Evolutionary Patterns of Non-Coding RNA in Cardiovascular Biology
Abstract
1. Introduction
2. Heart and Non-Coding RNAs
2.1. Long Non-Coding RNAs
2.2. MicroRNAs
2.3. Circular RNAs
3. RNA-Sequencing for Identification of Non-Coding RNA
4. Experimental Methodologies to Explore ncRNA Functionality
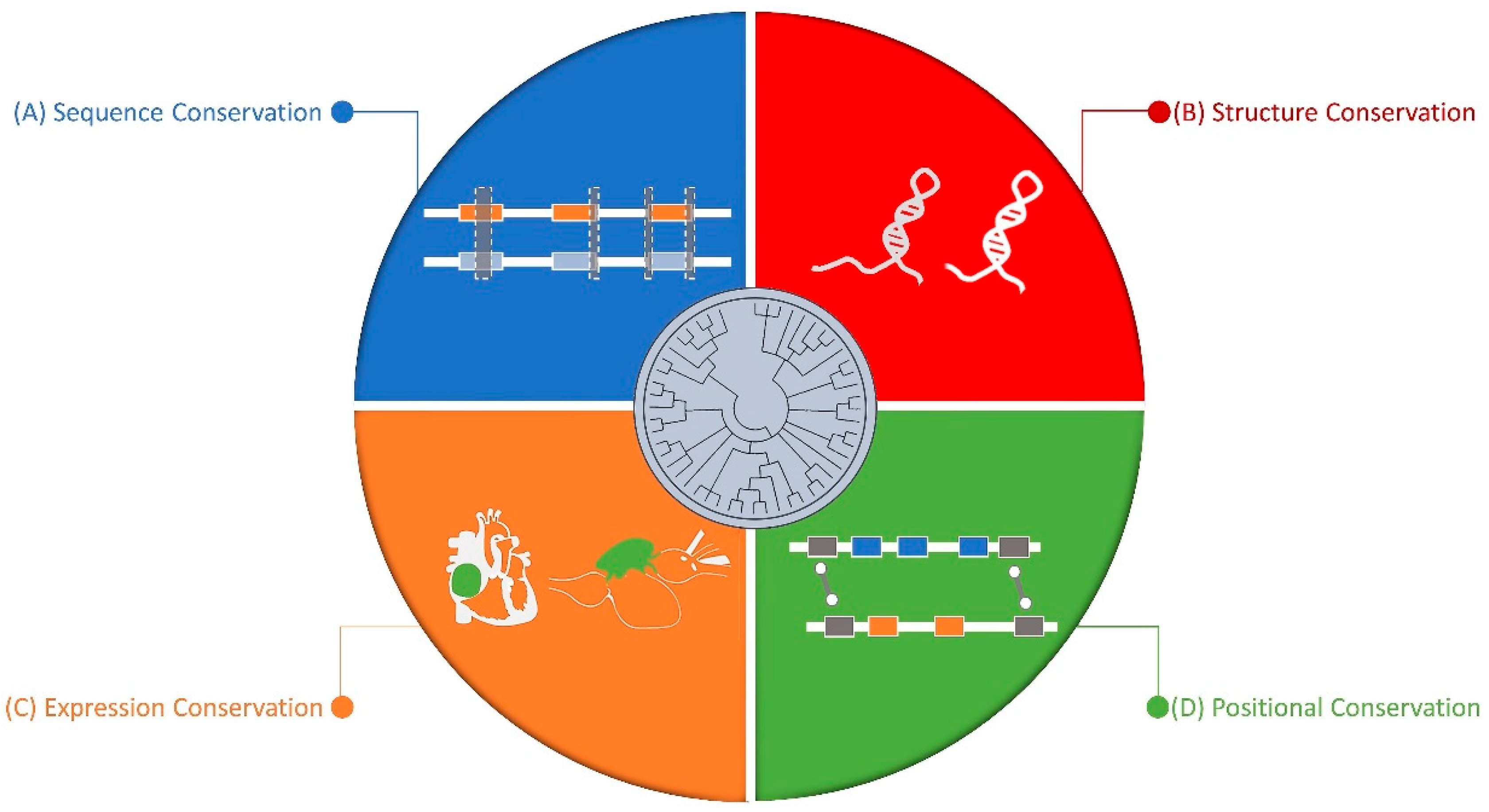
5. Conserved Nature of Non-Coding RNAs
5.1. Sequence Level
5.2. Structure Level
5.3. Synteny Level
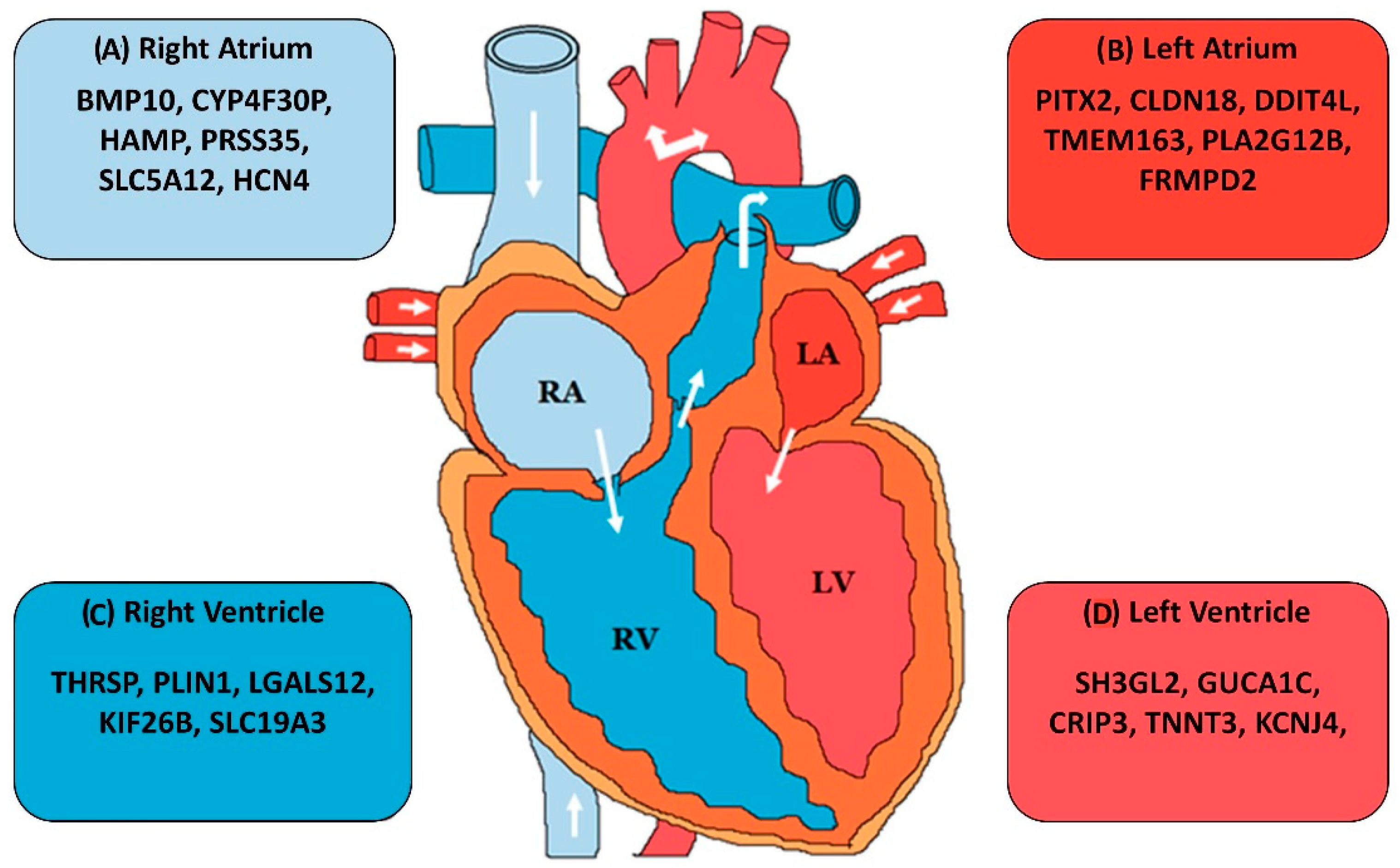
5.4. Expression Level
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Cardiovascular Diseases (CVDs). Available online: https://www.who.int/news-room/fact-sheets/detail/cardiovascular-diseases-(cvds) (accessed on 14 January 2019).
- Benjamin, E.J.; Virani, S.S.; Callaway, C.W.; Chamberlain, A.M.; Chang, A.R.; Cheng, S.; Chiuve, S.E.; Cushman, M.; Delling, F.N.; Deo, R.; et al. Heart disease and stroke statistics—2018 Update: A report from the American Heart Association. Circulation 2018, 137, e67–e492. [Google Scholar] [CrossRef] [PubMed]
- International Diabetes Federation. Diabetes and Cardiovascular Disease; International Diabetes Federation: Brussels, Belgium, 2016. [Google Scholar]
- Manolio, T.A.; Collins, F.S.; Cox, N.J.; Goldstein, D.B.; Hindorff, L.A.; Hunter, D.J.; McCarthy, M.I.; Ramos, E.M.; Cardon, L.R.; Chakravarti, A.; et al. Finding the missing heritability of complex diseases. Nature 2009, 461, 747–753. [Google Scholar] [CrossRef] [PubMed]
- Wu, P.Y.; Chandramohan, R.; Phan, J.H.; Mahle, W.T.; Gaynor, J.W.; Maher, K.O.; Wang, M.D. Cardiovascular transcriptomics and epigenomics using next-generation sequencing challenges, progress, and opportunities. Circ. Cardiovasc. Genet. 2014, 7, 701–710. [Google Scholar] [CrossRef] [PubMed]
- Pedrotty, D.M.; Morley, M.P.; Cappola, T.P. Transcriptomic biomarkers of cardiovascular disease. Prog. Cardiovasc. Dis. 2012, 55, 64–69. [Google Scholar] [CrossRef] [PubMed]
- Song, Y.; Stampfer, M.J.; Liu, S. Meta-analysis: Apolipoprotein E genotypes and risk for coronary heart disease. Ann. Intern. Med. 2004, 141, 137–147. [Google Scholar] [CrossRef] [PubMed]
- Boekholdt, S.M.; Bijsterveld, N.R.; Moons, A.H.M.; Levi, M.; Büller, H.R.; Peters, R.J.G. Genetic variation in coagulation and fibrinolytic proteins and their relation with acute myocardial infarction: A systematic review. Circulation 2001, 104, 3063–3068. [Google Scholar] [CrossRef] [PubMed]
- Agerholm-Larsen, B.; Nordestgaard, B.G.; Tybjaerg-Hansen, A. ACE gene polymorphism in cardiovascular disease: Meta-analyses of small and large studies in whites. Arterioscler. Thromb. Vasc. Biol. 2000, 20, 484–492. [Google Scholar] [CrossRef]
- Klerk, M.; Verhoef, P.; Clarke, R.; Blom, H.J.; Kok, F.J.; Schouten, E.G. MTHFR Studies Collaboration Group MTHFR 677C→T polymorphism and risk of coronary heart disease: A meta-analysis. JAMA 2002, 288, 2023–2031. [Google Scholar] [CrossRef]
- Jalali, S.; Gandhi, S.; Scaria, V. Navigating the dynamic landscape of long noncoding RNA and protein-coding gene annotations in GENCODE. Hum. Genom. 2016, 10, 35. [Google Scholar] [CrossRef]
- Pertea, M. The human transcriptome: An unfinished story. Genes 2012, 3, 344–360. [Google Scholar] [CrossRef]
- Young, R.S.; Ponting, C.P. Identification and function of long non-coding RNAs. Essays Biochem. 2013, 54, 113–126. [Google Scholar] [CrossRef] [PubMed]
- Tian, D.; Sun, S.; Lee, J.T. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 2010, 143, 390–403. [Google Scholar] [CrossRef] [PubMed]
- Chen, L.-L. Linking long noncoding RNA localization and function. Trends Biochem. Sci. 2016, 41, 761–772. [Google Scholar] [CrossRef] [PubMed]
- van Heesch, S.; van Iterson, M.; Jacobi, J.; Boymans, S.; Essers, P.B.; de Bruijn, E.; Hao, W.; MacInnes, A.W.; Cuppen, E.; Simonis, M. Extensive localization of long noncoding RNAs to the cytosol and mono- and polyribosomal complexes. Genome Biol. 2014, 15, R6. [Google Scholar] [CrossRef] [PubMed]
- Carlevaro-Fita, J.; Rahim, A.; Guigó, R.; Vardy, L.A.; Johnson, R. Cytoplasmic long noncoding RNAs are frequently bound to and degraded at ribosomes in human cells. RNA 2016, 22, 867–882. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.-Z.; Chen, M.; Chen, D.; Gao, X.-C.; Zhu, S.; Huang, H.; Hu, M.; Zhu, H.; Yan, G.-R. A peptide encoded by a putative lncRNA HOXB-AS3 suppresses colon cancer growth. Mol. Cell 2017, 68, 171–184. [Google Scholar] [CrossRef] [PubMed]
- Archer, K.; Broskova, Z.; Bayoumi, A.; Teoh, J.; Davila, A.; Tang, Y.; Su, H.; Kim, I. Long non-coding RNAs as master regulators in cardiovascular diseases. Int. J. Mol. Sci. 2015, 16, 23651–23667. [Google Scholar] [CrossRef] [PubMed]
- Devaux, Y.; Zangrando, J.; Schroen, B.; Creemers, E.E.; Pedrazzini, T.; Chang, C.-P.; Dorn, G.W.; Thum, T.; Heymans, S. Cardiolinc network Long noncoding RNAs in cardiac development and ageing. Nat. Rev. Cardiol. 2015, 12, 415–425. [Google Scholar] [PubMed]
- Haemmig, S.; Simion, V.; Yang, D.; Deng, Y.; Feinberg, M.W. Long noncoding RNAs in cardiovascular disease, diagnosis, and therapy. Curr. Opin. Cardiol. 2017, 32, 776–783. [Google Scholar] [CrossRef] [PubMed]
- Gomes, C.P.C.; Spencer, H.; Ford, K.L.; Michel, L.Y.M.; Baker, A.H.; Emanueli, C.; Balligand, J.-L.; Devaux, Y. Cardiolinc network the function and therapeutic potential of long non-coding RNAs in cardiovascular development and disease. Mol. Ther. Nucleic Acids 2017, 8, 494–507. [Google Scholar] [CrossRef]
- Boulberdaa, M.; Scott, E.; Ballantyne, M.; Garcia, R.; Descamps, B.; Angelini, G.D.; Brittan, M.; Hunter, A.; McBride, M.; McClure, J.; et al. A role for the long noncoding RNA SENCR in commitment and function of endothelial cells. Mol. Ther. 2016, 24, 978–990. [Google Scholar] [CrossRef] [PubMed]
- Deloukas, P.; Kanoni, S.; Willenborg, C.; Farrall, M.; Assimes, T.L.; Thompson, J.R.; Ingelsson, E.; Saleheen, D.; Erdmann, J.; Goldstein, B.A.; et al. Large-scale association analysis identifies new risk loci for coronary artery disease. Nat. Genet. 2013, 45, 25–33. [Google Scholar] [CrossRef] [PubMed]
- Xue, Z.; Hennelly, S.; Doyle, B.; Gulati, A.A.; Novikova, I.V.; Sanbonmatsu, K.Y.; Boyer, L.A. A G-Rich motif in the lncRNA braveheart interacts with a zinc-finger transcription factor to specify the cardiovascular lineage. Mol. Cell 2016, 64, 37–50. [Google Scholar] [CrossRef] [PubMed]
- Grote, P.; Wittler, L.; Hendrix, D.; Koch, F.; Währisch, S.; Beisaw, A.; Macura, K.; Bläss, G.; Kellis, M.; Werber, M.; et al. The tissue-specific lncRNA fendrr is an essential regulator of heart and body wall development in the mouse. Dev. Cell 2013, 24, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Ounzain, S.; Micheletti, R.; Beckmann, T.; Schroen, B.; Alexanian, M.; Pezzuto, I.; Crippa, S.; Nemir, M.; Sarre, A.; Johnson, R.; et al. Genome-wide profiling of the cardiac transcriptome after myocardial infarction identifies novel heart-specific long non-coding RNAs. Eur. Heart J. 2015, 36, 353–368. [Google Scholar] [CrossRef] [PubMed]
- Ounzain, S.; Micheletti, R.; Arnan, C.; Plaisance, I.; Cecchi, D.; Schroen, B.; Reverter, F.; Alexanian, M.; Gonzales, C.; Ng, S.Y.; et al. CARMEN, a human super enhancer-associated long noncoding RNA controlling cardiac specification, differentiation and homeostasis. J. Mol. Cell. Cardiol. 2015, 89, 98–112. [Google Scholar] [CrossRef] [PubMed]
- Korostowski, L.; Sedlak, N.; Engel, N. The Kcnq1ot1 long non-coding RNA affects chromatin conformation and expression of Kcnq1, but does not regulate its imprinting in the developing heart. PLoS Genet. 2012, 8, e1002956. [Google Scholar] [CrossRef]
- Bell, R.D.; Long, X.; Lin, M.; Bergmann, J.H.; Nanda, V.; Cowan, S.L.; Zhou, Q.; Han, Y.; Spector, D.L.; Zheng, D.; et al. Identification and initial functional characterization of a human vascular cell-enriched long noncoding RNA. Arterioscler. Thromb. Vasc. Biol. 2014, 34, 1249–1259. [Google Scholar] [CrossRef]
- Zhang, M.; Gu, H.; Chen, J.; Zhou, X. Involvement of long noncoding RNA MALAT1 in the pathogenesis of diabetic cardiomyopathy. Int. J. Cardiol. 2016, 202, 753–755. [Google Scholar] [CrossRef]
- Gao, W.; Zhu, M.; Wang, H.; Zhao, S.; Zhao, D.; Yang, Y.; Wang, Z.-M.; Wang, F.; Yang, Z.-J.; Lu, X.; et al. Association of polymorphisms in long non-coding RNA H19 with coronary artery disease risk in a Chinese population. Mutat. Res. Mol. Mech. Mutagen. 2015, 772, 15–22. [Google Scholar] [CrossRef]
- Shan, K.; Jiang, Q.; Wang, X.-Q.; Wang, Y.-N.-Z.; Yang, H.; Yao, M.-D.; Liu, C.; Li, X.-M.; Yao, J.; Liu, B.; et al. Role of long non-coding RNA-RNCR3 in atherosclerosis-related vascular dysfunction. Cell Death Dis. 2016, 7, e2248. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Zhang, X.-J.; Ji, Y.-X.; Zhang, P.; Deng, K.-Q.; Gong, J.; Ren, S.; Wang, X.; Chen, I.; Wang, H.; et al. The long noncoding RNA Chaer defines an epigenetic checkpoint in cardiac hypertrophy. Nat. Med. 2016, 22, 1131–1139. [Google Scholar] [CrossRef] [PubMed]
- Kumarswamy, R.; Bauters, C.; Volkmann, I.; Maury, F.; Fetisch, J.; Holzmann, A.; Lemesle, G.; de Groote, P.; Pinet, F.; Thum, T. Circulating long noncoding RNA, LIPCAR, predicts survival in patients with heart failure. Circ. Res. 2014, 114, 1569–1575. [Google Scholar] [CrossRef] [PubMed]
- Sun, C.; Huang, L.; Li, Z.; Leng, K.; Xu, Y.; Jiang, X.; Cui, Y. Long non-coding RNA MIAT in development and disease: A new player in an old game. J. Biomed. Sci. 2018, 25, 23. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Gao, C.; Meng, M.; Tang, H. Long noncoding RNA MHRT Protects cardiomyocytes against H2O2-Induced apoptosis. Biomol. Ther. 2016, 24, 19–24. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.-N.-Z.; Shan, K.; Yao, M.-D.; Yao, J.; Wang, J.-J.; Li, X.; Liu, B.; Zhang, Y.-Y.; Ji, Y.; Jiang, Q.; et al. Long noncoding RNA-GAS5. Hypertension 2016, 68, 736–748. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Yuan, J.; Gao, L.; Rao, J.; Hu, J. Long noncoding RNA MEG3 activation of p53 mediates ischemic neuronal death in stroke. Neuroscience 2016, 337, 191–199. [Google Scholar] [CrossRef]
- Yan, Y.; Zhang, B.; Liu, N.; Qi, C.; Xiao, Y.; Tian, X.; Li, T.; Liu, B. Circulating long noncoding RNA UCA1 as a novel biomarker of acute myocardial infarction. Biomed Res. Int. 2016, 2016, 1–7. [Google Scholar] [CrossRef]
- Wang, J.; Chen, L.; Li, H.; Yang, J.; Gong, Z.; Wang, B.; Zhao, X. Clopidogrel reduces apoptosis and promotes proliferation of human vascular endothelial cells induced by palmitic acid via suppression of the long non-coding RNA HIF1A-AS1 in vitro. Mol. Cell. Biochem. 2015, 404, 203–210. [Google Scholar] [CrossRef]
- Annilo, T.; Kepp, K.; Laan, M. Natural antisense transcript of natriuretic peptide precursor A (NPPA): Structural organization and modulation of NPPA expression. BMC Mol. Biol. 2009, 10, 81. [Google Scholar] [CrossRef]
- Wang, K.; Liu, F.; Zhou, L.-Y.; Long, B.; Yuan, S.-M.; Wang, Y.; Liu, C.-Y.; Sun, T.; Zhang, X.-J.; Li, P.-F. The long noncoding RNA CHRF regulates cardiac hypertrophy by targeting miR-489. Circ. Res. 2014, 114, 1377–1388. [Google Scholar] [CrossRef] [PubMed]
- Viereck, J.; Kumarswamy, R.; Foinquinos, A.; Xiao, K.; Avramopoulos, P.; Kunz, M.; Dittrich, M.; Maetzig, T.; Zimmer, K.; Remke, J.; et al. Long noncoding RNA Chast promotes cardiac remodeling. Sci. Transl. Med. 2016, 8, 326ra22. [Google Scholar] [CrossRef] [PubMed]
- Gore-Panter, S.R.; Hsu, J.; Barnard, J.; Moravec, C.S.; Van Wagoner, D.R.; Chung, M.K.; Smith, J.D. PANCR, the PITX2 adjacent noncoding RNA, is expressed in human left atria and regulates PITX2c expression. Circ. Arrhythm Electrophysiol. 2016, 9, e003197. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.-H.; Hu, Z.-Y.; Li, M.-H.; Li, B.; Wang, Z.-M.; Chen, S.-L. Cardiac hypertrophy is positively regulated by long non-coding RNA PVT1. Int. J. Clin. Exp. Pathol. 2015, 8, 2582–2589. [Google Scholar] [PubMed]
- Wang, K.; Long, B.; Zhou, L.-Y.; Liu, F.; Zhou, Q.-Y.; Liu, C.-Y.; Fan, Y.-Y.; Li, P.-F. CARL lncRNA inhibits anoxia-induced mitochondrial fission and apoptosis in cardiomyocytes by impairing miR-539-dependent PHB2 downregulation. Nat. Commun. 2014, 5, 3596. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Lu, Z.-N. Long non-coding RNA HOTAIR promotes ischemic infarct induced by hypoxia through up-regulating the expression of NOX2. Biochem. Biophys. Res. Commun. 2016, 479, 186–191. [Google Scholar] [CrossRef] [PubMed]
- Ha, M.; Kim, V.N. Regulation of microRNA biogenesis. Nat. Rev. Mol. Cell Biol. 2014, 15, 509–524. [Google Scholar] [CrossRef] [PubMed]
- Ebert, M.S.; Neilson, J.R.; Sharp, P.A. MicroRNA sponges: Competitive inhibitors of small RNAs in mammalian cells. Nat. Methods 2007, 4, 721–726. [Google Scholar] [CrossRef]
- Thum, T.; Gross, C.; Fiedler, J.; Fischer, T.; Kissler, S.; Bussen, M.; Galuppo, P.; Just, S.; Rottbauer, W.; Frantz, S.; et al. MicroRNA-21 contributes to myocardial disease by stimulating MAP kinase signalling in fibroblasts. Nature 2008, 456, 980–984. [Google Scholar] [CrossRef]
- Zhao, Y.; Samal, E.; Srivastava, D. Serum response factor regulates a muscle-specific microRNA that targets Hand2 during cardiogenesis. Nature 2005, 436, 214–220. [Google Scholar] [CrossRef]
- Montgomery, R.L.; Hullinger, T.G.; Semus, H.M.; Dickinson, B.A.; Seto, A.G.; Lynch, J.M.; Stack, C.; Latimer, P.A.; Olson, E.N.; van Rooij, E. Therapeutic inhibition of miR-208a improves cardiac function and survival during heart failure. Circulation 2011, 124, 1537–1547. [Google Scholar] [CrossRef] [PubMed]
- Shan, S.W.; Lee, D.Y.; Deng, Z.; Shatseva, T.; Jeyapalan, Z.; Du, W.W.; Zhang, Y.; Xuan, J.W.; Yee, S.-P.; Siragam, V.; et al. MicroRNA MiR-17 retards tissue growth and represses fibronectin expression. Nat. Cell Biol. 2009, 11, 1031–1038. [Google Scholar] [CrossRef] [PubMed]
- Peddibhotla, S.; Rosen, J.M. Chking and executing cell division to prevent genomic instability. Cell Cycle 2009, 8, 2339–2342. [Google Scholar] [CrossRef] [PubMed]
- Dirkx, E.; Gladka, M.M.; Philippen, L.E.; Armand, A.-S.; Kinet, V.; Leptidis, S.; el Azzouzi, H.; Salic, K.; Bourajjaj, M.; da Silva, G.J.J.; et al. Nfat and miR-25 cooperate to reactivate the transcription factor Hand2 in heart failure. Nat. Cell Biol. 2013, 15, 1282–1293. [Google Scholar] [CrossRef] [PubMed]
- Katz, M.G.; Fargnoli, A.S.; Kendle, A.P.; Hajjar, R.J.; Bridges, C.R. The role of microRNAs in cardiac development and regenerative capacity. Am. J. Physiol. Circ. Physiol. 2016, 310, H528–H541. [Google Scholar] [CrossRef] [PubMed]
- Eulalio, A.; Mano, M.; Ferro, M.D.; Zentilin, L.; Sinagra, G.; Zacchigna, S.; Giacca, M. Functional screening identifies miRNAs inducing cardiac regeneration. Nature 2012, 492, 376–381. [Google Scholar] [CrossRef] [PubMed]
- Aguirre, A.; Montserrat, N.; Zacchigna, S.; Nivet, E.; Hishida, T.; Krause, M.N.; Kurian, L.; Ocampo, A.; Vazquez-Ferrer, E.; Rodriguez-Esteban, C.; et al. In vivo activation of a conserved microRNA program induces mammalian heart regeneration. Cell Stem Cell 2014, 15, 589–604. [Google Scholar] [CrossRef] [PubMed]
- Hosoda, T.; Zheng, H.; Cabral-da-Silva, M.; Sanada, F.; Ide-Iwata, N.; Ogórek, B.; Ferreira-Martins, J.; Arranto, C.; D’Amario, D.; del Monte, F.; et al. Human cardiac stem cell differentiation is regulated by a mircrine mechanism. Circulation 2011, 123, 1287–1296. [Google Scholar] [CrossRef]
- Hullinger, T.G.; Montgomery, R.L.; Seto, A.G.; Dickinson, B.A.; Semus, H.M.; Lynch, J.M.; Dalby, C.M.; Robinson, K.; Stack, C.; Latimer, P.A.; et al. Inhibition of miR-15 protects against cardiac ischemic injury. Circ. Res. 2012, 110, 71–81. [Google Scholar] [CrossRef]
- Caruso, P.; Dempsie, Y.; Stevens, H.C.; McDonald, R.A.; Long, L.; Lu, R.; White, K.; Mair, K.M.; McClure, J.D.; Southwood, M.; et al. A role for miR-145 in pulmonary arterial hypertension. Circ. Res. 2012, 111, 290–300. [Google Scholar] [CrossRef]
- Ren, X.-P.; Wu, J.; Wang, X.; Sartor, M.A.; Qian, J.; Jones, K.; Nicolaou, P.; Pritchard, T.J.; Fan, G.-C. MicroRNA-320 is involved in the regulation of cardiac ischemia/reperfusion injury by targeting heat-shock protein 20. Circulation 2009, 119, 2357–2366. [Google Scholar] [CrossRef] [PubMed]
- Friese, R.S.; Altshuler, A.E.; Zhang, K.; Miramontes-Gonzalez, J.P.; Hightower, C.M.; Jirout, M.L.; Salem, R.M.; Gayen, J.R.; Mahapatra, N.R.; Biswas, N.; et al. MicroRNA-22 and promoter motif polymorphisms at the Chga locus in genetic hypertension: Functional and therapeutic implications for gene expression and the pathogenesis of hypertension. Hum. Mol. Genet. 2013, 22, 3624–3640. [Google Scholar] [CrossRef] [PubMed]
- Danan, M.; Schwartz, S.; Edelheit, S.; Sorek, R. Transcriptome-wide discovery of circular RNAs in Archaea. Nucleic Acids Res. 2012, 40, 3131–3142. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.L.; Bao, Y.; Yee, M.-C.; Barrett, S.P.; Hogan, G.J.; Olsen, M.N.; Dinneny, J.R.; Brown, P.O.; Salzman, J. Circular RNA is expressed across the eukaryotic tree of life. PLoS ONE 2014, 9, e90859. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. CircRNA biogenesis competes with Pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Abdelmohsen, K.; Panda, A.C.; Munk, R.; Grammatikakis, I.; Dudekula, D.B.; De, S.; Kim, J.; Noh, J.H.; Kim, K.M.; Martindale, J.L.; et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol. 2017, 14, 361–369. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef]
- Pamudurti, N.R.; Bartok, O.; Jens, M.; Ashwal-Fluss, R.; Stottmeister, C.; Ruhe, L.; Hanan, M.; Wyler, E.; Perez-Hernandez, D.; Ramberger, E.; et al. Translation of CircRNAs. Mol. Cell 2017, 66, 9–21. [Google Scholar] [CrossRef]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 is a circular RNA that can be translated and functions in myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Zhang, M.; Zheng, X.; Yi, P.; Lan, C.; Xu, M. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2017, 143, 17–27. [Google Scholar] [CrossRef] [PubMed]
- You, X.; Vlatkovic, I.; Babic, A.; Will, T.; Epstein, I.; Tushev, G.; Akbalik, G.; Wang, M.; Glock, C.; Quedenau, C.; et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 2015, 18, 603–610. [Google Scholar] [CrossRef] [PubMed]
- Zhou, M.; Yang, J.-M.; Xiong, X. The emerging landscape of circular RNA in cardiovascular diseases. J. Mol. Cell. Cardiol. 2018, 122, 134–139. [Google Scholar] [CrossRef] [PubMed]
- Werfel, S.; Nothjunge, S.; Schwarzmayr, T.; Strom, T.-M.; Meitinger, T.; Engelhardt, S. Characterization of circular RNAs in human, mouse and rat hearts. J. Mol. Cell. Cardiol. 2016, 98, 103–107. [Google Scholar] [CrossRef] [PubMed]
- Tan, W.L.W.; Lim, B.T.S.; Anene-Nzelu, C.G.O.; Ackers-Johnson, M.; Dashi, A.; See, K.; Tiang, Z.; Lee, D.P.; Chua, W.W.; Luu, T.D.A.; et al. A landscape of circular RNA expression in the human heart. Cardiovasc. Res. 2017, 113, 298–309. [Google Scholar] [CrossRef] [PubMed]
- Enuka, Y.; Lauriola, M.; Feldman, M.E.; Sas-Chen, A.; Ulitsky, I.; Yarden, Y. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acids Res. 2016, 44, 1370–1383. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Papavasileiou, P.; Peters, O.; Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS ONE 2015, 10, e0141214. [Google Scholar] [CrossRef]
- Li, T.; Shao, Y.; Fu, L.; Xie, Y.; Zhu, L.; Sun, W.; Yu, R.; Xiao, B.; Guo, J. Plasma circular RNA profiling of patients with gastric cancer and their droplet digital RT-PCR detection. J. Mol. Med. 2018, 96, 85–96. [Google Scholar] [CrossRef]
- Li, Y.; Zheng, Q.; Bao, C.; Li, S.; Guo, W.; Zhao, J.; Chen, D.; Gu, J.; He, X.; Huang, S. Circular RNA is enriched and stable in exosomes: A promising biomarker for cancer diagnosis. Cell Res. 2015, 25, 981–984. [Google Scholar] [CrossRef]
- Bahn, J.H.; Zhang, Q.; Li, F.; Chan, T.-M.; Lin, X.; Kim, Y.; Wong, D.T.W.; Xiao, X. the landscape of microRNA, piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 2015, 61, 221–230. [Google Scholar] [CrossRef] [PubMed]
- Wesselhoeft, R.A.; Kowalski, P.S.; Anderson, D.G. Engineering circular RNA for potent and stable translation in eukaryotic cells. Nat. Commun. 2018, 9, 2629. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Long, B.; Liu, F.; Wang, J.-X.; Liu, C.-Y.; Zhao, B.; Zhou, L.-Y.; Sun, T.; Wang, M.; Yu, T.; et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 2016, 37, 2602–2611. [Google Scholar] [CrossRef] [PubMed]
- Vausort, M.; Salgado-Somoza, A.; Zhang, L.; Leszek, P.; Scholz, M.; Teren, A.; Burkhardt, R.; Thiery, J.; Wagner, D.R.; Devaux, Y. Myocardial infarction-associated circular RNA predicting left ventricular dysfunction. J. Am. Coll. Cardiol. 2016, 68, 1247–1248. [Google Scholar] [CrossRef] [PubMed]
- Geng, H.-H.; Li, R.; Su, Y.-M.; Xiao, J.; Pan, M.; Cai, X.-X.; Ji, X.-P. The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLoS ONE 2016, 11, e0151753. [Google Scholar] [CrossRef] [PubMed]
- Du, W.W.; Yang, W.; Chen, Y.; Wu, Z.-K.; Foster, F.S.; Yang, Z.; Li, X.; Yang, B.B. Foxo3 circular RNA promotes cardiac senescence by modulating multiple factors associated with stress and senescence responses. Eur. Heart J. 2016, 38, 1402–1412. [Google Scholar] [CrossRef] [PubMed]
- Boeckel, J.-N.; Jaé, N.; Heumüller, A.W.; Chen, W.; Boon, R.A.; Stellos, K.; Zeiher, A.M.; John, D.; Uchida, S.; Dimmeler, S. Identification and characterization of hypoxia-regulated endothelial circular RNA. Circ. Res. 2015, 117, 884–890. [Google Scholar] [CrossRef] [PubMed]
- Holdt, L.M.; Stahringer, A.; Sass, K.; Pichler, G.; Kulak, N.A.; Wilfert, W.; Kohlmaier, A.; Herbst, A.; Northoff, B.H.; Nicolaou, A.; et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 2016, 7, 12429. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.R.; Baccarella, A.; Parrish, J.Z.; Kim, C.C. Empirical assessment of analysis workflows for differential expression analysis of human samples using RNA-Seq. BMC Bioinformatics 2017, 18, 38. [Google Scholar] [CrossRef]
- Housman, G.; Ulitsky, I. Methods for distinguishing between protein-coding and long noncoding RNAs and the elusive biological purpose of translation of long noncoding RNAs. Biochim. Biophys. Acta 2016, 1859, 31–40. [Google Scholar] [CrossRef]
- Zeng, X.; Lin, W.; Guo, M.; Zou, Q. A comprehensive overview and evaluation of circular RNA detection tools. PLoS Comput. Biol. 2017, 13, e1005420. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Venø, M.T.; Damgaard, C.K.; Kjems, J. Comparison of circular RNA prediction tools. Nucleic Acids Res. 2016, 44, e58. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Metge, F.; Dieterich, C. Specific identification and quantification of circular RNAs from sequencing data. Bioinformatics 2016, 32, 1094–1096. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Xie, X.; Zhou, J.; Sheng, M.; Yin, X.; Ko, E.-A.; Zhou, T.; Gu, W. Quantifying circular RNA expression from RNA-seq data using model-based framework. Bioinformatics 2017, 33, 2131–2139. [Google Scholar] [CrossRef] [PubMed]
- Rühle, F.; Stoll, M. Long non-coding RNA Databases in Cardiovascular Research. Genomics Proteomics Bioinformatics 2016, 14, 191–199. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.J.D.; Kim, V.; Muth, D.C.; Witwer, K.W. Validated microRNA target databases: An evaluation. Drug Dev. Res. 2015, 76, 389–396. [Google Scholar] [PubMed]
- Zhou, B.; Zhao, H.; Yu, J.; Guo, C.; Dou, X.; Song, F.; Hu, G.; Cao, Z.; Qu, Y.; Yang, Y.; et al. EVLncRNAs: A manually curated database for long non-coding RNAs validated by low-throughput experiments. Nucleic Acids Res. 2018, 46, D100–D105. [Google Scholar] [CrossRef] [PubMed]
- Pian, C.; Zhang, G.; Tu, T.; Ma, X.; Li, F. LncCeRBase: A database of experimentally validated human competing endogenous long non-coding RNAs. Database 2018, 2018, bay061. [Google Scholar] [CrossRef] [PubMed]
- Kaikkonen, M.U.; Adelman, K. Emerging roles of non-coding RNA transcription. Trends Biochem. Sci. 2018, 43, 654–667. [Google Scholar] [CrossRef]
- Salehi, S.; Taheri, M.N.; Azarpira, N.; Zare, A.; Behzad-Behbahani, A. State of the art technologies to explore long non-coding RNAs in cancer. J. Cell. Mol. Med. 2017, 21, 3120–3140. [Google Scholar] [CrossRef]
- Kashi, K.; Henderson, L.; Bonetti, A.; Carninci, P. Discovery and functional analysis of lncRNAs: Methodologies to investigate an uncharacterized transcriptome. Biochim. Biophys. Acta 2016, 1859, 3–15. [Google Scholar] [CrossRef] [PubMed]
- Bejerano, G.; Pheasant, M.; Makunin, I.; Stephen, S.; Kent, W.J.; Mattick, J.S.; Haussler, D. Ultraconserved elements in the human genome. Science 2004, 304, 1321–1325. [Google Scholar] [CrossRef] [PubMed]
- Calin, G.A.; Liu, C.; Ferracin, M.; Hyslop, T.; Spizzo, R.; Sevignani, C.; Fabbri, M.; Cimmino, A.; Lee, E.J.; Wojcik, S.E.; et al. Ultraconserved regions encoding ncRNAs are altered in human leukemias and carcinomas. Cancer Cell 2007, 12, 215–229. [Google Scholar] [CrossRef] [PubMed]
- Pollard, K.S.; Salama, S.R.; Lambert, N.; Lambot, M.-A.; Coppens, S.; Pedersen, J.S.; Katzman, S.; King, B.; Onodera, C.; Siepel, A.; et al. An RNA gene expressed during cortical development evolved rapidly in humans. Nature 2006, 443, 167–172. [Google Scholar] [CrossRef] [PubMed]
- Altenhoff, A.M.; Dessimoz, C. Inferring orthology and paralogy. Methods Mol. Biol. 2012, 855, 259–279. [Google Scholar] [PubMed]
- Altenhoff, A.M.; Boeckmann, B.; Capella-Gutierrez, S.; Dalquen, D.A.; DeLuca, T.; Forslund, K.; Huerta-Cepas, J.; Linard, B.; Pereira, C.; Pryszcz, L.P.; et al. Standardized benchmarking in the quest for orthologs. Nat. Methods 2016, 13, 425–430. [Google Scholar] [CrossRef] [PubMed]
- Hulsen, T.; Huynen, M.A.; de Vlieg, J.; Groenen, P.M. Benchmarking ortholog identification methods using functional genomics data. Genome Biol. 2006, 7, R31. [Google Scholar] [CrossRef] [PubMed]
- Chen, F.; Mackey, A.J.; Vermunt, J.K.; Roos, D.S. Assessing performance of orthology detection strategies applied to eukaryotic genomes. PLoS ONE 2007, 2, e383. [Google Scholar] [CrossRef]
- Gutschner, T.; Hämmerle, M.; Diederichs, S. MALAT1—A paradigm for long noncoding RNA function in cancer. J. Mol. Med. 2013, 91, 791–801. [Google Scholar] [CrossRef]
- Washietl, S.; Kellis, M.; Garber, M. Evolutionary dynamics and tissue specificity of human long noncoding RNAs in six mammals. Genome Res. 2014, 24, 616–628. [Google Scholar] [CrossRef]
- Necsulea, A.; Soumillon, M.; Warnefors, M.; Liechti, A.; Daish, T.; Zeller, U.; Baker, J.C.; Grützner, F.; Kaessmann, H. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 2014, 505, 635–640. [Google Scholar] [CrossRef] [PubMed]
- Hezroni, H.; Koppstein, D.; Schwartz, M.G.; Avrutin, A.; Bartel, D.P.; Ulitsky, I. Principles of long noncoding RNA evolution derived from direct comparison of transcriptomes in 17 species. Cell Rep. 2015, 11, 1110–1122. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Ezquerra, A.; Harrison, M.C.; Bornberg-Bauer, E. Comparative analysis of lincRNA in insect species. BMC Evol. Biol. 2017, 17, 155. [Google Scholar] [CrossRef] [PubMed]
- Gardner, P.P.; Fasold, M.; Burge, S.W.; Ninova, M.; Hertel, J.; Kehr, S.; Steeves, T.E.; Griffiths-Jones, S.; Stadler, P.F. Conservation and losses of non-coding RNAs in avian genomes. PLoS ONE 2015, 10, e0121797. [Google Scholar] [CrossRef] [PubMed]
- Engreitz, J.M.; Haines, J.E.; Perez, E.M.; Munson, G.; Chen, J.; Kane, M.; McDonel, P.E.; Guttman, M.; Lander, E.S. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 2016, 539, 452–455. [Google Scholar] [CrossRef] [PubMed]
- Kalvari, I.; Nawrocki, E.P.; Argasinska, J.; Quinones-Olvera, N.; Finn, R.D.; Bateman, A.; Petrov, A.I. Non-coding RNA analysis using the Rfam database. Curr. Protoc. Bioinformatics 2018, 62, e51. [Google Scholar] [CrossRef] [PubMed]
- Smith, M.A.; Gesell, T.; Stadler, P.F.; Mattick, J.S. Widespread purifying selection on RNA structure in mammals. Nucleic Acids Res. 2013, 41, 8220–8236. [Google Scholar] [CrossRef] [PubMed]
- Rivas, E.; Clements, J.; Eddy, S.R. A statistical test for conserved RNA structure shows lack of evidence for structure in lncRNAs. Nat. Methods 2017, 14, 45–48. [Google Scholar] [CrossRef] [PubMed]
- Managadze, D.; Rogozin, I.B.; Chernikova, D.; Shabalina, S.A.; Koonin, E.V. Negative correlation between expression level and evolutionary rate of long intergenic noncoding RNAs. Genome Biol. Evol. 2011, 3, 1390–1404. [Google Scholar] [CrossRef] [PubMed]
- Tavares, R.C.A.; Pyle, A.M.; Somarowthu, S. Covariation analysis with improved parameters reveals conservation in lncRNA structures. bioRxiv. 2018. bioRxiv: 364109. Available online: https://doi.org/10.1101/364109 (accessed on 21 December 2018).
- Lu, Z.; Zhang, Q.C.; Lee, B.; Flynn, R.A.; Smith, M.A.; Robinson, J.T.; Davidovich, C.; Gooding, A.R.; Goodrich, K.J.; Mattick, J.S.; et al. RNA duplex map in living cells reveals higher-order transcriptome structure. Cell 2016, 165, 1267–1279. [Google Scholar] [CrossRef] [PubMed]
- Kino, T.; Hurt, D.E.; Ichijo, T.; Nader, N.; Chrousos, G.P. Noncoding RNA Gas5 is a growth arrest- and starvation-associated repressor of the glucocorticoid receptor. Sci. Signal. 2010, 3, ra8. [Google Scholar] [CrossRef] [PubMed]
- Somarowthu, S.; Legiewicz, M.; Chillón, I.; Marcia, M.; Liu, F.; Pyle, A.M. HOTAIR forms an intricate and modular secondary structure. Mol. Cell 2015, 58, 353–361. [Google Scholar] [CrossRef] [PubMed]
- Nitsche, A.; Stadler, P.F. Evolutionary clues in lncRNAs. WIREs RNA 2017, 8, 1376. [Google Scholar] [CrossRef] [PubMed]
- Washietl, S.; Hofacker, I.L.; Lukasser, M.; Hüttenhofer, A.; Stadler, P.F. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat. Biotechnol. 2005, 23, 1383–1390. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, J.S.; Bejerano, G.; Siepel, A.; Rosenbloom, K.; Lindblad-Toh, K.; Lander, E.S.; Kent, J.; Miller, W.; Haussler, D. Identification and classification of conserved RNA secondary structures in the human genome. PLoS Comput. Biol. 2006, 2, e33. [Google Scholar] [CrossRef] [PubMed]
- Washietl, S.; Pedersen, J.S.; Korbel, J.O.; Stocsits, C.; Gruber, A.R.; Hackermuller, J.; Hertel, J.; Lindemeyer, M.; Reiche, K.; Tanzer, A.; et al. Structured RNAs in the ENCODE selected regions of the human genome. Genome Res. 2007, 17, 852–864. [Google Scholar] [CrossRef] [PubMed]
- Torarinsson, E.; Yao, Z.; Wiklund, E.D.; Bramsen, J.B.; Hansen, C.; Kjems, J.; Tommerup, N.; Ruzzo, W.L.; Gorodkin, J. Comparative genomics beyond sequence-based alignments: RNA structures in the ENCODE regions. Genome Res. 2008, 18, 242–251. [Google Scholar] [CrossRef] [PubMed]
- Rabani, M.; Kertesz, M.; Segal, E. Computational prediction of RNA structural motifs involved in post-transcriptional regulatory processes. In Methods in Molecular Biology (Clifton, N.J.); Humana Press: New York, NY, USA, 2011; Volume 714, pp. 467–479. [Google Scholar]
- Parker, B.J.; Moltke, I.; Roth, A.; Washietl, S.; Wen, J.; Kellis, M.; Breaker, R.; Pedersen, J.S. New families of human regulatory RNA structures identified by comparative analysis of vertebrate genomes. Genome Res. 2011, 21, 1929–1943. [Google Scholar] [CrossRef]
- Will, S.; Yu, M.; Berger, B. Structure-based whole-genome realignment reveals many novel noncoding RNAs. Genome Res. 2013, 23, 1018–1027. [Google Scholar] [CrossRef] [PubMed]
- Seemann, S.E.; Mirza, A.H.; Hansen, C.; Bang-Berthelsen, C.H.; Garde, C.; Christensen-Dalsgaard, M.; Torarinsson, E.; Yao, Z.; Workman, C.T.; Pociot, F.; et al. The identification and functional annotation of RNA structures conserved in vertebrates. Genome Res. 2017, 27, 1371–1383. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Tang, Y.; Kwok, C.K.; Zhang, Y.; Bevilacqua, P.C.; Assmann, S.M. In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features. Nature 2014, 505, 696–700. [Google Scholar] [CrossRef] [PubMed]
- Rouskin, S.; Zubradt, M.; Washietl, S.; Kellis, M.; Weissman, J.S. Genome-wide probing of RNA structure reveals active unfolding of mRNA structures in vivo. Nature 2014, 505, 701–705. [Google Scholar] [CrossRef] [PubMed]
- Wan, Y.; Qu, K.; Zhang, Q.C.; Flynn, R.A.; Manor, O.; Ouyang, Z.; Zhang, J.; Spitale, R.C.; Snyder, M.P.; Segal, E.; et al. Landscape and variation of RNA secondary structure across the human transcriptome. Nature 2014, 505, 706–709. [Google Scholar] [CrossRef] [PubMed]
- Aw, J.G.A.; Shen, Y.; Wilm, A.; Sun, M.; Lim, X.N.; Boon, K.-L.; Tapsin, S.; Chan, Y.-S.; Tan, C.-P.; Sim, A.Y.L.; et al. In vivo mapping of eukaryotic RNA interactomes reveals principles of higher-order organization and regulation. Mol. Cell 2016, 62, 603–617. [Google Scholar] [CrossRef] [PubMed]
- Strobel, E.J.; Yu, A.M.; Lucks, J.B. High-throughput determination of RNA structures. Nat. Rev. Genet. 2018, 19, 615–634. [Google Scholar] [CrossRef] [PubMed]
- Kutter, C.; Watt, S.; Stefflova, K.; Wilson, M.D.; Goncalves, A.; Ponting, C.P.; Odom, D.T.; Marques, A.C. Rapid turnover of long noncoding RNAs and the evolution of gene expression. PLoS Genet. 2012, 8, e1002841. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Shishkin, A.A.; Zhu, X.; Kadri, S.; Maza, I.; Guttman, M.; Hanna, J.H.; Regev, A.; Garber, M. Evolutionary analysis across mammals reveals distinct classes of long non-coding RNAs. Genome Biol. 2016, 17, 19. [Google Scholar] [CrossRef]
- Hezroni, H.; Ben-Tov Perry, R.; Meir, Z.; Housman, G.; Lubelsky, Y.; Ulitsky, I. A subset of conserved mammalian long non-coding RNAs are fossils of ancestral protein-coding genes. Genome Biol. 2017, 18, 162. [Google Scholar] [CrossRef] [PubMed]
- Ning, Q.; Li, Y.; Wang, Z.; Zhou, S.; Sun, H.; Yu, G. The evolution and expression pattern of human overlapping lncRNA and protein-coding gene pairs. Sci. Rep. 2017, 7, 42775. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Orera, J.; Messeguer, X.; Subirana, J.A.; Alba, M.M. Long non-coding RNAs as a source of new peptides. Elife 2014, 3, e03523. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.-Y.; Shen, Q.S.; Zhou, W.-Z.; Peng, J.; He, B.Z.; Li, Y.; Liu, C.-J.; Luan, X.; Ding, W.; Li, S.; et al. Emergence, retention and selection: A trilogy of origination for functional de novo proteins from ancestral LncRNAs in primates. PLoS Genet. 2015, 11, e1005391. [Google Scholar] [CrossRef] [PubMed]
- Villegas, V.; Zaphiropoulos, P. Neighboring gene regulation by antisense long non-coding RNAs. Int. J. Mol. Sci. 2015, 16, 3251–3266. [Google Scholar] [CrossRef] [PubMed]
- Amaral, P.P.; Leonardi, T.; Han, N.; Viré, E.; Gascoigne, D.K.; Arias-Carrasco, R.; Büscher, M.; Pandolfini, L.; Zhang, A.; Pluchino, S.; et al. Genomic positional conservation identifies topological anchor point RNAs linked to developmental loci. Genome Biol. 2018, 19, 32. [Google Scholar] [CrossRef] [PubMed]
- Brawand, D.; Soumillon, M.; Necsulea, A.; Julien, P.; Csárdi, G.; Harrigan, P.; Weier, M.; Liechti, A.; Aximu-Petri, A.; Kircher, M.; et al. The evolution of gene expression levels in mammalian organs. Nature 2011, 478, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Singh, A.R.; Sivadas, A.; Sabharwal, A.; Vellarikal, S.K.; Jayarajan, R.; Verma, A.; Kapoor, S.; Joshi, A.; Scaria, V.; Sivasubbu, S. Chamber specific gene expression landscape of the zebrafish heart. PLoS ONE 2016, 11, e0147823. [Google Scholar] [CrossRef]
- Johnson, E.K.; Matkovich, S.J.; Nerbonne, J.M. Regional differences in mRNA and lncRNA expression profiles in non-failing human atria and ventricles. Sci. Rep. 2018, 8, 13919. [Google Scholar] [CrossRef]
- van den Berg, C.W.; Okawa, S.; Chuva de Sousa Lopes, S.M.; van Iperen, L.; Passier, R.; Braam, S.R.; Tertoolen, L.G.; del Sol, A.; Davis, R.P.; Mummery, C.L. Transcriptome of human foetal heart compared with cardiomyocytes from pluripotent stem cells. Development 2015, 142, 3231–3238. [Google Scholar] [CrossRef]
- Kahr, P.C.; Piccini, I.; Fabritz, L.; Greber, B.; Schöler, H.; Scheld, H.H.; Hoffmeier, A.; Brown, N.A.; Kirchhof, P. Systematic analysis of gene expression differences between left and right atria in different mouse strains and in human atrial tissue. PLoS ONE 2011, 6, e26389. [Google Scholar] [CrossRef]
- Hsu, J.; Hanna, P.; Van Wagoner, D.R.; Barnard, J.; Serre, D.; Chung, M.K.; Smith, J.D. Whole genome expression differences in human left and right atria ascertained by RNA sequencing. Circ. Cardiovasc. Genet. 2012, 5, 327–335. [Google Scholar] [CrossRef]
| Transcript | Host Gene | Organism Studied | Mechanism | Disease | Reference |
|---|---|---|---|---|---|
| Anril | CDKN2B-AS1 | Human | Binds to CBX7, recruits PRC-1 and PRC-2 to INK4 locus-leading to the repression of p15 and p16 transcription | Genetic risk factor for coronary artery disease (CAD) and myocardial infarction (MI) | [24] |
| BVHT | BVHT | Mouse | Activation of mesoderm posterior 1 (MesP1) and interacts with SUZ12, a component of PRC2, during cardiomyocyte differentiation | Impairs cardiomyocyte differentiation | [25] |
| FENDRR | FENDRR | Human, Mouse, Rat | Binds to the histone-remodeling PRC2 complex and TrxG/MLL to modulate chromatin status | Low expression leads to cardiac hypoplasia | [26] |
| NOVLNC6 | Intergenic | Mouse | Modulates expression of MKX2.5 | Downregulated in dilated cardiomyopathy (DCM) | [27] |
| CARMEN | CARMN | Human, Mouse, Rat | Interacts with SUZ12 and EZH2 of PRC2 complex | Plays a critical role in maintaining a differentiated cardiac fate in mature cardiomyocytes in case of DCM and aortic stenosis (AOS) | [28] |
| KCNQ1OT1 | KCNQ1OT1 | Human, Mouse | Kcnq1 imprinted domain in heart development | Defects in KCNQ1 leads to cardiac arrhythmias, predicts left ventricular dysfunction | [29] |
| SENCR | SENCR | Human | Inhibitor of smooth muscle cell migration | Downregulated in CAD and MI | [30] |
| MALAT1 | MALAT1 | Human, Mouse | - | Involvement in the pathogenesis of diabetic cardiomyopathy | [31] |
| H19 | H19 | Human | Acts by targeting VDAC1 | Regulates cardiomyocyte apoptosis in diabetic cardiomyopathy | [32] |
| RNCR3 | RNCR3/LINC00599 | Human, Mouse | miR-185-5p sponge | RNCR3 is athero-protective | [33] |
| CHAER | CHAER1/GM42105 | Mouse | Interacts with PRC2 | Inhibition of Chaer expression in the heart before, but not after, the onset of pressure overload substantially attenuates cardiac hypertrophy and dysfunction | [34] |
| LIPCAR | JA760602 | Human | - | Elevated in patients with chronic heart failure | [35] |
| MIAT | MIAT/RNCR2 | Human | MIAT functioned as a ceRNA for miR-24 to modulate Furin and TGF-β1 expression | Involved in pathological angiogenesis and is suggested as a predictor of MI | [36] |
| MHRT | MHRT | Human | - | Protective factor for cardiomyocyte | [37] |
| GAS5 | GAS5 | Human, Mouse, Rat | Interacts with miR-290, Inhibits nuclear translocation of beta-catenin, inducing expression of downstream genes | GAS5 knockdown aggravate hypertension-induced microvascular dysfunction | [38] |
| MEG3 | MEG3 | Mouse | MEG3 directly binds with the p53 DNA binding domain | MEG3 is upregulated following ischemia and stroke | [39] |
| UCA1 | UCA1 | Human | Inhibit the expression of p27 | Upregulated in the plasma of patients after MI | [40] |
| HIF1A-AS1 | HIF1A-AS1 | Human | - | Plays an important role in the pathogenesis of cardiovascular disease (CVD) | [41] |
| NPPA-AS1 | NPPA | Human | Alternative splicing of the NPPA gene | Involved in CVD | [42] |
| CHRF | DCC | Human | Targeting miR-489 | Regulates cardiac hypertrophy | [43] |
| CHAST | - | Mouse, Human | CHAST negatively regulated Pleckstrin homology domain–containing protein family M member 1 | Potential target to prevent cardiac remodeling | [44] |
| PANCR | PITX2 | Human | miR-143 and miR-501 sponge | Affected in atrial fibrillation (AF) | [45] |
| PVT1 | PVT1 | Mouse | Essential for the maintenance of cell size of cardiomyocytes | Regulation of cardiac hypertrophy | [46] |
| Carl | CASC11 | Human | Targeting miR-539 and PHB2 | Regulates mitochondrial fission and apoptosis in MI | [47] |
| HOTAIR | HOTAIR | Human | Targets expression of NOX2 | Upregulated in ischemic heart failure | [48] |
| Transcript | Organism Studied | Mechanism | Disease | Reference |
|---|---|---|---|---|
| miR-133 | Mouse, Human | Targets HAND-2, de-repression of IRX5 | Regulates the balance between differentiation and proliferation during cardiogenesis | [52] |
| miR-208a | Mouse, Human | Regulates the balance between the a- and b-myosin heavy chains | MiR-208 inhibition is protective in heart failure | [53] |
| miR-17 | Mouse | target genes are BIM, BMP 2/4, ISL1, TBX1 | Regulation of cardiac progenitor genes, repression of fibronectin | [54] |
| miR-195 | Human | CHEK1 regulation | Associated with ventricular septal defect and right ventricular hypoplasia | [55] |
| miR-25 | Mouse, Human | Regulation of cardiac contractility through control of SERCA2a expression | Mitigation of heart failure | [56] |
| miR-302/367 | Mouse, Human | - | Role in differentiation and reprogramming in cardiac remodeling | [57] |
| miR-590 | Mouse, Rat | - | Stimulates cardiac regeneration | [58] |
| miR-99a | Mouse, Zebrafish | Regulates FNTB, SMARCA5 expression | Regulate cardiac regeneration | [59] |
| miR-499 | Mouse, Human | Regulates SOX6, ROD1, MYH7B | Reduces cell proliferation and enhances myocyte differentiation | [60] |
| miR-15a/b | Mouse, Pigs | Targets BCL2 and ARL2 in cardiomyocytes | Reduced infarct size and cardiac remodeling and enhances cardiac function in response to myocardial infarction (MI) | [61] |
| miR-145 | Mouse, Human | Regulates KLF4 and KLF5 expression | Prevented the development of pulmonary artery hypertension | [62] |
| miR-320a | Mouse, Rat | Reduced infarct size via antithetical regulation of heat-shock protein-20 | Potential therapeutic target for ischemic heart disease | [63] |
| miR-22 | Human, Mouse, Rat | Regulates MYH7 expression | Deregulated in human heart failure and also in animal models of cardiac hypertrophy and failure | [64] |
| miR-21 | Mouse | Regulates ER-MAP kinase activity | In vivo silencing of miR-21 in rodent model of cardiac fibrosis impacts global cardiac structure and ameliorates cardiac dysfunction | [51] |
| Transcript | Host Gene | Organism Studied | Mechanism | Disease | Reference |
|---|---|---|---|---|---|
| HRCR | PWWP2A | Mouse | miRNA sponge for miR-223 | Inhibits hypertrophic cardiomyopathy and heart failure | [85] |
| MICRA | ZNF609 | Human | - | Downregulated in heart failure | [86] |
| CDR1AS | CDR1 | Mouse | miRNA sponge for miR-7 | Upregulated in myocardial infarction | [87] |
| circFoxo3 | FOXO3 | Mouse | Retains ageing factors ID-1, E2F1, FAK, and HIF1α in cytoplasm | Upregulated in myocardial senescence | [88] |
| cZNF292 | ZNF292 | Human | - | Promotes angiogenesis | [89] |
| circANRIL | CDKN2B-AS1 | Human | Binds to PES1 | Protects against atherosclerosis | [90] |
| Technique | Used for | Throughput |
|---|---|---|
| Microarrays | Quantification of transcript expression | High |
| Serial analysis of gene expression (SAGE) | Transcript identification and quantification of expression | High |
| Next generation sequencing (NGS)-based transcriptome analysis methods (RNA-seq—RNA sequencing, CAGE—Cap Analysis of Gene Expression, GRO-Seq—Genomic run-on sequencing, etc.) | Transcript identification and quantification of expression | High |
| Quantitative RT-PCR—Real time- polymerase chain reaction | Validation of transcript existence and abundance in real time | Low |
| RNA-Fluorescence in-situ hybridization (RNA-FISH) | Transcript Localization in the cellular compartment and relative abundance | Low |
| Northern Blot | transcript size, the observation of alternate splice products, the use of probes with partial homology, the quality and quantity of transcript | Low |
| RNA immunoprecipitation (RIP) | RNA–protein interaction | Moderate |
| Crosslinking immunoprecipitation sequencing (CLIP-Seq) based methods (HITS-CLIP—High-throughput sequencing, PAR-CLIP—photoactivatable ribonucleoside, iCLIP—individual-nucleotide resolution etc.) | RNA–Protein interaction | High |
| Chromatin isolation by RNA purification (ChIRP) | RNA–DNA interaction | High |
| DNA-RNA fluorescence in-situ hybridization (FISH) | RNA–DNA interaction | Low |
| Capture hybridization analysis of RNA targets (CHART) | RNA–DNA interaction (localization in the genome) | Moderate |
| RNA antisense purification (RAP)-DNA | RNA–DNA interaction | High |
| RNA antisense purification (RAP)-RNA | lncRNA-RNA interactions that occur through protein intermediates or through direct RNA-RNA hybridization | High |
| Cross-linking, ligation and sequencing of hybrids (CLASH) | RNA–RNA interaction | High |
| Clustered regularly interspaced short palindromic repeats (CRISPR) based techniques | Overexpression/Knockdown, interactions, cellular compartment localization of particular transcript | Low |
| Transcription activator-like effector nucleases (TALENs) | Knockout/overexpression of the transcript | Low |
| Zinc-finger nucleases (ZFNs) | Knockout/overexpression of the transcript | Low |
| Antisense oligos (ASOs), Locked nucleic acids (LNAs) based methods | Silencing of transcript | Low |
| RNA interference (RNAi) | Silencing of transcript | Low |
| Luciferase Reporter Assays | Target sites of the transcript | Low |
| RNase protection assays | Transcription start-site localization | Low |
| Study | Species/Organisms | Details | Reference |
|---|---|---|---|
| Necsulea et al. | 11 vertebrates | RNA sequencing (RNA-seq) of multiple tissues | [113] |
| Washietl et al. | 6 mammals | RNA-seq of multiple tissues | [112] |
| PLAR | 17 vertebrates | RNA-seq of multiple tissues | [114] |
| Lopez-Ezquerra et al. | 7 insect species | Comparative analysis of long non-coding RNAs (lncRNAs) in insect species | [115] |
| Gardner et. al. | 48 avian species | Comparative analysis of non-coding RNAs (ncRNAs) in avian genomes | [116] |
| Study | Technique/Tool Used | Reference |
|---|---|---|
| Washietl et al., 2005 | RNAz | [127] |
| Pedersen et al., 2006 | EvoFold | [128] |
| Washietl et al., 2007 | AlifoldZ, RNAz, EvoFold | [129] |
| Torarinsson et al., 2008 | CMfinder, RNAz, EvoFold | [130] |
| Rabani et al., 2009 | RNApromo | [131] |
| Parker et al., 2011 | EvoFam | [132] |
| Smith et al., 2013 | RNAz and SISSIz | [119] |
| Will et al., 2013 | RE-Alignment for Prediction of structural ncRNA (REAPR) | [133] |
| Seemann et al., 2017 | CMfinder | [134] |
| Ding et al., 2014 | Structure-seq | [135] |
| Rouskin et al., 2014 | Dimethyl sulfate sequencing (DMS-seq) | [136] |
| Wan et al., 2014 | Parallel Analysis of RNA Structure (PARS) | [137] |
| Aw et al., 2016 | Sequencing of psoralen crosslinked, ligated, and selected hybrids (SPLASH) | [138] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gandhi, S.; Ruehle, F.; Stoll, M. Evolutionary Patterns of Non-Coding RNA in Cardiovascular Biology. Non-Coding RNA 2019, 5, 15. https://doi.org/10.3390/ncrna5010015
Gandhi S, Ruehle F, Stoll M. Evolutionary Patterns of Non-Coding RNA in Cardiovascular Biology. Non-Coding RNA. 2019; 5(1):15. https://doi.org/10.3390/ncrna5010015
Chicago/Turabian StyleGandhi, Shrey, Frank Ruehle, and Monika Stoll. 2019. "Evolutionary Patterns of Non-Coding RNA in Cardiovascular Biology" Non-Coding RNA 5, no. 1: 15. https://doi.org/10.3390/ncrna5010015
APA StyleGandhi, S., Ruehle, F., & Stoll, M. (2019). Evolutionary Patterns of Non-Coding RNA in Cardiovascular Biology. Non-Coding RNA, 5(1), 15. https://doi.org/10.3390/ncrna5010015





