Cell Type-Selective Expression of Circular RNAs in Human Pancreatic Islets
Abstract
:1. Introduction
2. Results
2.1. Wide-Spread Expression of circRNAs in α- and β-Cells
2.2. Genomic Features of circRNAs in α- and β-Cells
2.3. Alternative Circular Isoforms
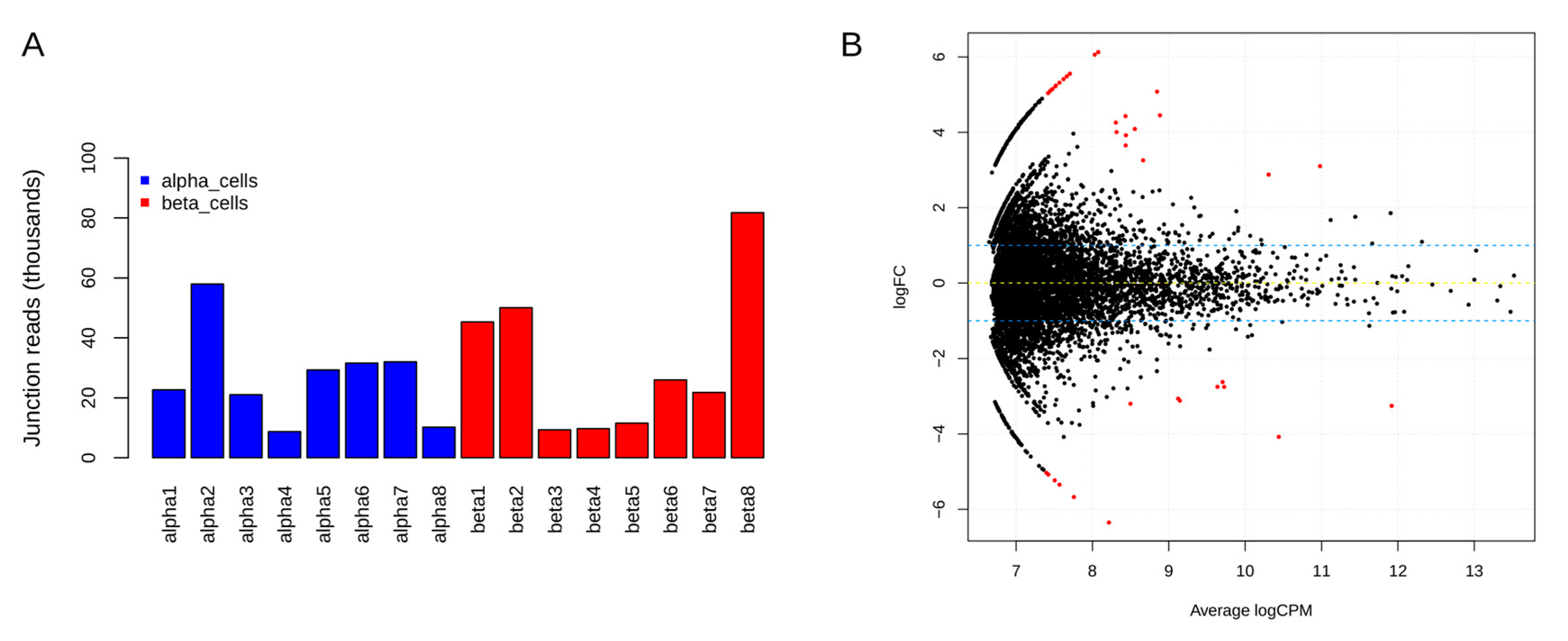
2.4. Differentially Expressed circRNAs in β-Cells
2.5. Selectively Expressed circRNAs in α- and β-Cells
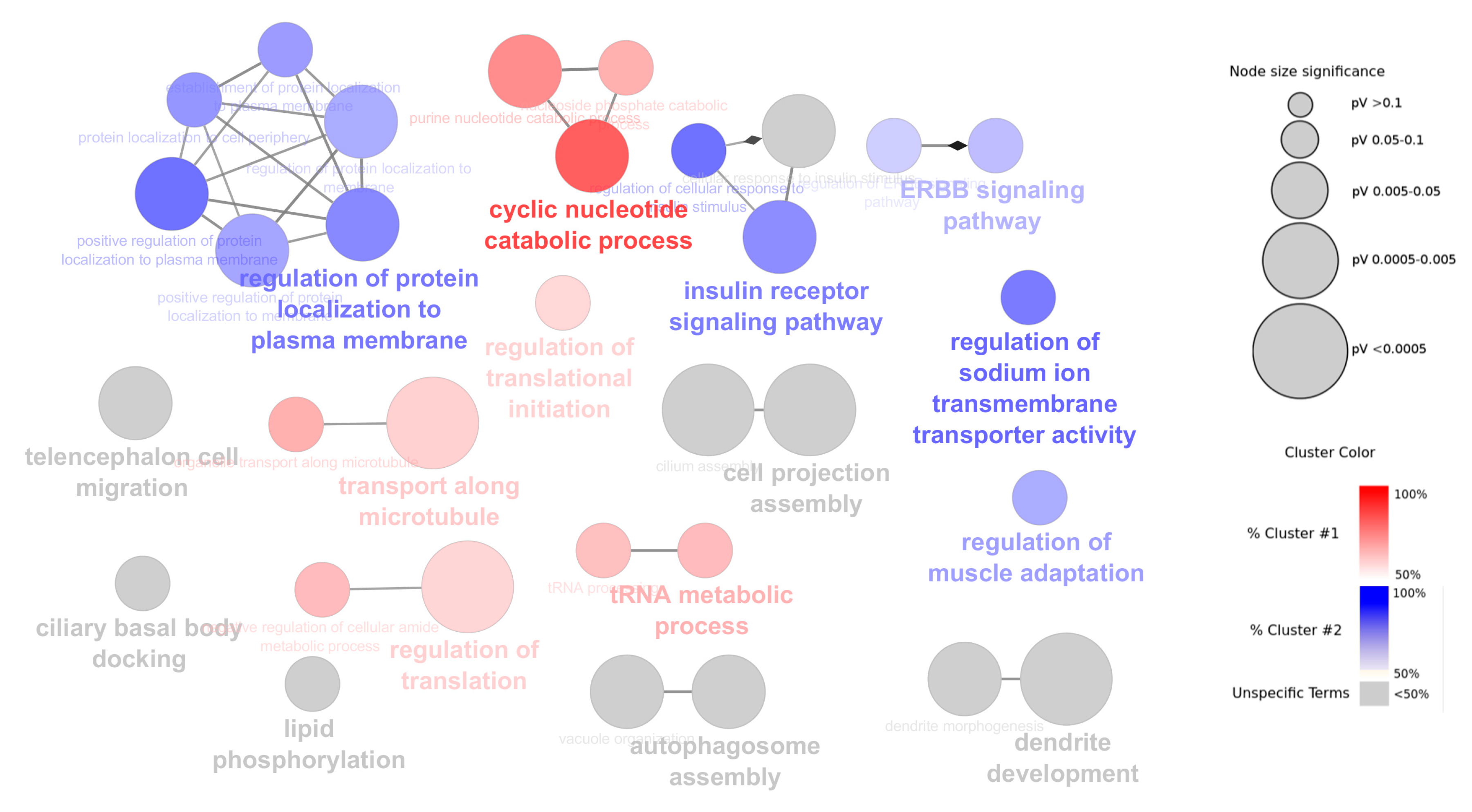
2.6. Functional Annotation of α- and β-Cell Selective circRNAs
3. Discussion
4. Materials and Methods
4.1. circRNA Identification and Analysis
4.2. Differential Expression Analysis
4.3. Functional Annotation of circRNAs
4.4. miRNA-circRNA Regulatory Network
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Morris, K.V.; Mattick, J.S. The rise of regulatory RNA. Nat. Rev. Genet. 2014, 15, 423–437. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Adams, B.D.; Parsons, C.; Walker, L.; Zhang, W.C.; Slack, F.J. Targeting noncoding RNAs in disease. J. Clin. Investig. 2017, 127, 761–771. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mishra, P.J. Non-coding RNAs as clinical biomarkers for cancer diagnosis and prognosis. Expert Rev. Mol. Diagn. 2014, 14, 917–919. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Maass, P.G.; Luft, F.C.; Bähring, S. Long non-coding RNA in health and disease. J. Mol. Med. 2014, 92, 337–346. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- De Planell-Saguer, M.; Rodicio, M.C. Analytical aspects of microRNA in diagnostics: A review. Anal. Chim. Acta 2011, 699, 134–152. [Google Scholar] [CrossRef] [PubMed]
- Kanwar, J.R.; Mahidhara, G.; Kanwar, R.K. MicroRNA in human cancer and chronic inflammatory diseases. Front. Biosci. 2010, 2, 1113–1126. [Google Scholar] [CrossRef]
- Salzman, J.; Gawad, C.; Wang, P.L.; Lacayo, N.; Brown, P.O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 2012, 7, e30733. [Google Scholar] [CrossRef] [PubMed]
- Barrett, S.P.; Wang, P.L.; Salzman, J. Circular RNA biogenesis can proceed through an exon-containing lariat precursor. eLife 2015, 4, e07540. [Google Scholar] [CrossRef] [PubMed]
- Schindewolf, C.; Braun, S.; Domdey, H. In vitro generation of a circular exon from a linear pre-mRNA transcript. Nucleic Acids Res. 1996, 24, 1260–1266. [Google Scholar] [CrossRef] [PubMed]
- Starke, S.; Jost, I.; Rossbach, O.; Schneider, T.; Schreiner, S.; Hung, L.-H.; Bindereif, A. Exon circularization requires canonical splice signals. Cell Rep. 2015, 10, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Nigro, J.M.; Cho, K.R.; Fearon, E.R.; Kern, S.E.; Ruppert, J.M.; Oliner, J.D.; Kinzler, K.W.; Vogelstein, B. Scrambled exons. Cell 1991, 64, 607–613. [Google Scholar] [CrossRef]
- Rybak-Wolf, A.; Stottmeister, C.; Glažar, P.; Jens, M.; Pino, N.; Giusti, S.; Hanan, M.; Behm, M.; Bartok, O.; Ashwal-Fluss, R.; et al. Circular RNAs in the Mammalian Brain Are Highly Abundant, Conserved, and Dynamically Expressed. Mol. Cell 2015, 58, 870–885. [Google Scholar] [CrossRef] [PubMed]
- Jeck, W.R.; Sorrentino, J.A.; Wang, K.; Slevin, M.K.; Burd, C.E.; Liu, J.; Marzluff, W.F.; Sharpless, N.E. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 2013, 19, 141–157. [Google Scholar] [CrossRef] [PubMed]
- Salzman, J.; Chen, R.E.; Olsen, M.N.; Wang, P.L.; Brown, P.O. Cell-type specific features of circular RNA expression. PLoS Genet. 2013, 9, e1003777. [Google Scholar] [CrossRef]
- Wang, P.L.; Bao, Y.; Yee, M.-C.; Barrett, S.P.; Hogan, G.J.; Olsen, M.N.; Dinneny, J.R.; Brown, P.O.; Salzman, J. Circular RNA is expressed across the eukaryotic tree of life. PLoS ONE 2014, 9, e90859. [Google Scholar] [CrossRef] [PubMed]
- Barrett, S.P.; Salzman, J. Circular RNAs: Analysis, expression and potential functions. Development 2016, 143, 1838–1847. [Google Scholar] [CrossRef] [PubMed]
- Szabo, L.; Morey, R.; Palpant, N.J.; Wang, P.L.; Afari, N.; Jiang, C.; Parast, M.M.; Murry, C.E.; Laurent, L.C.; Salzman, J. Statistically based splicing detection reveals neural enrichment and tissue-specific induction of circular RNA during human fetal development. Genome Biol. 2015, 16, 126. [Google Scholar] [CrossRef] [PubMed]
- You, X.; Vlatkovic, I.; Babic, A.; Will, T.; Epstein, I.; Tushev, G.; Akbalik, G.; Wang, M.; Glock, C.; Quedenau, C.; et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 2015, 18, 603–610. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Venø, M.T.; Hansen, T.B.; Venø, S.T.; Clausen, B.H.; Grebing, M.; Finsen, B.; Holm, I.E.; Kjems, J. Spatio-temporal regulation of circular RNA expression during porcine embryonic brain development. Genome Biol. 2015, 16, 245. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Westholm, J.O.; Miura, P.; Olson, S.; Shenker, S.; Joseph, B.; Sanfilippo, P.; Celniker, S.E.; Graveley, B.R.; Lai, E.C. Genome-wide analysis of drosophila circular RNAs reveals their structural and sequence properties and age-dependent neural accumulation. Cell Rep. 2014, 9, 1966–1980. [Google Scholar] [CrossRef] [PubMed]
- Izuogu, O.G.; Alhasan, A.A.; Mellough, C.; Collin, J.; Gallon, R.; Hyslop, J.; Mastrorosa, F.K.; Ehrmann, I.; Lako, M.; Elliott, D.J.; et al. Analysis of human ES cell differentiation establishes that the dominant isoforms of the lncRNAs RMST and FIRRE are circular. BMC Genom. 2018, 19, 276. [Google Scholar] [CrossRef] [PubMed]
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Jensen, T.I.; Clausen, B.H.; Bramsen, J.B.; Finsen, B.; Damgaard, C.K.; Kjems, J. Natural RNA circles function as efficient microRNA sponges. Nature 2013, 495, 384–388. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Guo, S.; Li, W.; Yu, P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci. Rep. 2015, 5, 12453. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, F.; Zhang, L.; Li, W.; Deng, J.; Zheng, J.; An, M.; Lu, J.; Zhou, Y. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/β-catenin pathway. Oncotarget 2015, 6, 6001–6013. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zheng, Q.; Bao, C.; Guo, W.; Li, S.; Chen, J.; Chen, B.; Luo, Y.; Lyu, D.; Li, Y.; Shi, G.; et al. Circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 2016, 7, 11215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peng, L.; Chen, G.; Zhu, Z.; Shen, Z.; Du, C.; Zang, R.; Su, Y.; Xie, H.; Li, H.; Xu, X.; et al. Circular RNA ZNF609 functions as a competitive endogenous RNA to regulate AKT3 expression by sponging miR-150-5p in Hirschsprung’s disease. Oncotarget 2017, 8, 808–818. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Long, B.; Liu, F.; Wang, J.-X.; Liu, C.-Y.; Zhao, B.; Zhou, L.-Y.; Sun, T.; Wang, M.; Yu, T.; et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 2016, 37, 2602–2611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Du, W.W.; Yang, W.; Liu, E.; Yang, Z.; Dhaliwal, P.; Yang, B.B. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 2016, 44, 2846–2858. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ashwal-Fluss, R.; Meyer, M.; Pamudurti, N.R.; Ivanov, A.; Bartok, O.; Hanan, M.; Evantal, N.; Memczak, S.; Rajewsky, N.; Kadener, S. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 2014, 56, 55–66. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Wiklund, E.D.; Bramsen, J.B.; Villadsen, S.B.; Statham, A.L.; Clark, S.J.; Kjems, J. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 2011, 30, 4414–4422. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ng, W.L.; Marinov, G.K.; Liau, E.S.; Lam, Y.L.; Lim, Y.-Y.; Ea, C.-K. Inducible RasGEF1B circular RNA is a positive regulator of ICAM-1 in the TLR4/LPS pathway. RNA Biol. 2016, 13, 861–871. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Li, Z.; Huang, C.; Bao, C.; Chen, L.; Lin, M.; Wang, X.; Zhong, G.; Yu, B.; Hu, W.; Dai, L.; et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 2015, 22, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, X.-O.; Chen, T.; Xiang, J.-F.; Yin, Q.-F.; Xing, Y.-H.; Zhu, S.; Yang, L.; Chen, L.-L. Circular intronic long noncoding RNAs. Mol. Cell 2013, 51, 792–806. [Google Scholar] [CrossRef] [PubMed]
- Legnini, I.; Di Timoteo, G.; Rossi, F.; Morlando, M.; Briganti, F.; Sthandier, O.; Fatica, A.; Santini, T.; Andronache, A.; Wade, M.; et al. Circ-ZNF609 Is a Circular RNA that Can Be Translated and Functions in Myogenesis. Mol. Cell 2017, 66, 22–37. [Google Scholar] [CrossRef] [PubMed]
- Pamudurti, N.R.; Bartok, O.; Jens, M.; Ashwal-Fluss, R.; Stottmeister, C.; Ruhe, L.; Hanan, M.; Wyler, E.; Perez-Hernandez, D.; Ramberger, E.; et al. Translation of CircRNAs. Mol. Cell 2017, 66, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Li, Y.; Zheng, Q.; Bao, C.; He, J.; Chen, B.; Lyu, D.; Zheng, B.; Xu, Y.; Long, Z.; et al. Circular RNA profile identifies circPVT1 as a proliferative factor and prognostic marker in gastric cancer. Cancer Lett. 2017, 388, 208–219. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Li, T.; Zhao, Q.; Xiao, B.; Guo, J. Using circular RNA hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin. Chim. Acta 2017, 466, 167–171. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.-N.; Li, J.; Zhu, C.-L.; Feng, W.-T.; Shao, J.-X.; Wan, L.; Huang, M.; He, J.-D. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation, migration, and invasion in colorectal cancer. Oncol. Targets Ther. 2016, 9, 7451–7458. [Google Scholar] [CrossRef] [PubMed]
- Dou, Y.; Cha, D.J.; Franklin, J.L.; Higginbotham, J.N.; Jeppesen, D.K.; Weaver, A.M.; Prasad, N.; Levy, S.; Coffey, R.J.; Patton, J.G.; et al. Circular RNAs are down-regulated in KRAS mutant colon cancer cells and can be transferred to exosomes. Sci. Rep. 2016, 6, 37982. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Wan, L.; Zhang, L.; Fan, K.; Cheng, Z.-X.; Sun, Q.-C.; Wang, J.-J. Circular RNA-ITCH Suppresses Lung Cancer Proliferation via Inhibiting the Wnt/β-Catenin Pathway. Biomed. Res. Int. 2016, 2016, 1579490. [Google Scholar] [CrossRef] [PubMed]
- Yao, J.-T.; Zhao, S.-H.; Liu, Q.-P.; Lv, M.-Q.; Zhou, D.-X.; Liao, Z.-J.; Nan, K.-J. Over-expression of CircRNA_100876 in non-small cell lung cancer and its prognostic value. Pathol. Res. Pract. 2017, 213, 453–456. [Google Scholar] [CrossRef] [PubMed]
- Guarnerio, J.; Bezzi, M.; Jeong, J.C.; Paffenholz, S.V.; Berry, K.; Naldini, M.M.; Lo-Coco, F.; Tay, Y.; Beck, A.H.; Pandolfi, P.P. Oncogenic Role of Fusion-circRNAs Derived from Cancer-Associated Chromosomal Translocations. Cell 2016, 166, 1055–1056. [Google Scholar] [CrossRef] [PubMed]
- Shang, X.; Li, G.; Liu, H.; Li, T.; Liu, J.; Zhao, Q.; Wang, C. Comprehensive Circular RNA Profiling Reveals That hsa_circ_0005075, a New Circular RNA Biomarker, Is Involved in Hepatocellular Crcinoma Development. Medicine (Baltimore) 2016, 95, e3811. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Qin, M.; Liu, G.; Huo, X.; Tao, X.; Sun, X.; Ge, Z.; Yang, J.; Fan, J.; Liu, L.; Qin, W. Hsa_circ_0001649: A circular RNA and potential novel biomarker for hepatocellular carcinoma. Cancer Biomark. 2016, 16, 161–169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Xu, L.; Zhang, M.; Zheng, X.; Yi, P.; Lan, C.; Xu, M. The circular RNA ciRS-7 (Cdr1as) acts as a risk factor of hepatic microvascular invasion in hepatocellular carcinoma. J. Cancer Res. Clin. Oncol. 2017, 143, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Shan, K.; Liu, C.; Liu, B.-H.; Chen, X.; Dong, R.; Liu, X.; Zhang, Y.-Y.; Liu, B.; Zhang, S.-J.; Wang, J.-J.; et al. Circular Noncoding RNA HIPK3 Mediates Retinal Vascular Dysfunction in Diabetes Mellitus. Circulation 2017, 136, 1629–1642. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Li, X.; Jian, D.; Hao, P.; Rao, L.; Li, M. Hsa_circ_0054633 in peripheral blood can be used as a diagnostic biomarker of pre-diabetes and type 2 diabetes mellitus. Acta Diabetol. 2017, 54, 237–245. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Wang, X.; Li, W.; Han, J.; Jin, J.; Su, F.; Zhang, J.; Huang, W.; Xiao, F.; Pan, Q.; et al. Screening of circular RNAs and validation of circANKRD36 associated with inflammation in patients with type 2 diabetes mellitus. Int. J. Mol. Med. 2018, 42, 1865–1874. [Google Scholar] [CrossRef] [PubMed]
- Yan, L.; Feng, J.; Cheng, F.; Cui, X.; Gao, L.; Chen, Y.; Wang, F.; Zhong, T.; Li, Y.; Liu, L. Circular RNA expression profiles in placental villi from women with gestational diabetes mellitus. Biochem. Biophys. Res. Commun. 2018, 498, 743–750. [Google Scholar] [CrossRef] [PubMed]
- Stoll, L.; Sobel, J.; Rodriguez-Trejo, A.; Guay, C.; Lee, K.; Venø, M.T.; Kjems, J.; Laybutt, D.R.; Regazzi, R. Circular RNAs as novel regulators of β-cell functions in normal and disease conditions. Mol. Metab. 2018, 9, 69–83. [Google Scholar] [CrossRef] [PubMed]
- Brissova, M.; Fowler, M.J.; Nicholson, W.E.; Chu, A.; Hirshberg, B.; Harlan, D.M.; Powers, A.C. Assessment of human pancreatic islet architecture and composition by laser scanning confocal microscopy. J. Histochem. Cytochem. 2005, 53, 1087–1097. [Google Scholar] [CrossRef] [PubMed]
- Cabrera, O.; Berman, D.M.; Kenyon, N.S.; Ricordi, C.; Berggren, P.-O.; Caicedo, A. The unique cytoarchitecture of human pancreatic islets has implications for islet cell function. Proc. Natl. Acad. Sci. USA 2006, 103, 2334–2339. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dorrell, C.; Schug, J.; Lin, C.F.; Canaday, P.S.; Fox, A.J.; Smirnova, O.; Bonnah, R.; Streeter, P.R.; Stoeckert, C.J.; Kaestner, K.H.; et al. Transcriptomes of the major human pancreatic cell types. Diabetologia 2011, 54, 2832–2844. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Blodgett, D.M.; Nowosielska, A.; Afik, S.; Pechhold, S.; Cura, A.J.; Kennedy, N.J.; Kim, S.; Kucukural, A.; Davis, R.J.; Kent, S.C.; et al. Novel Observations from Next-Generation RNA Sequencing of Highly Purified Human Adult and Fetal Islet Cell Subsets. Diabetes 2015, 64, 3172–3181. [Google Scholar] [CrossRef] [PubMed]
- Bramswig, N.C.; Everett, L.J.; Schug, J.; Dorrell, C.; Liu, C.; Luo, Y.; Streeter, P.R.; Naji, A.; Grompe, M.; Kaestner, K.H. Epigenomic plasticity enables human pancreatic α to β cell reprogramming. J. Clin. Investig. 2013, 123, 1275–1284. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ackermann, A.M.; Wang, Z.; Schug, J.; Naji, A.; Kaestner, K.H. Integration of ATAC-seq and RNA-seq identifies human α-cell and β-cell signature genes. Mol. Metab. 2016, 5, 233–244. [Google Scholar] [CrossRef] [PubMed]
- Nica, A.C.; Ongen, H.; Irminger, J.-C.; Bosco, D.; Berney, T.; Antonarakis, S.E.; Halban, P.A.; Dermitzakis, E.T. Cell-type, allelic, and genetic signatures in the human pancreatic β-cell transcriptome. Genome Res. 2013, 23, 1554–1562. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Trapnell, C. Single-cell transcriptome sequencing: Recent advances and remaining challenges. F1000Research 2016, 5. [Google Scholar] [CrossRef] [PubMed]
- Morán, I.; Akerman, I.; Van de Bunt, M.; Xie, R.; Benazra, M.; Nammo, T.; Arnes, L.; Nakić, N.; García-Hurtado, J.; Rodríguez-Seguí, S.; et al. Human β cell transcriptome analysis uncovers lncRNAs that are tissue-specific, dynamically regulated, and abnormally expressed in type 2 diabetes. Cell Metab. 2012, 16, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Eizirik, D.L.; Sammeth, M.; Bouckenooghe, T.; Bottu, G.; Sisino, G.; Igoillo-Esteve, M.; Ortis, F.; Santin, I.; Colli, M.L.; Barthson, J.; et al. The human pancreatic islet transcriptome: Expression of candidate genes for type 1 diabetes and the impact of pro-inflammatory cytokines. PLoS Genet. 2012, 8, e1002552. [Google Scholar] [CrossRef] [PubMed]
- Lopes, M.; Kutlu, B.; Miani, M.; Bang-Berthelsen, C.H.; Størling, J.; Pociot, F.; Goodman, N.; Hood, L.; Welsh, N.; Bontempi, G.; Eizirik, D.L. Temporal profiling of cytokine-induced genes in pancreatic β-cells by meta-analysis and network inference. Genomics 2014, 103, 264–275. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-O.; Dong, R.; Zhang, Y.; Zhang, J.-L.; Luo, Z.; Zhang, J.; Chen, L.-L.; Yang, L. Diverse alternative back-splicing and alternative splicing landscape of circular RNAs. Genome Res. 2016, 26, 1277–1287. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zeng, X.; Lin, W.; Guo, M.; Zou, Q. A comprehensive overview and evaluation of circular RNA detection tools. PLoS Comput. Biol. 2017, 13, e1005420. [Google Scholar] [CrossRef] [PubMed]
- Hansen, T.B.; Venø, M.T.; Damgaard, C.K.; Kjems, J. Comparison of circular RNA prediction tools. Nucleic Acids Res. 2016, 44, e58. [Google Scholar] [CrossRef] [PubMed]
- Glažar, P.; Papavasileiou, P.; Rajewsky, N. circBase: A database for circular RNAs. RNA 2014, 20, 1666–1670. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.-C.; Li, J.-R.; Sun, C.-H.; Andrews, E.; Chao, R.-F.; Lin, F.-M.; Weng, S.-L.; Hsu, S.-D.; Huang, C.-C.; Cheng, C.; et al. CircNet: A database of circular RNAs derived from transcriptome sequencing data. Nucleic Acids Res. 2016, 44, D209–D215. [Google Scholar] [CrossRef] [PubMed]
- Song, X.; Zhang, N.; Han, P.; Moon, B.-S.; Lai, R.K.; Wang, K.; Lu, W. Circular RNA profile in gliomas revealed by identification tool UROBORUS. Nucleic Acids Res. 2016, 44, e87. [Google Scholar] [CrossRef] [PubMed]
- Van de Bunt, M.; Gaulton, K.J.; Parts, L.; Moran, I.; Johnson, P.R.; Lindgren, C.M.; Ferrer, J.; Gloyn, A.L.; McCarthy, M.I. The miRNA profile of human pancreatic islets and β-cells and relationship to type 2 diabetes pathogenesis. PLoS ONE 2013, 8, e55272. [Google Scholar] [CrossRef] [PubMed]
- Klein, D.; Misawa, R.; Bravo-Egana, V.; Vargas, N.; Rosero, S.; Piroso, J.; Ichii, H.; Umland, O.; Zhijie, J.; Tsinoremas, N.; et al. MicroRNA expression in α- and β-cells of human pancreatic islets. PLoS ONE 2013, 8, e55064. [Google Scholar] [CrossRef] [PubMed]
- Pociot, F. Type 1 diabetes genome-wide association studies: Not to be lost in translation. Clin. Transl. Immunol. 2017, 6, e162. [Google Scholar] [CrossRef] [PubMed]
- Thurner, M.; Van de Bunt, M.; Torres, J.M.; Mahajan, A.; Nylander, V.; Bennett, A.J.; Gaulton, K.J.; Barrett, A.; Burrows, C.; Bell, C.G.; et al. Integration of human pancreatic islet genomic data refines regulatory mechanisms at Type 2 Diabetes susceptibility loci. eLife 2018, 7. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez, G.D.; Bender, A.S.; Cirulli, V.; Mastracci, T.L.; Kelly, S.M.; Tsirigos, A.; Kaestner, K.H.; Sussel, L. Pancreatic β cell identity requires continual repression of non-β cell programs. J. Clin. Investig. 2017, 127, 244–259. [Google Scholar] [CrossRef] [PubMed]
- Arnes, L.; Akerman, I.; Balderes, D.A.; Ferrer, J.; Sussel, L. βlinc1 encodes a long noncoding RNA that regulates islet β-cell formation and function. Genes Dev. 2016, 30, 502–507. [Google Scholar] [CrossRef] [PubMed]
- Mirza, A.H.; Kaur, S.; Pociot, F. Long non-coding RNAs as novel players in β cell function and type 1 diabetes. Hum. Genom. 2017, 11, 17. [Google Scholar] [CrossRef] [PubMed]
- Locke, J.M.; Da Silva Xavier, G.; Dawe, H.R.; Rutter, G.A.; Harries, L.W. Increased expression of miR-187 in human islets from individuals with type 2 diabetes is associated with reduced glucose-stimulated insulin secretion. Diabetologia 2014, 57, 122–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Szabo, L.; Salzman, J. Detecting circular RNAs: Bioinformatic and experimental challenges. Nat. Rev. Genet. 2016, 17, 679–692. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.-O.; Wang, H.-B.; Zhang, Y.; Lu, X.; Chen, L.-L.; Yang, L. Complementary sequence-mediated exon circularization. Cell 2014, 159, 134–147. [Google Scholar] [CrossRef] [PubMed]
- Sims, D.; Sudbery, I.; Ilott, N.E.; Heger, A.; Ponting, C.P. Sequencing depth and coverage: Key considerations in genomic analyses. Nat. Rev. Genet. 2014, 5, 121–132. [Google Scholar] [CrossRef]
- Panda, A.C.; De, S.; Grammatikakis, I.; Munk, R.; Yang, X.; Piao, Y.; Dudekula, D.B.; Abdelmohsen, K.; Gorospe, M. High-purity circular RNA isolation method (RPAD) reveals vast collection of intronic circRNAs. Nucleic Acids Res. 2017, 45, e116. [Google Scholar] [CrossRef] [PubMed]
- Dhawan, S.; Dirice, E.; Kulkarni, R.N.; Bhushan, A. Inhibition of TGF-β Signaling Promotes Human Pancreatic β-Cell Replication. Diabetes 2016, 65, 1208–1218. [Google Scholar] [CrossRef] [PubMed]
- Bagge, A.; Clausen, T.R.; Larsen, S.; Ladefoged, M.; Rosenstierne, M.W.; Larsen, L.; Vang, O.; Nielsen, J.H.; Dalgaard, L.T. MicroRNA-29a is up-regulated in β-cells by glucose and decreases glucose-stimulated insulin secretion. Biochem. Biophys. Res. Commun. 2012, 426, 266–272. [Google Scholar] [CrossRef] [PubMed]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Pertea, G.; Trapnell, C.; Pimentel, H.; Kelley, R.; Salzberg, S.L. TopHat2: Accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013, 14, R36. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, D.; Salzberg, S.L. TopHat-Fusion: An algorithm for discovery of novel fusion transcripts. Genome Biol. 2011, 12, R72. [Google Scholar] [CrossRef] [PubMed]
- Anders, S.; Pyl, P.T.; Huber, W. HTSeq—A Python framework to work with high-throughput sequencing data. Bioinformatics 2015, 31, 166–169. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- McCarthy, D.J.; Chen, Y.; Smyth, G.K. Differential expression analysis of multifactor RNA-Seq experiments with respect to biological variation. Nucleic Acids Res. 2012, 40, 4288–4297. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bindea, G.; Mlecnik, B.; Hackl, H.; Charoentong, P.; Tosolini, M.; Kirilovsky, A.; Fridman, W.-H.; Pagès, F.; Trajanoski, Z.; Galon, J. ClueGO: A Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 2009, 25, 1091–1093. [Google Scholar] [CrossRef] [PubMed]
- Chou, C.-H.; Shrestha, S.; Yang, C.-D.; Chang, N.-W.; Lin, Y.-L.; Liao, K.-W.; Huang, W.-C.; Sun, T.-H.; Tu, S.-J.; Lee, W.-H.; et al. miRTarBase update 2018: A resource for experimentally validated microRNA-target interactions. Nucleic Acids Res. 2018, 46, D296–D302. [Google Scholar] [CrossRef] [PubMed]





| circRNAs | Genes | |
|---|---|---|
| α-Cells | 7667 | 3141 |
| β-Cells | 8396 | 3291 |
| Exocrine-cells | 456 | 388 |
| Common circRNAs | 382 | 332 |
| Total | 10,830 | 3833 |
| circRNA ID | Gene Name | Average Junction Reads (α-Cells) | Average Junction Reads (β-Cells) | circBase ID |
|---|---|---|---|---|
| 1:117402185:117420649 | MAN1A2 | 267 | 354 | hsa_circ_0000118 |
| 12:97492460:97561047 | RMST | 354 | 220 | - |
| 11:33286412:33287511 | HIPK3 | 263 | 286 | hsa_circ_0000284 |
| 4:152411302:152412529 | FBXW7 | 285 | 241 | hsa_circ_0001451 |
| 5:95755395:95763620 | RHOBTB3 | 137 | 307 | hsa_circ_0007444 |
| 4:143543508:143543972 | SMARCA5 | 180 | 228 | hsa_circ_0001445 |
| 7:100023418:100024307 | ZKSCAN1 | 220 | 177 | hsa_circ_0001727 |
| 10:31908171:31910563 | ARHGAP12 | 174 | 167 | hsa_circ_0000231 |
| 7:24623665:24668660 | MPP6 | 134 | 144 | hsa_circ_0001686 |
| 18:9182381:9221999 | ANKRD12 | 101 | 123 | hsa_circ_0000826 |
| 2:50921868:50925955 | NRXN1 | 106 | 110 | hsa_circ_0001004 |
| circRNA | Gene Name | Isoform Name | logFC | p-Value | adj. p-Value |
|---|---|---|---|---|---|
| Down-Regulated | |||||
| 2:162183413:162203145 | FAP | ENST00000627638 | −6.34 | 4.40 × 10−8 | 7.23 × 10−5 |
| 2:190895151:190931637 | GLS | ENST00000320717 | −5.67 | 5.97 × 10−6 | 3.92 × 10−3 |
| 2:200329430:200419496 | SPATS2L | ENST00000409718 | −5.34 | 4.09 × 10−5 | 1.22 × 10−2 |
| 11:101062446:101126158 | PGR | ENST00000325455 | −5.22 | 7.83 × 10−5 | 1.98 × 10−2 |
| 5:95906522:95919545 | ELL2 | ENST00000237853 | −5.07 | 1.77 × 10−4 | 3.53 × 10−2 |
| 8:61633682:61653660 | ASPH | ENST00000379454 | −5.03 | 2.26 × 10−4 | 4.24 × 10−2 |
| 9:110972072:110973558 | LPAR1 | ENST00000374431 | −4.07 | 2.45 × 10−9 | 1.61 × 10−5 |
| 2:190900563:190924593 | GLS | ENST00000320717 | −3.25 | 4.03 × 10−8 | 7.23 × 10−5 |
| 2:190895151:190910321 | GLS | ENST00000320717 | −3.19 | 1.65 × 10−4 | 3.39 × 10−2 |
| 10:94030682:94032252 | PLCE1 | ENST00000260766 | −3.11 | 2.67 × 10−5 | 1.03 × 10−2 |
| 2:190900563:190931637 | GLS | ENST00000320717 | −3.06 | 3.98 × 10−5 | 1.22 × 10−2 |
| 5:16779544:16783469 | MYO10 | ENST00000274203 | −2.74 | 5.29 × 10−5 | 1.51 × 10−2 |
| 1:176594523:176595595 | PAPPA2 | ENST00000367661 | −2.74 | 5.93 × 10−5 | 1.56 × 10−2 |
| 1:95143890:95151419 | TMEM56 | ENST00000370203 | −2.61 | 9.18 × 10−5 | 2.08 × 10−2 |
| Up-regulated | |||||
| 1:91861470:91861644 | TGFBR3 | ENST00000212355 | 6.14 | 3.16 × 10−7 | 3.46 × 10−4 |
| 7:18666212:18666476 | HDAC9 | ENST00000461159 | 6.07 | 4.91 × 10−7 | 4.24 × 10−4 |
| 7:14607433:14673027 | DGKB | ENST00000399322 | 5.56 | 1.41 × 10−5 | 7.13 × 10−3 |
| 16:77816920:77825461 | VAT1L | ENST00000302536 | 5.49 | 2.17 × 10−5 | 8.90 × 10−3 |
| 4:42485495:42524847 | ATP8A1 | ENST00000381668 | 5.42 | 3.33 × 10−5 | 1.18 × 10−2 |
| 9:70761600:70784279 | TRPM3 | ENST00000377110 | 5.33 | 5.62 × 10−5 | 1.54 × 10−2 |
| 9:70681505:70784279 | TRPM3 | ENST00000377110 | 5.25 | 9.04 × 10−5 | 2.08 × 10−2 |
| 13:69940039:69975802 | KLHL1 | ENST00000377844 | 5.23 | 9.71 × 10−5 | 2.12 × 10−2 |
| 7:22291174:22318037 | RAPGEF5 | ENST00000344041 | 5.15 | 1.43 × 10−4 | 3.02 × 10−2 |
| 9:76703336:76713721 | PRUNE2 | ENST00000376718 | 5.11 | 1.85 × 10−4 | 3.58 × 10−2 |
| 1:65913244:65918835 | PDE4B | ENST00000329654 | 5.09 | 3.28 × 10−8 | 7.23 × 10−5 |
| X:154789953:154792285 | MPP1 | ENST00000369534 | 5.05 | 2.37 × 10−4 | 4.33 × 10−2 |
| 7:75943781:75943916 | SNORA14A | ENST00000364773 | 4.46 | 3.13 × 10−7 | 3.46 × 10−4 |
| 8:69755423:69761875 | SLCO5A1 | ENST00000260126 | 4.44 | 3.25 × 10−6 | 2.37 × 10−3 |
| 7:14574211:14673027 | DGKB | ENST00000399322 | 4.27 | 1.62 × 10−5 | 7.60 × 10−3 |
| 13:26337566:26353880 | CDK8 | ENST00000536792 | 4.10 | 6.81 × 10−6 | 4.06 × 10−3 |
| 2:144208602:144211579 | GTDC1 | ENST00000392869 | 4.02 | 3.42 × 10−5 | 1.18 × 10−2 |
| 1:58506059:58539310 | OMA1 | ENST00000371226 | 3.93 | 2.17 × 10−5 | 8.90 × 10−3 |
| 8:32595827:32616885 | NRG1 | ENST00000523534 | 3.66 | 4.06 × 10−5 | 1.22 × 10−2 |
| 18:62348167:62349937 | TNFRSF11A | ENST00000269485 | 3.27 | 8.98 × 10−5 | 2.08 × 10−2 |
| 4:186706562:186709845 | FAT1 | ENST00000441802 | 3.11 | 5.17 × 10−7 | 4.24 × 10−4 |
| 8:69738039:69761875 | SLCO5A1 | ENST00000260126 | 2.89 | 7.85 × 10−6 | 4.29 × 10−3 |
| Selectively Expressed circRNAs | Highly Selective circRNAs | |||
|---|---|---|---|---|
| circRNAs | Genes | circRNAs | Genes | |
| α-Cell | 392 | 343 | 7 | 5 |
| β-Cell | 417 | 359 | 1 | 1 |
| Exocrine cells | 57 | 52 | - | - |
| Gene Name | circ ID | Exon Count | circRNA Length | log2(RC) | Gene Biotype |
|---|---|---|---|---|---|
| α-cell selective circRNAs | |||||
| PLCE1 | 10:94030682:94032252 | 1 | 1570 | 4.43 | protein_coding |
| FAP | 2:162183413:162203145 | 9 | 822 | 3.38 | protein_coding |
| KANK1 | 9:710803:713464 | 1 | 2661 | 3.27 | protein_coding |
| MCU | 10:72715110:72715902 | 2 | 165 | 3.02 | protein_coding |
| C1orf168 | 1:56787174:56792803 | 3 | 944 | 2.95 | protein_coding |
| FRMPD3 | X:107526581:107533550 | 3 | 304 | 2.93 | protein_coding |
| PLCE1 | 10:94245945:94246621 | 1 | 676 | 2.88 | protein_coding |
| KIAA1217 | 10:24219625:24219909 | 1 | 284 | 2.70 | protein_coding |
| SYTL5 | X:38033533:38034008 | 1 | 475 | 2.70 | protein_coding |
| FAM126A | 7:22960255:22991139 | 9 | 1019 | 2.64 | protein_coding |
| SPG21 | 15:64974601:64983593 | 4 | 476 | 2.55 | protein_coding |
| POU6F2 | 7:39204234:39207620 | 2 | 321 | 2.55 | protein_coding |
| PITPNB | 22:27894554:27914347 | 6 | 436 | 2.52 | protein_coding |
| PRKCE | 2:46001403:46010517 | 4 | 614 | 2.49 | protein_coding |
| SPATS2L | 2:200329430:200419496 | 5 | 517 | 2.49 | protein_coding |
| β-cell selective circRNAs | |||||
| SNORA14A | 7:75943781:75943916 | 1 | 135 | 4.96 | snoRNA |
| PDE4B | 1:65913244:65918835 | 2 | 351 | 4.37 | protein_coding |
| TNFRSF11A | 18:62348167:62349937 | 2 | 208 | 4.21 | protein_coding |
| NRG1 | 8:32595827:32616885 | 4 | 402 | 3.95 | protein_coding |
| SNORD9 | 14:21392149:21392253 | 1 | 104 | 3.63 | snoRNA |
| DGKB | 7:14574211:14673027 | 8 | 738 | 3.58 | protein_coding |
| GTDC1 | 2:144208602:144211579 | 2 | 267 | 3.48 | protein_coding |
| SYNE2 | 14:63998913:64022863 | 12 | 2284 | 3.39 | protein_coding |
| HDAC9 | 7:18666212:18666476 | 1 | 264 | 3.36 | protein_coding |
| TNRC6B | 22:40117054:40156182 | 3 | 233 | 3.36 | protein_coding |
| TGFBR3 | 1:91861470:91861644 | 1 | 174 | 3.29 | protein_coding |
| NRD1 | 1:51827795:51834170 | 2 | 228 | 3.29 | protein_coding |
| GLCCI1 | 7:8003907:8060248 | 4 | 509 | 3.21 | protein_coding |
| ROR1 | 1:64009304:64050716 | 3 | 391 | 3.04 | protein_coding |
| TSNAX | 1:231537212:231542611 | 2 | 246 | 2.88 | protein_coding |
| Exocrine-cell selective circRNAs | |||||
| SNORD26 | 11:62855291:62855366 | 1 | 75 | 5.82 | snoRNA |
| PNLIPRP2 | 10:116625945:116627874 | 3 | 373 | 4.78 | polymorphic_pseudogene |
| CFTR | 7:117504252:117559655 | 10 | 1531 | 4.29 | protein_coding |
| C15orf38-AP3S2 | 15:89903215:89910819 | 4 | 580 | 4.29 | protein_coding |
| PNLIPRP2 | 10:116625945:116631357 | 5 | 613 | 4.21 | polymorphic_pseudogene |
| RAB3D | 19:11335446:11335783 | 2 | 244 | 4.13 | protein_coding |
| SEL1L | 14:81492479:81495137 | 2 | 126 | 3.81 | protein_coding |
| TRHDE | 12:72562164:72575542 | 5 | 533 | 3.64 | protein_coding |
| TRHDE | 12:72562852:72575542 | 4 | 467 | 3.52 | protein_coding |
| SLC7A6 | 16:68266592:68266721 | 1 | 129 | 3.09 | protein_coding |
| IKBKB | 8:42325969:42329214 | 2 | 219 | 3.09 | protein_coding |
| SLC43A1 | 11:57491223:57491862 | 3 | 322 | 3.09 | protein_coding |
| C3orf52 | 3:112109542:112113145 | 2 | 253 | 2.91 | protein_coding |
| ARF4 | 3:57583897:57584464 | 2 | 191 | 2.91 | protein_coding |
| SNORD58B | 18:49491663:49491729 | 1 | 66 | 2.91 | snoRNA |
| α-Cell-Selective circRNAs | ||||
|---|---|---|---|---|
| Gene Name | CHR | circ ID | circBase or CircNet Id | log2(RC) |
| FAP | 2 | 2:162183413:162203145 | hsa-circ-FAP.15 | 3.4 |
| SYTL5 | X | X:38033533:38034008 | hsa-circ-SYTL5.3 | 2.7 |
| PTPRT | 20 | 20:42448219:42472562 | hsa-circ-PTPRT.2, hsa_circ_0060424 | 1.9 |
| PTPRT | 20 | 20:42350627:42352285 | hsa-circ-PTPRT.5 | 1 |
| PTPRT | 20 | 20:42756461:42791466 | hsa-circ-PTPRT.15 | 0.2 |
| STK32B | 4 | 4:5139904:5168450 | hsa-circ-STK32B.6, hsa_circ_0004536 | 0.5 |
| BVES | 6 | 6:105115685:105125578 | hsa-circ-BVES.2, hsa_circ_0077527 | 0.3 |
| β-cell-selective circRNAs | ||||
| TGFBR3 | 1 | 1:91861470:91861644 | hsa-circ-TGFBR3.25, hsa_circ_0006622 | 3.3 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kaur, S.; Mirza, A.H.; Pociot, F. Cell Type-Selective Expression of Circular RNAs in Human Pancreatic Islets. Non-Coding RNA 2018, 4, 38. https://doi.org/10.3390/ncrna4040038
Kaur S, Mirza AH, Pociot F. Cell Type-Selective Expression of Circular RNAs in Human Pancreatic Islets. Non-Coding RNA. 2018; 4(4):38. https://doi.org/10.3390/ncrna4040038
Chicago/Turabian StyleKaur, Simranjeet, Aashiq H. Mirza, and Flemming Pociot. 2018. "Cell Type-Selective Expression of Circular RNAs in Human Pancreatic Islets" Non-Coding RNA 4, no. 4: 38. https://doi.org/10.3390/ncrna4040038
APA StyleKaur, S., Mirza, A. H., & Pociot, F. (2018). Cell Type-Selective Expression of Circular RNAs in Human Pancreatic Islets. Non-Coding RNA, 4(4), 38. https://doi.org/10.3390/ncrna4040038







