Upregulation of Long Non-Coding RNA DRAIC Correlates with Adverse Features of Breast Cancer
Abstract
:1. Introduction
2. Results
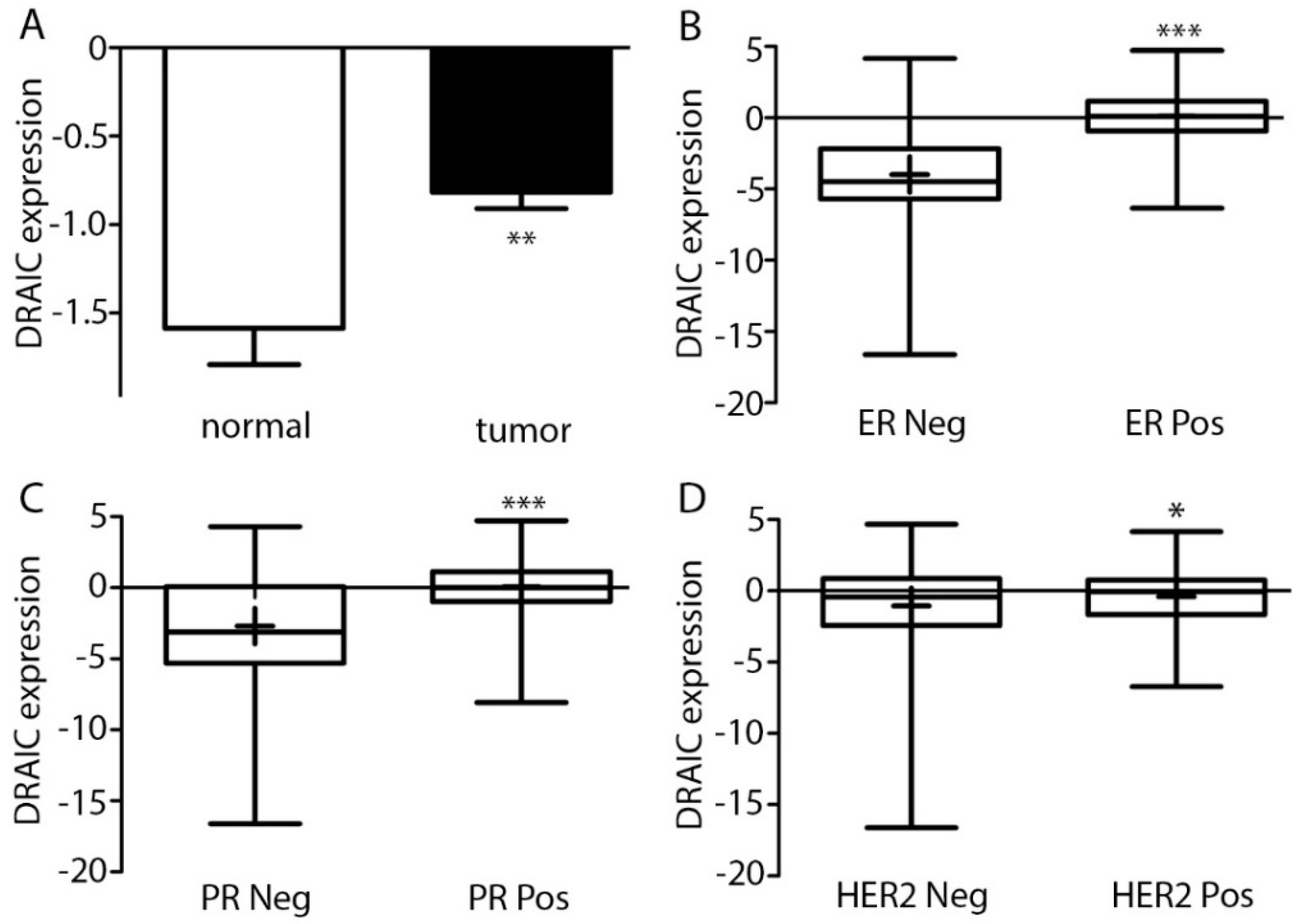
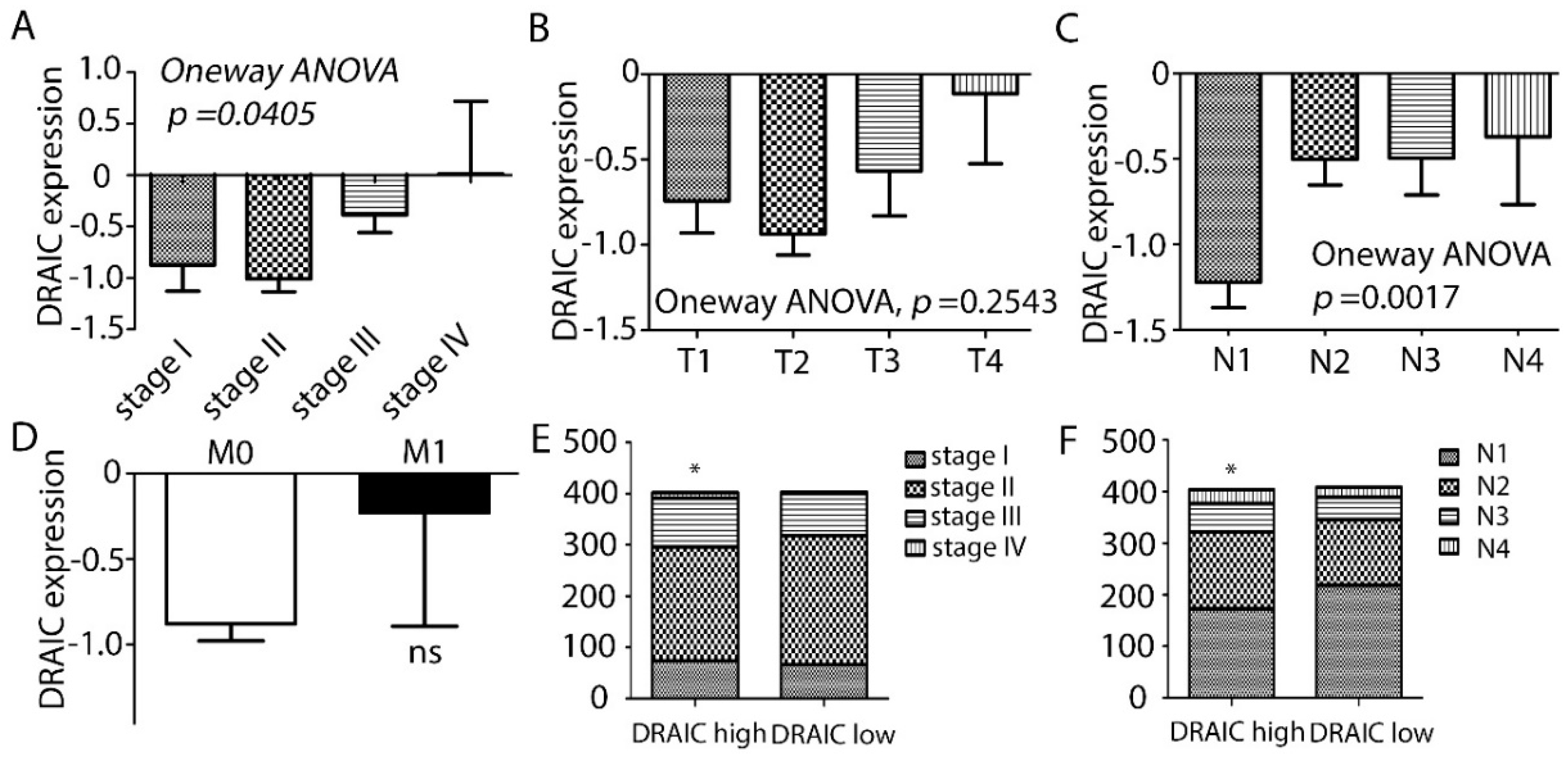
2.1. DRAIC Expression Increased in Breast Cancer and Correlated with Estrogen Receptor, Progesterone Receptor, and Human Epidermal Growth Factor 2 and Tumor Stages and Lymph Node Metastasis
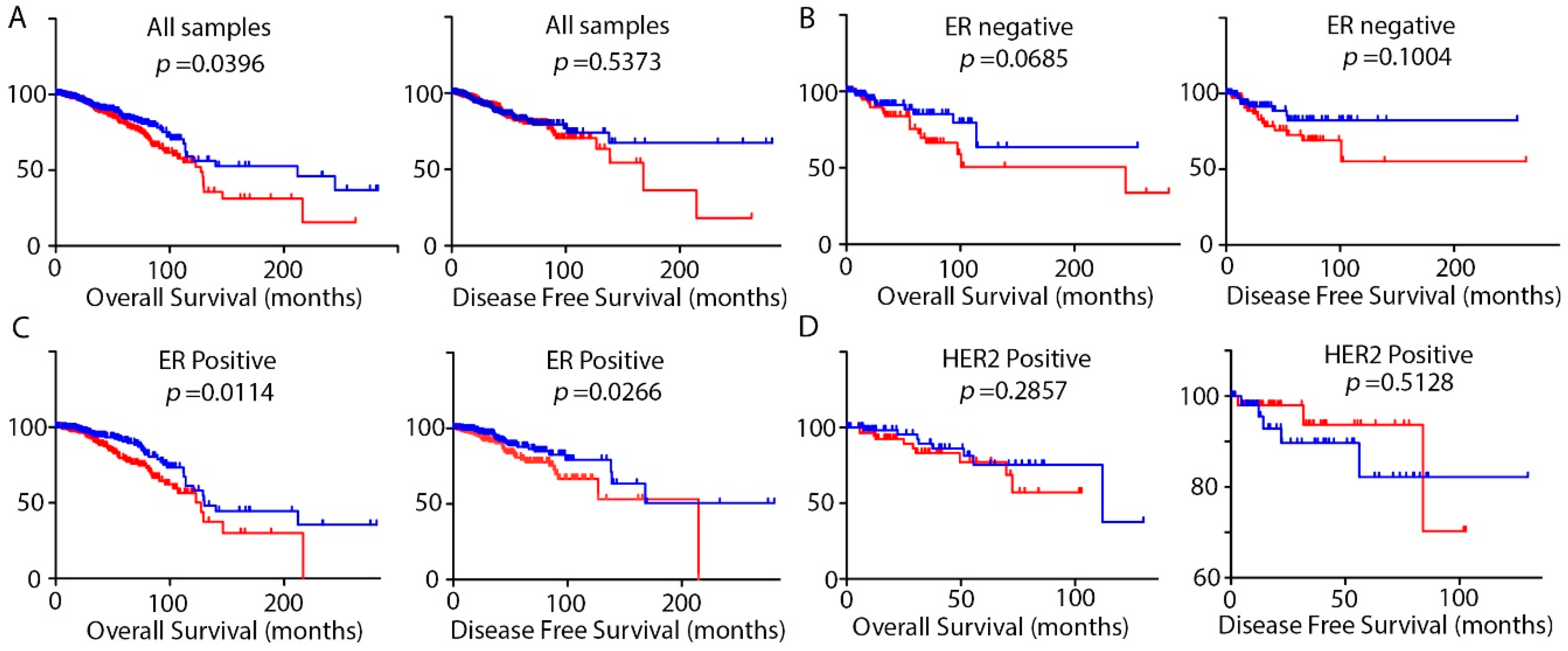
2.2. DRAIC Expression Impact Patients Survival
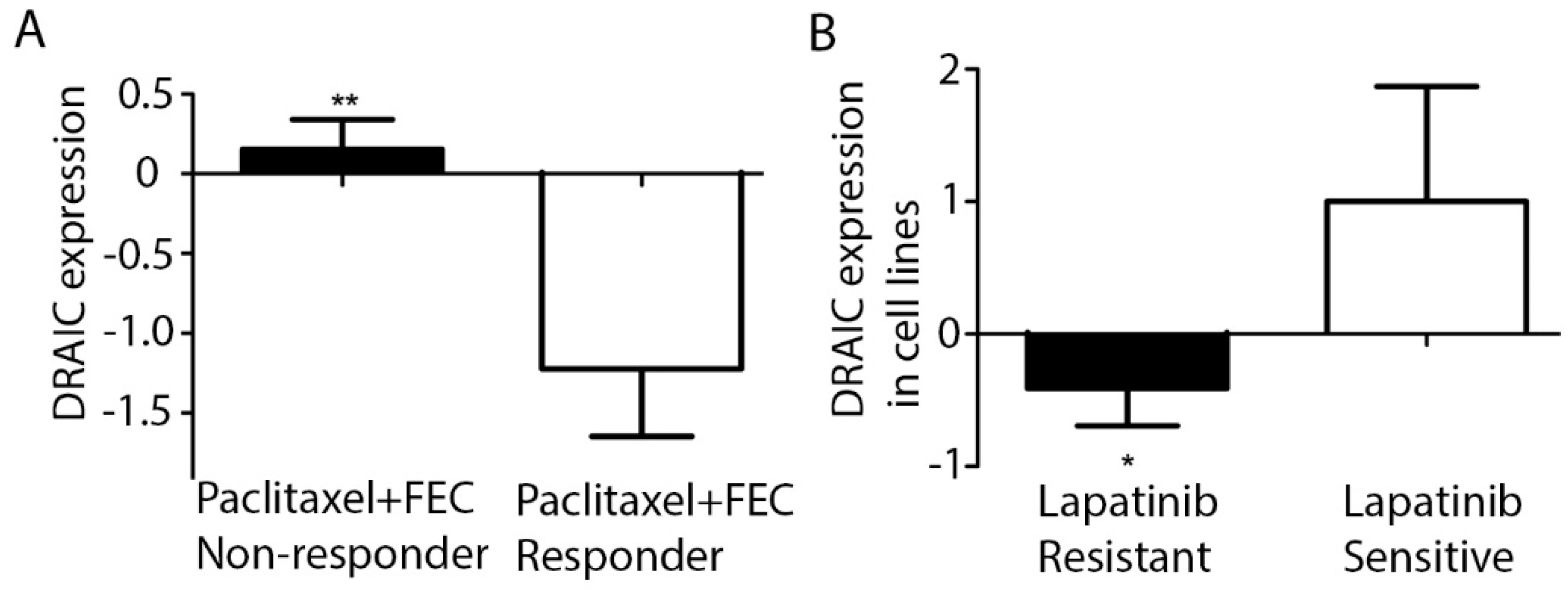
2.3. DRAIC Expression Predicts Patients’ Response to Chemotherapy Treatments
3. Discussion
4. Materials and Methods
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Ponting, C.P.; Oliver, P.L.; Reik, W. Evolution and functions of long noncoding RNAs. Cell 2009, 136, 629–641. [Google Scholar] [CrossRef] [PubMed]
- Wilusz, J.E.; Sunwoo, H.; Spector, D.L. Long noncoding RNAs: Functional surprises from the RNA world. Genes Dev. 2009, 23, 1494–1504. [Google Scholar] [CrossRef] [PubMed]
- Tsai, M.C.; Spitale, R.C.; Chang, H.Y. Long intergenic noncoding RNAs: New links in cancer progression. Cancer Res. 2011, 71, 3–7. [Google Scholar] [CrossRef] [PubMed]
- Wapinski, O.; Chang, H.Y. Long noncoding RNAs and human disease. Trends Cell Biol. 2011, 21, 354–361. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, A.M.; Chang, H.Y. Long noncoding RNAs in cancer pathways. Cancer Cell 2016, 29, 452–463. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Tudoran, O.M.; Calin, G.A.; Ivan, M. The many faces of long noncoding RNAs in cancer. Antioxid. Redox Signal. 2017. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.C.; Chang, H.Y. Molecular mechanisms of long noncoding RNAs. Mol. Cell 2011, 43, 904–914. [Google Scholar] [CrossRef] [PubMed]
- Nagano, T.; Fraser, P. No-nonsense functions for long noncoding RNAs. Cell 2011, 145, 178–181. [Google Scholar] [CrossRef]
- Sakurai, K.; Reon, B.J.; Anaya, J.; Dutta, A. The lncRNA DRAIC/PCAT29 locus constitutes a tumor-suppressive nexus. Mol. Cancer Res. 2015, 13, 828–838. [Google Scholar] [CrossRef]
- Sun, M.; Gadad, S.S.; Kim, D.S.; Kraus, W.L. Discovery, annotation, and functional analysis of long noncoding RNAs controlling cell-cycle gene expression and proliferation in breast cancer cells. Mol. Cell 2015, 59, 698–711. [Google Scholar] [CrossRef]
- Eroles, P.; Bosch, A.; Perez-Fidalgo, J.A.; Lluch, A. Molecular biology in breast cancer: Intrinsic subtypes and signaling pathways. Cancer Treat. Rev. 2012, 38, 698–707. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.R.; Yu, J.J.; Shanker, K.; Deshpande, N.; Varambally, R.; Ghosh, D.; Barrette, T.; Pandey, A.; Chinnaiyan, A.M. Oncomine: A cancer microarray database and integrated data-mining platform. Neoplasia 2004, 6, 1–6. [Google Scholar] [CrossRef]
- Miyake, T.; Nakayama, T.; Naoi, Y.; Yamamoto, N.; Otani, Y.; Kim, S.J.; Shimazu, K.; Shimomura, A.; Maruyama, N.; Tamaki, Y.; et al. GSTP1 expression predicts poor pathological complete response to neoadjuvant chemotherapy in ER-negative breast cancer. Cancer Sci. 2012, 103, 913–920. [Google Scholar] [CrossRef] [PubMed]
- Barretina, J.; Caponigro, G.; Stransky, N.; Venkatesan, K.; Margolin, A.A.; Kim, S.; Wilson, C.J.; Lehar, J.; Kryukov, G.V.; Sonkin, D.; et al. The cancer cell line encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 2012, 483, 603–607. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.F.; Alyamani, M.; Li, J.N.; Rogacki, K.; Abazeed, M.; Upadhyay, S.K.; Balk, S.P.; Taplin, M.E.; Auchus, R.J.; Sharifi, N. Redirecting abiraterone metabolism to fine-tune prostate cancer anti-androgen therapy. Nature 2016, 533, 547. [Google Scholar] [CrossRef]
- Colditz, J.; Rupf, B.; Maiwald, C.; Baniahmad, A. Androgens induce a distinct response of epithelial-mesenchymal transition factors in human prostate cancer cells. Mol. Cell Biochem. 2016, 421, 139–147. [Google Scholar] [CrossRef]
- Agemy, L.; Kela, I.; Waks, T.; Pfeffer, R.M.; Bar-Shira, A.; Orr-Urtreger, A.; Domany, E.; Eshhar, Z. Gene expression profiles predict sensitivity of prostate cancer to radiotherapy. J. Cancer Ther. 2013, 4, 11. [Google Scholar] [CrossRef]
- Li, S.Q.; Li, B.; Zheng, Y.T.; Li, M.L.; Shi, L.M.; Pu, X.M. Exploring functions of long noncoding RNAs across multiple cancers through co-expression network. Sci. Rep. 2017, 7, 754. [Google Scholar] [CrossRef]
- Chakraborty, S. In silico analysis identifies genes common between five primary gastrointestinal cancer sites with potential clinical applications. Ann. Gastroenterol. Q. Publ. Hell. Soc. Gastroenterol. 2014, 27, 231. [Google Scholar]
- Li, L.X.; Shang, J.; Zhang, Y.P.; Liu, S.; Peng, Y.N.; Zhou, Z.; Pan, H.Q.; Wang, X.B.; Chen, L.P.; Zhao, Q. MEG3 is a prognostic factor for CRC and promotes chemosensitivity by enhancing oxaliplatin-induced cell apoptosis. Oncol. Rep. 2017, 38, 1383–1392. [Google Scholar] [CrossRef]
- Yu, H.; Xu, Q.H.; Liu, F.; Ye, X.; Wang, J.L.; Meng, X. Identification and validation of long noncoding RNA biomarkers in human non-small-cell lung carcinomas. J. Thorac. Oncol. 2015, 10, 645–654. [Google Scholar] [CrossRef] [PubMed]
- Molina-Pinelo, S.; Gutierrez, G.; Pastor, M.D.; Hergueta, M.; Moreno-Bueno, G.; Garcia-Carbonero, R.; Nogal, A.; Suarez, R.; Salinas, A.; Pozo-Rodriguez, F.; et al. MicroRNA-dependent regulation of transcription in non-small cell lung cancer. PLoS ONE 2014, 9, e90524. [Google Scholar] [CrossRef]
- Zhang, Y.; Wagner, E.K.; Guo, X.; May, I.; Cai, Q.; Zheng, W.; He, C.; Long, J. Long intergenic non-coding RNA expression signature in human breast cancer. Sci. Rep. 2016, 6, 37821. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qian, C.; Guan, M.; Si, C.; Shen, H.; Jin, T.; Zhang, T. Identification of differentially expressed profiles of lncRNAs and mRNAs in ER-negative and HER-2 positive breast cancer. Arch. Med. Sci.-Civ. Dis. 2017, 2, 148–160. [Google Scholar] [CrossRef] [Green Version]
- Van Grembergen, O.; Bizet, M.; de Bony, E.J.; Calonne, E.; Putmans, P.; Brohee, S.; Olsen, C.; Guo, M.Z.; Bontempi, G.; Sotiriou, C.; et al. Portraying breast cancers with long noncoding RNAs. Sci. Adv. 2016, 2, e1600220. [Google Scholar] [CrossRef] [PubMed]
- Varley, K.E.; Gertz, J.; Roberts, B.S.; Davis, N.S.; Bowling, K.M.; Kirby, M.K.; Nesmith, A.S.; Oliver, P.G.; Grizzle, W.E.; Forero, A.; et al. Recurrent read-through fusion transcripts in breast cancer. Breast Cancer Res. Treat. 2014, 146, 287–297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, T.; Fan, J.; Wang, B.; Traugh, N.; Chen, Q.; Liu, J.S.; Li, B.; Liu, X.S. Timer: A web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017, 77, e108–e110. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Han, L.; Roebuck, P.; Diao, L.X.; Liu, L.X.; Yuan, Y.; Weinstein, J.N.; Liang, H. Tanric: An interactive open platform to explore the function of lncRNAs in cancer. Cancer Res. 2015, 75, 3728–3737. [Google Scholar] [CrossRef] [PubMed]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Gao, J.J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.C.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioportal. Sci. Signal. 2013, 6. [Google Scholar] [CrossRef]
- Zhao, D.; Ma, G.; Zhang, X.; He, Y.; Li, M.; Han, X.; Fu, L.; Dong, X.Y.; Nagy, T.; Zhao, Q.; et al. Zinc finger homeodomain factor Zfhx3 is essential for mammary lactogenic differentiation by maintaining prolactin signaling activity. J. Biol. Chem. 2016, 291, 12809–12820. [Google Scholar] [CrossRef] [PubMed]
- Yang, Y.G.; Leonard, M.; Zhang, Y.J.; Zhao, D.; Mahmoud, C.; Khan, S.; Wang, J.; Lower, E.E.; Zhang, X.T. HER2-driven breast tumorigenesis relies upon interactions of the estrogen receptor with coactivator MED1. Cancer Res. 2018, 78, 422–435. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhao, D.; Dong, J.-T. Upregulation of Long Non-Coding RNA DRAIC Correlates with Adverse Features of Breast Cancer. Non-Coding RNA 2018, 4, 39. https://doi.org/10.3390/ncrna4040039
Zhao D, Dong J-T. Upregulation of Long Non-Coding RNA DRAIC Correlates with Adverse Features of Breast Cancer. Non-Coding RNA. 2018; 4(4):39. https://doi.org/10.3390/ncrna4040039
Chicago/Turabian StyleZhao, Dan, and Jin-Tang Dong. 2018. "Upregulation of Long Non-Coding RNA DRAIC Correlates with Adverse Features of Breast Cancer" Non-Coding RNA 4, no. 4: 39. https://doi.org/10.3390/ncrna4040039



