Alternative Non-Mammalian Animal and Cellular Methods for the Study of Host–Fungal Interactions
Abstract
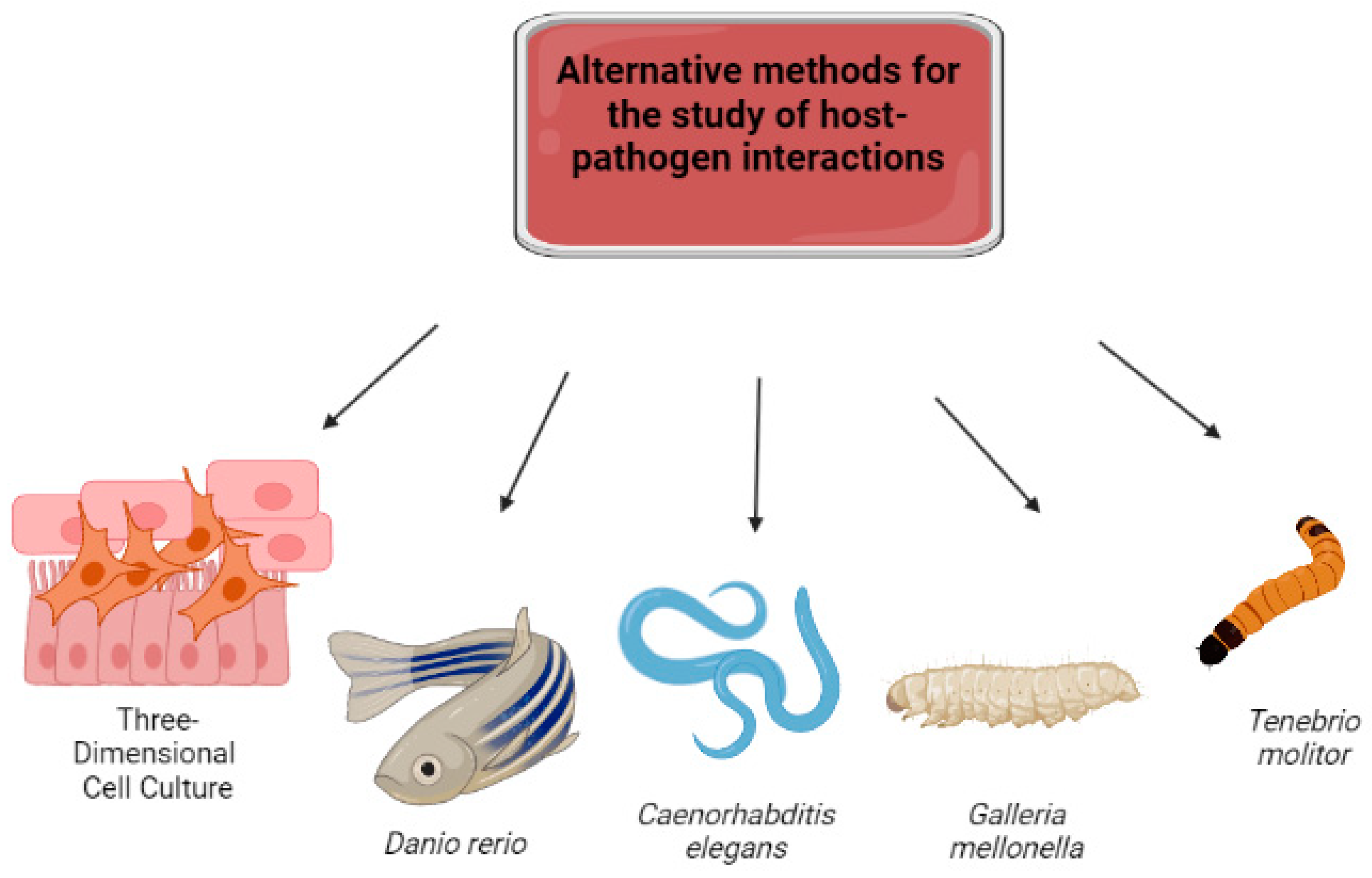
1. Introduction
2. Cell Culture
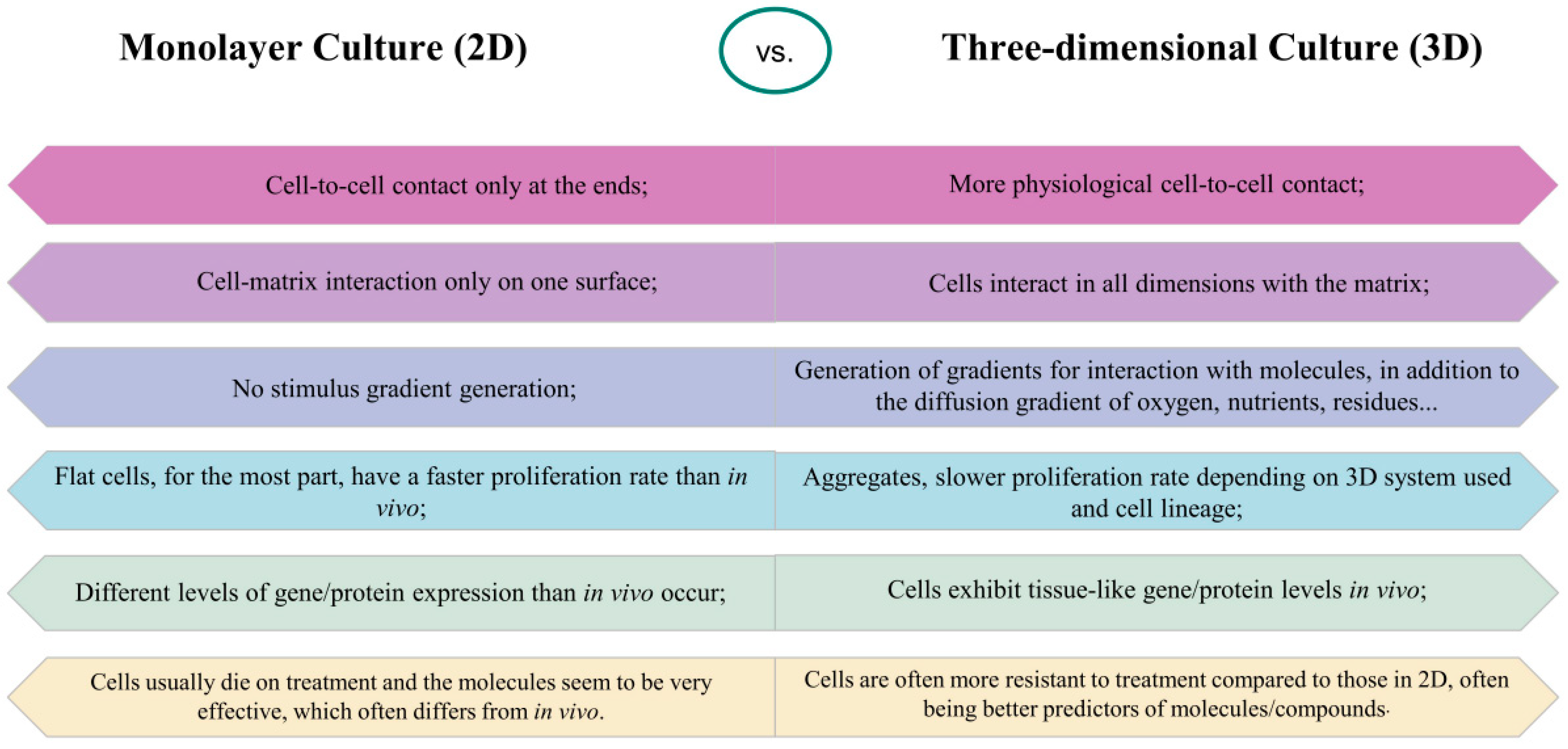
2.1. Three-Dimensional Cultures
2.2. Three-Dimensional Cultures in Fungal Pathogenesis
2.2.1. Organoids
2.2.2. Skin and Mucous Membrane Model
3. The Danio rerio Model
D. rerio as a Model of Fungal Infections
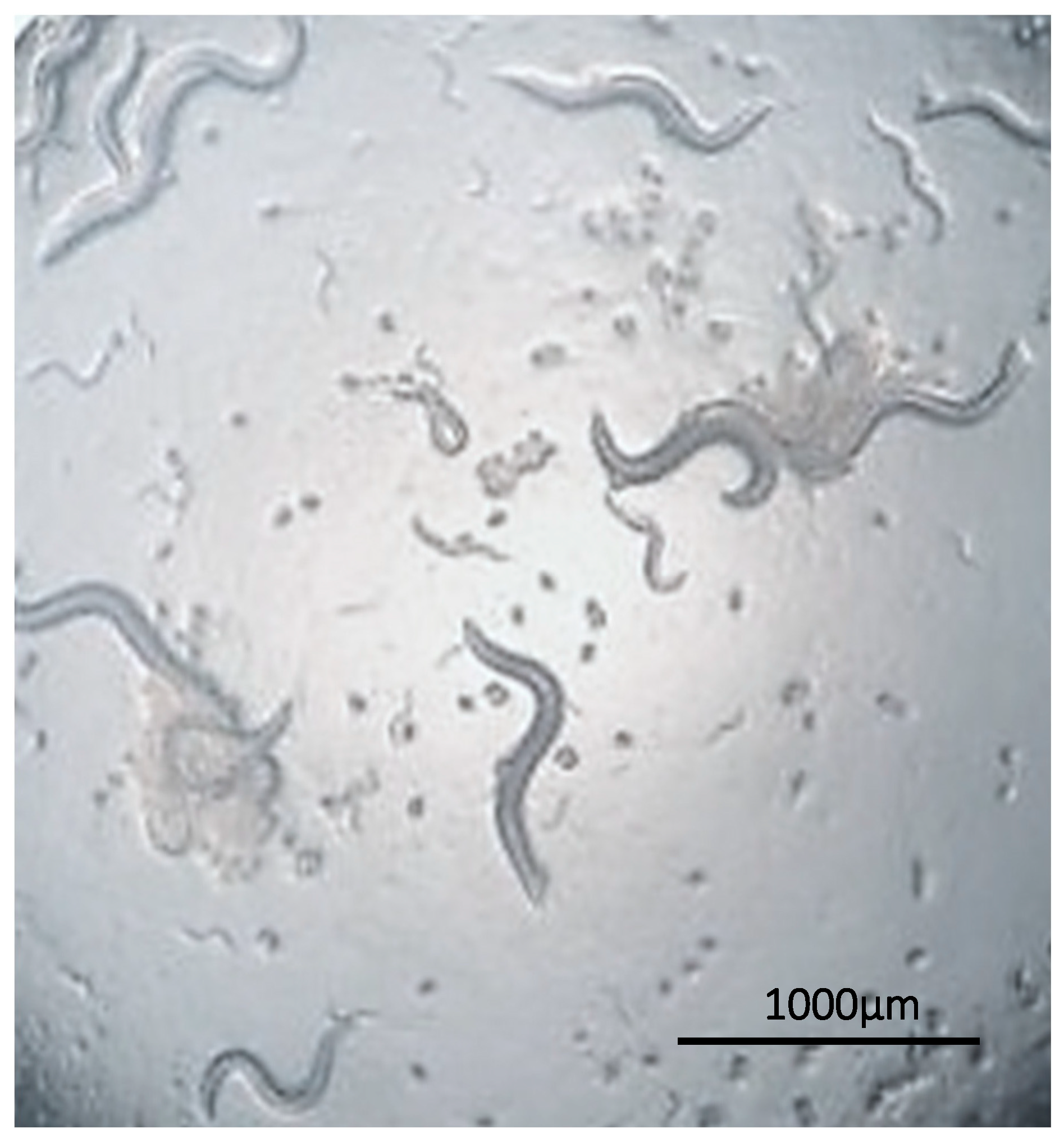
4. The Caenorhabditis elegans
4.1. C. elegans as an Infection Model of Pathogenic Fungi
4.2. C. elegans as a Model for New Antifungal Assessments
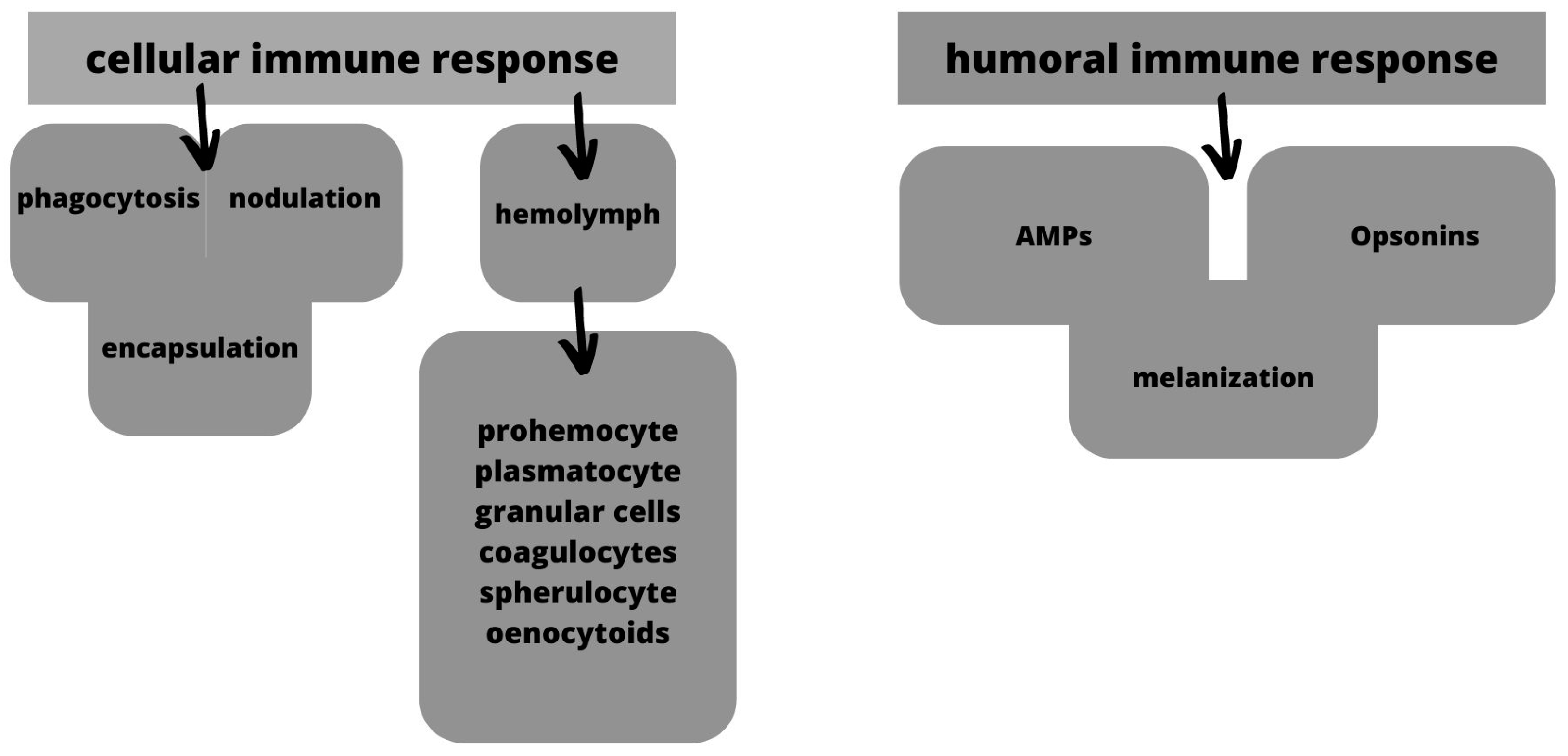
5. The Galleria mellonella Model
G. mellonella as a Host–Fungi Interaction Model
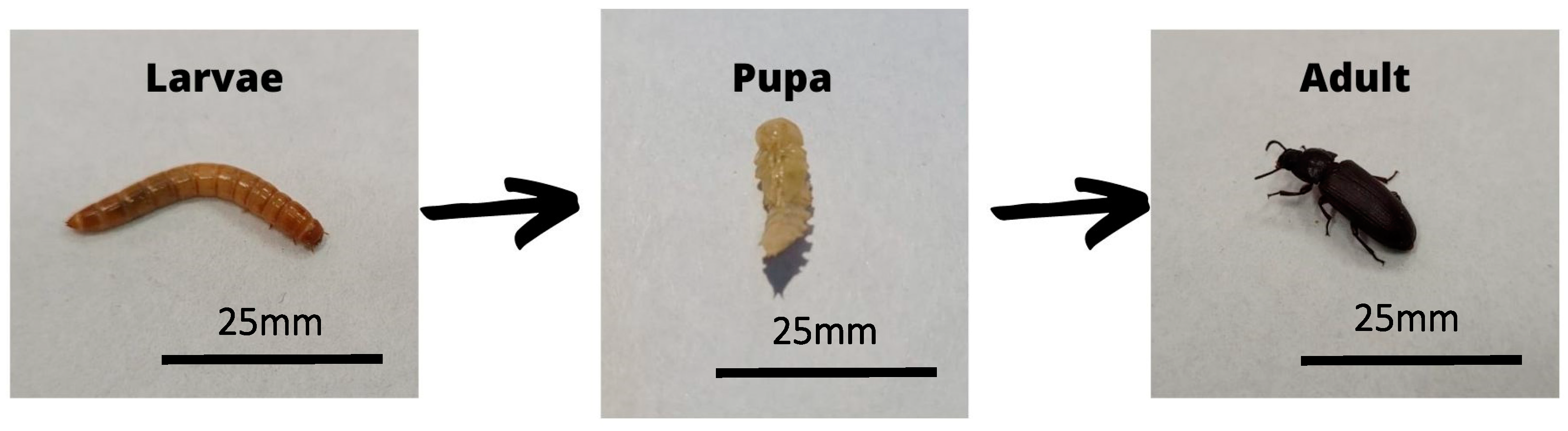
6. The Tenebrio molitor Model
7. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Bongomin, F.; Gago, S.; Oladele, R.; Denning, D. Global and Multi-National Prevalence of Fungal Diseases—Estimate Precision. J. Fungi 2017, 3, 57. [Google Scholar] [CrossRef] [PubMed]
- Kwon-Chung, K.J. Taxonomy of Fungi Causing Mucormycosis and Entomophthoramycosis (Zygomycosis) and Nomenclature of the Disease: Molecular Mycologic Perspectives. Clin. Infect. Dis. 2012, 54, S8–S15. [Google Scholar] [CrossRef] [PubMed]
- Rubin, L.G.; Levin, M.J.; Ljungman, P.; Davies, E.G.; Avery, R.; Tomblyn, M.; Bousvaros, A.; Dhanireddy, S.; Sung, L.; Keyserling, H.; et al. 2013 IDSA Clinical Practice Guideline for Vaccination of the Immunocompromised Host. Clin. Infect. Dis. 2014, 58, e44–e100. [Google Scholar] [CrossRef]
- Peng, J.; Wang, Q.; Mei, H.; Zheng, H.; Liang, G.; She, X.; Liu, W. Fungal Co-Infection in COVID-19 Patients: Evidence from a Systematic Review and Meta-Analysis. Aging 2021, 13, 7745–7757. [Google Scholar] [CrossRef] [PubMed]
- Moskaluk, A.E.; VandeWoude, S. Current Topics in Dermatophyte Classification and Clinical Diagnosis. Pathogens 2022, 11, 957. [Google Scholar] [CrossRef]
- Dias, M.F.R.G.; Quaresma-Santos, M.V.P.; Bernardes-Filho, F.; Amorim, A.G.d.F.; Schechtman, R.C.; Azulay, D.R. Update on Therapy for Superficial Mycoses: Review Article Part I. An. Bras. Dermatol. 2013, 88, 764–774. [Google Scholar] [CrossRef]
- Rajendran, R.; Sherry, L.; Nile, C.J.; Sherriff, A.; Johnson, E.M.; Hanson, M.F.; Williams, C.; Munro, C.A.; Jones, B.J.; Ramage, G. Biofilm Formation Is a Risk Factor for Mortality in Patients with Candida albicans Bloodstream Infection—Scotland, 2012–2013. Clin. Microbiol. Infect. 2016, 22, 87–93. [Google Scholar] [CrossRef]
- Vichare, A.S.; Kamath, S.U.; Leist, M.; Hayes, A.W.; Mahadevan, B. Application of the 3Rs Principles in the Development of Pharmaceutical Generics. Regul. Toxicol. Pharmacol. 2021, 125, 105016. [Google Scholar] [CrossRef]
- Kim, J.H.; Cheng, L.W.; Chan, K.L.; Tam, C.C.; Mahoney, N.; Friedman, M.; Shilman, M.M.; Land, K.M. Antifungal Drug Repurposing. Antibiotics 2020, 9, 812. [Google Scholar] [CrossRef]
- Sneddon, L.U.; Halsey, L.G.; Bury, N.R. Considering Aspects of the 3Rs Principles within Experimental Animal Biology. J. Exp. Biol. 2017, 220, 3007–3016. [Google Scholar] [CrossRef]
- MacArthur Clark, J. The 3Rs in Research: A Contemporary Approach to Replacement, Reduction and Refinement. Br. J. Nutr. 2018, 120, S1–S7. [Google Scholar] [CrossRef]
- Wange, R.L.; Brown, P.C.; Davis-Bruno, K.L. Implementation of the Principles of the 3Rs of Animal Testing at CDER: Past, Present and Future. Regul. Toxicol. Pharmacol. 2021, 123, 104953. [Google Scholar] [CrossRef] [PubMed]
- Arck, P.C. When 3 Rs Meet a Forth R: Replacement, Reduction and Refinement of Animals in Research on Reproduction. J. Reprod. Immunol. 2019, 132, 54–59. [Google Scholar] [CrossRef]
- Molinaro, E.M.; Caputo, L.F.G.; Amendoeira, M.R.R. Conceitos e Métodos Para a Formação de Profissionais Em Laboratórios de Saúde, v. 2; Molinaro, E.M., Caputo, L.F.G., Amendoeira, M.R.R., Eds.; EPSJV: Rio de Janeiro, Brazil, 2010; 254p. [Google Scholar]
- Traversa, D.; Joachim, A. The 3Rs Concept: Time to Change How We Evaluate the Efficacy of Anthelmintics in Companion Animals. Trends Parasitol. 2018, 34, 41–52. [Google Scholar] [CrossRef] [PubMed]
- Grässer, U.; Bubel, M.; Sossong, D.; Oberringer, M.; Pohlemann, T.; Metzger, W. Dissociation of Mono- and Co-Culture Spheroids into Single Cells for Subsequent Flow Cytometric Analysis. Ann. Anat. Anat. Anz. 2018, 216, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Braz, J.D.; Sardi, J.d.C.O.; Pitangui, N.d.S.; Voltan, A.R.; Almeida, A.M.F.; Mendes-Giannini, M.J.S. Gene Expression of Paracoccidioides Virulence Factors after Interaction with Macrophages and Fibroblasts. Mem. Inst. Oswaldo Cruz 2021, 116, e200592. [Google Scholar] [CrossRef]
- Alonso, M.; Gow, N.; Erwig, L.; Bain, J. Macrophage Migration Is Impaired within Candida albicans Biofilms. J. Fungi 2017, 3, 31. [Google Scholar] [CrossRef]
- Chandra, J.; McCormick, T.S.; Imamura, Y.; Mukherjee, P.K.; Ghannoum, M.A. Interaction of Candida albicans with Adherent Human Peripheral Blood Mononuclear Cells Increases C. Albicans Biofilm Formation and Results in Differential Expression of Pro- and Anti-Inflammatory Cytokines. Infect. Immun. 2007, 75, 2612–2620. [Google Scholar] [CrossRef]
- Almeida Donanzam, D.d.F.; Donato, T.A.G.; dos Reis, K.H.; da Silva, A.P.; Finato, A.C.; dos Santos, A.R.; Cavalcante, R.S.; Mendes, R.P.; Venturini, J. Exoantigens of Paracoccidioides spp. Promote Proliferation and Modulation of Human and Mouse Pulmonary Fibroblasts. Front. Cell Infect. Microbiol. 2020, 10, 590025. [Google Scholar] [CrossRef]
- Lopez, C.M.; Wallich, R.; Riesbeck, K.; Skerka, C.; Zipfel, P.F. Candida albicans Uses the Surface Protein Gpm1 to Attach to Human Endothelial Cells and to Keratinocytes via the Adhesive Protein Vitronectin. PLoS ONE 2014, 9, e90796. [Google Scholar] [CrossRef]
- Hernández-Santos, N.; Wiesner, D.L.; Fites, J.S.; McDermott, A.J.; Warner, T.; Wüthrich, M.; Klein, B.S. Lung Epithelial Cells Coordinate Innate Lymphocytes and Immunity against Pulmonary Fungal Infection. Cell Host Microbe 2018, 23, 511–522.e5. [Google Scholar] [CrossRef] [PubMed]
- Rodríguez-Echeverri, C.; Gómez, B.L.; González, Á. Histoplasma capsulatum Activates Hematopoietic Stem Cells and Their Progenitors through a Mechanism Dependent on TLR2, TLR4, and Dectin-1. J. Fungi 2022, 8, 1108. [Google Scholar] [CrossRef] [PubMed]
- Friedrich, J.; Seidel, C.; Ebner, R.; Kunz-Schughart, L.A. Spheroid-Based Drug Screen: Considerations and Practical Approach. Nat. Protoc. 2009, 4, 309–324. [Google Scholar] [CrossRef]
- dos Santos, K.S.; Oliveira, L.T.; de Lima Fontes, M.; Migliato, K.F.; Fusco-Almeida, A.M.; Mendes Giannini, M.J.S.; Moroz, A. Alginate-Based 3D A549 Cell Culture Model to Study Paracoccidioides Infection. J. Fungi 2023, 9, 634. [Google Scholar] [CrossRef] [PubMed]
- Vaso, C.O.; Bila, N.M.; Pandolfi, F.; De Vita, D.; Bortolami, M.; Bonatti, J.L.C.; De Moraes Silva, R.A.; Gonçalves, L.N.C.; Tudino, V.; Costi, R.; et al. Evaluation of the Anti-Histoplasma capsulatum Activity of Indole and Nitrofuran Derivatives and Their Pharmacological Safety in Three-Dimensional Cell Cultures. Pharmaceutics 2022, 14, 1043. [Google Scholar] [CrossRef]
- Cardoso, B.D.; Castanheira, E.M.S.; Lanceros-Méndez, S.; Cardoso, V.F. Recent Advances on Cell Culture Platforms for In Vitro Drug Screening and Cell Therapies: From Conventional to Microfluidic Strategies. Adv. Healthc. Mater. 2023, 12, e2202936. [Google Scholar] [CrossRef]
- Justice, B.A.; Badr, N.A.; Felder, R.A. 3D Cell Culture Opens New Dimensions in Cell-Based Assays. Drug. Discov. Today 2009, 14, 102–107. [Google Scholar] [CrossRef]
- DesRochers, T.M.; Palma, E.; Kaplan, D.L. Tissue-Engineered Kidney Disease Models. Adv. Drug Deliv. Rev. 2014, 69–70, 67–80. [Google Scholar] [CrossRef]
- Weaver, V.M.; Petersen, O.W.; Wang, F.; Larabell, C.A.; Briand, P.; Damsky, C.; Bissell, M.J. Reversion of the Malignant Phenotype of Human Breast Cells in Three-Dimensional Culture and In Vivo by Integrin Blocking Antibodies. J. Cell Biol. 1997, 137, 231–245. [Google Scholar] [CrossRef]
- Wilson, H.V. A New Method by Which Sponges May Be Artificially Reared. Science 1907, 25, 912–915. [Google Scholar] [CrossRef]
- Elsdale, T.; Bard, J. Collagen Substrata for Studies on Cell Behavior. J. Cell Biol. 1972, 54, 626–637. [Google Scholar] [CrossRef] [PubMed]
- Nature, J. Goodbye, Flat Biology? Nature 2003, 424, 861. [Google Scholar] [CrossRef]
- Abbott, A. Biology’s New Dimension. Nature 2003, 424, 870–872. [Google Scholar] [CrossRef] [PubMed]
- Fennema, E.; Rivron, N.; Rouwkema, J.; van Blitterswijk, C.; de Boer, J. Spheroid culture as a tool for creating 3D complex tissues. Trends Biotechnol. 2013, 31, 108–115. [Google Scholar] [CrossRef]
- Ravi, M.; Paramesh, V.; Kaviya, S.R.; Anuradha, E.; Solomon, F.D.P. 3D Cell Culture Systems: Advantages and Applications. J. Cell Physiol. 2015, 230, 16–26. [Google Scholar] [CrossRef]
- Kim, J. Bin Three-Dimensional Tissue Culture Models in Cancer Biology. Semin. Cancer Biol. 2005, 15, 365–377. [Google Scholar] [CrossRef]
- Gurski, L.A.; Petrelli, N.J.; Jia, X.; Farach-Carson, M.C. 3D Matrices for Anti-Cancer Drug Testing and Development. Oncol. Issues 2010, 25, 20–25. [Google Scholar] [CrossRef]
- Xu, X.; Gurski, L.A.; Zhang, C.; Harrington, D.A.; Farach-Carson, M.C.; Jia, X. Recreating the Tumor Microenvironment in a Bilayer, Hyaluronic Acid Hydrogel Construct for the Growth of Prostate Cancer Spheroids. Biomaterials 2012, 33, 9049–9060. [Google Scholar] [CrossRef]
- Edmondson, R.; Broglie, J.J.; Adcock, A.F.; Yang, L. Three-Dimensional Cell Culture Systems and Their Applications in Drug Discovery and Cell-Based Biosensors. Assay Drug Dev. Technol. 2014, 12, 207–218. [Google Scholar] [CrossRef]
- Marei, I.; Abu Samaan, T.; Al-Quradaghi, M.A.; Farah, A.A.; Mahmud, S.H.; Ding, H.; Triggle, C.R. 3D Tissue-Engineered Vascular Drug Screening Platforms: Promise and Considerations. Front. Cardiovasc. Med. 2022, 9, 847554. [Google Scholar] [CrossRef]
- Ryu, N.-E.; Lee, S.-H.; Park, H. Spheroid Culture System Methods and Applications for Mesenchymal Stem Cells. Cells 2019, 8, 1620. [Google Scholar] [CrossRef] [PubMed]
- Decarli, M.C.; Amaral, R.; dos Santos, D.P.; Tofani, L.B.; Katayama, E.; Rezende, R.A.; da Silva, J.V.L.; Swiech, K.; Suazo, C.A.T.; Mota, C.; et al. Cell Spheroids as a Versatile Research Platform: Formation Mechanisms, High Throughput Production, Characterization and Applications. Biofabrication 2021, 13, 032002. [Google Scholar] [CrossRef] [PubMed]
- El-Sherbiny, I.M.; Yacoub, M.H. Hydrogel Scaffolds for Tissue Engineering: Progress and Challenges. Glob. Cardiol. Sci. Pract. 2013, 2013, 38. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Shim, K.Y.; Kim, B.; Sung, J.H. Hydrogel-based Three-dimensional Cell Culture for Organ-on-a-chip Applications. Biotechnol. Prog. 2017, 33, 580–589. [Google Scholar] [CrossRef]
- Pezzulo, A.A.; Starner, T.D.; Scheetz, T.E.; Traver, G.L.; Tilley, A.E.; Harvey, B.-G.; Crystal, R.G.; McCray, P.B.; Zabner, J. The Air-Liquid Interface and Use of Primary Cell Cultures Are Important to Recapitulate the Transcriptional Profile of in Vivo Airway Epithelia. Am. J. Physiol. Lung Cell. Mol. Physiol. 2011, 300, L25–L31. [Google Scholar] [CrossRef]
- Ramasamy, S.; Davoodi, P.; Vijayavenkataraman, S.; Teoh, J.H.; Thamizhchelvan, A.M.; Robinson, K.S.; Wu, B.; Fuh, J.Y.H.; DiColandrea, T.; Zhao, H.; et al. Optimized Construction of a Full Thickness Human Skin Equivalent Using 3D Bioprinting and a PCL/Collagen Dermal Scaffold. Bioprinting 2021, 21, e00123. [Google Scholar] [CrossRef]
- Chambers, E.S.; Vukmanovic-Stejic, M. Skin Barrier Immunity and Ageing. Immunology 2020, 160, 116–125. [Google Scholar] [CrossRef]
- Gao, C.; Lu, C.; Jian, Z.; Zhang, T.; Chen, Z.; Zhu, Q.; Tai, Z.; Liu, Y. 3D Bioprinting for Fabricating Artificial Skin Tissue. Colloids Surf. B Biointerfaces 2021, 208, 112041. [Google Scholar] [CrossRef]
- Singh, D.; Singh, D.; Han, S. 3D Printing of Scaffold for Cells Delivery: Advances in Skin Tissue Engineering. Polymers 2016, 8, 19. [Google Scholar] [CrossRef]
- Wang, X.; Zhang, X.; Dai, X.; Wang, X.; Li, X.; Diao, J.; Xu, T. Tumor-like Lung Cancer Model Based on 3D Bioprinting. 3 Biotech 2018, 8, 501. [Google Scholar] [CrossRef]
- Jiang, H.; Wang, D. The Microbial Zoo in the C. Elegans Intestine: Bacteria, Fungi and Viruses. Viruses 2018, 10, 85. [Google Scholar] [CrossRef] [PubMed]
- Brohem, C.A.; da Silva Cardeal, L.B.; Tiago, M.; Soengas, M.S.; de Moraes Barros, S.B.; Maria-Engler, S.S. Artificial Skin in Perspective: Concepts and Applications. Pigment. Cell Melanoma Res. 2011, 24, 35–50. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.S.; Sun, X.; Lee, J.-H.; Kim, H.-W.; Fu, X.; Leong, K.W. Advanced Drug Delivery Systems and Artificial Skin Grafts for Skin Wound Healing. Adv. Drug Deliv. Rev. 2019, 146, 209–239. [Google Scholar] [CrossRef] [PubMed]
- Low, Z.W.K.; Li, Z.; Owh, C.; Chee, P.L.; Ye, E.; Dan, K.; Chan, S.Y.; Young, D.J.; Loh, X.J. Recent Innovations in Artificial Skin. Biomater. Sci. 2020, 8, 776–797. [Google Scholar] [CrossRef]
- Wang, M.; Luo, Y.; Wang, T.; Wan, C.; Pan, L.; Pan, S.; He, K.; Neo, A.; Chen, X. Artificial Skin Perception. Adv. Mater. 2021, 33, 2003014. [Google Scholar] [CrossRef] [PubMed]
- Dongari-Bagtzoglou, A.; Kashleva, H. Development of a Novel Three-Dimensional in Vitro Model of Oral Candida Infection. Microb. Pathog. 2006, 40, 271–278. [Google Scholar] [CrossRef]
- Rouabhia, M.; Deslauriers, N. Production and Characterization of an in Vitro Engineered Human Oral Mucosa. Biochem. Cell Biol. 2002, 80, 189–195. [Google Scholar] [CrossRef]
- Claveau, I.; Mostefaoui, Y.; Rouabhia, M. Basement Membrane Protein and Matrix Metalloproteinase Deregulation in Engineered Human Oral Mucosa Following Infection with Candida albicans. Matrix Biol. 2004, 23, 477–486. [Google Scholar] [CrossRef]
- de Carvalho Dias, K.; de Sousa, D.L.; Barbugli, P.A.; Cerri, P.S.; Salih, V.M.; Vergani, C.E. Development and Characterization of a 3D Oral Mucosa Model as a Tool for Host-Pathogen Interactions. J. Microbiol. Methods 2018, 152, 52–60. [Google Scholar] [CrossRef]
- Lancaster, M.A.; Renner, M.; Martin, C.-A.; Wenzel, D.; Bicknell, L.S.; Hurles, M.E.; Homfray, T.; Penninger, J.M.; Jackson, A.P.; Knoblich, J.A. Cerebral Organoids Model Human Brain Development and Microcephaly. Nature 2013, 501, 373–379. [Google Scholar] [CrossRef]
- Driggers, R.W.; Ho, C.-Y.; Korhonen, E.M.; Kuivanen, S.; Jääskeläinen, A.J.; Smura, T.; Rosenberg, A.; Hill, D.A.; DeBiasi, R.L.; Vezina, G.; et al. Zika Virus Infection with Prolonged Maternal Viremia and Fetal Brain Abnormalities. N. Engl. J. Med. 2016, 374, 2142–2151. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Saucedo-Cuevas, L.; Shresta, S.; Gleeson, J.G. The Neurobiology of Zika Virus. Neuron 2016, 92, 949–958. [Google Scholar] [CrossRef] [PubMed]
- Garcez, P.P.; Loiola, E.C.; Madeiro da Costa, R.; Higa, L.M.; Trindade, P.; Delvecchio, R.; Nascimento, J.M.; Brindeiro, R.; Tanuri, A.; Rehen, S.K. Zika Virus Impairs Growth in Human Neurospheres and Brain Organoids. Science 2016, 352, 816–818. [Google Scholar] [CrossRef] [PubMed]
- Xu, M.; Lee, E.M.; Wen, Z.; Cheng, Y.; Huang, W.-K.; Qian, X.; TCW, J.; Kouznetsova, J.; Ogden, S.C.; Hammack, C.; et al. Identification of Small-Molecule Inhibitors of Zika Virus Infection and Induced Neural Cell Death via a Drug Repurposing Screen. Nat. Med. 2016, 22, 1101–1107. [Google Scholar] [CrossRef]
- Watanabe, M.; Buth, J.E.; Vishlaghi, N.; de la Torre-Ubieta, L.; Taxidis, J.; Khakh, B.S.; Coppola, G.; Pearson, C.A.; Yamauchi, K.; Gong, D.; et al. Self-Organized Cerebral Organoids with Human-Specific Features Predict Effective Drugs to Combat Zika Virus Infection. Cell Rep. 2017, 21, 517–532. [Google Scholar] [CrossRef] [PubMed]
- Zhou, T.; Tan, L.; Cederquist, G.Y.; Fan, Y.; Hartley, B.J.; Mukherjee, S.; Tomishima, M.; Brennand, K.J.; Zhang, Q.; Schwartz, R.E.; et al. High-Content Screening in HPSC-Neural Progenitors Identifies Drug Candidates That Inhibit Zika Virus Infection in Fetal-like Organoids and Adult Brain. Cell Stem Cell 2017, 21, 274–283.e5. [Google Scholar] [CrossRef] [PubMed]
- Finkbeiner, S.R.; Zeng, X.-L.; Utama, B.; Atmar, R.L.; Shroyer, N.F.; Estes, M.K. Stem Cell-Derived Human Intestinal Organoids as an Infection Model for Rotaviruses. mBio 2012, 3, e00159-12. [Google Scholar] [CrossRef]
- Yin, Y.; Bijvelds, M.; Dang, W.; Xu, L.; van der Eijk, A.A.; Knipping, K.; Tuysuz, N.; Dekkers, J.F.; Wang, Y.; de Jonge, J.; et al. Modeling Rotavirus Infection and Antiviral Therapy Using Primary Intestinal Organoids. Antivir. Res. 2015, 123, 120–131. [Google Scholar] [CrossRef]
- McCracken, K.W.; Catá, E.M.; Crawford, C.M.; Sinagoga, K.L.; Schumacher, M.; Rockich, B.E.; Tsai, Y.-H.; Mayhew, C.N.; Spence, J.R.; Zavros, Y.; et al. Modelling Human Development and Disease in Pluripotent Stem-Cell-Derived Gastric Organoids. Nature 2014, 516, 400–404. [Google Scholar] [CrossRef]
- Bertaux-Skeirik, N.; Feng, R.; Schumacher, M.A.; Li, J.; Mahe, M.M.; Engevik, A.C.; Javier, J.E.; Peek, R.M., Jr.; Ottemann, K.; Orian-Rousseau, V.; et al. CD44 Plays a Functional Role in Helicobacter Pylori-Induced Epithelial Cell Proliferation. PLoS Pathog. 2015, 11, e1004663. [Google Scholar] [CrossRef]
- Ettayebi, K.; Crawford, S.E.; Murakami, K.; Broughman, J.R.; Karandikar, U.; Tenge, V.R.; Neill, F.H.; Blutt, S.E.; Zeng, X.-L.; Qu, L.; et al. Replication of Human Noroviruses in Stem Cell-Derived Human Enteroids. Science 2016, 353, 1387–1393. [Google Scholar] [CrossRef] [PubMed]
- Jose, S.S.; De Zuani, M.; Tidu, F.; Hortová Kohoutková, M.; Pazzagli, L.; Forte, G.; Spaccapelo, R.; Zelante, T.; Frič, J. Comparison of Two Human Organoid Models of Lung and Intestinal Inflammation Reveals Toll-like Receptor Signalling Activation and Monocyte Recruitment. Clin. Transl. Immunol. 2020, 9, e1131. [Google Scholar] [CrossRef] [PubMed]
- Nam, S.Y.; Lee, S.J.; Lim, H.J.; Park, J.Y.; Jeon, S.W. Clinical Risk Factors and Pattern of Initial Fungal Contamination in Endoscopic Biopsy-Derived Gastrointestinal Cancer Organoid Culture. Korean J. Intern. Med. 2021, 36, 878–887. [Google Scholar] [CrossRef] [PubMed]
- Tisdale-Macioce, N.; Green, J.; Perl, A.-K.T.; Ashbaugh, A.; Wiederhold, N.P.; Patterson, T.F.; Cushion, M.T. The Promise of Lung Organoids for Growth and Investigation of Pneumocystis Species. Front. Fungal Biol. 2021, 2, 740845. [Google Scholar] [CrossRef]
- Zhao, Z.; Chen, X.; Dowbaj, A.M.; Sljukic, A.; Bratlie, K.; Lin, L.; Fong, E.L.S.; Balachander, G.M.; Chen, Z.; Soragni, A.; et al. Organoids. Nat. Rev. Methods Primers 2022, 2, 94. [Google Scholar] [CrossRef]
- Corrò, C.; Novellasdemunt, L.; Li, V.S.W. A Brief History of Organoids. Am. J. Physiol. Cell Physiol. 2020, 319, C151–C165. [Google Scholar] [CrossRef]
- Chandorkar, P.; Posch, W.; Zaderer, V.; Blatzer, M.; Steger, M.; Ammann, C.G.; Binder, U.; Hermann, M.; Hörtnagl, P.; Lass-Flörl, C.; et al. Fast-Track Development of an in Vitro 3D Lung/Immune Cell Model to Study Aspergillus Infections. Sci. Rep. 2017, 7, 11644. [Google Scholar] [CrossRef]
- Pimentel, B.N.A.d.S.; Marin-Dett, F.H.; Assis, M.; Barbugli, P.A.; Longo, E.; Vergani, C.E. Antifungal Activity and Biocompatibility of α-AgVO3, α-Ag2WO4, and β-Ag2MoO4 Using a Three-Dimensional Coculture Model of the Oral Mucosa. Front. Bioeng. Biotechnol. 2022, 10, 826123. [Google Scholar] [CrossRef]
- Creaser, C.W. The Technic of Handling the Zebra Fish (Brachydanio rerio) for the Production of Eggs Which Are Favorable for Embryological Research and Are Available at Any Specified Time Throughout the Year. Copeia 1934, 1934, 159. [Google Scholar] [CrossRef]
- Meyers, J.R. Zebrafish: Development of a Vertebrate Model Organism. Curr. Protoc. Essent. Lab. Tech. 2018, 16, e19. [Google Scholar] [CrossRef]
- Baird, S.F.; Hamilton, F.; Swaine, J. An Account of the Fishes Found in the River Ganges and Its Branches; Hurst, Robinson, and Co.: Edinburgh, UK, 1822. [Google Scholar]
- Sundin, J.; Morgan, R.; Finnøen, M.H.; Dey, A.; Sarkar, K.; Jutfelt, F. On the Observation of Wild Zebrafish (Danio rerio) in India. Zebrafish 2019, 16, 546–553. [Google Scholar] [CrossRef] [PubMed]
- Suriyampola, P.S.; Shelton, D.S.; Shukla, R.; Roy, T.; Bhat, A.; Martins, E.P. Zebrafish Social Behavior in the Wild. Zebrafish 2016, 13, 1–8. [Google Scholar] [CrossRef] [PubMed]
- McClure, M.M.; McIntyre, P.B.; McCune, A.R. Notes on the Natural Diet and Habitat of Eight Danionin Fishes, Including the Zebrafish Danio rerio. J. Fish Biol. 2006, 69, 553–570. [Google Scholar] [CrossRef]
- Streisinger, G.; Walker, C.; Dower, N.; Knauber, D.; Singer, F. Production of Clones of Homozygous Diploid Zebra Fish (Brachydanio rerio). Nature 1981, 291, 293–296. [Google Scholar] [CrossRef] [PubMed]
- Kimmel, C.B.; Ballard, W.W.; Kimmel, S.R.; Ullmann, B.; Schilling, T.F. Stages of Embryonic Development of the Zebrafish. Dev. Dyn. 1995, 203, 253–310. [Google Scholar] [CrossRef]
- Parichy, D.M.; Elizondo, M.R.; Mills, M.G.; Gordon, T.N.; Engeszer, R.E. Normal Table of Postembryonic Zebrafish Development: Staging by Externally Visible Anatomy of the Living Fish. Dev. Dyn. 2009, 238, 2975–3015. [Google Scholar] [CrossRef]
- Schneider, A.C.R.; Maurer, R.L.; Matte, U.; Porawski, M.; Schaefer, P.G.; dos Santos, J.L.; da Silveira, T.R. Implementação de Um Novo Modelo de Experimentação Animal: Zebrafish. HCPA 2009, 29, 100–102. [Google Scholar]
- Strähle, U.; Grabher, C. The Zebrafish Embryo as a Model for Assessing Off-Target Drug Effects. Dis. Models Mech. 2010, 3, 689–692. [Google Scholar] [CrossRef][Green Version]
- MacRae, C.A.; Peterson, R.T. Zebrafish as Tools for Drug Discovery. Nat. Rev. Drug Discov. 2015, 14, 721–731. [Google Scholar] [CrossRef]
- Stainier, D.Y.R.; Raz, E.; Lawson, N.D.; Ekker, S.C.; Burdine, R.D.; Eisen, J.S.; Ingham, P.W.; Schulte-Merker, S.; Yelon, D.; Weinstein, B.M.; et al. Guidelines for Morpholino Use in Zebrafish. PLoS Genet. 2017, 13, e1007000. [Google Scholar] [CrossRef]
- Torraca, V.; Mostowy, S. Zebrafish Infection: From Pathogenesis to Cell Biology. Trends Cell Biol. 2018, 28, 143–156. [Google Scholar] [CrossRef]
- Galeane, M.C.; Gomes, P.C.; Singulani, J.d.L.; de Souza, B.M.; Palma, M.S.; Mendes-Giannini, M.J.; Almeida, A.M. Study of Mastoparan Analog Peptides against Candida albicans and Safety in Zebrafish Embryos (Danio rerio). Future Microbiol. 2019, 14, 1087–1097. [Google Scholar] [CrossRef] [PubMed]
- Gratacap, R.L.; Scherer, A.K.; Seman, B.G.; Wheeler, R.T. Control of Mucosal Candidiasis in the Zebrafish Swim Bladder Depends on Neutrophils That Block Filament Invasion and Drive Extracellular-Trap Production. Infect. Immun. 2017, 85, e00276-17. [Google Scholar] [CrossRef] [PubMed]
- Ho, J.; Yang, X.; Nikou, S.-A.; Kichik, N.; Donkin, A.; Ponde, N.O.; Richardson, J.P.; Gratacap, R.L.; Archambault, L.S.; Zwirner, C.P.; et al. Candidalysin Activates Innate Epithelial Immune Responses via Epidermal Growth Factor Receptor. Nat. Commun. 2019, 10, 2297. [Google Scholar] [CrossRef] [PubMed]
- Scherer, A.K.; Blair, B.A.; Park, J.; Seman, B.G.; Kelley, J.B.; Wheeler, R.T. Redundant Trojan Horse and Endothelial-Circulatory Mechanisms for Host-Mediated Spread of Candida albicans Yeast. PLoS Pathog. 2020, 16, e1008414. [Google Scholar] [CrossRef]
- Archambault, L.S.; Trzilova, D.; Gonia, S.; Gale, C.; Wheeler, R.T. Intravital Imaging Reveals Divergent Cytokine and Cellular Immune Responses to Candida albicans and Candida Parapsilosis. mBio 2019, 10, e00266-19. [Google Scholar] [CrossRef]
- Johnson, C.J.; Davis, J.M.; Huttenlocher, A.; Kernien, J.F.; Nett, J.E. Emerging Fungal Pathogen Candida Auris Evades Neutrophil Attack. mBio 2018, 9, e01403-18. [Google Scholar] [CrossRef]
- Koch, B.E.V.; Hajdamowicz, N.H.; Lagendijk, E.; Ram, A.F.J.; Meijer, A.H. Aspergillus fumigatus Establishes Infection in Zebrafish by Germination of Phagocytized Conidia, While Aspergillus Niger Relies on Extracellular Germination. Sci. Rep. 2019, 9, 12791. [Google Scholar] [CrossRef]
- Lim, J.-Y.; Kim, Y.J.; Woo, S.A.; Jeong, J.W.; Lee, Y.-R.; Kim, C.-H.; Park, H.-M. The LAMMER Kinase, LkhA, Affects Aspergillus fumigatus Pathogenicity by Modulating Reproduction and Biosynthesis of Cell Wall PAMPs. Front. Cell Infect. Microbiol. 2021, 11, 756206. [Google Scholar] [CrossRef]
- Rosowski, E.E.; Raffa, N.; Knox, B.P.; Golenberg, N.; Keller, N.P.; Huttenlocher, A. Macrophages Inhibit Aspergillus fumigatus Germination and Neutrophil-Mediated Fungal Killing. PLoS Pathog. 2018, 14, e1007229. [Google Scholar] [CrossRef]
- Schoen, T.J.; Rosowski, E.E.; Knox, B.P.; Bennin, D.; Keller, N.P.; Huttenlocher, A. Neutrophil Phagocyte Oxidase Activity Controls Invasive Fungal Growth and Inflammation in Zebrafish. J. Cell Sci. 2020, 133, jcs236539. [Google Scholar] [CrossRef] [PubMed]
- Pazhakh, V.; Ellett, F.; Croker, B.A.; O’Donnell, J.A.; Pase, L.; Schulze, K.E.; Greulich, R.S.; Gupta, A.; Reyes-Aldasoro, C.C.; Andrianopoulos, A.; et al. β-Glucan–Dependent Shuttling of Conidia from Neutrophils to Macrophages Occurs during Fungal Infection Establishment. PLoS Biol. 2019, 17, e3000113. [Google Scholar] [CrossRef] [PubMed]
- Ellett, F.; Pazhakh, V.; Pase, L.; Benard, E.L.; Weerasinghe, H.; Azabdaftari, D.; Alasmari, S.; Andrianopoulos, A.; Lieschke, G.J. Macrophages Protect Talaromyces Marneffei Conidia from Myeloperoxidase-Dependent Neutrophil Fungicidal Activity during Infection Establishment in Vivo. PLoS Pathog. 2018, 14, e1007063. [Google Scholar] [CrossRef]
- Singulani, J.d.L.; Galeane, M.C.; Ramos, M.D.; Gomes, P.C.; dos Santos, C.T.; de Souza, B.M.; Palma, M.S.; Fusco Almeida, A.M.; Mendes Giannini, M.J.S. Antifungal Activity, Toxicity, and Membranolytic Action of a Mastoparan Analog Peptide. Front. Cell Infect. Microbiol. 2019, 9, 419. [Google Scholar] [CrossRef]
- Singulani, J.d.L.; Oliveira, L.T.; Ramos, M.D.; Fregonezi, N.F.; Gomes, P.C.; Galeane, M.C.; Palma, M.S.; Fusco Almeida, A.M.; Mendes Giannini, M.J.S. The Antimicrobial Peptide MK58911-NH 2 Acts on Planktonic, Biofilm, and Intramacrophage Cells of Cryptococcus Neoformans. Antimicrob. Agents Chemother. 2021, 65, e0090421. [Google Scholar] [CrossRef]
- Tenor, J.L.; Oehlers, S.H.; Yang, J.L.; Tobin, D.M.; Perfect, J.R. Live Imaging of Host-Parasite Interactions in a Zebrafish Infection Model Reveals Cryptococcal Determinants of Virulence and Central Nervous System Invasion. mBio 2015, 6, e01425-15. [Google Scholar] [CrossRef] [PubMed]
- Bojarczuk, A.; Miller, K.A.; Hotham, R.; Lewis, A.; Ogryzko, N.V.; Kamuyango, A.A.; Frost, H.; Gibson, R.H.; Stillman, E.; May, R.C.; et al. Cryptococcus Neoformans Intracellular Proliferation and Capsule Size Determines Early Macrophage Control of Infection. Sci. Rep. 2016, 6, 21489. [Google Scholar] [CrossRef]
- Inglesfield, S.; Jasiulewicz, A.; Hopwood, M.; Tyrrell, J.; Youlden, G.; Mazon-Moya, M.; Millington, O.R.; Mostowy, S.; Jabbari, S.; Voelz, K. Robust Phagocyte Recruitment Controls the Opportunistic Fungal Pathogen Mucor Circinelloides in Innate Granulomas In Vivo. mBio 2018, 9, e02010-17. [Google Scholar] [CrossRef]
- López-Muñoz, A.; Nicolás, F.E.; García-Moreno, D.; Pérez-Oliva, A.B.; Navarro-Mendoza, M.I.; Hernández-Oñate, M.A.; Herrera-Estrella, A.; Torres-Martínez, S.; Ruiz-Vázquez, R.M.; Garre, V.; et al. An Adult Zebrafish Model Reveals That Mucormycosis Induces Apoptosis of Infected Macrophages. Sci. Rep. 2018, 8, 12802. [Google Scholar] [CrossRef]
- Lips, K.R. Overview of Chytrid Emergence and Impacts on Amphibians. Philos. Trans. R. Soc. B Biol. Sci. 2016, 371, 20150465. [Google Scholar] [CrossRef]
- Liew, N.; Mazon Moya, M.J.; Wierzbicki, C.J.; Hollinshead, M.; Dillon, M.J.; Thornton, C.R.; Ellison, A.; Cable, J.; Fisher, M.C.; Mostowy, S. Chytrid Fungus Infection in Zebrafish Demonstrates That the Pathogen Can Parasitize Non-Amphibian Vertebrate Hosts. Nat. Commun. 2017, 8, 15048. [Google Scholar] [CrossRef] [PubMed]
- Schulenburg, H.; Félix, M.-A. The Natural Biotic Environment of Caenorhabditis Elegans. Genetics 2017, 206, 55–86. [Google Scholar] [CrossRef] [PubMed]
- Hunt, P.R. The C. Elegans Model in Toxicity Testing. J. Appl. Toxicol. 2017, 37, 50–59. [Google Scholar] [CrossRef]
- Ballestriero, F.; Thomas, T.; Burke, C.; Egan, S.; Kjelleberg, S. Identification of Compounds with Bioactivity against the Nematode Caenorhabditis Elegans by a Screen Based on the Functional Genomics of the Marine Bacterium Pseudoalteromonas Tunicata D2. Appl. Environ. Microbiol. 2010, 76, 5710–5717. [Google Scholar] [CrossRef]
- Huang, X.; Li, D.; Xi, L.; Mylonakis, E. Caenorhabditis Elegans: A Simple Nematode Infection Model for Penicillium Marneffei. PLoS ONE 2014, 9, e108764. [Google Scholar] [CrossRef] [PubMed]
- Elkabti, A.; Issi, L.; Rao, R. Caenorhabditis Elegans as a Model Host to Monitor the Candida Infection Processes. J. Fungi 2018, 4, 123. [Google Scholar] [CrossRef]
- Mylonakis, E.; Aballay, A. Worms and Flies as Genetically Tractable Animal Models To Study Host-Pathogen Interactions. Infect. Immun. 2005, 73, 3833–3841. [Google Scholar] [CrossRef]
- Tan, M.-W.; Mahajan-Miklos, S.; Ausubel, F.M. Killing of Caenorhabditis Elegans by Pseudomonas Aeruginosa Used to Model Mammalian Bacterial Pathogenesis. Proc. Natl. Acad. Sci. USA 1999, 96, 715–720. [Google Scholar] [CrossRef]
- Tan, M.-W.; Rahme, L.G.; Sternberg, J.A.; Tompkins, R.G.; Ausubel, F.M. Pseudomonas Aeruginosa Killing of Caenorhabditis Elegans Used to Identify P. Aeruginosa Virulence Factors. Proc. Natl. Acad. Sci. USA 1999, 96, 2408–2413. [Google Scholar] [CrossRef]
- Mylonakis, E.; Idnurm, A.; Moreno, R.; El Khoury, J.; Rottman, J.B.; Ausubel, F.M.; Heitman, J.; Calderwood, S.B. Cryptococcus Neoformans Kin1 Protein Kinase Homologue, Identified through a Caenorhabditis Elegans Screen, Promotes Virulence in Mammals. Mol. Microbiol. 2004, 54, 407–419. [Google Scholar] [CrossRef]
- Okoli, I.; Bignell, E. Caenorhabditis Elegans-Aspergillus fumigatus (Nematode-Mould) Model for Study of Fungal Pathogenesis. Br. Microbiol. Res. J. 2015, 7, 93–99. [Google Scholar] [CrossRef]
- Okoli, I.; Coleman, J.J.; Tempakakis, E.; An, W.F.; Holson, E.; Wagner, F.; Conery, A.L.; Larkins-Ford, J.; Wu, G.; Stern, A.; et al. Identification of Antifungal Compounds Active against Candida albicans Using an Improved High-Throughput Caenorhabditis Elegans Assay. PLoS ONE 2009, 4, e7025. [Google Scholar] [CrossRef] [PubMed]
- Brenner, S. The Genetics of Caenorhabditis Elegans. Genetics 1974, 77, 71–94. [Google Scholar] [CrossRef] [PubMed]
- Lewis, K. Riddle of Biofilm Resistance. Antimicrob. Agents Chemother. 2001, 45, 999–1007. [Google Scholar] [CrossRef]
- Madende, M.; Albertyn, J.; Sebolai, O.; Pohl, C.H. Caenorhabditis Elegans as a Model Animal for Investigating Fungal Pathogenesis. Med. Microbiol. Immunol. 2020, 209, 1–13. [Google Scholar] [CrossRef]
- Irazoqui, J.E.; Troemel, E.R.; Feinbaum, R.L.; Luhachack, L.G.; Cezairliyan, B.O.; Ausubel, F.M. Distinct Pathogenesis and Host Responses during Infection of C. Elegans by P. Aeruginosa and S. Aureus. PLoS Pathog. 2010, 6, e1000982. [Google Scholar] [CrossRef]
- Sem, X.; Rhen, M. Pathogenicity of Salmonella Enterica in Caenorhabditis Elegans Relies on Disseminated Oxidative Stress in the Infected Host. PLoS ONE 2012, 7, e45417. [Google Scholar] [CrossRef]
- Mylonakis, E.; Ausubel, F.M.; Perfect, J.R.; Heitman, J.; Calderwood, S.B. Killing of Caenorhabditiselegansby Cryptococcus Neoformans as a Model of Yeast Pathogenesis. Proc. Natl. Acad. Sci. USA 2002, 99, 15675–15680. [Google Scholar] [CrossRef]
- Desalermos, A.; Tan, X.; Rajamuthiah, R.; Arvanitis, M.; Wang, Y.; Li, D.; Kourkoumpetis, T.K.; Fuchs, B.B.; Mylonakis, E. A Multi-Host Approach for the Systematic Analysis of Virulence Factors in Cryptococcus neoformans. J. Infect. Dis. 2015, 211, 298–305. [Google Scholar] [CrossRef]
- Jain, C.; Pastor, K.; Gonzalez, A.Y.; Lorenz, M.C.; Rao, R.P. The Role of Candida albicans AP-1 Protein against Host Derived ROS in in Vivo Models of Infection. Virulence 2013, 4, 67–76. [Google Scholar] [CrossRef]
- Couillault, C.; Fourquet, P.; Pophillat, M.; Ewbank, J.J. A UPR-Independent Infection-Specific Role for a BiP/GRP78 Protein in the Control of Antimicrobial Peptide Expression in C. elegans Epidermis. Virulence 2012, 3, 299–308. [Google Scholar] [CrossRef] [PubMed]
- Zugasti, O.; Thakur, N.; Belougne, J.; Squiban, B.; Kurz, C.L.; Soulé, J.; Omi, S.; Tichit, L.; Pujol, N.; Ewbank, J.J. A Quantitative Genome-Wide RNAi Screen in C. Elegans for Antifungal Innate Immunity Genes. BMC Biol. 2016, 14, 35. [Google Scholar] [CrossRef]
- Ahamefule, C.S.; Ezeuduji, B.C.; Ogbonna, J.C.; Moneke, A.N.; Ike, A.C.; Jin, C.; Wang, B.; Fang, W. Caenorhabditis Elegans as an Infection Model for Pathogenic Mold and Dimorphic Fungi: Applications and Challenges. Front. Cell Infect. Microbiol. 2021, 11, 751947. [Google Scholar] [CrossRef] [PubMed]
- Ahamefule, C.S.; Qin, Q.; Odiba, A.S.; Li, S.; Moneke, A.N.; Ogbonna, J.C.; Jin, C.; Wang, B.; Fang, W. Caenorhabditis Elegans-Based Aspergillus fumigatus Infection Model for Evaluating Pathogenicity and Drug Efficacy. Front. Cell Infect. Microbiol. 2020, 10, 320. [Google Scholar] [CrossRef]
- Snelders, E.; Huis in’t Veld, R.A.G.; Rijs, A.J.M.M.; Kema, G.H.J.; Melchers, W.J.G.; Verweij, P.E. Possible Environmental Origin of Resistance of Aspergillus fumigatus to Medical Triazoles. Appl. Environ. Microbiol. 2009, 75, 4053–4057. [Google Scholar] [CrossRef]
- Fang, W.; Latgé, J.-P. Microbe Profile: Aspergillus fumigatus: A Saprotrophic and Opportunistic Fungal Pathogen. Microbiology 2018, 164, 1009–1011. [Google Scholar] [CrossRef]
- Geißel, B.; Loiko, V.; Klugherz, I.; Zhu, Z.; Wagener, N.; Kurzai, O.; van den Hondel, C.A.M.J.J.; Wagener, J. Azole-Induced Cell Wall Carbohydrate Patches Kill Aspergillus fumigatus. Nat. Commun. 2018, 9, 3098. [Google Scholar] [CrossRef]
- Pukkila-Worley, R.; Holson, E.; Wagner, F.; Mylonakis, E. Antifungal Drug Discovery through the Study of Invertebrate Model Hosts. Curr. Med. Chem. 2009, 16, 1588–1595. [Google Scholar] [CrossRef]
- Souza, A.C.R.; Fuchs, B.B.; Alves, V.d.S.; Jayamani, E.; Colombo, A.L.; Mylonakis, E. Pathogenesis of the Candida Parapsilosis Complex in the Model Host Caenorhabditis Elegans. Genes 2018, 9, 401. [Google Scholar] [CrossRef]
- Jansson, H.B. Adhesion of Conidia of Drechmeria Coniospora to Caenorhabditis Elegans Wild Type and Mutants. J. Nematol. 1994, 26, 430–435. [Google Scholar]
- Zhang, X.; Harding, B.W.; Aggad, D.; Courtine, D.; Chen, J.-X.; Pujol, N.; Ewbank, J.J. Antagonistic Fungal Enterotoxins Intersect at Multiple Levels with Host Innate Immune Defences. PLoS Genet. 2021, 17, e1009600. [Google Scholar] [CrossRef]
- Mylonakis, E.; Flanigan, T.P. Editorial Response: Antifungal Prophylaxis with Weekly Fluconazole for Patients with AIDS. Clin. Infect. Dis. 1998, 27, 1376–1378. [Google Scholar] [CrossRef][Green Version]
- Gumbo, T.; Kadzirange, G.; Mielke, J.; Gangaidzo, I.T.; Hakim, J.G. Cryptococcus Neoformans Meningoencephalitis in African Children with Acquired Immunodeficiency Syndrome. Pediatr. Infect. Dis. J. 2002, 21, 54–56. [Google Scholar] [CrossRef] [PubMed]
- Perfect, J.R.; Casadevall, A. Cryptococcosis. Infect. Dis. Clin. N. Am. 2002, 16, 837–874. [Google Scholar] [CrossRef]
- Merryman, M.; Crigler, J.; Seipelt-Thiemann, R.; McClelland, E. A Mutation in C. Neoformans Mitochondrial NADH Dehydrogenase Results in Increased Virulence in Mice. Virulence 2020, 11, 1366–1378. [Google Scholar] [CrossRef] [PubMed]
- Mittal, J.; Ponce, M.G.; Gendlina, I.; Nosanchuk, J.D. Histoplasma capsulatum: Mechanisms for Pathogenesis. Curr. Top. Microbiol. Immunol. 2019, 422, 157–191. [Google Scholar] [CrossRef]
- Johnson, C.H.; Ayyadevara, S.; Mcewen, J.E.; Shmookler Reis, R.J. Histoplasma capsulatum and Caenorhabditis Elegans: A Simple Nematode Model for an Innate Immune Response to Fungal Infection. Med. Mycol. 2009, 47, 808–813. [Google Scholar] [CrossRef]
- Scorzoni, L.; de Lucas, M.P.; Singulani, J.d.L.; de Oliveira, H.C.; Assato, P.A.; Fusco-Almeida, A.M.; Mendes-Giannini, M.J.S. Evaluation of Caenorhabditis Elegans as a Host Model for Paracoccidioides brasiliensis and Paracoccidioides lutzii. Pathog. Dis. 2018, 76, fty004. [Google Scholar] [CrossRef]
- Scorzoni, L.; Alves de Paula e Silva, A.C.; de Oliveira, H.C.; Tavares dos Santos, C.; de Lacorte Singulani, J.; Akemi Assato, P.; Maria Marcos, C.; Teodoro Oliveira, L.; Ferreira Fregonezi, N.; Rossi, D.C.P.; et al. In Vitro and In Vivo Effect of Peptides Derived from 14-3-3 Paracoccidioides spp. Protein. J. Fungi 2021, 7, 52. [Google Scholar] [CrossRef]
- Navas, A.; Cobas, G.; Talavera, M.; Ayala, J.A.; López, J.A.; Martínez, J.L. Experimental Validation of Haldane’s Hypothesis on the Role of Infection as an Evolutionary Force for Metazoans. Proc. Natl. Acad. Sci. USA 2007, 104, 13728–13731. [Google Scholar] [CrossRef]
- Abdolrasouli, A.; Scourfield, A.; Rhodes, J.; Shah, A.; Elborn, J.S.; Fisher, M.C.; Schelenz, S.; Armstrong-James, D. High Prevalence of Triazole Resistance in Clinical Aspergillus fumigatus Isolates in a Specialist Cardiothoracic Centre. Int. J. Antimicrob. Agents 2018, 52, 637–642. [Google Scholar] [CrossRef] [PubMed]
- Prigitano, A.; Esposto, M.C.; Biffi, A.; De Lorenzis, G.; Favuzzi, V.; Koncan, R.; Lo Cascio, G.; Barao Ocampo, M.; Colombo, C.; Pizzamiglio, G.; et al. Triazole Resistance in Aspergillus fumigatus Isolates from Patients with Cystic Fibrosis in Italy. J. Cyst. Fibros. 2017, 16, 64–69. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ashu, E.E.; Korfanty, G.A.; Samarasinghe, H.; Pum, N.; You, M.; Yamamura, D.; Xu, J. Widespread Amphotericin B-Resistant Strains of Em Aspergillus fumigatus Em in Hamilton, Canada. Infect. Drug Resist. 2018, 11, 1549–1555. [Google Scholar] [CrossRef] [PubMed]
- Hagiwara, D.; Arai, T.; Takahashi, H.; Kusuya, Y.; Watanabe, A.; Kamei, K. Non- Cyp51A Azole-Resistant Aspergillus fumigatus Isolates with Mutation in HMG-CoA Reductase. Emerg. Infect. Dis. 2018, 24, 1889–1897. [Google Scholar] [CrossRef]
- Breger, J.; Fuchs, B.B.; Aperis, G.; Moy, T.I.; Ausubel, F.M.; Mylonakis, E. Antifungal Chemical Compounds Identified Using a C. Elegans Pathogenicity Assay. PLoS Pathog. 2007, 3, e18. [Google Scholar] [CrossRef]
- Smith, A.C.; Dinh, J.; Hickman, M.A. Two Infection Assays to Study Non-Lethal Virulence Phenotypes in C. albicans using C. elegans. J. Vis. Exp. 2021, 117, e62170. [Google Scholar] [CrossRef]
- Eldesouky, H.E.; Li, X.; Abutaleb, N.S.; Mohammad, H.; Seleem, M.N. Synergistic Interactions of Sulfamethoxazole and Azole Antifungal Drugs against Emerging Multidrug-Resistant Candida Auris. Int. J. Antimicrob. Agents 2018, 52, 754–761. [Google Scholar] [CrossRef]
- Peleg, A.Y.; Tampakakis, E.; Fuchs, B.B.; Eliopoulos, G.M.; Moellering, R.C.; Mylonakis, E. Prokaryote–Eukaryote Interactions Identified by Using Caenorhabditis Elegans. Proc. Natl. Acad. Sci. USA 2008, 105, 14585–14590. [Google Scholar] [CrossRef]
- de Barros, P.P.; Scorzoni, L.; Ribeiro, F.d.C.; Fugisaki, L.R.d.O.; Fuchs, B.B.; Mylonakis, E.; Jorge, A.O.C.; Junqueira, J.C.; Rossoni, R.D. Lactobacillus Paracasei 28.4 Reduces in Vitro Hyphae Formation of Candida albicans and Prevents the Filamentation in an Experimental Model of Caenorhabditis Elegans. Microb. Pathog. 2018, 117, 80–87. [Google Scholar] [CrossRef]
- Ramarao, N.; Nielsen-Leroux, C.; Lereclus, D. The Insect Galleria mellonella as a Powerful Infection Model to Investigate Bacterial Pathogenesis. J. Vis. Exp. 2012, 70, e4392. [Google Scholar] [CrossRef]
- Lange, A.; Beier, S.; Huson, D.H.; Parusel, R.; Iglauer, F.; Frick, J.-S. Genome Sequence of Galleria mellonella (Greater Wax Moth). Genome Announc. 2018, 6, e01220-17. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, K.; Vilcinskas, A. Development and Immunity-Related MicroRNAs of the Lepidopteran Model Host Galleria mellonella. BMC Genom. 2014, 15, 705. [Google Scholar] [CrossRef] [PubMed]
- Vogel, H.; Altincicek, B.; Glöckner, G.; Vilcinskas, A. A Comprehensive Transcriptome and Immune-Gene Repertoire of the Lepidopteran Model Host Galleria mellonella. BMC Genom. 2011, 12, 308. [Google Scholar] [CrossRef] [PubMed]
- Smith, T.L. External Morphology of the Larva, Pupa, and Adult of the Wax Moth, Galleria mellonella L. EurekaMag 1965, 38, 287–310. [Google Scholar]
- Stephens, J.M. Protective Effects of Several Immunizing Preparations That Produce Active Immunity in Galleria mellonella (Linnaeus). J. Insect Pathol. 1963, 5, 129. [Google Scholar]
- Fuchs, B.B.; O’Brien, E.; El Khoury, J.B.; Mylonakis, E. Methods for Using Galleria mellonella as a Model Host to Study Fungal Pathogenesis. Virulence 2010, 1, 475–482. [Google Scholar] [CrossRef]
- Kwadha, C.A.; Ong’amo, G.O.; Ndegwa, P.N.; Raina, S.K.; Fombong, A.T. The Biology and Control of the Greater Wax Moth, Galleria mellonella. Insects 2017, 8, 61. [Google Scholar] [CrossRef]
- Scully, L.R.; Bidochka, M.J. Serial Passage of the Opportunistic Pathogen Aspergillus Flavus through an Insect Host Yields Decreased Saprobic Capacity. Can. J. Microbiol. 2005, 51, 185–189. [Google Scholar] [CrossRef]
- Singkum, P.; Suwanmanee, S.; Pumeesat, P.; Luplertlop, N. A Powerful in Vivo Alternative Model in Scientific Research: Galleria mellonella. Acta Microbiol. Immunol. Hung. 2019, 66, 31–55. [Google Scholar] [CrossRef]
- Wittig, G. Phagocytosis by Blood Cells in Healthy and Diseased Caterpillars. J. Invertebr. Pathol. 1966, 8, 461–477. [Google Scholar] [CrossRef]
- Junqueira, J.C.; Fuchs, B.B.; Muhammed, M.; Coleman, J.J.; Suleiman, J.M.; Vilela, S.F.; Costa, A.C.; Rasteiro, V.M.; Jorge, A.O.; Mylonakis, E. Oral Candida albicans Isolates from HIV-Positive Individuals Have Similar in Vitro Biofilm-Forming Ability and Pathogenicity as Invasive Candida Isolates. BMC Microbiol. 2011, 11, 247. [Google Scholar] [CrossRef]
- Lavine, M.D.; Strand, M.R. Insect Hemocytes and Their Role in Immunity. Insect Biochem. Mol. Biol. 2002, 32, 1295–1309. [Google Scholar] [CrossRef]
- Pereira, T.; de Barros, P.; Fugisaki, L.; Rossoni, R.; Ribeiro, F.; de Menezes, R.; Junqueira, J.; Scorzoni, L. Recent Advances in the Use of Galleria mellonella Model to Study Immune Responses against Human Pathogens. J. Fungi 2018, 4, 128. [Google Scholar] [CrossRef]
- Ellis, J.D.; Graham, J.R.; Mortensen, A. Standard Methods for Wax Moth Research. J. Apic. Res. 2013, 52, 1–17. [Google Scholar] [CrossRef]
- Sheehan, G.; Kavanagh, K. Analysis of the Early Cellular and Humoral Responses of Galleria mellonella Larvae to Infection by Candida albicans. Virulence 2018, 9, 163–172. [Google Scholar] [CrossRef]
- Browne, N.; Heelan, M.; Kavanagh, K. An Analysis of the Structural and Functional Similarities of Insect Hemocytes and Mammalian Phagocytes. Virulence 2013, 4, 597–603. [Google Scholar] [CrossRef]
- Nett, J. Special Issue: Candida and Candidiasis. J. Fungi 2018, 4, 74. [Google Scholar] [CrossRef]
- Strand, M.R. The Insect Cellular Immune Response. Insect Sci. 2008, 15, 1–14. [Google Scholar] [CrossRef]
- Brennan, M. Correlation between Virulence of Candida albicans Mutants in Mice and Galleria mellonella Larvae. FEMS Immunol. Med. Microbiol. 2002, 34, 153–157. [Google Scholar] [CrossRef]
- Arteaga Blanco, L.A.; Crispim, J.S.; Fernandes, K.M.; de Oliveira, L.L.; Pereira, M.F.; Bazzolli, D.M.S.; Martins, G.F. Differential Cellular Immune Response of Galleria mellonella to Actinobacillus Pleuropneumoniae. Cell Tissue Res. 2017, 370, 153–168. [Google Scholar] [CrossRef]
- Bistoni, F.; Vecchiarelli, A.; Cenci, E.; Sbaraglia, G.; Perito, S.; Cassone, A. A Comparison of Experimental Pathogenicity of Candida Species in Cyclophosphamide-Immunodepressed Mice. Med. Mycol. 1984, 22, 409–418. [Google Scholar] [CrossRef]
- Loh, J.M.; Adenwalla, N.; Wiles, S.; Proft, T. Galleria mellonella Larvae as an Infection Model for Group A Streptococcus. Virulence 2013, 4, 419–428. [Google Scholar] [CrossRef]
- Melloul, E.; Roisin, L.; Durieux, M.-F.; Woerther, P.-L.; Jenot, D.; Risco, V.; Guillot, J.; Dannaoui, E.; Decousser, J.-W.; Botterel, F. Interactions of Aspergillus fumigatus and Stenotrophomonas Maltophilia in an in Vitro Mixed Biofilm Model: Does the Strain Matter? Front. Microbiol. 2018, 9, 2850. [Google Scholar] [CrossRef]
- Mowlds, P.; Kavanagh, K. Effect of Pre-Incubation Temperature on Susceptibility of Galleria mellonella Larvae to Infection by Candida albicans. Mycopathologia 2008, 165, 5–12. [Google Scholar] [CrossRef]
- Arendrup, M.; Horn, T.; Frimodt-Møller, N. In Vivo Pathogenicity of Eight Medically Relevant Candida Species in an Animal Model. Infection 2002, 30, 286–291. [Google Scholar] [CrossRef]
- Cotter, G.; Doyle, S.; Kavanagh, K. Development of an Insect Model for the in Vivo Pathogenicity Testing of Yeasts. FEMS Immunol. Med. Microbiol. 2000, 27, 163–169. [Google Scholar] [CrossRef]
- Jacobsen, I.D.; Brunke, S.; Seider, K.; Schwarzmüller, T.; Firon, A.; d’Enfért, C.; Kuchler, K.; Hube, B. Candida glabrata Persistence in Mice Does Not Depend on Host Immunosuppression and Is Unaffected by Fungal Amino Acid Auxotrophy. Infect. Immun. 2010, 78, 1066–1077. [Google Scholar] [CrossRef]
- Hernando-Ortiz, A.; Mateo, E.; Perez-Rodriguez, A.; de Groot, P.W.J.; Quindós, G.; Eraso, E. Virulence of Candida auris from Different Clinical Origins in Caenorhabditis elegans and Galleria mellonella Host Models. Virulence 2021, 12, 1063–1075. [Google Scholar] [CrossRef]
- Garcia-Bustos, V.; Ruiz-Saurí, A.; Ruiz-Gaitán, A.; Sigona-Giangreco, I.A.; Cabañero-Navalon, M.D.; Sabalza-Baztán, O.; Salavert-Lletí, M.; Tormo, M.Á.; Pemán, J. Characterization of the Differential Pathogenicity of Candida Auris in a Galleria mellonella Infection Model. Microbiol. Spectr. 2021, 9, e0001321. [Google Scholar] [CrossRef]
- Romera, D.; Aguilera-Correa, J.J.; Gadea, I.; Viñuela-Sandoval, L.; García-Rodríguez, J.; Esteban, J. Candida Auris: A Comparison between Planktonic and Biofilm Susceptibility to Antifungal Drugs. J. Med. Microbiol. 2019, 68, 1353–1358. [Google Scholar] [CrossRef]
- Romera, D.; Aguilera-Correa, J.-J.; García-Coca, M.; Mahillo-Fernández, I.; Viñuela-Sandoval, L.; García-Rodríguez, J.; Esteban, J. The Galleria mellonella Infection Model as a System to Investigate the Virulence of Candida Auris Strains. Pathog. Dis. 2020, 78, ftaa067. [Google Scholar] [CrossRef]
- Kousha, M.; Tadi, R.; Soubani, A.O. Pulmonary Aspergillosis: A Clinical Review. Eur. Respir. Rev. 2011, 20, 156–174. [Google Scholar] [CrossRef]
- Kamei, K.; Watanabe, A. Aspergillus Mycotoxins and Their Effect on the Host. Med. Mycol. 2005, 43, 95–99. [Google Scholar] [CrossRef]
- Hoenigl, M.; Seidel, D.; Carvalho, A.; Rudramurthy, S.M.; Arastehfar, A.; Gangneux, J.-P.; Nasir, N.; Bonifaz, A.; Araiza, J.; Klimko, N.; et al. The Emergence of COVID-19 Associated Mucormycosis: A Review of Cases from 18 Countries. Lancet Microbe 2022, 3, e543–e552. [Google Scholar] [CrossRef]
- Chatterjee, S.; Alampalli, S.V.; Nageshan, R.K.; Chettiar, S.T.; Joshi, S.; Tatu, U.S. Draft Genome of a Commonly Misdiagnosed Multidrug Resistant Pathogen Candida Auris. BMC Genom. 2015, 16, 686. [Google Scholar] [CrossRef]
- Maurer, E.; Hörtnagl, C.; Lackner, M.; Grässle, D.; Naschberger, V.; Moser, P.; Segal, E.; Semis, M.; Lass-Flörl, C.; Binder, U. Galleria mellonella as a Model System to Study Virulence Potential of Mucormycetes and Evaluation of Antifungal Treatment. Med. Mycol. 2019, 57, 351–362. [Google Scholar] [CrossRef]
- Prado, M.; da Silva, M.B.; Laurenti, R.; Travassos, L.R.; Taborda, C.P. Mortality Due to Systemic Mycoses as a Primary Cause of Death or in Association with AIDS in Brazil: A Review from 1996 to 2006. Mem. Inst. Oswaldo Cruz 2009, 104, 513–521. [Google Scholar] [CrossRef]
- Thomaz, L.; García-Rodas, R.; Guimarães, A.J.; Taborda, C.P.; Zaragoza, O.; Nosanchuk, J.D. Galleria mellonella as a Model Host to Study Paracoccidioides lutzii and Histoplasma capsulatum. Virulence 2013, 4, 139–146. [Google Scholar] [CrossRef]
- Cordero, R.J.B.; Liedke, S.C.; Araújo, G.R.d.S.; Martinez, L.R.; Nimrichter, L.; Frases, S.; Peralta, J.M.; Casadevall, A.; Rodrigues, M.L.; Nosanchuk, J.D.; et al. Enhanced Virulence of Histoplasma capsulatum through Transfer and Surface Incorporation of Glycans from Cryptococcus Neoformans during Co-Infection. Sci. Rep. 2016, 6, 21765. [Google Scholar] [CrossRef]
- de Lacorte Singulani, J.; Scorzoni, L.; de Paula e Silva, A.C.A.; Fusco-Almeida, A.M.; Mendes-Giannini, M.J.S. Evaluation of the Efficacy of Antifungal Drugs against Paracoccidioides brasiliensis and Paracoccidioides lutzii in a Galleria mellonella Model. Int. J. Antimicrob. Agents 2016, 48, 292–297. [Google Scholar] [CrossRef]
- Fregonezi, N.F.; Oliveira, L.T.; Singulani, J.d.L.; Marcos, C.M.; dos Santos, C.T.; Taylor, M.L.; Mendes-Giannini, M.J.S.; de Oliveira, H.C.; Fusco-Almeida, A.M. Heat Shock Protein 60, Insights to Its Importance in Histoplasma capsulatum: From Biofilm Formation to Host-Interaction. Front. Cell Infect. Microbiol. 2021, 10, 591950. [Google Scholar] [CrossRef] [PubMed]
- Ghaly, A.E.; Alkoaik, F.N. The Yellow Mealworm as a Novel Source of Protein. Am. J. Agric. Biol. Sci. 2009, 4, 319–331. [Google Scholar] [CrossRef]
- Park, J.B.; Choi, W.H.; Kim, S.H.; Jin, H.J.; Han, Y.S.; Lee, Y.S.; Kim, N.J. Developmental Characteristics of Tenebrio molitor Larvae (Coleoptera: Tenebrionidae) in Different Instars. Int. J. Ind. Entomol. 2014, 28, 5–9. [Google Scholar] [CrossRef]
- Oppert, B.; Dowd, S.E.; Bouffard, P.; Li, L.; Conesa, A.; Lorenzen, M.D.; Toutges, M.; Marshall, J.; Huestis, D.L.; Fabrick, J.; et al. Transcriptome Profiling of the Intoxication Response of Tenebrio molitor Larvae to Bacillus Thuringiensis Cry3Aa Protoxin. PLoS ONE 2012, 7, e34624. [Google Scholar] [CrossRef] [PubMed]
- Canteri de Souza, P.; Custódio Caloni, C.; Wilson, D.; Sergio Almeida, R. An Invertebrate Host to Study Fungal Infections, Mycotoxins and Antifungal Drugs: Tenebrio molitor. J. Fungi 2018, 4, 125. [Google Scholar] [CrossRef] [PubMed]
- Moret, Y.; Siva-Jothy, M.T. Adaptive Innate Immunity? Responsive-Mode Prophylaxis in the Mealworm Beetle, Tenebrio molitor. Proc. R. Soc. Lond. B Biol. Sci. 2003, 270, 2475–2480. [Google Scholar] [CrossRef] [PubMed]
- Johnston, P.R.; Makarova, O.; Rolff, J. Inducible Defenses Stay Up Late: Temporal Patterns of Immune Gene Expression in Tenebrio molitor. G3 Genes|Genomes|Genet. 2014, 4, 947–955. [Google Scholar] [CrossRef] [PubMed]
- El-Kamand, S.; Steiner, M.; Ramirez, C.; Halliday, C.; Chen, S.C.-A.; Papanicolaou, A.; Morton, C.O. Assessing Differences between Clinical Isolates of Aspergillus fumigatus from Cases of Proven Invasive Aspergillosis and Colonizing Isolates with Respect to Phenotype (Virulence in Tenebrio molitor Larvae) and Genotype. Pathogens 2022, 11, 428. [Google Scholar] [CrossRef]
- Silva, A.P.R.; Valério, A.D.; Viana, R.O.; dos Santos Ricoy, A.C.; Johann, S.; de Souza Alves, V. Caenorhabditis elegans and Tenebrio molitor—New tools to investigate Malassezia species. Preprints.Org. 2018, 2018100001. [Google Scholar] [CrossRef]
- Lozoya-Pérez, N.E.; García-Carnero, L.C.; Martínez-Álvarez, J.A.; Martínez-Duncker, I.; Mora-Montes, H.M. Tenebrio molitor as an Alternative Model to Analyze the Sporothrix Species Virulence. Infect. Drug Resist. 2021, 14, 2059–2072. [Google Scholar] [CrossRef]
- Fornari, G.; Gomes, R.R.; Degenhardt-Goldbach, J.; dos Santos, S.S.; de Almeida, S.R.; dos Santos, G.D.; Muro, M.D.; Bona, C.; Scola, R.H.; Trindade, E.S.; et al. A Model for Trans-Kingdom Pathogenicity in Fonsecaea Agents of Human Chromoblastomycosis. Front. Microbiol. 2018, 9, 2211. [Google Scholar] [CrossRef] [PubMed]
- Cordeiro, B.M.P.C.; Carvalho Junior, A.R.; Santos, J.R.A.; Araújo, A.D.; Silva, A.G.; Correia, M.T.S.; Silva, M.V.; Napoleão, T.H.; Silva, L.C.N.; Santos, N.D.L.; et al. Anticryptococcal Activity of Hexane Extract from Spondias Tuberosa Arruda and Associated Cellular Events. J. Mycol. Med. 2020, 30, 100965. [Google Scholar] [CrossRef] [PubMed]
- Silva, M.G.; de Curcio, J.S.; Silva-Bailão, M.G.; Lima, R.M.; Tomazett, M.V.; de Souza, A.F.; Cruz-Leite, V.R.M.; Sbaraini, N.; Bailão, A.M.; Rodrigues, F.; et al. Molecular Characterization of Siderophore Biosynthesis in Paracoccidioides brasiliensis. IMA Fungus 2020, 11, 11. [Google Scholar] [CrossRef]
- Kim, S.G.; Jo, Y.H.; Seong, J.H.; Park, K.B.; Noh, M.Y.; Cho, J.H.; Ko, H.J.; Kim, C.E.; Tindwa, H.; Patnaik, B.B.; et al. Tm SR-C, Scavenger Receptor Class C, Plays a Pivotal Role in Antifungal and Antibacterial Immunity in the Coleopteran Insect Tenebrio molitor. Insect Biochem. Mol. Biol. 2017, 89, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Seong, J.H.; Jo, Y.H.; Seo, G.W.; Park, S.; Park, K.B.; Cho, J.H.; Ko, H.J.; Kim, C.E.; Patnaik, B.B.; Jun, S.A.; et al. Molecular Cloning and Effects of Tm14-3-3ζ-Silencing on Larval Survivability against E. coli and C. albicans in Tenebrio molitor. Genes 2018, 9, 330. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fusco-Almeida, A.M.; de Matos Silva, S.; dos Santos, K.S.; de Lima Gualque, M.W.; Vaso, C.O.; Carvalho, A.R.; Medina-Alarcón, K.P.; Pires, A.C.M.d.S.; Belizario, J.A.; de Souza Fernandes, L.; et al. Alternative Non-Mammalian Animal and Cellular Methods for the Study of Host–Fungal Interactions. J. Fungi 2023, 9, 943. https://doi.org/10.3390/jof9090943
Fusco-Almeida AM, de Matos Silva S, dos Santos KS, de Lima Gualque MW, Vaso CO, Carvalho AR, Medina-Alarcón KP, Pires ACMdS, Belizario JA, de Souza Fernandes L, et al. Alternative Non-Mammalian Animal and Cellular Methods for the Study of Host–Fungal Interactions. Journal of Fungi. 2023; 9(9):943. https://doi.org/10.3390/jof9090943
Chicago/Turabian StyleFusco-Almeida, Ana Marisa, Samanta de Matos Silva, Kelvin Sousa dos Santos, Marcos William de Lima Gualque, Carolina Orlando Vaso, Angélica Romão Carvalho, Kaila Petrolina Medina-Alarcón, Ana Carolina Moreira da Silva Pires, Jenyffie Araújo Belizario, Lígia de Souza Fernandes, and et al. 2023. "Alternative Non-Mammalian Animal and Cellular Methods for the Study of Host–Fungal Interactions" Journal of Fungi 9, no. 9: 943. https://doi.org/10.3390/jof9090943
APA StyleFusco-Almeida, A. M., de Matos Silva, S., dos Santos, K. S., de Lima Gualque, M. W., Vaso, C. O., Carvalho, A. R., Medina-Alarcón, K. P., Pires, A. C. M. d. S., Belizario, J. A., de Souza Fernandes, L., Moroz, A., Martinez, L. R., Ruiz, O. H., González, Á., & Mendes-Giannini, M. J. S. (2023). Alternative Non-Mammalian Animal and Cellular Methods for the Study of Host–Fungal Interactions. Journal of Fungi, 9(9), 943. https://doi.org/10.3390/jof9090943







