicalvi, M.C.; Sena, K.X.; Nascimento, S.C. Antimicrobial activity of prodigiosin isolated from Serratia marcescens UFPEDA 398. World J. Microbiol. Biotechnol. 2015, 31, 399–406. [Google Scholar] [CrossRef]
- Kimyon, O.; Das, T.; Ibugo, A.I.; Kutty, S.K.; Ho, K.K.; Tebben, J.; Kumar, N.; Manefield, M. Serratia Secondary Metabolite Prodigiosin Inhibits Pseudomonas aeruginosa Biofilm Development by Producing Reactive Oxygen Species that Damage Biological Molecules. Front. Microbiol. 2016, 7, 972. [Google Scholar] [CrossRef]
- Montaner, B.; Navarro, S.; Pique, M.; Vilaseca, M.; Martinell, M.; Giralt, E.; Gil, J.; Perez-Tomas, R. Prodigiosin from the supernatant of Serratia marcescens induces apoptosis in haematopoietic cancer cell lines. Br. J. Pharmacol. 2000, 131, 585–593. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.-C.; Yu, W.-J.; Chang, C.-C.; Chang, J.-S.; Huang, S.-H.; Chang, C.-H.; Chen, S.-Y.; Chien, C.-C.; Yao, C.-L.; Chen, W.-M.; et al. Enhancing production of prodigiosin from Serratia marcescens C3 by statistical experimental design and porous carrier addition strategy. Biochem. Eng. J. 2013, 78, 93–100. [Google Scholar] [CrossRef]
- Li, X.; Zhang, G.; Zhu, T.; Li, D.; Gu, Q. Strain and Culture Medium Optimization for Production Enhancement of Prodiginines from Marine-Derived Streptomyces sp. GQQ-10. J. Ocean Univ. China 2012, 11, 361–365. [Google Scholar] [CrossRef]
- Kim, S.J.; Lee, H.K.; Lee, Y.K.; Yim, J.H. Mutant selection of Hahella chejuensis KCTC 2396 and statistical optimization of medium components for prodigiosin yield-up. J. Microbiol. 2008, 46, 183–188. [Google Scholar] [CrossRef]
- Su, W.-T.; Tsou, T.-Y.; Liu, H.-L. Response surface optimization of microbial prodigiosin production from Serratia marcescens. J. Taiwan Inst. Chem. E 2011, 42, 217–222. [Google Scholar] [CrossRef]
- Giri, A.V.; Anandkumar, N.; Muthukumaran, G.; Pennathur, G. A novel medium for the enhanced cell growth and production of prodigiosin from Serratia marcescens isolated from soil. BMC Microbiol. 2004, 4, 11. [Google Scholar] [CrossRef]
- Lin, C.; Jia, X.; Fang, Y.; Chen, L.; Zhang, H.; Lin, R.; Chen, J. Enhanced production of prodigiosin by Serratia marcescens FZSF02 in the form of pigment pellets. Electron. J. Biotechnol. 2019, 40, 58–64. [Google Scholar] [CrossRef]
- Fiorentini, D.; Cappadone, C.; Farruggia, G.; Prata, C. Magnesium: Biochemistry, Nutrition, Detection, and Social Impact of Diseases Linked to Its Deficiency. Nutrients 2021, 13, 1136. [Google Scholar] [CrossRef]
- Waldron, K.J.; Rutherford, J.C.; Ford, D.; Robinson, N.J. Metalloproteins and metal sensing. Nature 2009, 460, 823–830. [Google Scholar] [CrossRef]
- Dong, L.; Fu, J.; Li, S.; Wang, F.; Lin, X.; Huang, Z. The Optimization of Fermentation Medium Formulation for Six Hydrogen Cycloheptane Prodigiosin (ShcPG) Production from Spartinivicinus ruber Strain S2-4-1HT in Shake-Flask Fermentation. J. Quanzhou Norm. Univ. 2023, 41, 9–18. [Google Scholar]
- Fu, J.; Li, S.; Dong, L.; Lin, X.; Huang, Z.; Wang, F. Optimization of Condition Parameter for Six Hydrogen Cycloheptane Prodigiosin (ShcPG) Production from Spartinivicinus ruber Strain S2-4-1HT in Shake-Flask Fermentation. Food Ind. 2024, 45, 39–44. [Google Scholar]
- Xia, Y.; Wang, G.; Lin, X.; Song, X.; Ai, L. Solid-state fermentation with Serratia marcescens Xd-1 enhanced production of prodigiosin by using bagasse as an inertia matrix. Ann. Microbiol. 2016, 66, 1239–1247. [Google Scholar] [CrossRef]
- de Araujo, H.W.; Fukushima, K.; Takaki, G.M. Prodigiosin production by Serratia marcescens UCP 1549 using renewable-resources as a low cost substrate. Molecules 2010, 15, 6931–6940. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Lee, J.S.; Park, Y.K.; Kim, J.F.; Jeong, H.; Oh, T.K.; Kim, B.S.; Lee, C.H. Biosynthesis of antibiotic prodiginines in the marine bacterium Hahella chejuensis KCTC 2396. J. Appl. Microbiol. 2007, 102, 937–944. [Google Scholar] [CrossRef]
- Lee, J.S.; Kim, Y.S.; Park, S.; Kim, J.; Kang, S.J.; Lee, M.H.; Ryu, S.; Choi, J.M.; Oh, T.K.; Yoon, J.H. Exceptional production of both prodigiosin and cycloprodigiosin as major metabolic constituents by a novel marine bacterium, Zooshikella rubidus S1-1. Appl. Environ. Microbiol. 2011, 77, 4967–4973. [Google Scholar] [CrossRef]
- HJ 668-2013; Water Quality-Determination of Total Nitrogen by Flow Injection Analysis (FIA) and N-(1-naphthyl)ethylenediamine Dihydrochloride Spectrophotometry. China Environment Publishing Group Co., Ltd.: Beijing, China, 2013.
- HJ 671-2013; Water Quality-Determination of Total Phosphorus-Flow Injection Analysis (FIA) and Ammonium Molybdate Spectrophotometry. China Environment Publishing Group Co., Ltd.: Beijing, China, 2013.
- HJ 501-2009; Water Quality-Determination of Total Organic Carbon-Combustion Oxidation Nondispersive Infrared Absorption Method. China Environmental Science Press: Beijing, China, 2009.





| Factors | Low Level (−1) | High Level (+1) |
|---|---|---|
| A-soybean oil | 0 mL/L | 3.333 mL/L |
| B-peptone | 7 g/L | 11 g/L |
| C-MgCl2·6H2O | 2 g/L | 4 g/L |
| D-yeast extract | 1 g/L | 2 g/L |
| Run | A- Soybean Oil | B- Peptone | C- MgCl2·6H2O | D- Yeast Extract | Prodiginine Concentration (mg/L) |
|---|---|---|---|---|---|
| 1 | +1 | +1 | +1 | −1 | 15.22 |
| 2 | −1 | −1 | −1 | −1 | 4.15 |
| 3 | +1 | −1 | +1 | −1 | 8.55 |
| 4 | −1 | +1 | +1 | −1 | 6.52 |
| 5 | +1 | −1 | −1 | −1 | 10.14 |
| 6 | −1 | +1 | −1 | −1 | 6.09 |
| 7 | +1 | +1 | −1 | +1 | 12.37 |
| 8 | +1 | +1 | −1 | −1 | 14.06 |
| 9 | −1 | −1 | −1 | +1 | 3.57 |
| 10 | +1 | +1 | +1 | +1 | 10.05 |
| 11 | −1 | +1 | −1 | +1 | 6.14 |
| 12 | −1 | +1 | +1 | +1 | 4.40 |
| 13 | +1 | −1 | +1 | +1 | 7.58 |
| 14 | −1 | −1 | +1 | −1 | 4.69 |
| 15 | −1 | −1 | +1 | +1 | 4.25 |
| 16 | +1 | −1 | −1 | +1 | 7.29 |
| Step | A-Soybean Oil (mL/L) | B-Peptone (g/L) | D-Yeast Extract (g/L) |
|---|---|---|---|
| −1 | 0 | 8.00 | 1.667 |
| 0 | 1.667 | 9.00 | 1.500 |
| 1 | 3.333 | 10.00 | 1.333 |
| 2 | 5.000 | 11.00 | 1.167 |
| 3 | 6.667 | 12.00 | 1.000 |
| 4 | 8.333 | 13.00 | 0.833 |
| Step | A-Soybean Oil (mL/L) | B-Peptone (g/L) | D-Yeast Extract (g/L) |
|---|---|---|---|
| 2 | 5.000 | 11.000 | 1.167 |
| 2.5 | 5.833 | 11.500 | 1.083 |
| 3 | 6.667 | 12.000 | 1.000 |
| 3.5 | 7.500 | 12.500 | 0.917 |
| 4 | 8.333 | 13.000 | 0.833 |
| Level | Factors | ||
|---|---|---|---|
| A-Peptone (g/L) | B-Yeast Extract (g/L) | C-Soybean Oil (mL/L) | |
| 1 | 11.0 | 1.167 | 5.000 |
| 2 | 11.5 | 1.083 | 5.833 |
| 3 | 12.0 | 1.000 | 6.667 |
| Run | Factors | Prodiginine Concentration (mg/L) | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| A | B | A × B | A × B | C | A × C | A × C | B × C | Blank | Blank | B × C | Blank | Blank | ||
| 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 1 | 14.82 |
| 2 | 1 | 1 | 1 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 15.14 |
| 3 | 1 | 1 | 1 | 1 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 13.10 |
| 4 | 1 | 2 | 2 | 2 | 1 | 1 | 1 | 2 | 2 | 2 | 3 | 3 | 3 | 15.25 |
| 5 | 1 | 2 | 2 | 2 | 2 | 2 | 2 | 3 | 3 | 3 | 1 | 1 | 1 | 15.20 |
| 6 | 1 | 2 | 2 | 2 | 3 | 3 | 3 | 1 | 1 | 1 | 2 | 2 | 2 | 15.68 |
| 7 | 1 | 3 | 3 | 3 | 1 | 1 | 1 | 3 | 3 | 3 | 2 | 2 | 2 | 15.46 |
| 8 | 1 | 3 | 3 | 3 | 2 | 2 | 2 | 1 | 1 | 1 | 3 | 3 | 3 | 16.40 |
| 9 | 1 | 3 | 3 | 3 | 3 | 3 | 3 | 2 | 2 | 2 | 1 | 1 | 1 | 15.27 |
| 10 | 2 | 1 | 2 | 3 | 1 | 2 | 3 | 1 | 2 | 3 | 1 | 2 | 3 | 14.49 |
| 11 | 2 | 1 | 2 | 3 | 2 | 3 | 1 | 2 | 3 | 1 | 2 | 3 | 1 | 12.16 |
| 12 | 2 | 1 | 2 | 3 | 3 | 1 | 2 | 3 | 1 | 2 | 3 | 1 | 2 | 11.90 |
| 13 | 2 | 2 | 3 | 1 | 1 | 2 | 3 | 2 | 3 | 1 | 3 | 1 | 2 | 15.26 |
| 14 | 2 | 2 | 3 | 1 | 2 | 3 | 1 | 3 | 1 | 2 | 1 | 2 | 3 | 15.15 |
| 15 | 2 | 2 | 3 | 1 | 3 | 1 | 2 | 1 | 2 | 3 | 2 | 3 | 1 | 14.22 |
| 16 | 2 | 3 | 1 | 2 | 1 | 2 | 3 | 3 | 1 | 2 | 2 | 3 | 1 | 13.75 |
| 17 | 2 | 3 | 1 | 2 | 2 | 3 | 1 | 1 | 2 | 3 | 3 | 1 | 2 | 15.38 |
| 18 | 2 | 3 | 1 | 2 | 3 | 1 | 2 | 2 | 3 | 1 | 1 | 2 | 3 | 15.56 |
| 19 | 3 | 1 | 3 | 2 | 1 | 3 | 2 | 1 | 3 | 2 | 1 | 3 | 2 | 10.54 |
| 20 | 3 | 1 | 3 | 2 | 2 | 1 | 3 | 2 | 1 | 3 | 2 | 1 | 3 | 9.60 |
| 21 | 3 | 1 | 3 | 2 | 3 | 2 | 1 | 3 | 2 | 1 | 3 | 2 | 1 | 11.55 |
| 22 | 3 | 2 | 1 | 3 | 1 | 3 | 2 | 2 | 1 | 3 | 3 | 2 | 1 | 12.57 |
| 23 | 3 | 2 | 1 | 3 | 2 | 1 | 3 | 3 | 2 | 1 | 1 | 3 | 2 | 11.90 |
| 24 | 3 | 2 | 1 | 3 | 3 | 2 | 1 | 1 | 3 | 2 | 2 | 1 | 3 | 10.42 |
| 25 | 3 | 3 | 2 | 1 | 1 | 3 | 2 | 3 | 2 | 1 | 2 | 1 | 3 | 14.39 |
| 26 | 3 | 3 | 2 | 1 | 2 | 1 | 3 | 1 | 3 | 2 | 3 | 2 | 1 | 13.26 |
| 27 | 3 | 3 | 2 | 1 | 3 | 2 | 1 | 2 | 1 | 3 | 1 | 3 | 2 | 12.79 |
| Source | Sum of Squares | df | Mean Square | F-Value | p-Value | |
|---|---|---|---|---|---|---|
| Model | 190.86 a | 5 | 38.17 | 48.02 | <0.0001 | significant |
| A-soybean oil | 129.16 | 1 | 129.16 | 162.48 | <0.0001 | significant |
| B-peptone | 37.79 | 1 | 37.79 | 47.54 | <0.0001 | significant |
| D-yeast extract | 11.85 | 1 | 11.85 | 14.90 | 0.0032 | significant |
| AB | 8.47 | 1 | 8.47 | 10.66 | 0.0085 | significant |
| AD | 3.60 | 1 | 3.60 | 4.52 | 0.0593 | |
| Residual | 7.95 | 10 | 0.7949 | |||
| Cor Total | 198.81 | 15 |
| Source | Type III Sum of Square | df | Mean Square | F Statistic | Significance |
|---|---|---|---|---|---|
| Corrected Model | 73.200 a | 6 | 12.200 | 14.056 | <0.001 |
| Intercept | 5103.525 | 1 | 5103.525 | 5880.081 | <0.001 |
| A-peptone | 50.530 | 2 | 25.265 | 29.110 | <0.001 |
| B-yeast extract | 20.606 | 2 | 10.303 | 11.871 | <0.001 |
| C-soybean oil | 2.064 | 2 | 1.032 | 1.189 | 0.325 |
| Error | 17.359 | 20 | 0.868 | ||
| Total | 5194.084 | 27 | |||
| Corrected Total | 90.559 | 26 |
| Level | N | Subset | ||
|---|---|---|---|---|
| 1 | 2 | 3 | ||
| 3 | 9 | 11.8911 | ||
| 2 | 9 | 14.2080 | ||
| 1 | 9 | 15.1462 | ||
| Level | N | Subset | |
|---|---|---|---|
| 1 | 2 | ||
| 1 | 9 | 12.5883 | |
| 2 | 9 | 13.9606 | |
| 3 | 9 | 14.6964 | |
| Level | N | Subset |
|---|---|---|
| 1 | ||
| 3 | 9 | 13.3872 |
| 2 | 9 | 13.7995 |
| 1 | 9 | 14.0586 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, X.; Wu, P.; Huang, Y.; Dai, C. Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T. Fermentation 2025, 11, 629. https://doi.org/10.3390/fermentation11110629
Lin X, Wu P, Huang Y, Dai C. Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T. Fermentation. 2025; 11(11):629. https://doi.org/10.3390/fermentation11110629
Chicago/Turabian StyleLin, Xiaosi, Peiyun Wu, Yajue Huang, and Congjie Dai. 2025. "Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T" Fermentation 11, no. 11: 629. https://doi.org/10.3390/fermentation11110629
APA StyleLin, X., Wu, P., Huang, Y., & Dai, C. (2025). Optimization of Fermentation Parameters and Medium Composition for Producing Prodiginines from Marine Bacterium Spartinivicinus ruber MCCC 1K03745T. Fermentation, 11(11), 629. https://doi.org/10.3390/fermentation11110629