Assessment of Fungal Succession in Decomposing Swine Carcasses (Sus scrofa L.) Using DNA Metabarcoding
Abstract
1. Introduction
1.1. Decomposition in Forensic Investigations
1.2. Fungi in Animal Decomposition
1.3. DNA Metabarcoding of Fungi
2. Materials and Methods
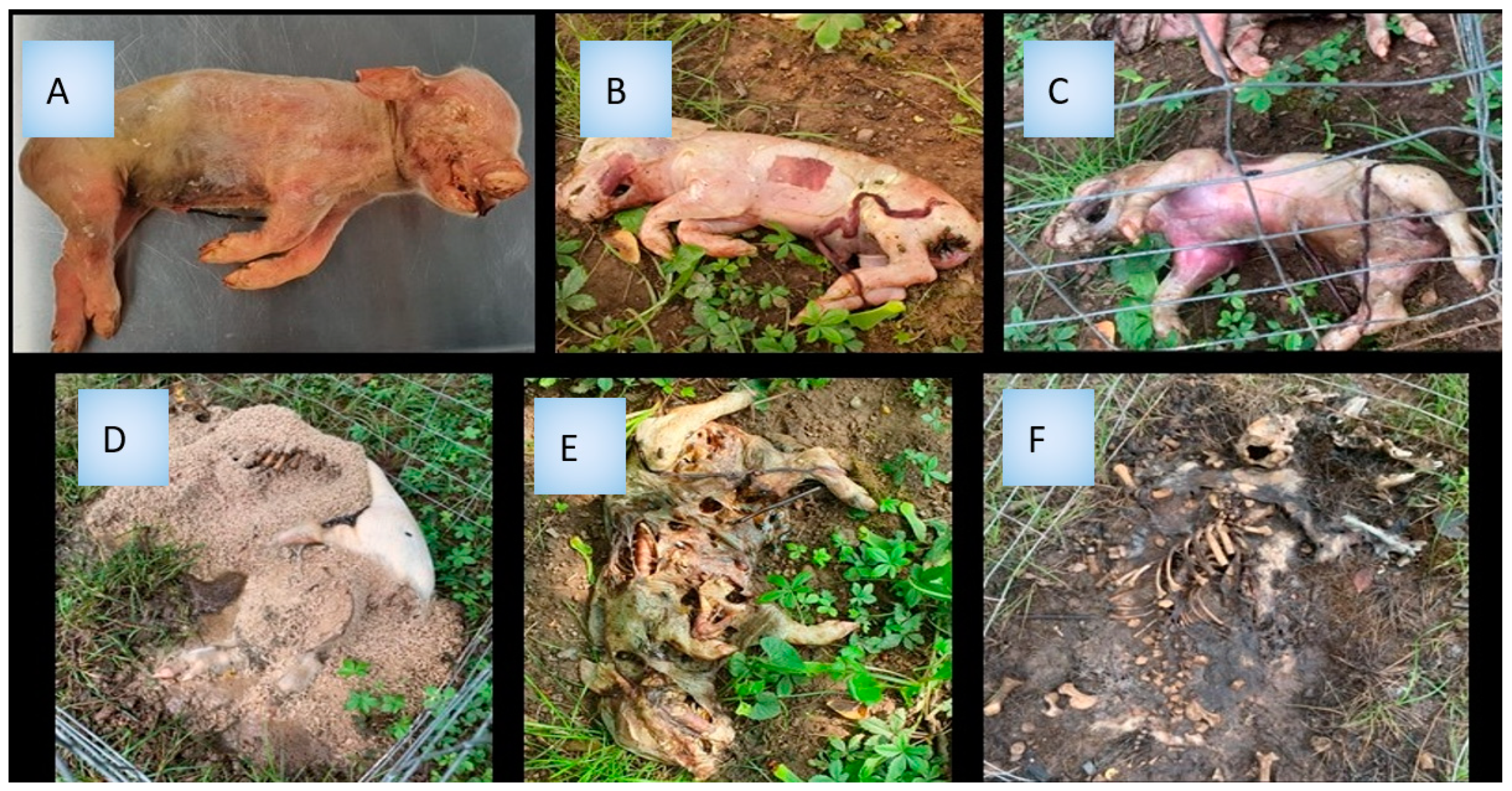
2.1. Field Experiment Design
2.2. DNA Extraction
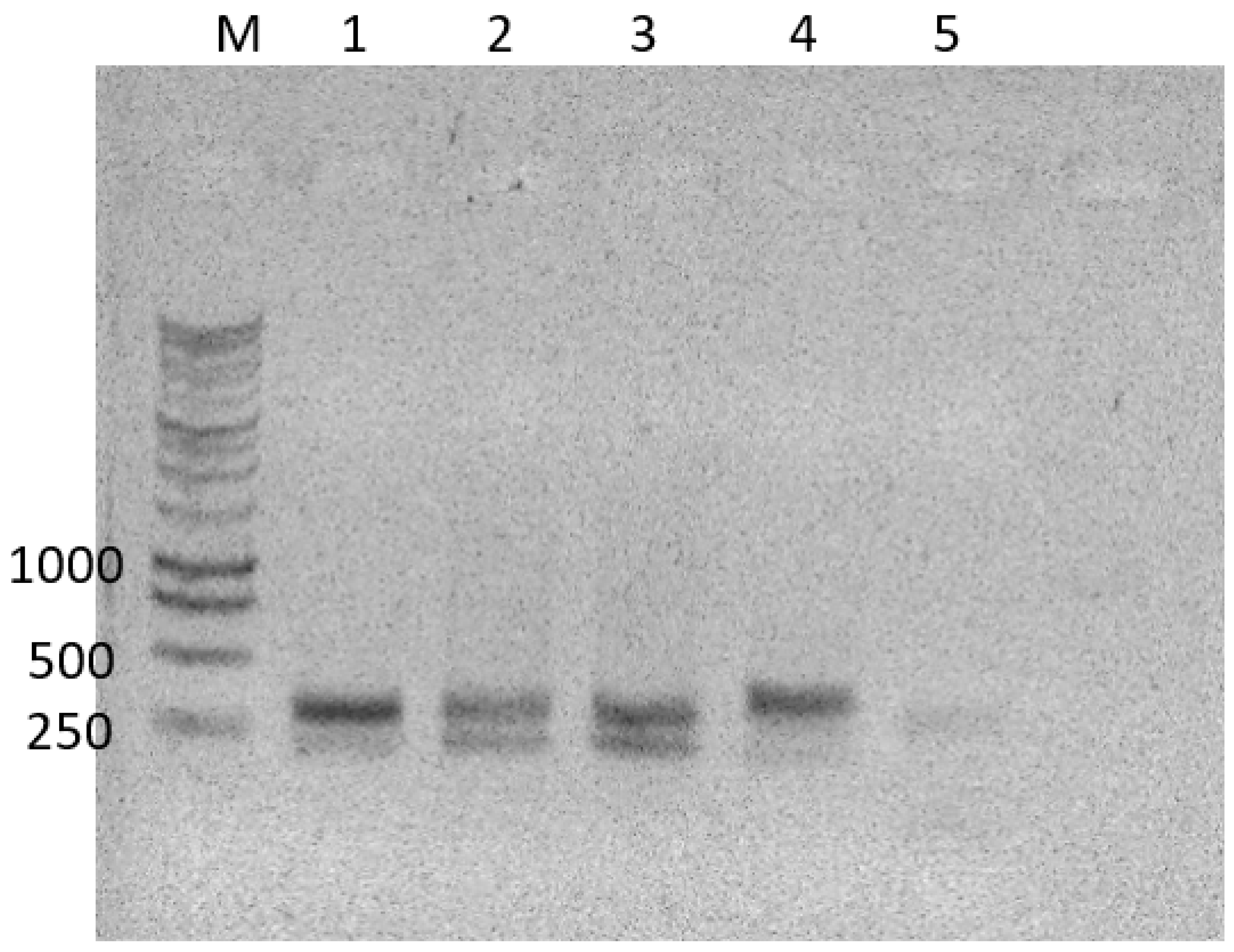
2.3. PCR
2.4. High-Throughput Sequencing (Next Generation Sequencing, NGS)
2.5. Data Analysis
3. Results
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Roy, D.; Tomo, S.; Purohit, P.; Setia, P. Microbiome in Death and Beyond: Current Vistas and Future Trends. Front. Ecol. Evol. 2021, 9, 630397. [Google Scholar] [CrossRef]
- Cai, J.; Guo, Y.; Zha, L.; Guo, J. The role of the microbiome in PMI estimation. In Forensic Ecogenomics; Academic Press: Cambridge, MA, USA, 2018; pp. 113–131. [Google Scholar] [CrossRef]
- Niemela, T.; Renvall, P.; Penttila, R. Interactions of fungi at late stages of wood decomposition. Ann. Bot. Fenn. 1995, 32, 141–152. [Google Scholar]
- Bobiec, A.; Gutowski, J.M.; Zub, K.; Pawlaczyk, P.; Laudenslayer, W.F. The Afterlife of a Tree; WWF Poland: Warsaw, Poland, 2005; p. 252. [Google Scholar]
- Valentin, L.; Rajala, T.; Peltoniemi, M.; Heinonsalo, J.; Pennanen, T.; Makipaa, R. Loss of diversity in wood-inhabiting fungal communities affects decomposition activity in Norway spruce wood. Front. Microbiol. 2014, 5, 11. [Google Scholar] [CrossRef] [PubMed]
- Fukasawa, Y.; Matsukura, K. Decay stages of wood and associated fungal communities characterise diversity-decomposition relationships. Sci. Rep. 2021, 11, 8972. [Google Scholar] [CrossRef]
- Venugopal, P.; Junninen, K.; Edman, M.; Kouki, J. Assemblage composition of fungal wood-decay species has a major influence on how climate and wood quality modify decomposition. FEMS Microbiol. Ecol. 2017, 93, 8. [Google Scholar] [CrossRef]
- Goff, M.L. Early post-mortem changes and stages of decomposition in exposed cadavers. Exp. Appl. Acarol. 2009, 49, 21–36. [Google Scholar] [CrossRef]
- Goff, M.L. Estimation of Postmortem Interval Using Arthropod Development and Successional Patterns. Forensic Sci. Rev. 1993, 5, 81–94. [Google Scholar]
- Vass, A.A.; Barshick, S.A.; Sega, G.; Caton, J.; Skeen, J.T.; Love, J.C.; Synstelien, J.A. Decomposition chemistry of human remains: A new methodology for determining the postmortem interval. J. Forensic Sci. 2002, 47, 542–553. [Google Scholar] [CrossRef]
- Caplan, M.; Koontz, F. Cumitech: Postmortem Microbiology; McCurdy, B., Ed.; ASM Press: Washington, DC, USA, 2001; Volume 35. [Google Scholar]
- Clark, M.; Worell, M.; Pless, J. Postmortem changes in soft tissues. In Forensic Taphonomy: The Postmortem Fate of Human Remains; Sorg, M., Haglulnd, W., Eds.; CRC Press: Boca Raton, FL, USA, 1997; pp. 151–164. [Google Scholar]
- Swann, L.; Chidlow, G.E.; Forbes, S.; Lewis, S.W. Preliminary Studies into the Characterization of Chemical Markers of Decomposition for Geoforensics. J. Forensic Sci. 2010, 55, 308–314. [Google Scholar] [CrossRef]
- Dangerfield, C.R.; Frehner, E.H.; Buechley, E.R.; Sekercioglu, C.H.; Brazelton, W.J. Succession of bacterial communities on carrion is independent of vertebrate scavengers. Peerj 2020, 8, e9307. [Google Scholar] [CrossRef]
- Pechal, J.L.; Crippen, T.L.; Tarone, A.M.; Lewis, A.J.; Tomberlin, J.K.; Benbow, M.E. Microbial Community Functional Change during Vertebrate Carrion Decomposition. PLoS ONE 2013, 8, 11. [Google Scholar] [CrossRef] [PubMed]
- Klein, N.S. A Comparative Study of Human Decomposition Research Facilities in the United States: The Role of “Body Farms”; Louisiana State University and Agricultural & Mechanical College: Baton Rouge, LA, USA, 2014. [Google Scholar]
- Wolff, B. A Review of ‘Body Farm’ Research Facilities across America with a Focus on Policy and the Impacts When Dealing with Decompositional Changes in Human Remains. Masters Thesis, University of Texas at Arlington, Arlington, TX, USA, 2015. [Google Scholar]
- Forbes, S. Body farms. Forensic Sci. Med. Pathol. 2017, 13, 477–479. [Google Scholar] [CrossRef]
- Payne, J. A summer carrion study of the baby pig Sus scrofa Linnaeus. Ecology 1965, 46, 592–602. [Google Scholar] [CrossRef]
- Payne, J.; King, E.; Beinhart, G. Arthropod succession and decomposition of buried pigs. Nature 1968, 219, 1180–1181. [Google Scholar] [CrossRef]
- Dautartas, A.; Kenyhercz, M.W.; Vidoli, G.M.; Jantz, L.M.; Mundorff, A.; Steadman, D.W. Differential Decomposition Among Pig, Rabbit, and Human Remains. J. Forensic Sci. 2018, 63, 1673–1683. [Google Scholar] [CrossRef]
- Matuszewski, S.; Hall, M.J.R.; Moreau, G.; Schoenly, K.G.; Tarone, A.M.; Villet, M.H. Pigs vs people: The use of pigs as analogues for humans in forensic entomology and taphonomy research. Int. J. Leg. Med. 2020, 134, 793–810. [Google Scholar] [CrossRef] [PubMed]
- Jordan, H.R.; Tomberlin, J.K. Abiotic and Biotic Factors Regulating Inter-Kingdom Engagement between Insects and Microbe Activity on Vertebrate Remains. Insects 2017, 8, 54. [Google Scholar] [CrossRef] [PubMed]
- Noriega, J.A.; Hortal, J.; Azcarate, F.M.; Berg, M.P.; Bonada, N.; Briones, M.J.I.; Del Toro, I.; Goulson, D.; Ibanez, S.; Landis, D.A.; et al. Research trends in ecosystem services provided by insects. Basic Appl. Ecol. 2018, 26, 8–23. [Google Scholar] [CrossRef]
- Huntington, T.; Weidner, L.M.; Hall, R. Introduction: Current perceptions and status of forensic entomology. In Forensic Entomology: The Utility of Arthropods in Legal Investigations; CRC Press: Boca Raton, FL, USA, 2020; pp. 1–16. [Google Scholar]
- Rodriguez, W.C.; Bass, W.M. Decomposition of buried bodies and methods that may aid in their location. J. Forensic Sci. 1985, 30, 836–852. [Google Scholar] [CrossRef]
- Anderson, G.S.; VanLaerhoven, S.L. Initial studies on insect succession on carrion in southwestern British Columbia. J. Forensic Sci. 1996, 41, 617–625. [Google Scholar] [CrossRef]
- Anderson, G. Insect succession on carrion and its relationship to determining time of death. In Forensic Entomology: The Utility of Arthropods in Legal Investigations; Byrd, J., Castner, J., Eds.; CRC Press: Boca Raton, FL, USA, 2000; pp. 143–175. [Google Scholar]
- Greenberg, B.; Kunich, J. Entomology and the Law: Flies as Forensic Indicators; Cambridge University Press: Cambridge, UK, 2002. [Google Scholar]
- Introna, F.; Campobasso, C.P.; Goff, M.L. Entomotoxicology. Forensic Sci. Int. 2001, 120, 42–47. [Google Scholar] [CrossRef] [PubMed]
- Gemmellaro, M.D.; Bucolo, C.; Musumeci, E.; Hamilton, G.C.; Weidner, L.M. First Observations of Initial Blow Fly (Diptera: Calliphoridae) Activity on Lava Fields and in Subterranean Environments in Sicily in Cool Temperatures. J. Med. Entomol. 2018, 55, 1622–1626. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, B. Flies as forensic indicators. J. Med. Entomol. 1991, 28, 565–577. [Google Scholar] [CrossRef]
- Koffi, A.F.; Aboua, L.R.N.; Djodjo, M.; Dao, H.; Koffi-Tebele, J.; Yapo, C. Contribution of different groups of necrophagous insects, in the process of decomposition of a pig corpse (Sus scrofa domesticus L.) exposed to the open air, in the guinean zone of Côte d’Ivoire. Int. J. Sci. Eng. Appl. Sci. 2017, 3, 14–22. [Google Scholar]
- Damann, F.E.; Carter, D.O. Human decomposition ecology and postmortem microbiology. In Manual of Forensic Taphonomy; CRC Press: Boca Raton, FL, USA, 2013; pp. 37–49. [Google Scholar]
- Howard, G.T.; Duos, B.; Watson-Horzelski, E.J. Characterization of the soil microbial community associated with the decomposition of a swine carcass. Int. Biodeterior. Biodegrad. 2010, 64, 300–304. [Google Scholar] [CrossRef]
- Blackwell, M. The fungi: 1, 2, 3... 5.1 million species? Am. J. Bot. 2011, 98, 426–438. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Luecking, R. Fungal diversity revisited: 2.2 to 3.8 million species. Microbiol. Spectr. 2017, 5, 5.4.10. [Google Scholar] [CrossRef] [PubMed]
- Hawksworth, D.L.; Wiltshire, P.E.J. Forensic mycology: The use of fungi in criminal investigations. Forensic Sci. Int. 2011, 206, 1–11. [Google Scholar] [CrossRef]
- Hitosugi, M.; Ishii, K.; Yaguchi, T.; Chigusa, Y.; Kurosu, A.; Kido, M.; Nagai, T.; Tokudome, S. Fungi can be a useful forensic tool. Leg. Med. 2006, 8, 240–242. [Google Scholar] [CrossRef]
- Tranchida, M.C.; Pelizza, S.A.; Eliades, L.A. The use of fungi in forensic science, a brief overview. Can. Soc. Forensic Sci. J. 2021, 54, 35–48. [Google Scholar] [CrossRef]
- Hawksworth, D.L.; Wiltshire, P.E. Forensic mycology: Current perspective. Res. Rep. Forensic Med. Sci. 2015, 5, 75–83. [Google Scholar] [CrossRef]
- Vandevoorde, H.; Vandijck, P.J. Determination of the time of death by fungal growth. Z. Fur Rechtsmed. J. Leg. Med. 1982, 89, 75–80. [Google Scholar] [CrossRef] [PubMed]
- Menezes, R.G.; Kanchan, T.; Lobo, S.W.; Jain, A.; Bhat, N.B.; Rao, N.G. Cadaveric fungi: Not yet an established forensic tool. J. Forensic Leg. Med. 2008, 15, 124–125. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, P.; Dannaoui, E.; Gehl, A.; Felske-Zech, H.; Birngruber, C.G.; Dettmeyer, R.B.; Verhoff, M.A. Molecular identification of fungi found on decomposed human bodies in forensic autopsy cases. Int. J. Leg. Med. 2015, 129, 785–791. [Google Scholar] [CrossRef] [PubMed]
- Nilsson, R.H.; Larsson, K.H.; Taylor, A.F.S.; Bengtsson-Palme, J.; Jeppesen, T.S.; Schigel, D.; Kennedy, P.; Picard, K.; Glockner, F.O.; Tedersoo, L.; et al. The UNITE database for molecular identification of fungi: Handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res. 2019, 47, D259–D264. [Google Scholar] [CrossRef]
- Ruppert, K.M.; Kline, R.J.; Rahman, M.S. Past, present, and future perspectives of environmental DNA (eDNA) metabarcoding: A systematic review in methods, monitoring, and applications of global eDNA. Glob. Ecol. Conserv. 2019, 17, e00547. [Google Scholar] [CrossRef]
- Taberlet, P.; Bonin, A.; Zinger, L.; Coissac, E. Environmental DNA: For Biodiversity Research and Monitoring; Oxford University Press: New York, NY, USA, 2018; p. 253. [Google Scholar]
- Giampaoli, S.; De Vittori, E.; Frajese, G.V.; Paytuvi, A.; Sanseverino, W.; Anselmo, A.; Barni, F.; Berti, A. A semi-automated protocol for NGS metabarcoding and fungal analysis in forensic. Forensic Sci. Int. 2020, 306, 110052. [Google Scholar] [CrossRef]
- Shumskaya, M.; Lorusso, N.; Patel, U.; Leigh, M.; Somervuo, P.; Schigel, D. MycoPins: A metabarcoding-based method to monitor fungal colonization of fine woody debris. Mycokeys 2023, 96, 77–95. [Google Scholar] [CrossRef]
- Tedersoo, L.; Bahram, M.; Polme, S.; Koljalg, U.; Yorou, N.S.; Wijesundera, R.; Ruiz, L.V.; Vasco-Palacios, A.M.; Thu, P.Q.; Suija, A.; et al. Global diversity and geography of soil fungi. Science 2014, 346, 1256688. [Google Scholar] [CrossRef]
- Xu, J. Fungal DNA barcoding. Genome 2016, 59, 913–932. [Google Scholar] [CrossRef]
- Eberhardt, U. Methods for DNA barcoding of fungi. In Methods in Molecular Biology; Humana Press: Totowa, NJ, USA, 2012; Volume 858, pp. 183–205. [Google Scholar] [CrossRef]
- Fu, X.L.; Guo, J.J.; Zhu, Z.Y.; Ding, Z.Y.; Zha, L.; Cai, J.F. The potential use of fungi community in postmortem interval estimation in China. Forensic Sci. Int. Genet. Suppl. Ser. 2015, 5, e476–e478. [Google Scholar] [CrossRef][Green Version]
- Fu, X.L.; Guo, J.J.; Finkelbergs, D.; He, J.; Zha, L.; Guo, Y.D.; Cai, J.F. Fungal succession during mammalian cadaver decomposition and potential forensic implications. Sci. Rep. 2019, 9, 12907. [Google Scholar] [CrossRef]
- van der Wal, A.; Geydan, T.D.; Kuyper, T.W.; de Boer, W. A thready affair: Linking fungal diversity and community dynamics to terrestrial decomposition processes. FEMS Microbiol. Rev. 2013, 37, 477–494. [Google Scholar] [CrossRef] [PubMed]
- Clemmensen, K.E.; Ihrmark, K.; Durling, M.B.; Lindahl, B.D. Sample Preparation for Fungal Community Analysis by High-Throughput Sequencing of Barcode Amplicons. In Methods in Molecular Biology; Humana Press: New York, NY, USA, 2016; Volume 1399, pp. 61–88. [Google Scholar] [CrossRef]
- Palmer, J.M.; Jusino, M.A.; Banik, M.T.; Lindner, D.L. Non-biological synthetic spike-in controls and the AMPtk software pipeline improve mycobiome data. Peerj 2018, 6, e4925. [Google Scholar] [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing. 2020. Available online: https://www.R-project.org/ (accessed on 16 July 2023).
- Korkmaz, S.; Goksuluk, D.; Zararsiz, G. MVN: An R Package for Assessing Multivariate Normality. R J. 2014, 6, 151–162. [Google Scholar] [CrossRef]
- Roberts, D.W.; Roberts, M.D.W. Package ‘labdsv’. Version 2.0-1. Ordination and multivariate analysis for ecology. 2016, Volume 775, pp. 1–68. Available online: https://CRAN.R-project.org/package=labdsv (accessed on 16 July 2023).
- Oksanen, J.; Blanchet, G.F.; Friendly, M.; Kindt, R.; Legendre, P.; McGlinn, D.; Minchin, P.R.; O’Hara, R.B.; Simpson, G.L.; Solymos, P.; et al. Vegan: Community Ecology Package. R Package Version 2.5-7. 2020. Available online: https://CRAN.R-project.org/package=vegan (accessed on 16 July 2023).
- Nguyen, N.H.; Song, Z.; Bates, S.T.; Branco, S.; Tedersoo, L.; Menke, J.; Schilling, J.S.; Kennedy, P.G. FUNGuild: An open annotation tool for parsing fungal community datasets by ecological guild. Fungal Ecol. 2016, 20, 241–248. [Google Scholar] [CrossRef]
- Simpson, H.J.; Schilling, J.S. Using aggregated field collection data and the novel r package fungarium to investigate fungal fire association. Mycologia 2021, 113, 842–855. [Google Scholar] [CrossRef]
- Gemmellaro, M.D.; Lorusso, N.S.; Shumskaya, M. Dataset of fungal communities observed on decomposing pig carcasses in New Jersey. 2023. Available online: https://zenodo.org/record/8145408 (accessed on 13 July 2023).
- Metcalf, J.L.; Carter, D.O.; Knight, R. Microbiology of death. Curr. Biol. 2016, 26, R561–R563. [Google Scholar] [CrossRef]
- Cavka, M.; Glasnovic, A.; Jankovic, I.; Sikanjic, P.R.; Peric, B.; Brkljacic, B.; Mlinaric-Missoni, E.; Skrlin, J. Microbiological Analysis of a Mummy from the Archeological Museum in Zagreb. Coll. Antropol. 2010, 34, 803–805. [Google Scholar]
- López-Martínez, R.; Hernández-Hernández, F.; Millán-Chiu, B.E.; Manzano-Gayosso, P.; Méndez-Tovar, L.J. Effectiveness of imazalil to control the effect of fungal deterioration on mummies at the Mexico City Museum “El Carmen”. Rev. Iberoam. Micol. 2007, 24, 283–288. [Google Scholar] [CrossRef]
- Shumskaya, M.; Filippova, N.; Lorentzen, L.; Blue, S.; Andrew, C.; Lorusso, N.S. Citizen science helps in the study of fungal diversity in New Jersey. Sci. Data 2023, 10, 10. [Google Scholar] [CrossRef] [PubMed]
- Sidrim, J.J.C.; Moreira, R.E.; Cordeiro, R.A.; Rocha, M.F.G.; Caetano, E.P.; Monteiro, A.J.; Brilhante, R.S.N. Fungal microbiota dynamics as a postmortem investigation tool: Focus on Aspergillus, Penicillium and Candida species. J. Appl. Microbiol. 2010, 108, 1751–1756. [Google Scholar] [CrossRef] [PubMed]

| Primer Name | Primer Sequence, from 5′ |
|---|---|
| tag_1F | CACACGATCT GTGARTCATCGAATCTTTG |
| tag_1R | CACACGCTGT TCCTCCGCTTATTGATATGC |
| tag_4F | CACATAGTCT GTGARTCATCGAATCTTTG |
| tag_4R | CACATGTCGT TCCTCCGCTTATTGATATGC |
| tag_5F | CACATGACTT GTGARTCATCGAATCTTTG |
| tag_5R | CACGCAGCAT TCCTCCGCTTATTGATATGC |
| tag_6F | CACGATCAGT GTGARTCATCGAATCTTTG |
| tag_6R | CACTAGCGCT TCCTCCGCTTATTGATATGC |
| tag_7F | CACGTGCTCT GTGARTCATCGAATCTTTG |
| tag_7R | CACTATGCAT TCCTCCGCTTATTGATATGC |
| tag_8F | CACTATAGCT GTGARTCATCGAATCTTTG |
| tag_8R | CACTCACACT TCCTCCGCTTATTGATATGC |
| tag_9F | CACTATGTGT GTGARTCATCGAATCTTTG |
| tag_9R | CACTCTGAGT TCCTCCGCTTATTGATATGC |
| tag_10F | CACTCAGAGT GTGARTCATCGAATCTTTG |
| tag_10R | CACTGATCAT TCCTCCGCTTATTGATATGC |
| tag_11F | CACTCTCACT GTGARTCATCGAATCTTTG |
| tag_11R | CACTGTATGT TCCTCCGCTTATTGATATGC |
| tag_12F | CACTGCTACT GTGARTCATCGAATCTTTG |
| tag_12R | CAGACATAGT TCCTCCGCTTATTGATATGC |
| tag_13F | CAGACAGTGT GTGARTCATCGAATCTTTG |
| tag_13R | CAGACTATGT TCCTCCGCTTATTGATATGC |
| tag_14F | CAGACATCTT GTGARTCATCGAATCTTTG |
| tag_14R | CAGAGCTCAT TCCTCCGCTTATTGATATGC |
| tag_15F | CAGAGACGCT GTGARTCATCGAATCTTTG |
| tag_15R | CAGATACACT TCCTCCGCTTATTGATATGC |
| tag_16F | CAGAGCTCGT GTGARTCATCGAATCTTTG |
| tag_16R | CAGATGCTAT TCCTCCGCTTATTGATATGC |
| tag_17F | CAGAGTATGT GTGARTCATCGAATCTTTG |
| tag_17R | CAGCACGACT TCCTCCGCTTATTGATATGC |
| tag_18F | CAGATACAGT GTGARTCATCGAATCTTTG |
| tag_18R | CAGCACTCGT TCCTCCGCTTATTGATATGC |
| tag_20F | CAGCACTATT GTGARTCATCGAATCTTTG |
| tag_20R | CAGCTGACGT TCCTCCGCTTATTGATATGC |
| tag_21F | CAGCGATACT GTGARTCATCGAATCTTTG |
| tag_21R | CAGTATCTCT TCCTCCGCTTATTGATATGC |
| tag_22F | CAGCTAGATT GTGARTCATCGAATCTTTG |
| tag_22R | CAGTCAGTCT TCCTCCGCTTATTGATATGC |
| tag_23F | CAGCTCACTT GTGARTCATCGAATCTTTG |
| tag_23R | CAGTCGTGCT TCCTCCGCTTATTGATATGC |
| Fungal Species | Stage | Indicator Value | p-Value | Guild |
|---|---|---|---|---|
| Alternaria metachromatica | Zero | 0.450013 | 0.029 | |
| Apiotrichum mycotoxinovorans | Zero | 0.3204 | 0.005 | |
| Ascomycota sp. | Zero | 0.30861 | 0.004 | |
| Beauveria bassiana | Zero | 0.381168 | 0.044 | Animal pathogen |
| Cladosporium tenuissimum | Zero | 0.279405 | 0.029 | Endophyte |
| Cryptococcus uniguttulatus | Zero | 0.3 | 0.037 | |
| Cystobasidium pallidum | Zero | 0.616563 | 0.016 | |
| Cystofilobasidium capitatum | Fresh | 0.24132 | 0.002 | |
| Erysiphe corylacearum | Fresh | 0.558174 | 0.004 | |
| Exophiala salmonis | Bloat | 0.223837 | 0.021 | Endophyte |
| Filobasidium sp. | Bloat | 0.173077 | 0.032 | |
| Gibellulopsis piscis | Bloat | 0.370083 | 0.038 | |
| Jaminaea sp. | Bloat | 0.404757 | 0.003 | |
| Knufia tsunedae | Bloat | 0.329308 | 0.028 | Soil Saprotroph |
| Lachancea thermotolerans | Bloat | 0.325791 | 0.003 | |
| Leucosporidium intermedium | Bloat | 0.278229 | 0.016 | |
| Malassezia restricta | Bloat | 0.341931 | 0.035 | Animal pathogen |
| Metarhizium anisopliae | Bloat | 0.229167 | 0.003 | |
| Mortierella alpina | Decay | 0.352564 | 0.001 | |
| Mortierella elongata | Decay | 0.526913 | 0.016 | |
| Mucor racemosus | Decay | 0.25907 | 0.006 | Endophyte |
| Neophaeococcomyces catenatus | Decay | 0.264503 | 0.043 | |
| Phallus rugulosus | Decay | 0.214286 | 0.001 | |
| Pleosporales sp. | Decay | 0.25 | 0.011 | |
| Powellomyces sp. | Decay | 0.351021 | 0.004 | Plant pathogen |
| Pseudomicrostroma phylloplanum | Decay | 0.361429 | 0.03 | |
| Pyrenochaetopsis leptospora | Decay | 0.242138 | 0.001 | |
| Schwanniomyces occidentalis | Decay | 0.734502 | 0.019 | |
| Tausonia pullulans | Decay | 0.296296 | 0.003 | Saprotroph |
| Trichoderma harzianum | Decay | 0.480422 | 0.036 | Endophyte-Fungal Parasite-Plant Pathogen |
| Trichomeriaceae sp. | Decay | 0.333857 | 0.015 | Epiphyte |
| Ustilago nunavutica | Decay | 0.457515 | 0.001 | |
| Wallemia muriae | Decay | 0.257678 | 0.001 | |
| Wallemia tropicalis | Decay | 0.654491 | 0.002 | |
| Xylariales sp. | Active decay | 0.267912 | 0.027 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gemmellaro, M.D.; Lorusso, N.S.; Domke, R.; Kovalska, K.M.; Hashim, A.; Arevalo Mojica, M.; O’Connor, A.J.; Patel, U.; Pate, O.; Raise, G.; et al. Assessment of Fungal Succession in Decomposing Swine Carcasses (Sus scrofa L.) Using DNA Metabarcoding. J. Fungi 2023, 9, 866. https://doi.org/10.3390/jof9090866
Gemmellaro MD, Lorusso NS, Domke R, Kovalska KM, Hashim A, Arevalo Mojica M, O’Connor AJ, Patel U, Pate O, Raise G, et al. Assessment of Fungal Succession in Decomposing Swine Carcasses (Sus scrofa L.) Using DNA Metabarcoding. Journal of Fungi. 2023; 9(9):866. https://doi.org/10.3390/jof9090866
Chicago/Turabian StyleGemmellaro, M. Denise, Nicholas Steven Lorusso, Rachel Domke, Kristina M. Kovalska, Ayesha Hashim, Maria Arevalo Mojica, Amanda Joy O’Connor, Urvi Patel, Olivia Pate, Gloria Raise, and et al. 2023. "Assessment of Fungal Succession in Decomposing Swine Carcasses (Sus scrofa L.) Using DNA Metabarcoding" Journal of Fungi 9, no. 9: 866. https://doi.org/10.3390/jof9090866
APA StyleGemmellaro, M. D., Lorusso, N. S., Domke, R., Kovalska, K. M., Hashim, A., Arevalo Mojica, M., O’Connor, A. J., Patel, U., Pate, O., Raise, G., & Shumskaya, M. (2023). Assessment of Fungal Succession in Decomposing Swine Carcasses (Sus scrofa L.) Using DNA Metabarcoding. Journal of Fungi, 9(9), 866. https://doi.org/10.3390/jof9090866






