Multifunctional miR-155 Pathway in Avian Oncogenic Virus-Induced Neoplastic Diseases
Abstract
1. Introduction
2. MicroRNA-155
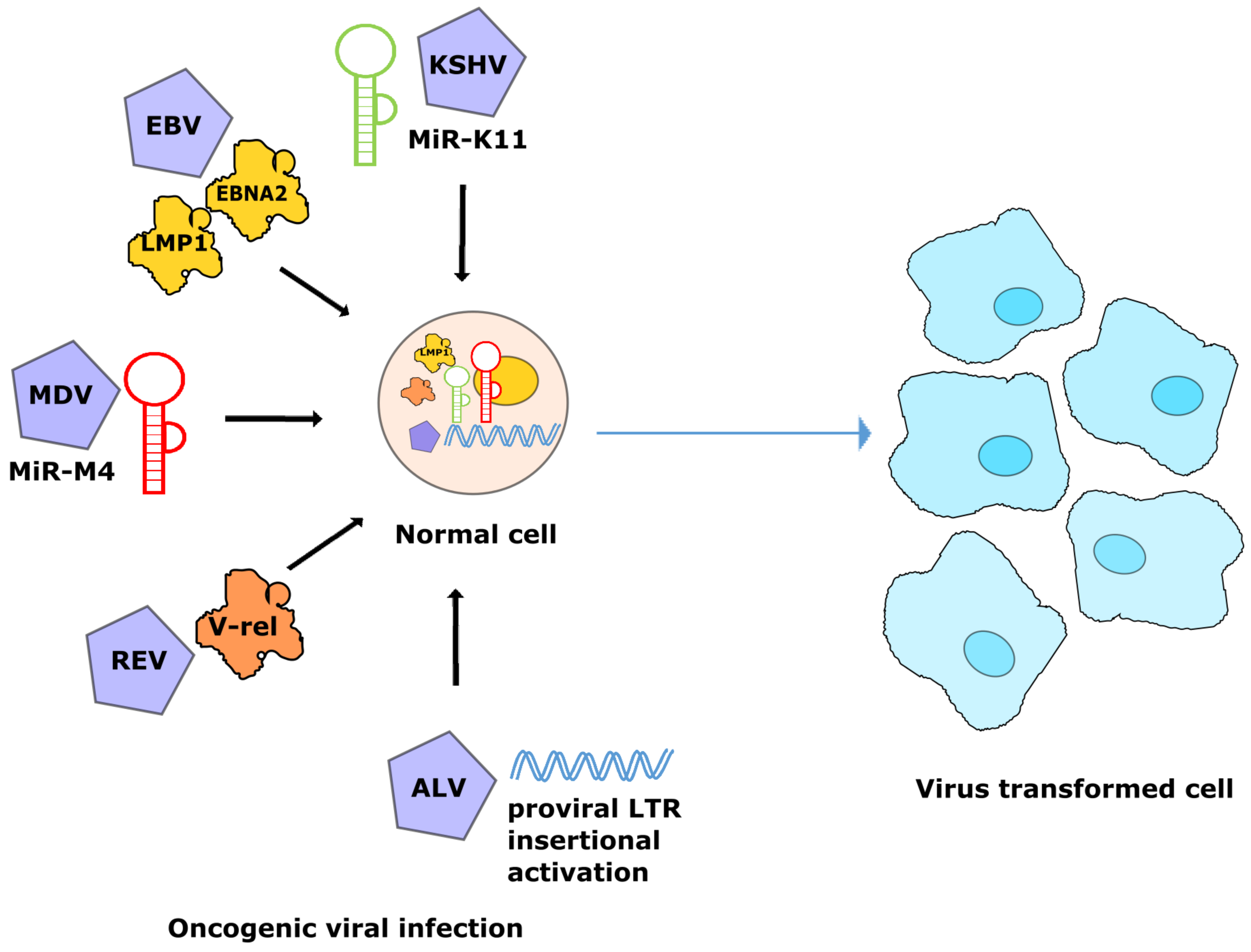
3. Modulation of miR-155 by Oncogenic Viruses
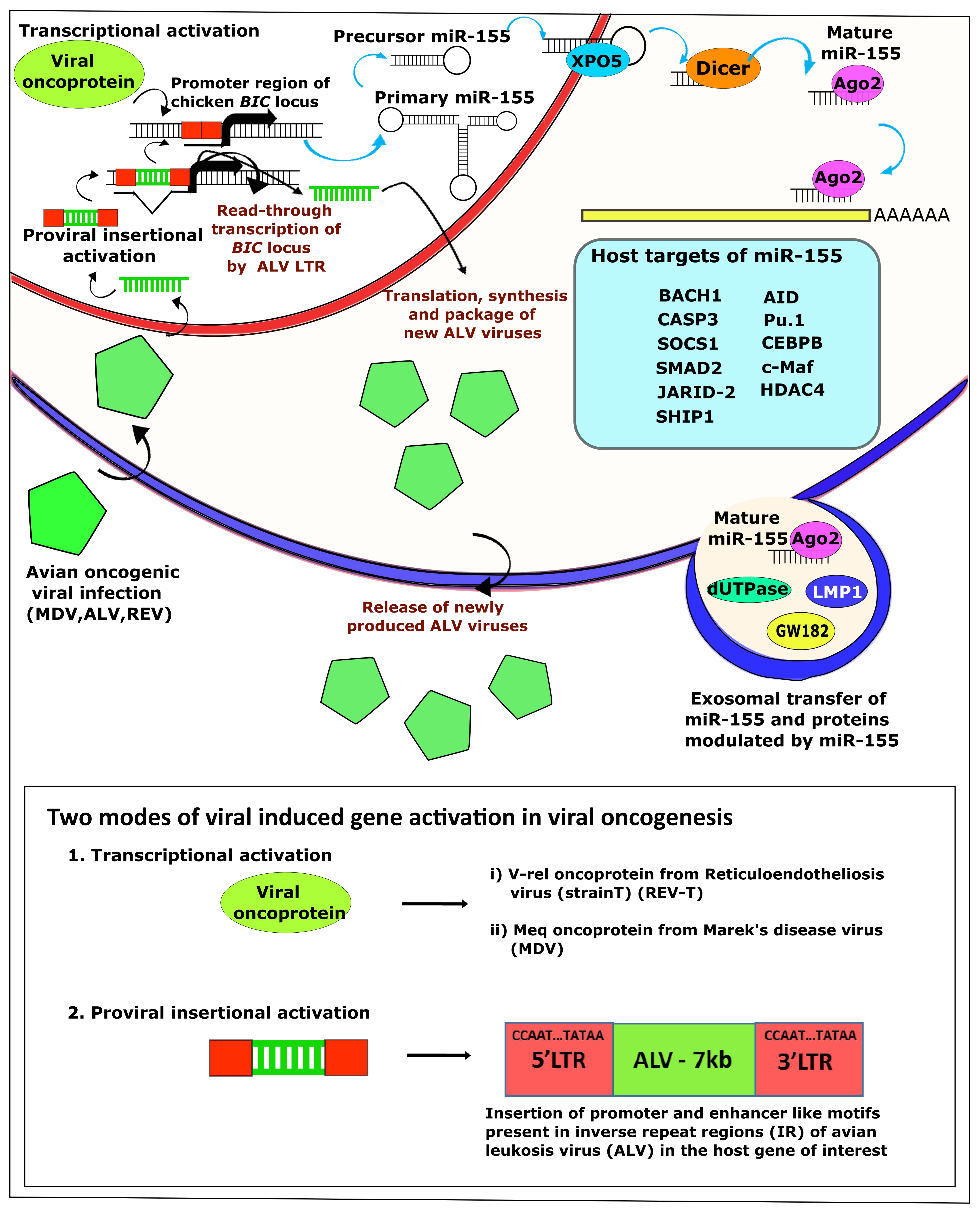
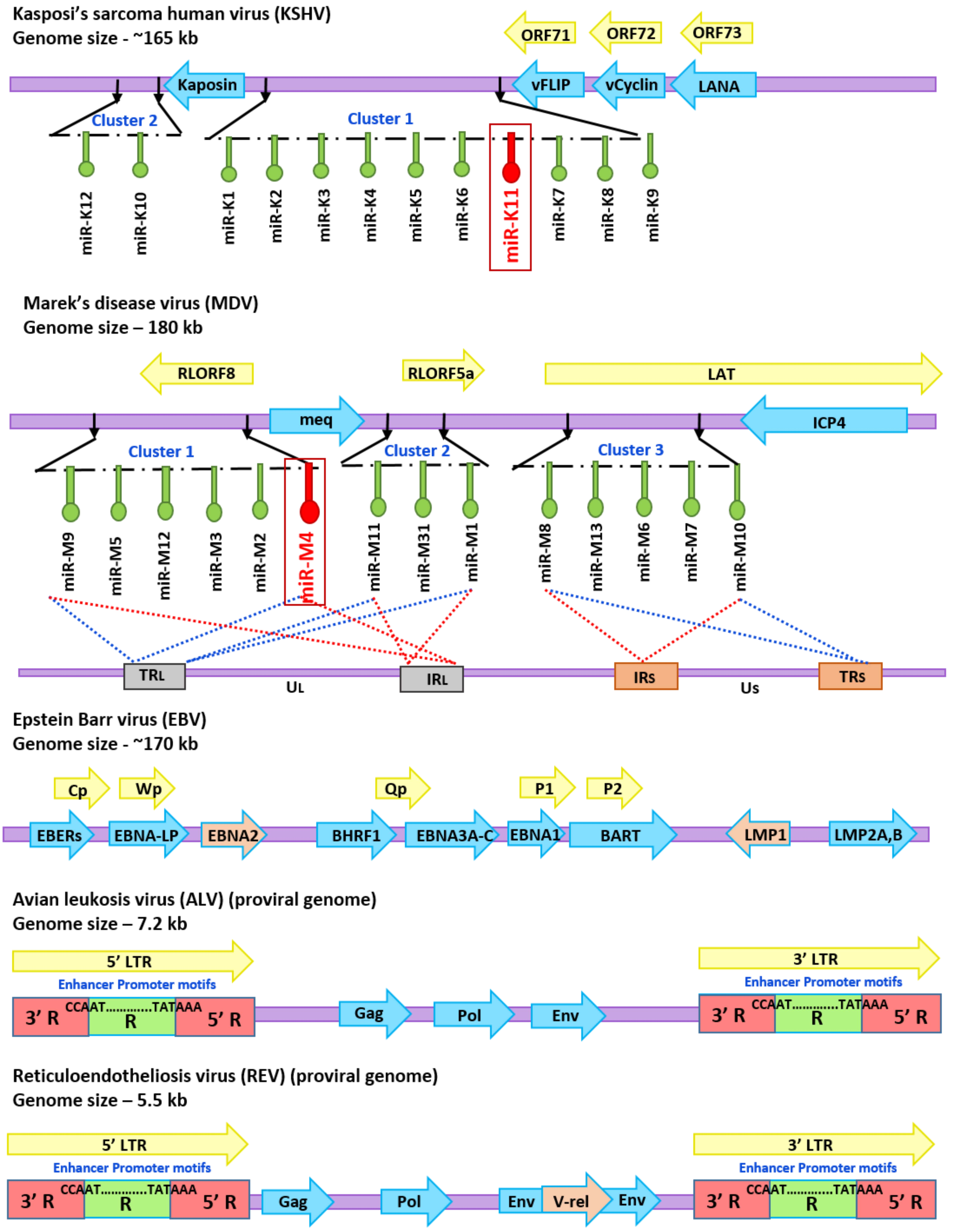
4. Avian Oncogenic Viruses and miR-155
4.1. MDV-1 and miRNAs
4.2. ALV and miRNAs
4.3. REV and miRNAs
5. Discussion and Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Weiss, R.A.; Vogt, P.K. 100 years of Rous sarcoma virus. J. Exp. Med. 2011, 208, 2351. [Google Scholar] [CrossRef]
- Clurman, B.E.; Hayward, W.S. Multiple proto-oncogene activations in avian leukosis virus-induced lymphomas: Evidence for stage-specific events. Mol. Cell. Biol. 1989, 9, 2657. [Google Scholar] [CrossRef] [PubMed]
- Rubin, H. The early history of tumor virology: Rous, RIF, and RAV. Proc. Natl. Acad. Sci. USA 2011, 108, 14389. [Google Scholar] [CrossRef] [PubMed]
- Tornesello, M.L.; Annunziata, C.; Tornesello, A.L.; Buonaguro, L.; Buonaguro, F.M. Human Oncoviruses and p53 Tumor Suppressor Pathway Deregulation at the Origin of Human Cancers. Cancers 2018, 10, 213. [Google Scholar] [CrossRef]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: Are the answers in sight? Nat. Rev. Genet. 2008, 9, 102–114. [Google Scholar] [CrossRef]
- Volinia, S.; Galasso, M.; Costinean, S.; Tagliavini, L.; Gamberoni, G.; Drusco, A.; Marchesini, J.; Mascellani, N.; Sana, M.E.; Abu Jarour, R.; et al. Reprogramming of miRNA networks in cancer and leukemia. Genome Res. 2010, 20, 589–599. [Google Scholar] [CrossRef]
- Calin, G.A.; Sevignani, C.; Dumitru, C.D.; Hyslop, T.; Noch, E.; Yendamuri, S.; Shimizu, M.; Rattan, S.; Bullrich, F.; Negrini, M.; et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl. Acad. Sci. USA 2004, 101, 2999–3004. [Google Scholar] [CrossRef]
- Lin, Z.; Flemington, E.K. miRNAs in the pathogenesis of oncogenic human viruses. Cancer Lett. 2011, 305, 186–199. [Google Scholar] [CrossRef]
- Lin, S.; Gregory, R.I. MicroRNA biogenesis pathways in cancer. Nat. Rev. Cancer 2015, 15, 321. [Google Scholar] [CrossRef]
- Van Duyne, R.; Guendel, I.; Klase, Z.; Narayanan, A.; Coley, W.; Jaworski, E.; Roman, J.; Popratiloff, A.; Mahieux, R.; Kehn-Hall, K.; et al. Localization and sub-cellular shuttling of HTLV-1 tax with the miRNA machinery. PLoS ONE 2012, 7, e40662. [Google Scholar] [CrossRef] [PubMed]
- Hsin, J.P.; Lu, Y.; Loeb, G.B.; Leslie, C.S.; Rudensky, A.Y. The effect of cellular context on miR-155-mediated gene regulation in four major immune cell types. Nat. Immunol. 2018, 19, 1137–1145. [Google Scholar] [CrossRef] [PubMed]
- Svoronos, A.A.; Engelman, D.M.; Slack, F.J. OncomiR or Tumor Suppressor? The Duplicity of MicroRNAs in Cancer. Cancer Res. 2016, 76, 3666–3670. [Google Scholar] [CrossRef]
- Pekarsky, Y.; Croce, C.M. Is miR-29 an oncogene or tumor suppressor in CLL? Oncotarget 2010, 1, 224–227. [Google Scholar] [CrossRef] [PubMed]
- Costa-Pinheiro, P.; Ramalho-Carvalho, J.; Vieira, F.Q.; Torres-Ferreira, J.; Oliveira, J.; Goncalves, C.S.; Costa, B.M.; Henrique, R.; Jeronimo, C. MicroRNA-375 plays a dual role in prostate carcinogenesis. Clin. Epigenet. 2015, 7, 42. [Google Scholar] [CrossRef] [PubMed]
- Gebert, L.F.R.; MacRae, I.J. Regulation of microRNA function in animals. Nat. Rev. Mol. Cell Biol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Anastasiadou, E.; Jacob, L.S.; Slack, F.J. Non-coding RNA networks in cancer. Nat. Rev. Cancer 2018, 18, 5–18. [Google Scholar] [CrossRef] [PubMed]
- Park, H.J.; Ji, P.; Kim, S.; Xia, Z.; Rodriguez, B.; Li, L.; Su, J.; Chen, K.; Masamha, C.P.; Baillat, D.; et al. 3’ UTR shortening represses tumor-suppressor genes in trans by disrupting ceRNA crosstalk. Nat. Genet. 2018, 50, 783–789. [Google Scholar] [CrossRef]
- Due, H.; Svendsen, P.; Bodker, J.S.; Schmitz, A.; Bogsted, M.; Johnsen, H.E.; El-Galaly, T.C.; Roug, A.S.; Dybkaer, K. miR-155 as a Biomarker in B-Cell Malignancies. Biomed. Res. Int. 2016, 2016, 9513037. [Google Scholar] [CrossRef]
- Narayan, N.; Bracken, C.P.; Ekert, P.G. MicroRNA-155 expression and function in AML: An evolving paradigm. Exp. Hematol. 2018, 62, 1–6. [Google Scholar] [CrossRef]
- Tam, W. Identification and characterization of human BIC, a gene on chromosome 21 that encodes a noncoding RNA. Gene 2001, 274, 157–167. [Google Scholar] [CrossRef]
- Eis, P.S.; Tam, W.; Sun, L.; Chadburn, A.; Li, Z.; Gomez, M.F.; Lund, E.; Dahlberg, J.E. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc. Natl. Acad. Sci. USA 2005, 102, 3627–3632. [Google Scholar] [CrossRef] [PubMed]
- Kluiver, J.; Poppema, S.; de Jong, D.; Blokzijl, T.; Harms, G.; Jacobs, S.; Kroesen, B.J.; van den Berg, A. BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 2005, 207, 243–249. [Google Scholar] [CrossRef]
- Van den Berg, A.; Kroesen, B.J.; Kooistra, K.; de Jong, D.; Briggs, J.; Blokzijl, T.; Jacobs, S.; Kluiver, J.; Diepstra, A.; Maggio, E.; et al. High expression of B-cell receptor inducible gene BIC in all subtypes of Hodgkin lymphoma. Genes Chromosomes Cancer 2003, 37, 20–28. [Google Scholar] [CrossRef] [PubMed]
- Costinean, S.; Sandhu, S.K.; Pedersen, I.M.; Tili, E.; Trotta, R.; Perrotti, D.; Ciarlariello, D.; Neviani, P.; Harb, J.; Kauffman, L.R.; et al. Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein beta are targeted by miR-155 in B cells of Emicro-MiR-155 transgenic mice. Blood 2009, 114, 1374–1382. [Google Scholar] [CrossRef] [PubMed]
- Costinean, S.; Zanesi, N.; Pekarsky, Y.; Tili, E.; Volinia, S.; Heerema, N.; Croce, C.M. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in E(mu)-miR155 transgenic mice. Proc. Natl. Acad. Sci. USA 2006, 103, 7024–7029. [Google Scholar] [CrossRef] [PubMed]
- Lu, L.F.; Gasteiger, G.; Yu, I.S.; Chaudhry, A.; Hsin, J.P.; Lu, Y.; Bos, P.D.; Lin, L.L.; Zawislak, C.L.; Cho, S.; et al. A Single miRNA-mRNA Interaction Affects the Immune Response in a Context- and Cell-Type-Specific Manner. Immunity 2015, 43, 52–64. [Google Scholar] [CrossRef]
- Vigorito, E.; Perks, K.L.; Abreu-Goodger, C.; Bunting, S.; Xiang, Z.; Kohlhaas, S.; Das, P.P.; Miska, E.A.; Rodriguez, A.; Bradley, A.; et al. microRNA-155 regulates the generation of immunoglobulin class-switched plasma cells. Immunity 2007, 27, 847–859. [Google Scholar] [CrossRef] [PubMed]
- Seddiki, N.; Brezar, V.; Ruffin, N.; Levy, Y.; Swaminathan, S. Role of miR-155 in the regulation of lymphocyte immune function and disease. Immunology 2014, 142, 32–38. [Google Scholar] [CrossRef] [PubMed]
- Vojtechova, Z.; Tachezy, R. The Role of miRNAs in Virus-Mediated Oncogenesis. Int. J. Mol. Sci. 2018, 19, 1217. [Google Scholar] [CrossRef]
- Yao, Y.; Nair, V. Role of virus-encoded microRNAs in Avian viral diseases. Viruses 2014, 6, 1379–1394. [Google Scholar] [CrossRef] [PubMed]
- Fan, C.; Tang, Y.; Wang, J.; Xiong, F.; Guo, C.; Wang, Y.; Xiang, B.; Zhou, M.; Li, X.; Wu, X.; et al. The emerging role of Epstein-Barr virus encoded microRNAs in nasopharyngeal carcinoma. J. Cancer 2018, 9, 2852–2864. [Google Scholar] [CrossRef] [PubMed]
- Kincaid, R.P.; Sullivan, C.S. Virus-encoded microRNAs: An overview and a look to the future. PLoS Pathog. 2012, 8, e1003018. [Google Scholar] [CrossRef] [PubMed]
- Piedade, D.; Azevedo-Pereira, J.M. The Role of microRNAs in the Pathogenesis of Herpesvirus Infection. Viruses 2016, 8, 156. [Google Scholar] [CrossRef]
- Gottwein, E.; Mukherjee, N.; Sachse, C.; Frenzel, C.; Majoros, W.H.; Chi, J.T.; Braich, R.; Manoharan, M.; Soutschek, J.; Ohler, U.; et al. A viral microRNA functions as an orthologue of cellular miR-155. Nature 2007, 450, 1096–1099. [Google Scholar] [CrossRef] [PubMed]
- Skalsky, R.L.; Samols, M.A.; Plaisance, K.B.; Boss, I.W.; Riva, A.; Lopez, M.C.; Baker, H.V.; Renne, R. Kaposi’s sarcoma-associated herpesvirus encodes an ortholog of miR-155. J. Virol. 2007, 81. [Google Scholar] [CrossRef] [PubMed]
- Sin, S.H.; Kim, Y.B.; Dittmer, D.P. Latency locus complements MicroRNA 155 deficiency in vivo. J. Virol. 2013, 87, 11908–11911. [Google Scholar] [CrossRef]
- Liu, Y.; Sun, R.; Lin, X.; Liang, D.; Deng, Q.; Lan, K. Kaposi’s sarcoma-associated herpesvirus-encoded microRNA miR-K12-11 attenuates transforming growth factor beta signaling through suppression of SMAD5. J. Virol. 2012, 86, 1372–1381. [Google Scholar] [CrossRef] [PubMed]
- Gay, L.A.; Sethuraman, S.; Thomas, M.; Turner, P.C.; Renne, R. Modified Cross-Linking, Ligation, and Sequencing of Hybrids (qCLASH) Identifies Kaposi’s Sarcoma-Associated Herpesvirus MicroRNA Targets in Endothelial Cells. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Zhao, Y.; Yao, Y.; Xu, H.; Lambeth, L.; Smith, L.P.; Kgosana, L.; Wang, X.; Nair, V. A functional MicroRNA-155 ortholog encoded by the oncogenic Marek’s disease virus. J. Virol. 2009, 83, 489–492. [Google Scholar] [CrossRef]
- Parnas, O.; Corcoran, D.L.; Cullen, B.R. Analysis of the mRNA targetome of microRNAs expressed by Marek’s disease virus. mBio 2014, 5, e01060-01013. [Google Scholar] [CrossRef] [PubMed]
- Gottwein, E.; Corcoran, D.L.; Mukherjee, N.; Skalsky, R.L.; Hafner, M.; Nusbaum, J.D.; Shamulailatpam, P.; Love, C.L.; Dave, S.S.; Tuschl, T.; et al. Viral microRNA targetome of KSHV-infected primary effusion lymphoma cell lines. Cell Host Microbe 2011, 10, 515–526. [Google Scholar] [CrossRef] [PubMed]
- Linnstaedt, S.D.; Gottwein, E.; Skalsky, R.L.; Luftig, M.A.; Cullen, B.R. Virally induced cellular microRNA miR-155 plays a key role in B-cell immortalization by Epstein-Barr virus. J. Virol. 2010, 84, 11670–11678. [Google Scholar] [CrossRef]
- Wood, C.D.; Carvell, T.; Gunnell, A.; Ojeniyi, O.O.; Osborne, C.; West, M.J. Enhancer control of miR-155 expression in Epstein-Barr virus infected B cells. J. Virol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Read, A.F.; Baigent, S.J.; Powers, C.; Kgosana, L.B.; Blackwell, L.; Smith, L.P.; Kennedy, D.A.; Walkden-Brown, S.W.; Nair, V.K. Imperfect Vaccination Can Enhance the Transmission of Highly Virulent Pathogens. PLoS Biol. 2015, 13, e1002198. [Google Scholar] [CrossRef] [PubMed]
- Burnside, J.; Bernberg, E.; Anderson, A.; Lu, C.; Meyers, B.C.; Green, P.J.; Jain, N.; Isaacs, G.; Morgan, R.W. Marek’s disease virus encodes MicroRNAs that map to meq and the latency-associated transcript. J. Virol. 2006, 80, 8778–8786. [Google Scholar] [CrossRef]
- Yao, Y.; Zhao, Y.; Xu, H.; Smith, L.P.; Lawrie, C.H.; Watson, M.; Nair, V. MicroRNA profile of Marek’s disease virus-transformed T-cell line MSB-1: Predominance of virus-encoded microRNAs. J. Virol. 2008, 82, 4007–4015. [Google Scholar] [CrossRef]
- Morgan, R.; Anderson, A.; Bernberg, E.; Kamboj, S.; Huang, E.; Lagasse, G.; Isaacs, G.; Parcells, M.; Meyers, B.C.; Green, P.J.; et al. Sequence conservation and differential expression of Marek’s disease virus microRNAs. J. Virol. 2008, 82, 12213–12220. [Google Scholar] [CrossRef]
- Burnside, J.; Morgan, R. Emerging roles of chicken and viral microRNAs in avian disease. BMC Proc. 2011, 5 (Suppl. 4), S2. [Google Scholar] [CrossRef]
- Coupeau, D.; Dambrine, G.; Rasschaert, D. Kinetic expression analysis of the cluster mdv1-mir-M9-M4, genes meq and vIL-8 differs between the lytic and latent phases of Marek’s disease virus infection. J. Gen. Virol. 2012, 93, 1519–1529. [Google Scholar] [CrossRef]
- Zhuang, G.; Sun, A.; Teng, M.; Luo, J. A Tiny RNA that Packs a Big Punch: The critical role of a viral miR-155 ortholog in lymphomagenesis in Marek’s disease. Front. Microbiol. 2017. [Google Scholar] [CrossRef]
- Zhao, Y.; Xu, H.; Yao, Y.; Smith, L.P.; Kgosana, L.; Green, J.; Petherbridge, L.; Baigent, S.J.; Nair, V. Critical role of the virus-encoded microRNA-155 ortholog in the induction of Marek’s disease lymphomas. PLoS Pathog. 2011, 7, e1001305. [Google Scholar] [CrossRef]
- Brown, A.C.; Nair, V.; Allday, M.J. Epigenetic regulation of the latency-associated region of Marek’s disease virus in tumor-derived T-cell lines and primary lymphoma. J. Virol. 2012, 86, 1683–1695. [Google Scholar] [CrossRef] [PubMed]
- Yu, Z.H.; Teng, M.; Sun, A.J.; Yu, L.L.; Hu, B.; Qu, L.H.; Ding, K.; Cheng, X.C.; Liu, J.X.; Cui, Z.Z.; et al. Virus-encoded miR-155 ortholog is an important potential regulator but not essential for the development of lymphomas induced by very virulent Marek’s disease virus. Virology 2014, 448, 55–64. [Google Scholar] [CrossRef] [PubMed]
- Nair, V.; Fadly, A. Leukosis/Sarcoma group. In Diseases of Poultry, 13th ed.; Swayne, D.E., Glisson, J.R., McDougald, L.R., Nolan, L.K., Suarez, D.L., Nair, V., Eds.; John Wiley & Sons, Inc.: Ames, IA, USA, 2013; pp. 553–592. [Google Scholar]
- Yao, Y.; Charlesworth, J.; Nair, V.; Watson, M. MicroRNA expression profiles in avian haemopoietic cells. Front. Genet. 2013, 4. [Google Scholar] [CrossRef]
- Tam, W.; Ben-Yehuda, D.; Hayward, W.S. BIC, a novel gene activated by proviral insertions in avian leukosis virus-induced lymphomas, is likely to function through its noncoding RNA. Mol. Cell. Biol. 1997, 17, 1490. [Google Scholar] [CrossRef]
- Yao, Y.; Smith, L.P.; Nair, V.; Watson, M. An avian retrovirus uses canonical expression and processing mechanisms to generate viral microRNA. J. Virol. 2014, 88, 2–9. [Google Scholar] [CrossRef] [PubMed]
- Justice, J.F.; Morgan, R.W.; Beemon, K.L. Common Viral Integration Sites Identified in Avian Leukosis Virus-Induced B-Cell Lymphomas. mBio 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Nair, V.; Zavala, G.; Fadly, A.M. Reticuloendotheliosis. In Diseases of Poultry, 13th ed.; Swayne, D.E., Glisson, J.R., McDougald, L.R., Nolan, L.K., Suarez, D.L., Nair, V., Eds.; Wiley-Blackwell: Ames, IA, USA, 2013; pp. 593–604. [Google Scholar]
- Niewiadomska, A.M.; Gifford, R.J. The extraordinary evolutionary history of the reticuloendotheliosis viruses. PLoS Biol. 2013, 11, e1001642. [Google Scholar] [CrossRef]
- Bolisetty, M.T.; Dy, G.; Tam, W.; Beemon, K.L. Reticuloendotheliosis virus strain T induces miR-155, which targets JARID2 and promotes cell survival. J. Virol. 2009, 83, 12009–12017. [Google Scholar] [CrossRef]
- Yao, Y.; Vasoya, D.; Kgosana, L.; Smith, L.P.; Gao, Y.; Wang, X.; Watson, M.; Nair, V. Activation of gga-miR-155 by reticuloendotheliosis virus T strain and its contribution to transformation. J. Gen. Virol. 2017, 98, 810–820. [Google Scholar] [CrossRef] [PubMed]
- Quarton, T.; Ehrhardt, K.; Lee, J.; Kannan, S.; Li, Y.; Ma, L.; Bleris, L. Mapping the operational landscape of microRNAs in synthetic gene circuits. NPJ Syst. Biol. Appl. 2018, 4, 6. [Google Scholar] [CrossRef]
- Stelekati, E.; Chen, Z.; Manne, S.; Kurachi, M.; Ali, M.A.; Lewy, K.; Cai, Z.; Nzingha, K.; McLane, L.M.; Hope, J.L.; et al. Long-Term Persistence of Exhausted CD8 T Cells in Chronic Infection Is Regulated by MicroRNA-155. Cell Rep. 2018, 23, 2142–2156. [Google Scholar] [CrossRef] [PubMed]
- Mikamori, M.; Yamada, D.; Eguchi, H.; Hasegawa, S.; Kishimoto, T.; Tomimaru, Y.; Asaoka, T.; Noda, T.; Wada, H.; Kawamoto, K.; et al. MicroRNA-155 Controls Exosome Synthesis and Promotes Gemcitabine Resistance in Pancreatic Ductal Adenocarcinoma. Sci. Rep. 2017, 7, 42339. [Google Scholar] [CrossRef]
- Pegtel, D.M.; Cosmopoulos, K.; Thorley-Lawson, D.A.; van Eijndhoven, M.A.J.; Hopmans, E.S.; Lindenberg, J.L.; de Gruijl, T.D.; Würdinger, T.; Middeldorp, J.M. Functional delivery of viral miRNAs via exosomes. Proc. Natl. Acad. Sci. USA 2010, 107, 6328. [Google Scholar] [CrossRef] [PubMed]
- Nath Neerukonda, S.; Egan, N.A.; Patria, J.; Assakhi, I.; Tavlarides-Hontz, P.; Modla, S.; Munoz, E.R.; Hudson, M.B.; Parcells, M.S. Comparison of exosomes purified via ultracentrifugation (UC) and Total Exosome Isolation (TEI) reagent from the serum of Marek’s disease virus (MDV)-vaccinated and tumor-bearing chickens. J. Virol. Methods 2019, 263, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Passacantilli, I.; Panzeri, V.; Bielli, P.; Farini, D.; Pilozzi, E.; Fave, G.D.; Capurso, G.; Sette, C. Alternative polyadenylation of ZEB1 promotes its translation during genotoxic stress in pancreatic cancer cells. Cell Death Dis. 2017, 8, e3168. [Google Scholar] [CrossRef] [PubMed]
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bondada, M.S.; Yao, Y.; Nair, V. Multifunctional miR-155 Pathway in Avian Oncogenic Virus-Induced Neoplastic Diseases. Non-Coding RNA 2019, 5, 24. https://doi.org/10.3390/ncrna5010024
Bondada MS, Yao Y, Nair V. Multifunctional miR-155 Pathway in Avian Oncogenic Virus-Induced Neoplastic Diseases. Non-Coding RNA. 2019; 5(1):24. https://doi.org/10.3390/ncrna5010024
Chicago/Turabian StyleBondada, Megha Sravani, Yongxiu Yao, and Venugopal Nair. 2019. "Multifunctional miR-155 Pathway in Avian Oncogenic Virus-Induced Neoplastic Diseases" Non-Coding RNA 5, no. 1: 24. https://doi.org/10.3390/ncrna5010024
APA StyleBondada, M. S., Yao, Y., & Nair, V. (2019). Multifunctional miR-155 Pathway in Avian Oncogenic Virus-Induced Neoplastic Diseases. Non-Coding RNA, 5(1), 24. https://doi.org/10.3390/ncrna5010024





