Manipulation of Auxin Signaling by Smut Fungi during Plant Colonization
Abstract
:1. Smut Fungi, Their Development, and Disease Symptoms
2. Auxin, the Master Regulator of Growth and Development in Plants
3. Auxin in Plant–Pathogen Interactions
4. Auxin Biosynthesis in Smut Fungi
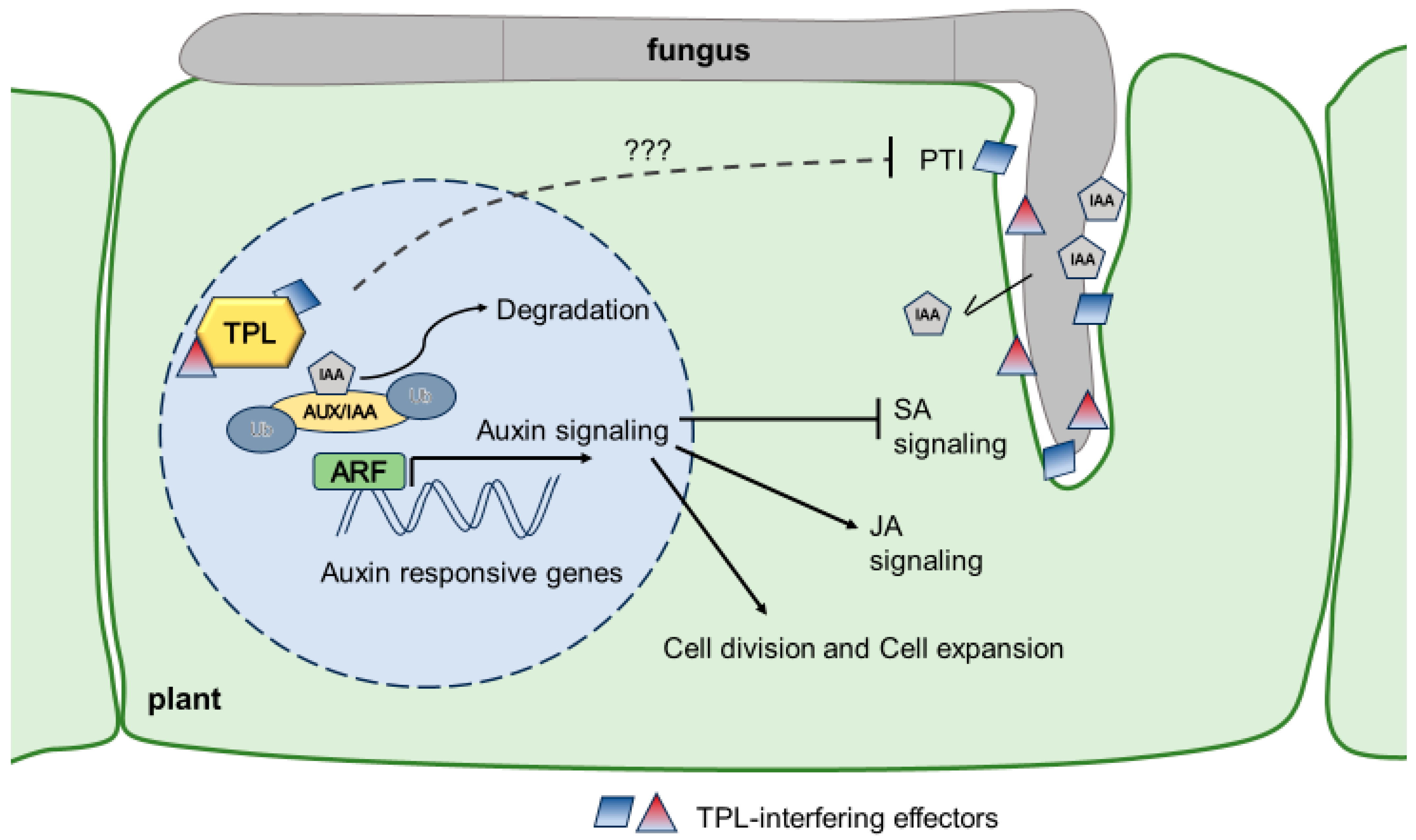
5. Transcriptional Reprogramming of Auxin Signaling by Smuts
6. Plants’ Counter-Defense to Auxin Signaling Manipulation?
7. Conclusions and Outlook
Author Contributions
Funding
Conflicts of Interest
References
- Begerow, D.; Stoll, M.; Bauer, R. A Phylogenetic Hypothesis of Ustilaginomycotina Based on Multiple Gene Analyses and Morphological Data. Mycologia 2006, 98, 906–916. [Google Scholar] [CrossRef]
- Steinberg, G.; Perez-Martin, J. Ustilago maydis, a new fungal model system for cell biology. Trends Cell Biol. 2008, 18, 61–67. [Google Scholar] [CrossRef]
- Djamei, A.; Depotter, J.; Saridis, G.; Prokchorchik, M.; Barghahn, S.; Silva, N.D.S.T.E.; Zuo, W.; Villamil, J.M.; Doehlemann, G. Modulation of Host Immunity and Development by Ustilago maydis. In Plant Relationships: Fungal-Plant Interactions; Scott, B., Mesarich, C., Eds.; Springer International Publishing: Cham, Switzerland, 2023; pp. 3–30. [Google Scholar] [CrossRef]
- Djamei, A. Ustilago maydis. Curr. Biol. 2023, 33, R458–R460. [Google Scholar] [CrossRef] [PubMed]
- Uhse, S.; Djamei, A. Effectors of plant-colonizing fungi and beyond. PLoS Pathog. 2018, 14, e1006992. [Google Scholar] [CrossRef] [PubMed]
- Ghareeb, H.; Drechsler, F.; Löfke, C.; Teichmann, T.; Schirawski, J. SUPPRESSOR OF APICAL DOMINANCE1 of Sporisorium reilianum Modulates Inflorescence Branching Architecture in Maize and Arabidopsis. Plant Physiol. 2015, 169, 2789–2804. [Google Scholar] [CrossRef] [PubMed]
- Navarrete, F.; Grujic, N.; Stirnberg, A.; Aleksza, D.; Gallei, M.; Adi, H.; Bindics, J.; Trujillo, M.; Djamei, A. The Pleiades cluster of fungal effector genes inhibit host defenses. bioRxiv 2019, 827600. [Google Scholar] [CrossRef]
- Navarrete, F.; Gallei, M.; Kornienko, A.E.; Saado, I.; Khan, M.; Chia, K.S.; Darino, M.A.; Bindics, J.; Djamei, A. TOPLESS promotes plant immunity by repressing auxin signaling and is targeted by the fungal effector Naked1. Plant Commun. 2022, 3, 100269. [Google Scholar] [CrossRef] [PubMed]
- Saado, I.; Chia, K.-S.; Betz, R.; Alcântara, A.; Pettkó-Szandtner, A.; Navarrete, F.; D’Auria, J.C.; Kolomiets, M.V.; Melzer, M.; Feussner, I.; et al. Effector-mediated relocalization of a maize lipoxygenase protein triggers susceptibility to Ustilago maydis. Plant Cell 2022, 34, 2785–2805. [Google Scholar] [CrossRef] [PubMed]
- Redkar, A.; Hoser, R.; Schilling, L.; Zechmann, B.; Krzymowska, M.; Walbot, V.; Doehlemann, G. A Secreted Effector Protein of Ustilago maydis Guides Maize Leaf Cells to Form Tumors. Plant Cell 2015, 27, 1332–1351. [Google Scholar] [CrossRef]
- Bindics, J.; Khan, M.; Uhse, S.; Kogelmann, B.; Baggely, L.; Reumann, D.; Ingole, K.D.; Stirnberg, A.; Rybecky, A.; Darino, M.; et al. Many ways to TOPLESS—Manipulation of plant auxin signalling by a cluster of fungal effectors. New Phytol. 2022, 236, 1455–1470. [Google Scholar] [CrossRef]
- Djamei, A.; Schipper, K.; Rabe, F.; Ghosh, A.; Vincon, V.; Kahnt, J.; Osorio, S.; Tohge, T.; Fernie, A.R.; Feussner, I.; et al. Metabolic priming by a secreted fungal effector. Nature 2011, 478, 395–398. [Google Scholar] [CrossRef] [PubMed]
- Khan, M.; Uhse, S.; Bindics, J.; Kogelmann, B.; Nagarajan, N.; Ingole, K.D.; Djamei, A. Tip of the iceberg? Three novel TOPLESS interacting effectors of the gall-inducing fungus Ustilago maydis. bioRxiv 2023. [Google Scholar] [CrossRef]
- Darino, M.; Chia, K.S.; Marques, J.; Aleksza, D.; Soto-Jimenez, L.M.; Saado, I.; Uhse, S.; Borg, M.; Betz, R.; Bindics, J.; et al. Ustilago maydis effector Jsi1 interacts with Topless corepressor, hijacking plant jasmonate/ethylene signaling. New Phytol. 2021, 229, 3393–3407. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Ökmen, B.; Stolze, S.C.; Kastl, M.; Khan, M.; Hilbig, D.; Nakagami, H.; Djamei, A.; Doehlemann, G. The fungal pathogen Ustilago maydis targets the maize corepressor TPL2 to modulate host transcription for tumorigenesis. bioRxiv 2023. [Google Scholar] [CrossRef]
- Zuo, W.; Depotter, J.R.L.; Stolze, S.C.; Nakagami, H.; Doehlemann, G. A transcriptional activator effector of Ustilago maydis regulates hyperplasia in maize during pathogen-induced tumor formation. Nat. Commun. 2023, 14, 6722. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Cao, Q.; Hu, P.; Cui, H.; Yu, X.; Ye, Z. Investigation on the differentiation of two Ustilago esculenta strains—Implications of a relationship with the host phenotypes appearing in the fields. BMC Microbiol. 2017, 17, 228. [Google Scholar] [CrossRef] [PubMed]
- Halisky, P.M.; Barbe, G.D. A Study of Melanopsichium pennsylvanicum Causing Gall Smut on Polygonum. Bull. Torrey Bot. Club 1962, 89, 181–186. [Google Scholar] [CrossRef]
- Sharma, R.; Mishra, B.; Runge, F.; Thines, M. Gene loss rather than gene gain is associated with a host jump from monocots to dicots in the Smut Fungus Melanopsichium pennsylvanicum. Genome Biol. Evol. 2014, 6, 2034–2049. [Google Scholar] [CrossRef]
- Went, F. Wuchsstoff und Wachstum. Recl. Trav. Bot. Néerl. 1928, 25, 1–16. [Google Scholar]
- Caumon, H.; Vernoux, T. A matter of time: Auxin signaling dynamics and the regulation of auxin responses during plant development. J. Exp. Bot. 2023, 74, 3887–3902. [Google Scholar] [CrossRef]
- Yu, Z.; Zhang, F.; Friml, J.; Ding, Z. Auxin signaling: Research advances over the past 30 years. J. Integr Plant Biol. 2022, 64, 371–392. [Google Scholar] [CrossRef] [PubMed]
- Enders, T.A.; Strader, L.C. Auxin activity: Past, present, and future. Am. J. Bot. 2015, 102, 180–196. [Google Scholar] [CrossRef] [PubMed]
- Gomes, G.L.B.; Scortecci, K.C. Auxin and its role in plant development: Structure, signalling, regulation and response mechanisms. Plant Biol. 2021, 23, 894–904. [Google Scholar] [CrossRef]
- Cao, X.; Yang, H.; Shang, C.; Ma, S.; Liu, L.; Cheng, J. The Roles of Auxin Biosynthesis YUCCA Gene Family in Plants. Int. J. Mol. Sci. 2019, 20, 6343. [Google Scholar] [CrossRef] [PubMed]
- Leyser, O. Auxin Signaling. Plant Physiol. 2018, 176, 465–479. [Google Scholar] [CrossRef]
- Weijers, D.; Wagner, D. Transcriptional Responses to the Auxin Hormone. Annu. Rev. Plant Biol. 2016, 67, 539–574. [Google Scholar] [CrossRef]
- Ang, A.C.H.; Ostergaard, L. Save your TIRs—More to auxin than meets the eye. New Phytol. 2023, 238, 971–976. [Google Scholar] [CrossRef]
- Szemenyei, H.; Hannon, M.; Long, J.A. TOPLESS mediates auxin-dependent transcriptional repression during Arabidopsis embryogenesis. Science 2008, 319, 1384–1386. [Google Scholar] [CrossRef]
- Ke, J.; Ma, H.; Gu, X.; Thelen, A.; Brunzelle, J.S.; Li, J.; Xu, H.E.; Melcher, K. Structural basis for recognition of diverse transcriptional repressors by the TOPLESS family of corepressors. Sci. Adv. 2015, 1, e1500107. [Google Scholar] [CrossRef]
- Ulmasov, T.; Murfett, J.; Hagen, G.; Guilfoyle, T.J. Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 1997, 9, 1963–1971. [Google Scholar] [CrossRef]
- Eckardt, N.A. Auxin and the power of the proteasome in plants. Plant Cell 2001, 13, 2161–2163. [Google Scholar] [CrossRef]
- Huot, B.; Yao, J.; Montgomery, B.L.; He, S.Y. Growth-defense tradeoffs in plants: A balancing act to optimize fitness. Mol. Plant 2014, 7, 1267–1287. [Google Scholar] [CrossRef] [PubMed]
- Iglesias, M.J.; Terrile, M.C.; Casalongué, C.A. Auxin and salicylic acid signalings counteract the regulation of adaptive responses to stress. Plant Signal. Behav. 2011, 6, 452–454. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.; Dhiman, V.K.; Pandey, H.; Dhiman, V.K.; Pandey, D. Crosstalk Between Salicylic Acid and Auxins, Cytokinins and Gibberellins Under Biotic Stress. In Auxins, Cytokinins and Gibberellins Signaling in Plants; Aftab, T., Ed.; Springer International Publishing: Cham, Switzerland, 2022; pp. 249–262. [Google Scholar] [CrossRef]
- Kunkel, B.N.; Johnson, J.M.B. Auxin Plays Multiple Roles during Plant-Pathogen Interactions. Cold Spring Harb. Perspect. Biol. 2021, 13, a040022. [Google Scholar] [CrossRef] [PubMed]
- Oosterbeek, M.; Lozano-Torres, J.L.; Bakker, J.; Goverse, A. Sedentary Plant-Parasitic Nematodes Alter Auxin Homeostasis via Multiple Strategies. Front. Plant Sci. 2021, 12, 668548. [Google Scholar] [CrossRef]
- Chen, Z.; Agnew, J.L.; Cohen, J.D.; He, P.; Shan, L.; Sheen, J.; Kunkel, B.N. Pseudomonas syringae type III effector AvrRpt2 alters Arabidopsis thaliana auxin physiology. Proc. Natl. Acad. Sci. USA 2007, 104, 20131–20136. [Google Scholar] [CrossRef] [PubMed]
- Hilbert, M.; Nostadt, R.; Zuccaro, A. Exogenous auxin affects the oxidative burst in barley roots colonized by Piriformospora indica. Plant Signal. Behav. 2013, 8, e23572. [Google Scholar] [CrossRef]
- Tang, J.; Li, Y.; Zhang, L.; Mu, J.; Jiang, Y.; Fu, H.; Zhang, Y.; Cui, H.; Yu, X.; Ye, Z. Biosynthetic Pathways and Functions of Indole-3-Acetic Acid in Microorganisms. Microorganisms 2023, 11, 2077. [Google Scholar] [CrossRef]
- Grunewald, W.; Cannoot, B.; Friml, J.; Gheysen, G. Parasitic nematodes modulate PIN-mediated auxin transport to facilitate infection. PLoS Pathog. 2009, 5, e1000266. [Google Scholar] [CrossRef]
- Yang, L.; Meng, D.; Wang, Y.; Wu, Y.; Lang, C.; Jin, T.; Zhou, X. The viral suppressor HCPro decreases DNA methylation and activates auxin biosynthesis genes. Virology 2020, 546, 133–140. [Google Scholar] [CrossRef]
- Padmanabhan, M.S.; Goregaoker, S.P.; Golem, S.; Shiferaw, H.; Culver, J.N. Interaction of the tobacco mosaic virus replicase protein with the Aux/IAA protein PAP1/IAA26 is associated with disease development. J. Virol. 2005, 79, 2549–2558. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Qin, Q.; Wang, Y.; Pu, Y.; Liu, L.; Wen, X.; Ji, S.; Wu, J.; Wei, C.; Ding, B.; et al. Rice Dwarf Virus P2 Protein Hijacks Auxin Signaling by Directly Targeting the Rice OsIAA10 Protein, Enhancing Viral Infection and Disease Development. PLoS Pathog. 2016, 12, e1005847. [Google Scholar] [CrossRef] [PubMed]
- Cui, F.; Wu, S.; Sun, W.; Coaker, G.; Kunkel, B.; He, P.; Shan, L. The Pseudomonas syringae type III effector AvrRpt2 promotes pathogen virulence via stimulating Arabidopsis auxin/indole acetic acid protein turnover. Plant Physiol. 2013, 162, 1018–1029. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Li, L.; He, Y.; Qin, Q.; Chen, C.; Wei, Z.; Tan, X.; Xie, K.; Zhang, R.; Hong, G.; et al. Distinct modes of manipulation of rice auxin response factor OsARF17 by different plant RNA viruses for infection. Proc. Natl. Acad. Sci. USA 2020, 117, 9112–9121. [Google Scholar] [CrossRef]
- Cerboneschi, M.; Decorosi, F.; Biancalani, C.; Ortenzi, M.V.; Macconi, S.; Giovannetti, L.; Viti, C.; Campanella, B.; Onor, M.; Bramanti, E.; et al. Indole-3-acetic acid in plant-pathogen interactions: A key molecule for in planta bacterial virulence and fitness. Res. Microbiol. 2016, 167, 774–787. [Google Scholar] [CrossRef] [PubMed]
- Qiao, L.; Zheng, L.; Sheng, C.; Zhao, H.; Jin, H.; Niu, D. Rice siR109944 suppresses plant immunity to sheath blight and impacts multiple agronomic traits by affecting auxin homeostasis. Plant J. 2020, 102, 948–964. [Google Scholar] [CrossRef]
- Qi, L.; Yan, J.; Li, Y.; Jiang, H.; Sun, J.; Chen, Q.; Li, H.; Chu, J.; Yan, C.; Sun, X.; et al. Arabidopsis thaliana plants differentially modulate auxin biosynthesis and transport during defense responses to the necrotrophic pathogen Alternaria brassicicola. New Phytol. 2012, 195, 872–882. [Google Scholar] [CrossRef]
- Wang, B.; Chu, J.; Yu, T.; Xu, Q.; Sun, X.; Yuan, J.; Xiong, G.; Wang, G.; Wang, Y.; Li, J. Tryptophan-independent auxin biosynthesis contributes to early embryogenesis in Arabidopsis. Proc. Natl. Acad. Sci. USA 2015, 112, 4821–4826. [Google Scholar] [CrossRef]
- Fu, J.; Wang, S. Insights into auxin signaling in plant-pathogen interactions. Front. Plant Sci. 2011, 2, 74. [Google Scholar] [CrossRef]
- Woodward, A.W.; Bartel, B. Auxin: Regulation, action, and interaction. Ann. Bot. 2005, 95, 707–735. [Google Scholar] [CrossRef]
- Turian, G.; Hamilton, R.H. Chemical detection of 3-indolylacetic acid in Ustilago zeae tumors. Biochim. Biophys. Acta 1960, 41, 148–150. [Google Scholar] [CrossRef]
- Wolf, F.T. The Production of Indole Acetic Acid by Ustilago Zeae, and Its Possible Significance in Tumor Formation. Proc. Natl. Acad. Sci. USA 1952, 38, 106–111. [Google Scholar] [CrossRef]
- Basse, C.W.; Lottspeich, F.; Steglich, W.; Kahmann, R. Two potential indole-3-acetaldehyde dehydrogenases in the phytopathogenic fungus Ustilago maydis. Eur. J. Biochem. 1996, 242, 648–656. [Google Scholar] [CrossRef]
- Reineke, G.; Heinze, B.; Schirawski, J.; Buettner, H.; Kahmann, R.; Basse, C.W. Indole-3-acetic acid (IAA) biosynthesis in the smut fungus Ustilago maydis and its relevance for increased IAA levels in infected tissue and host tumour formation. Mol. Plant Pathol. 2008, 9, 339–355. [Google Scholar] [CrossRef]
- Cui, G.; Huang, C.; Bi, X.; Wang, Y.; Yin, K.; Zhu, L.; Jiang, Z.; Chen, B.; Deng, Y.Z. Aminotransferase SsAro8 Regulates Tryptophan Metabolism Essential for Filamentous Growth of Sugarcane Smut Fungus Sporisorium scitamineum. Microbiol. Spectr. 2022, 10, e0057022. [Google Scholar] [CrossRef] [PubMed]
- Chung, K.R.; Tzeng, D. Biosynthesis of Indole-3-Acetic Acid by the Gall-inducing Fungus Ustilago esculenta. J. Biol. Sci. 2004, 4, 744–750. [Google Scholar] [CrossRef]
- Dong, L.; Shen, Q.; Chen, C.-Y.; Shen, L.; Yang, F.; Naqvi, N.I.; Deng, Y.Z. Fungal auxin is a quorum-based modulator of blast disease severity. bioRxiv 2021. [Google Scholar] [CrossRef]
- Doehlemann, G.; Wahl, R.; Horst, R.J.; Voll, L.M.; Usadel, B.; Poree, F.; Stitt, M.; Pons-Kuhnemann, J.; Sonnewald, U.; Kahmann, R.; et al. Reprogramming a maize plant: Transcriptional and metabolic changes induced by the fungal biotroph Ustilago maydis. Plant J. Cell Mol. Biol. 2008, 56, 181–195. [Google Scholar] [CrossRef]
- Hiratsu, K.; Matsui, K.; Koyama, T.; Ohme-Takagi, M. Dominant repression of target genes by chimeric repressors that include the EAR motif, a repression domain, in Arabidopsis. Plant J. Cell Mol. Biol. 2003, 34, 733–739. [Google Scholar] [CrossRef] [PubMed]
- Kacprzyk, J.; Burke, R.; Schwarze, J.; McCabe, P.F. Plant programmed cell death meets auxin signalling. FEBS J. 2022, 289, 1731–1745. [Google Scholar] [CrossRef] [PubMed]
- Gawehns, F.K.K. Function and Targets of Fusarium Oxysporum Effectors. Ph.D. Thesis, Universiteit van Amsterdam, Amsterdam, The Netherlands, 2014. [Google Scholar]
- Segonzac, C.; Newman, T.E.; Choi, S.; Jayaraman, J.; Choi, D.S.; Jung, G.Y.; Cho, H.; Lee, Y.K.; Sohn, K.H. A Conserved EAR Motif Is Required for Avirulence and Stability of the Ralstonia solanacearum Effector PopP2 In Planta. Front. Plant Sci. 2017, 8, 1330. [Google Scholar] [CrossRef] [PubMed]

| Percentage Identity | |||||
|---|---|---|---|---|---|
| Scientific Name | Gall Formation | Tam1 | Tam2 | Iad1 | Iad2 |
| Ustilago maydis 521 | + | 100 | 100 | 100 | 100 |
| Pseudozyma hubeiensis | - | 85.53 | 91.05 | 95.98 | 90.66 |
| Sporisorium scitamineum | - | 86.17 | 89.86 | 96.38 | 37.15 |
| Sporisorium graminicola | - | 86.32 | 89.26 | 96.78 | 87.55 |
| Sporisorium reilianum | - | 83.58 | 89.46 | 95.17 | 36.52 |
| Ustilago sp., UG-2017a | - | 80.13 | 84.26 | 94.57 | 65.49 |
| Ustilago hordei | - | 79.92 | 84.46 | 93.96 | 36.94 |
| Ustilago trichophora | + | 82.24 | 89.46 | 96.18 | 84.41 |
| Ustilago nuda | - | 79.96 | 84.06 | 93.76 | 36.73 |
| Ustilago tritici | - | 79.50 | 83.47 | 93.96 | 59.63 |
| Moesziomyces antarcticus | - | 77.12 | 84.69 | 94.16 | 85.86 |
| Melanopsichium pennsylvanicum | + | 74.23 | 87.08 | 94.57 | 36.17 |
| Kalmanozyma brasiliensis GHG001 | - | 74.04 | 86.48 | 92.96 | 82.37 |
| Ustilago bromivora | - | 74.27 | 83.86 | 94.57 | 67.98 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nagarajan, N.; Khan, M.; Djamei, A. Manipulation of Auxin Signaling by Smut Fungi during Plant Colonization. J. Fungi 2023, 9, 1184. https://doi.org/10.3390/jof9121184
Nagarajan N, Khan M, Djamei A. Manipulation of Auxin Signaling by Smut Fungi during Plant Colonization. Journal of Fungi. 2023; 9(12):1184. https://doi.org/10.3390/jof9121184
Chicago/Turabian StyleNagarajan, Nithya, Mamoona Khan, and Armin Djamei. 2023. "Manipulation of Auxin Signaling by Smut Fungi during Plant Colonization" Journal of Fungi 9, no. 12: 1184. https://doi.org/10.3390/jof9121184
APA StyleNagarajan, N., Khan, M., & Djamei, A. (2023). Manipulation of Auxin Signaling by Smut Fungi during Plant Colonization. Journal of Fungi, 9(12), 1184. https://doi.org/10.3390/jof9121184






