Independent Evolution Has Led to Distinct Genomic Signatures in Dutch Elm Disease-Causing Fungi and Other Vascular Wilts-Causing Fungal Pathogens
Abstract
1. Introduction
2. Materials and Methods
2.1. Wilt-Causing Species and Genome Assemblies
2.2. Gene Molecular Functions
2.3. Phylogeny
2.4. Ancestral State Reconstructions
2.5. Statistical Analyses
3. Results
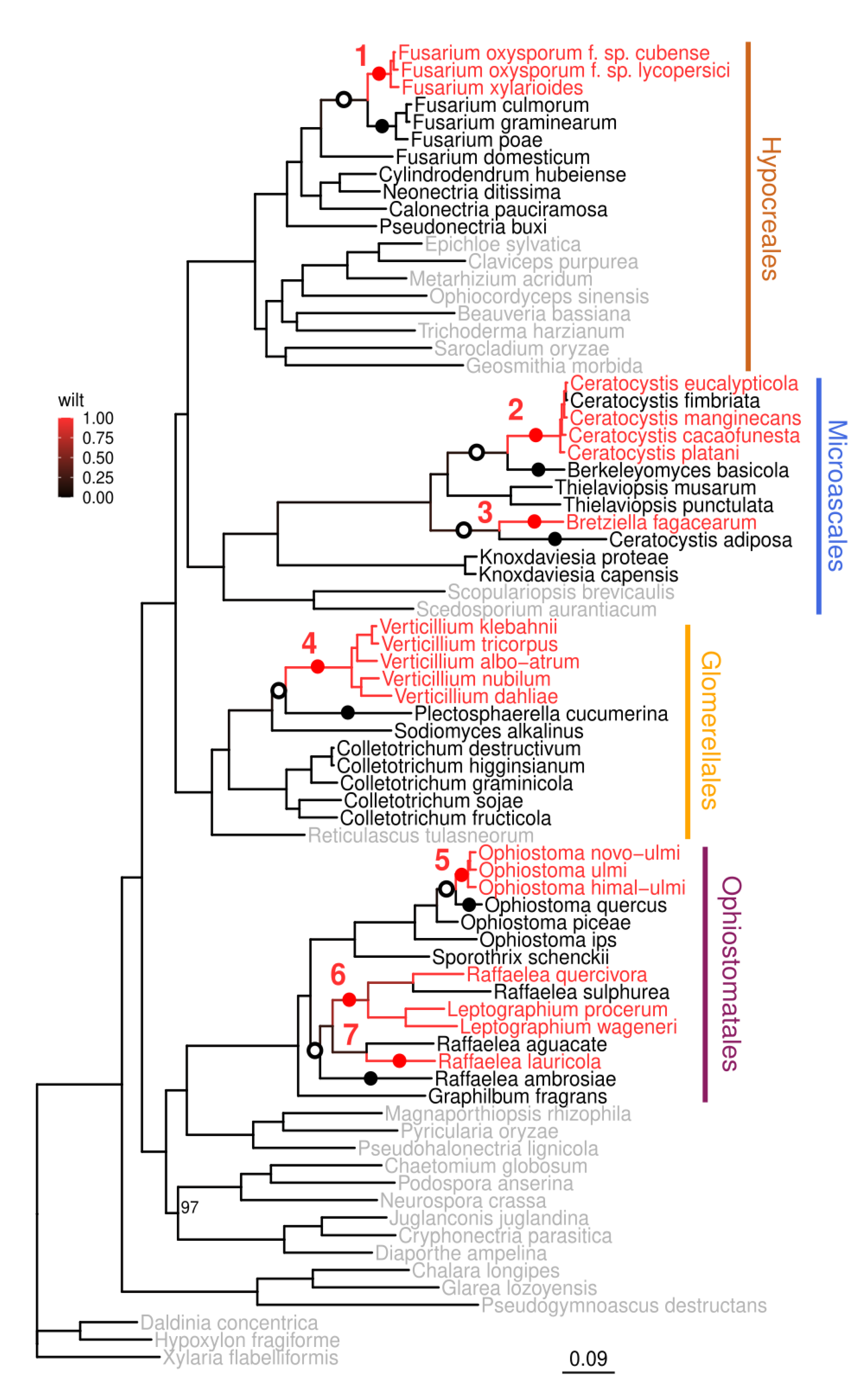
3.1. Wilt Pathogens Evolved Multiple Times
3.2. Pathogen-Related Gene Repertoires Do Not Distinguish Wilt from Non-Wilt-Causing Species
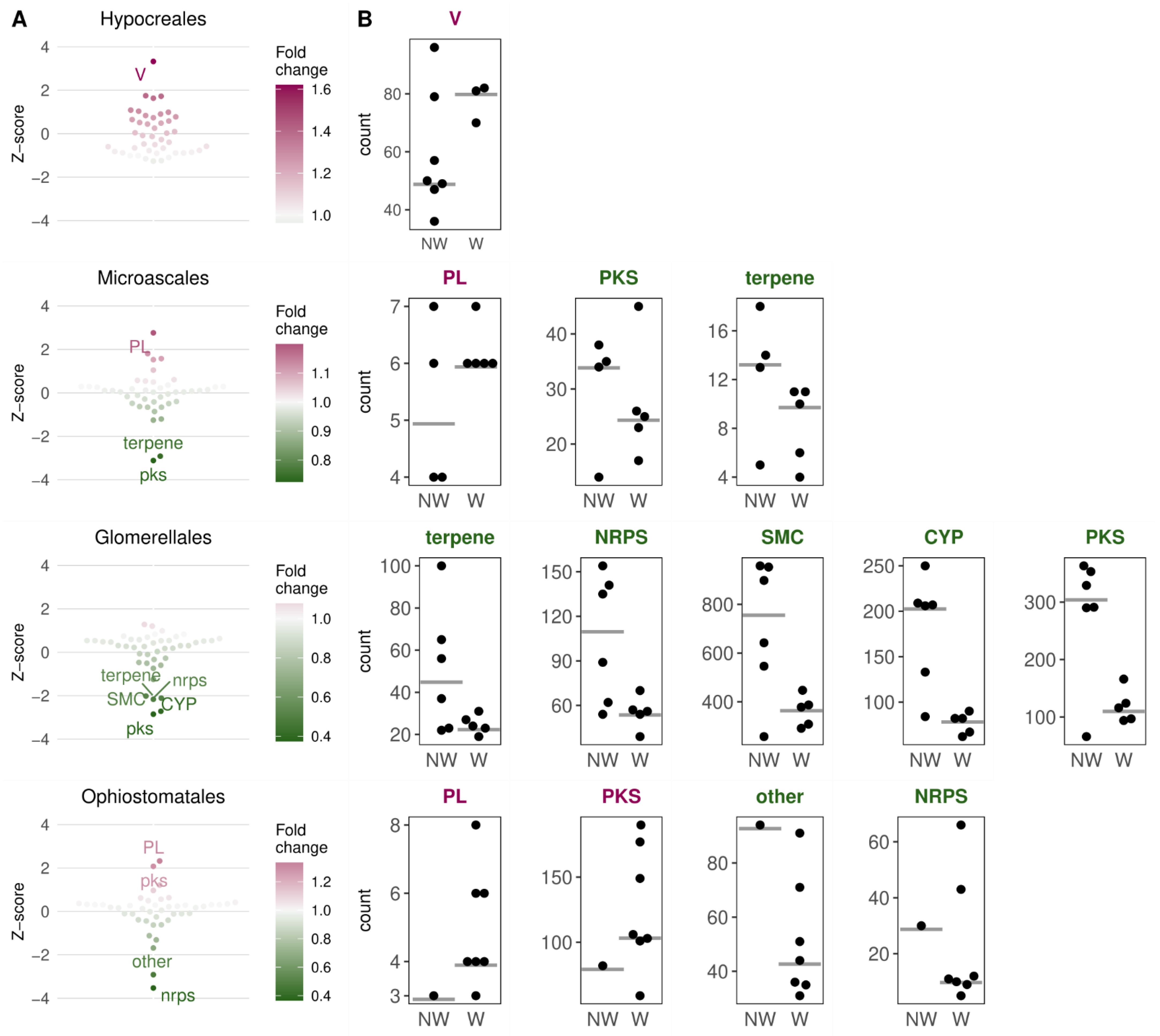
3.3. Secondary Metabolite Cluster (SMC) Diversification in Distinct Wilt Groups
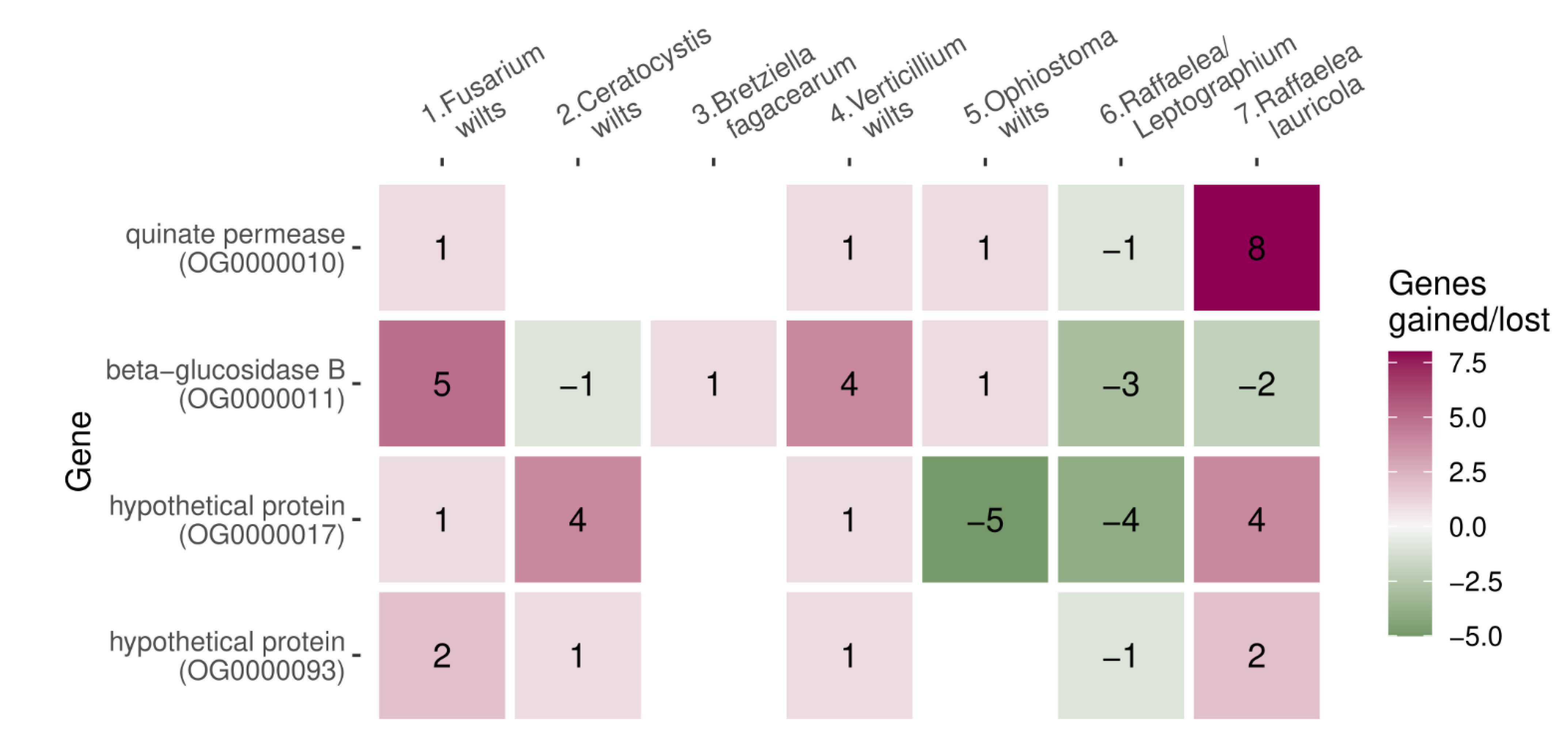
3.4. Gene Diversifications Shared among Wilt Groups
3.5. Dutch Elm Disease
4. Discussion
4.1. Genomics of Wilt Pathogens
4.2. Differences in Gene Repertoires across Wilt Pathogens
4.3. Common Patterns of Gene Diversification between Wilt Groups
4.4. No Large-Scale Expansions in DED Pathogens
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Harper, C.J.; Bomfleur, B.; Decombeix, A.-L.; Taylor, E.L.; Taylor, T.N.; Krings, M. Tylosis Formation and Fungal Interactions in an Early Jurassic Conifer from Northern Victoria Land, Antarctica. Rev. Palaeobot. Palynol. 2012, 175, 25–31. [Google Scholar] [CrossRef]
- Yadeta, K.A.; Thomma, B.P.H. The Xylem as Battleground for Plant Hosts and Vascular Wilt Pathogens. Front. Plant Sci. 2013, 4, 97. [Google Scholar] [CrossRef] [PubMed]
- Clérivet, A.; Déon, V.; Alami, I.; Lopez, F.; Geiger, J.-P.; Nicole, M. Tyloses and Gels Associated with Cellulose Accumulation in Vessels Are Responses of Plane Tree Seedlings (Platanus × acerifolia) to the Vascular Fungus Ceratocystis fimbriata f. sp. platani. Trees 2000, 15, 25–31. [Google Scholar] [CrossRef]
- Martín, J.A.; Solla, A.; Domingues, M.R.; Coimbra, M.A.; Gil, L. Exogenous Phenol Increase Resistance of Ulmus minor to Dutch Elm Disease through Formation of Suberin-like Compounds on Xylem Tissues. Environ. Exp. Bot. 2008, 64, 97–104. [Google Scholar] [CrossRef]
- Et-Touil, A.; Rioux, D.; Mathieu, F.M.; Bernier, L. External Symptoms and Histopathological Changes Following Inoculation of Elms Putatively Resistant to Dutch Elm Disease with Genetically Close Strains of Ophiostoma. Can. J. Bot. 2005, 83, 656–667. [Google Scholar] [CrossRef]
- Kashyap, A.; Planas-Marquès, M.; Capellades, M.; Valls, M.; Coll, N.S. Blocking Intruders: Inducible Physico-Chemical Barriers against Plant Vascular Wilt Pathogens. J. Exp. Bot. 2021, 72, 184–198. [Google Scholar] [CrossRef]
- Kashyap, A.; Jiménez-Jiménez, Á.L.; Zhang, W.; Capellades, M.; Srinivasan, S.; Laromaine, A.; Serra, O.; Figueras, M.; Rencoret, J.; Gutiérrez, A.; et al. Induced Ligno-Suberin Vascular Coating and Tyramine-Derived Hydroxycinnamic Acid Amides Restrict Ralstonia solanacearum Colonization in Resistant Tomato. New Phytol. 2022, 234, 1411–1429. [Google Scholar] [CrossRef]
- Schwarz, M. Das Zweigensterben Der Olmen, Trauerweiden Und Pfirschbäume [The Twig Dying of the Elms, Willows, and Peach Trees]. Meded. Phytopath. Lab. Willie Commelin Sch. 1922, 5, 1–73. [Google Scholar]
- Gibbs, J.N. Intercontinental Epidemiology of Dutch Elm Disease. Annu. Rev. Phytopathol. 1978, 16, 287–307. [Google Scholar] [CrossRef]
- Gibbs, J.N.; Heybroek, H.M.; Holmes, F.W. Aggressive Strain of Ceratocystis ulmi in Britain. Nature 1972, 236, 121–122. [Google Scholar] [CrossRef]
- Brasier, C.M.; Buck, K.W. Rapid Evolutionary Changes in a Globally Invading Fungal Pathogen (Dutch Elm Disease). Biol. Invasions 2001, 3, 223–233. [Google Scholar] [CrossRef]
- Brasier, C.M. Ophiostoma novo-ulmi sp. nov., Causative Agent of Current Dutch Elm Disease Pandemics. Mycopathologia 1991, 115, 151–161. [Google Scholar] [CrossRef]
- Juzwik, J.; Appel, D.N.; MacDonald, W.L.; Burks, S. Challenges and Successes in Managing Oak Wilt in the United States. Plant Dis. 2011, 95, 888–900. [Google Scholar] [CrossRef] [PubMed]
- Sinclair, W.A.; Lyon, H.H. Diseases of Trees and Shrubs; Cornell University Press: Ithaca, NY, USA, 1987. [Google Scholar]
- Harrington, T.C.; Cobb, F.W. Pathogenicity of Leptographium and Verticicladiella spp. Isolated from Roots of Western North American Conifers. Phytopathology 1983, 73, 596–599. [Google Scholar] [CrossRef]
- Witcosky, J.J.; Schowalter, T.D.; Hansen, E.M. Hylastes nigrinus (Coleoptera: Scolytidae), Pissodes fasciatus and Steremnius carinatus (Coleoptera: Curculionidae) as Vectors of Black-Stain Root Disease of Douglas-Fir. Environ. Entomol. 1986, 15, 1090–1095. [Google Scholar] [CrossRef]
- De Beer, Z.W.; Procter, M.; Wingfield, M.J.; Marincowitz, S.; Duong, T.A. Generic Boundaries in the Ophiostomatales Reconsidered and Revised. Stud. Mycol. 2022, 101, 57–120. [Google Scholar] [CrossRef]
- Fraedrich, S.W.; Harrington, T.C.; Rabaglia, R.J. Laurel Wilt: A New and Devastating Disease of Redbay Caused by a Fungal Symbiont of the Exotic Redbay Ambrosia Beetle. Newsl. Mich. Entomol. Soc. 2007, 52, 15–16. [Google Scholar]
- Harrington, T.C.; Fraedrich, S.W.; Aghayeva, D.N. Raffaelea lauricola, a New Ambrosia Beetle Symbiont and Pathogen on the Lauraceae. Mycotaxon 2008, 104, 399–404. [Google Scholar]
- Laurel Wilt. Available online: https://www.invasivespeciesinfo.gov/terrestrial/pathogens-and-diseases/laurel-wilt (accessed on 12 July 2022).
- Cannon, P.; Friday, J.B.; Harrington, T.; Keith, L.; Hughes, M.; Hauff, R.; Hughes, F.; Perroy, R.; Benitez, D.; Roy, K.; et al. Rapid ‘Ōhi‘a Death in Hawai‘i. In Forest Microbiology: Volume 2: Forest Tree Health; Asiegbu, F., Kovalchuk, A., Eds.; Academic Press: London, UK, 2022; Volume 2, pp. 267–289. [Google Scholar]
- Przybyl, K.; Dahm, H.; Ciesielska, A.; Molinski, K. Cellulolytic Activity and Virulence of Ophiostoma ulmi and O. novo-ulmi Isolates. For. Pathol. 2006, 36, 58–67. [Google Scholar] [CrossRef]
- Svaldi, R.; Elgersma, D.M. Further Studies on the Activity of Cell Wall Degrading Enzymes of Aggressive and Non-Aggressive Isolates of Ophiostoma ulmi. For. Pathol. 1982, 12, 29–36. [Google Scholar] [CrossRef]
- Binz, T.; Canevascini, G. Xylanases from the Dutch Elm Disease pathogens Ophiostoma ulmi and Ophiostoma novo-ulmi. Physiol. Mol. Plant Pathol. 1996, 49, 159–175. [Google Scholar] [CrossRef]
- Scheffer, R.J.; Elgersma, D.M. A Scanning Electron Microscope Study of Cell Wall Degradation in Elm Wood by Aggressive and Non-Aggressive Isolates of Ophiostoma ulmi. For. Pathol. 1982, 12, 25–28. [Google Scholar] [CrossRef]
- Ma, L.-J.; van der Does, H.C.; Borkovich, K.A.; Coleman, J.J.; Daboussi, M.-J.; Di Pietro, A.; Dufresne, M.; Freitag, M.; Grabherr, M.; Henrissat, B.; et al. Comparative Genomics Reveals Mobile Pathogenicity Chromosomes in Fusarium. Nature 2010, 464, 367–373. [Google Scholar] [CrossRef] [PubMed]
- Klosterman, S.J.; Subbarao, K.V.; Kang, S.; Veronese, P.; Gold, S.E.; Thomma, B.P.H.J.; Chen, Z.; Henrissat, B.; Lee, Y.-H.; Park, J.; et al. Comparative Genomics Yields Insights into Niche Adaptation of Plant Vascular Wilt Pathogens. PLoS Pathog. 2011, 7, e1002137. [Google Scholar] [CrossRef] [PubMed]
- Hamelin, R.C.; Roe, A.D. Genomic Biosurveillance of Forest Invasive Alien Enemies: A Story Written in Code. Evol. Appl. 2019, 13, 95–115. [Google Scholar] [CrossRef] [PubMed]
- Hessenauer, P.; Fijarczyk, A.; Martin, H.; Prunier, J.; Charron, G.; Chapuis, J.; Bernier, L.; Tanguay, P.; Hamelin, R.C.; Landry, C.R. Hybridization and Introgression Drive Genome Evolution of Dutch Elm Disease Pathogens. Nat. Ecol. Evol. 2020, 4, 626–638. [Google Scholar] [CrossRef]
- Takahashi, Y.; Matsushita, N.; Hogetsu, T. Spatial Distribution of Raffaelea quercivora in Xylem of Naturally Infested and Inoculated Oak Trees. Phytopathology 2010, 100, 747–755. [Google Scholar] [CrossRef]
- Lieutier, F.; Yart, A.; Salle, A. Stimulation of Tree Defenses by Ophiostomatoid Fungi Can Explain Attack Success of Bark Beetles on Conifers. Ann. For. Sci. 2009, 66, 801. [Google Scholar] [CrossRef]
- O’Connell, R.J.; Thon, M.R.; Hacquard, S.; Amyotte, S.G.; Kleemann, J.; Torres, M.F.; Damm, U.; Buiate, E.A.; Epstein, L.; Alkan, N.; et al. Lifestyle Transitions in Plant Pathogenic Colletotrichum Fungi Deciphered by Genome and Transcriptome Analyses. Nat. Genet. 2012, 44, 1060–1065. [Google Scholar] [CrossRef]
- Zampounis, A.; Pigné, S.; Dallery, J.-F.; Wittenberg, A.H.J.; Zhou, S.; Schwartz, D.C.; Thon, M.R.; O’Connell, R.J. Genome Sequence and Annotation of Colletotrichum higginsianum, a Causal Agent of Crucifer Anthracnose Disease. Genome Announc. 2016, 4, e00821-16. [Google Scholar] [CrossRef]
- Urban, M.; King, R.; Andongabo, A.; Maheswari, U.; Pedro, H.; Kersey, P.; Hammond-Kosack, K. First Draft Genome Sequence of a UK Strain (UK99) of Fusarium culmorum. Genome Announc. 2016, 4, e00771-16. [Google Scholar] [CrossRef] [PubMed]
- Cuomo, C.A.; Güldener, U.; Xu, J.-R.; Trail, F.; Turgeon, B.G.; Di Pietro, A.; Walton, J.D.; Ma, L.-J.; Baker, S.E.; Rep, M.; et al. The Fusarium graminearum Genome Reveals a Link between Localized Polymorphism and Pathogen Specialization. Science 2007, 317, 1400–1402. [Google Scholar] [CrossRef] [PubMed]
- DeIulio, G.A.; Guo, L.; Zhang, Y.; Goldberg, J.M.; Kistler, H.C.; Ma, L.-J. Kinome Expansion in the Fusarium oxysporum Species Complex Driven by Accessory Chromosomes. mSphere 2018, 3, e00231-18. [Google Scholar] [CrossRef]
- Gómez-Cortecero, A.; Harrison, R.J.; Armitage, A.D. Draft Genome Sequence of a European Isolate of the Apple Canker Pathogen Neonectria ditissima. Genome Announc. 2015, 3, e01243-15. [Google Scholar] [CrossRef] [PubMed]
- Forgetta, V.; Leveque, G.; Dias, J.; Grove, D.; Lyons, R., Jr.; Genik, S.; Wright, C.; Singh, S.; Peterson, N.; Zianni, M.; et al. Sequencing of the Dutch Elm Disease Fungus Genome Using the Roche/454 GS-FLX Titanium System in a Comparison of Multiple Genomics Core Facilities. J. Biomol. Tech. 2013, 24, 39–49. [Google Scholar] [CrossRef] [PubMed]
- Haridas, S.; Wang, Y.; Lim, L.; Massoumi Alamouti, S.; Jackman, S.; Docking, R.; Robertson, G.; Birol, I.; Bohlmann, J.; Breuil, C. The Genome and Transcriptome of the Pine Saprophyte Ophiostoma piceae, and a Comparison with the Bark Beetle-Associated Pine Pathogen Grosmannia clavigera. BMC Genom. 2013, 14, 373. [Google Scholar] [CrossRef]
- Fijarczyk, A.; Hessenauer, P.; Hamelin, R.C.; Landry, C.R. Lifestyles Shape Genome Size and Gene Content in Fungal Pathogens. bioRxiv 2022. [Google Scholar] [CrossRef]
- Waterhouse, R.M.; Seppey, M.; Simão, F.A.; Manni, M.; Ioannidis, P.; Klioutchnikov, G.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO Applications from Quality Assessments to Gene Prediction and Phylogenomics. Mol. Biol. Evol. 2018, 35, 543–548. [Google Scholar] [CrossRef]
- Stanke, M.; Morgenstern, B. AUGUSTUS: A Web Server for Gene Prediction in Eukaryotes That Allows User-Defined Constraints. Nucleic Acids Res. 2005, 33, W465–W467. [Google Scholar] [CrossRef]
- Emms, D.M.; Kelly, S. OrthoFinder: Phylogenetic Orthology Inference for Comparative Genomics. Genome Biol. 2019, 20, 238. [Google Scholar] [CrossRef]
- Cantarel, B.L.; Coutinho, P.M.; Rancurel, C.; Bernard, T.; Lombard, V.; Henrissat, B. The Carbohydrate-Active EnZymes Database (CAZy): An Expert Resource for Glycogenomics. Nucleic Acids Res. 2009, 37, D233–D238. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Barrett, A.J.; Thomas, P.D.; Huang, X.; Bateman, A.; Finn, R.D. The MEROPS Database of Proteolytic Enzymes, Their Substrates and Inhibitors in 2017 and a Comparison with Peptidases in the PANTHER Database. Nucleic Acids Res. 2018, 46, D624–D632. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Détry, N.; Kim, K.-T.; Asiegbu, F.O.; Valkonen, J.P.T.; Lee, Y.-H. fPoxDB: Fungal Peroxidase Database for Comparative Genomics. BMC Microbiol. 2014, 14, 117. [Google Scholar] [CrossRef] [PubMed]
- Urban, M.; Cuzick, A.; Seager, J.; Wood, V.; Rutherford, K.; Venkatesh, S.Y.; De Silva, N.; Martinez, M.C.; Pedro, H.; Yates, A.D.; et al. PHI-Base: The Pathogen-Host Interactions Database. Nucleic Acids Res. 2020, 48, D613–D620. [Google Scholar] [CrossRef] [PubMed]
- Tatusov, R.L.; Fedorova, N.D.; Jackson, J.D.; Jacobs, A.R.; Kiryutin, B.; Koonin, E.V.; Krylov, D.M.; Mazumder, R.; Mekhedov, S.L.; Nikolskaya, A.N.; et al. The COG Database: An Updated Version Includes Eukaryotes. BMC Bioinform. 2003, 4, 41. [Google Scholar] [CrossRef]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The Protein Families Database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef]
- Buchfink, B.; Xie, C.; Huson, D.H. Fast and Sensitive Protein Alignment Using DIAMOND. Nat. Methods 2015, 12, 59–60. [Google Scholar] [CrossRef]
- Mistry, J.; Finn, R.D.; Eddy, S.R.; Bateman, A.; Punta, M. Challenges in Homology Search: HMMER3 and Convergent Evolution of Coiled-Coil Regions. Nucleic Acids Res. 2013, 41, e121. [Google Scholar] [CrossRef]
- Blin, K.; Pascal Andreu, V.; de Los Santos, E.L.C.; Del Carratore, F.; Lee, S.Y.; Medema, M.H.; Weber, T. The antiSMASH Database Version 2: A Comprehensive Resource on Secondary Metabolite Biosynthetic Gene Clusters. Nucleic Acids Res. 2019, 47, D625–D630. [Google Scholar] [CrossRef]
- Katoh, K.; Standley, D.M. MAFFT Multiple Sequence Alignment Software Version 7: Improvements in Performance and Usability. Mol. Biol. Evol. 2013, 30, 772–780. [Google Scholar] [CrossRef]
- Capella-Gutiérrez, S.; Silla-Martínez, J.M.; Gabaldón, T. trimAl: A Tool for Automated Alignment Trimming in Large-Scale Phylogenetic Analyses. Bioinformatics 2009, 25, 1972–1973. [Google Scholar] [CrossRef]
- Nguyen, L.-T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A Fast and Effective Stochastic Algorithm for Estimating Maximum-Likelihood Phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Kalyaanamoorthy, S.; Minh, B.Q.; Wong, T.K.F.; von Haeseler, A.; Jermiin, L.S. ModelFinder: Fast Model Selection for Accurate Phylogenetic Estimates. Nat. Methods 2017, 14, 587–589. [Google Scholar] [CrossRef]
- Hoang, D.T.; Chernomor, O.; von Haeseler, A.; Minh, B.Q.; Vinh, L.S. UFBoot2: Improving the Ultrafast Bootstrap Approximation. Mol. Biol. Evol. 2018, 35, 518–522. [Google Scholar] [CrossRef]
- Sanderson, M.J. r8s: Inferring Absolute Rates of Molecular Evolution and Divergence Times in the Absence of a Molecular Clock. Bioinformatics 2003, 19, 301–302. [Google Scholar] [CrossRef] [PubMed]
- Hedges, B.; Kumar, S. TIMETREE 5: The Timescale of Life. Available online: http://www.timetree.org/ (accessed on 7 December 2021).
- Paradis, E.; Claude, J.; Strimmer, K. APE: Analyses of Phylogenetics and Evolution in R Language. Bioinformatics 2004, 20, 289–290. [Google Scholar] [CrossRef]
- Revell, L.J. Phytools: An R Package for Phylogenetic Comparative Biology (and Other Things). Methods Ecol. Evol. 2011, 3, 217–223. [Google Scholar] [CrossRef]
- Ho, L.; Si, T.; Ané, C. A Linear-Time Algorithm for Gaussian and Non-Gaussian Trait Evolution Models. Syst. Biol. 2014, 63, 397–408. [Google Scholar]
- Mendes, F.K.; Vanderpool, D.; Fulton, B.; Hahn, M.W. CAFE 5 Models Variation in Evolutionary Rates among Gene Families. Bioinformatics 2020, 36, 5516–5518. [Google Scholar] [CrossRef]
- Soanes, D.M.; Chakrabarti, A.; Paszkiewicz, K.H.; Dawe, A.L.; Talbot, N.J. Genome-Wide Transcriptional Profiling of Appressorium Development by the Rice Blast Fungus Magnaporthe oryzae. PLoS Pathog. 2012, 8, e1002514. [Google Scholar] [CrossRef]
- Nigg, M.; Laroche, J.; Landry, C.R.; Bernier, L. RNAseq Analysis Highlights Specific Transcriptome Signatures of Yeast and Mycelial Growth Phases in the Dutch Elm Disease Fungus Ophiostoma novo-ulmi. G3 Genes Genomes Genet. 2015, 5, 2487–2495. [Google Scholar] [CrossRef] [PubMed]
- Nigg, M.; Bernier, L. From Yeast to Hypha: Defining Transcriptomic Signatures of the Morphological Switch in the Dimorphic Fungal Pathogen Ophiostoma novo-ulmi. BMC Genom. 2016, 17, 920. [Google Scholar] [CrossRef] [PubMed]
- Nigg, M.; de Oliveira, T.C.; Sarmiento-Villamil, J.L.; de la Bastide, P.Y.; Hintz, W.E.; Sherif, S.M.; Shukla, M.; Bernier, L.; Saxena, P.K. Comparative Analysis of Transcriptomes of Ophiostoma novo-ulmi ssp. americana Colonizing Resistant or Sensitive Genotypes of American Elm. J. Fungi 2022, 8, 637. [Google Scholar] [CrossRef] [PubMed]
- Brasier, C.M. Intercontinental Spread and Continuing Evolution of the Dutch Elm Disease Pathogens. In The Elms: Breeding, Conservation, and Disease Management; Dunn, C.P., Ed.; Springer: Boston, MA, USA, 2000; pp. 61–72. ISBN 9781461545071. [Google Scholar]
- Bernier, L. Dutch Elm Disease. In Forest Microbiology; Asiegbu, F., Kovalchuk, A., Eds.; Elsevier: Amsterdam, The Netherlands, 2022; Volume 2, pp. 291–309. [Google Scholar]
- Martín, J.A.; Domínguez, J.; Solla, A.; Brasier, C.M.; Webber, J.F.; Santini, A.; Martínez-Arias, C.; Bernier, L.; Gil, L. Complexities Underlying the Breeding and Deployment of Dutch Elm Disease Resistant Elms. New For. 2021. [Google Scholar] [CrossRef]
- Paoletti, M.; Buck, K.W.; Brasier, C.M. Selective Acquisition of Novel Mating Type and Vegetative Incompatibility Genes via Interspecies Gene Transfer in the Globally Invading Eukaryote Ophiostoma novo-ulmi. Mol. Ecol. 2006, 15, 249–262. [Google Scholar] [CrossRef]
- Ahmadi, A.; Kavosi, M.R.; Soltanloo, H. Zelkova carpinifolia Reservoir from Hyrcanian Forests, Northern Iran, a New Sacrifice of Ophiostoma novo-ulmi. Biodiversitas J. Biol. Divers. 2014, 15, 48–52. [Google Scholar] [CrossRef]
- Khoshraftar, S.; Hung, S.; Khan, S.; Gong, Y.; Tyagi, V.; Parkinson, J.; Sain, M.; Moses, A.M.; Christendat, D. Sequencing and Annotation of the Ophiostoma ulmi Genome. BMC Genom. 2013, 14, 162. [Google Scholar] [CrossRef]
- Comeau, A.M.; Dufour, J.; Bouvet, G.F.; Jacobi, V.; Nigg, M.; Henrissat, B.; Laroche, J.; Levesque, R.C.; Bernier, L. Functional Annotation of the Ophiostoma novo-ulmi Genome: Insights into the Phytopathogenicity of the Fungal Agent of Dutch Elm Disease. Genome Biol. Evol. 2014, 7, 410–430. [Google Scholar] [CrossRef]
- Sarmiento-Villamil, J.L.; de Oliveira, T.C.; Naruzawa, E.S.; Bernier, L. An Efficient Strategy for Obtaining Mutants by Targeted Gene Deletion in Ophiostoma novo-ulmi. Front. Microbiol. 2021, 12, 699783. [Google Scholar] [CrossRef]
- Miyauchi, S.; Kiss, E.; Kuo, A.; Drula, E.; Kohler, A.; Sánchez-García, M.; Morin, E.; Andreopoulos, B.; Barry, K.W.; Bonito, G.; et al. Large-Scale Genome Sequencing of Mycorrhizal Fungi Provides Insights into the Early Evolution of Symbiotic Traits. Nat. Commun. 2020, 11, 5125. [Google Scholar] [CrossRef]
- Gan, P.; Ikeda, K.; Irieda, H.; Narusaka, M.; O’Connell, R.J.; Narusaka, Y.; Takano, Y.; Kubo, Y.; Shirasu, K. Comparative Genomic and Transcriptomic Analyses Reveal the Hemibiotrophic Stage Shift of Colletotrichum Fungi. New Phytol. 2013, 197, 1236–1249. [Google Scholar] [CrossRef] [PubMed]
- Ibarra Caballero, J.R.; Jeon, J.; Lee, Y.-H.; Fraedrich, S.; Klopfenstein, N.B.; Kim, M.-S.; Stewart, J.E. Genomic Comparisons of the Laurel Wilt Pathogen, Raffaelea lauricola, and Related Tree Pathogens Highlight an Arsenal of Pathogenicity Related Genes. Fungal Genet. Biol. 2019, 125, 84–92. [Google Scholar] [CrossRef] [PubMed]
- Fox, E.M.; Howlett, B.J. Secondary Metabolism: Regulation and Role in Fungal Biology. Curr. Opin. Microbiol. 2008, 11, 481–487. [Google Scholar] [CrossRef]
- Macheleidt, J.; Mattern, D.J.; Fischer, J.; Netzker, T.; Weber, J.; Schroeckh, V.; Valiante, V.; Brakhage, A.A. Regulation and Role of Fungal Secondary Metabolites. Annu. Rev. Genet. 2016, 50, 371–392. [Google Scholar] [CrossRef] [PubMed]
- Buser, C.C.; Newcomb, R.D.; Gaskett, A.C.; Goddard, M.R. Niche Construction Initiates the Evolution of Mutualistic Interactions. Ecol. Lett. 2014, 17, 1257–1264. [Google Scholar] [CrossRef] [PubMed]
- Mayer, V.E.; de Hoog, S.; Cristescu, S.M.; Vera, L.; Prenafeta-Boldú, F.X. Volatile Organic Compounds in the Azteca/Cecropia Ant-Plant Symbiosis and the Role of Black Fungi. J. Fungi 2021, 7, 836. [Google Scholar] [CrossRef]
- Lamping, E.; Baret, P.V.; Holmes, A.R.; Monk, B.C.; Goffeau, A.; Cannon, R.D. Fungal PDR Transporters: Phylogeny, Topology, Motifs and Function. Fungal Genet. Biol. 2010, 47, 127–142. [Google Scholar] [CrossRef]
- Kovalchuk, A.; Driessen, A.J.M. Phylogenetic Analysis of Fungal ABC Transporters. BMC Genom. 2010, 11, 177. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fijarczyk, A.; Bernier, L.; Sakalidis, M.L.; Medina-Mora, C.M.; Porth, I. Independent Evolution Has Led to Distinct Genomic Signatures in Dutch Elm Disease-Causing Fungi and Other Vascular Wilts-Causing Fungal Pathogens. J. Fungi 2023, 9, 2. https://doi.org/10.3390/jof9010002
Fijarczyk A, Bernier L, Sakalidis ML, Medina-Mora CM, Porth I. Independent Evolution Has Led to Distinct Genomic Signatures in Dutch Elm Disease-Causing Fungi and Other Vascular Wilts-Causing Fungal Pathogens. Journal of Fungi. 2023; 9(1):2. https://doi.org/10.3390/jof9010002
Chicago/Turabian StyleFijarczyk, Anna, Louis Bernier, Monique L. Sakalidis, Carmen M. Medina-Mora, and Ilga Porth. 2023. "Independent Evolution Has Led to Distinct Genomic Signatures in Dutch Elm Disease-Causing Fungi and Other Vascular Wilts-Causing Fungal Pathogens" Journal of Fungi 9, no. 1: 2. https://doi.org/10.3390/jof9010002
APA StyleFijarczyk, A., Bernier, L., Sakalidis, M. L., Medina-Mora, C. M., & Porth, I. (2023). Independent Evolution Has Led to Distinct Genomic Signatures in Dutch Elm Disease-Causing Fungi and Other Vascular Wilts-Causing Fungal Pathogens. Journal of Fungi, 9(1), 2. https://doi.org/10.3390/jof9010002





