Morphological and Phylogenetic Analyses Reveal Four New Species of Acrodictys (Acrodictyaceae) in China
Abstract
:1. Introduction
2. Materials and Methods
2.1. Isolation and Morphological Observation
2.2. DNA Extraction and Amplification
2.3. Phylogenetic Analyses
3. Results
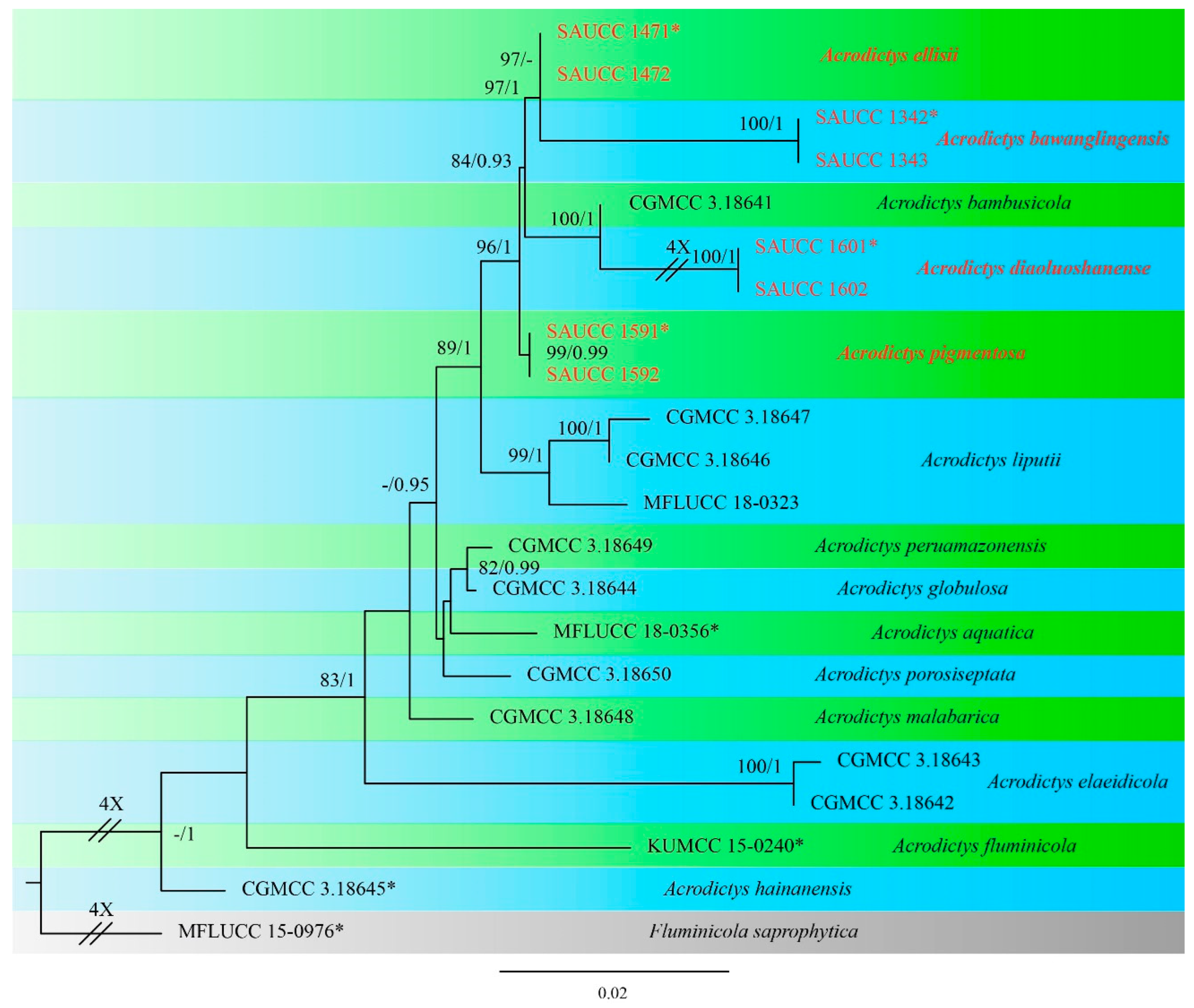
3.1. Phylogenetic Analyses
3.2. Taxonomy
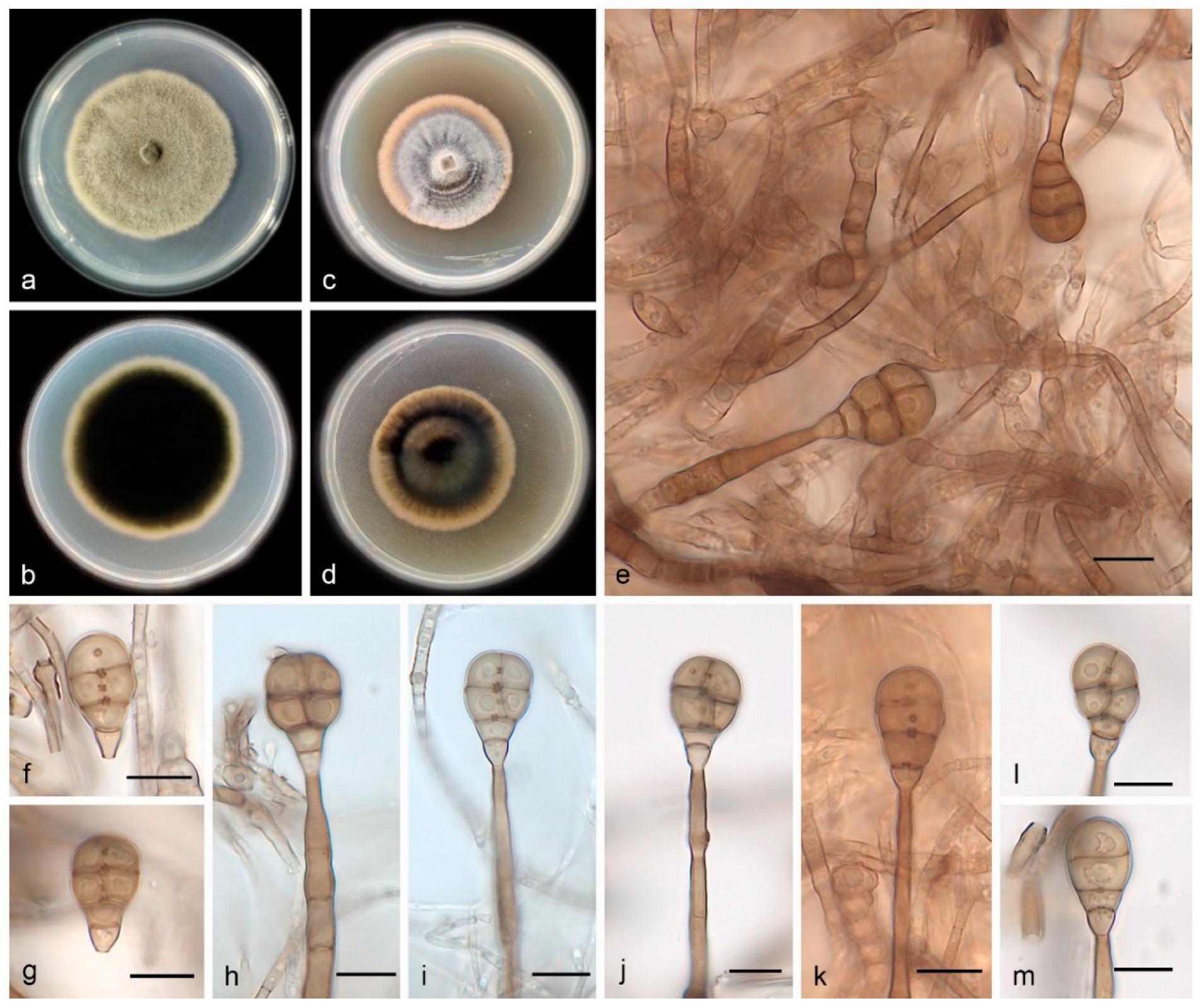
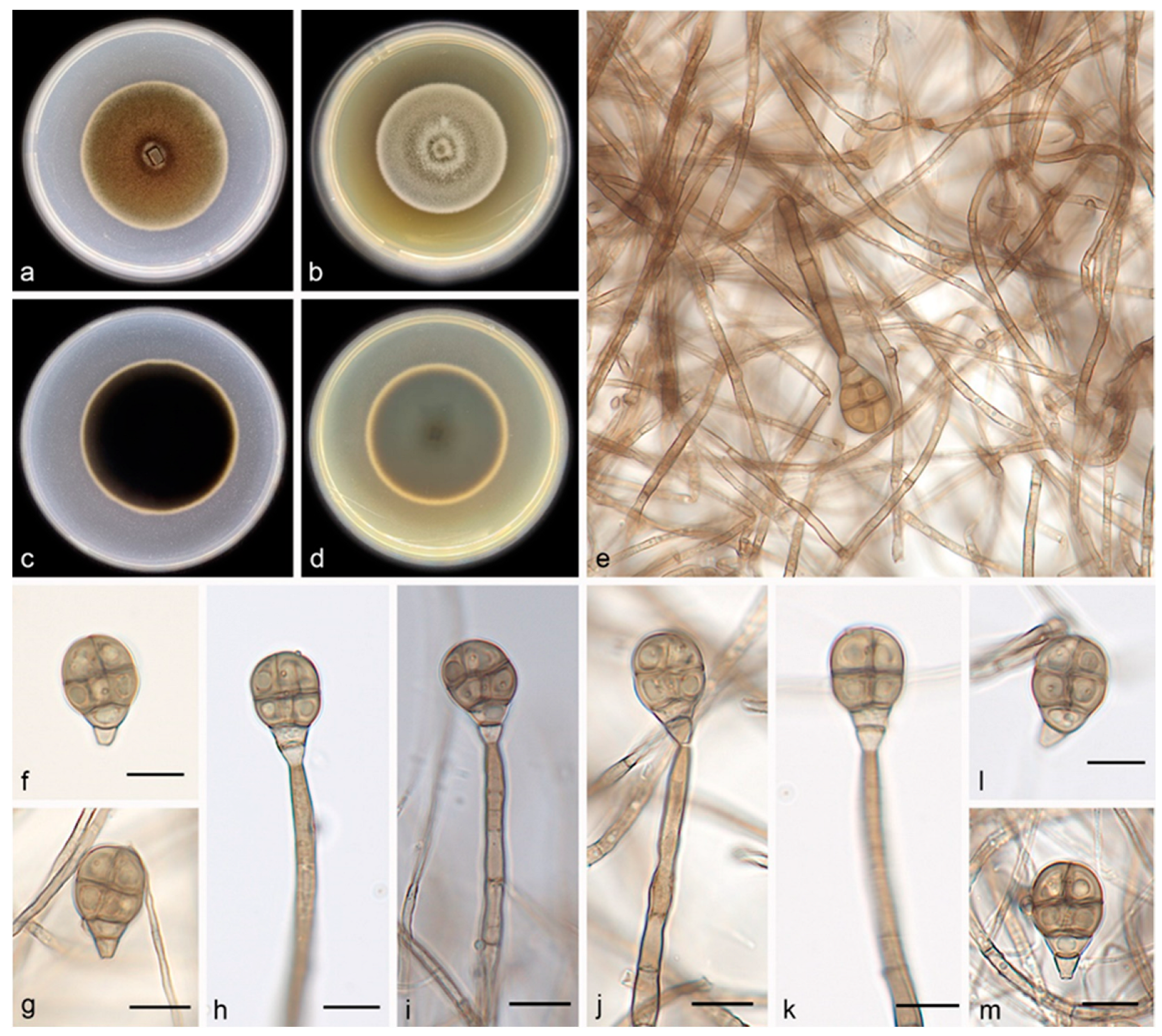
3.2.1. Acrodictys bawanglingensis S. Wang, J.W. Xia, X.Y. Liu, and X.G. Zhang, sp. nov.
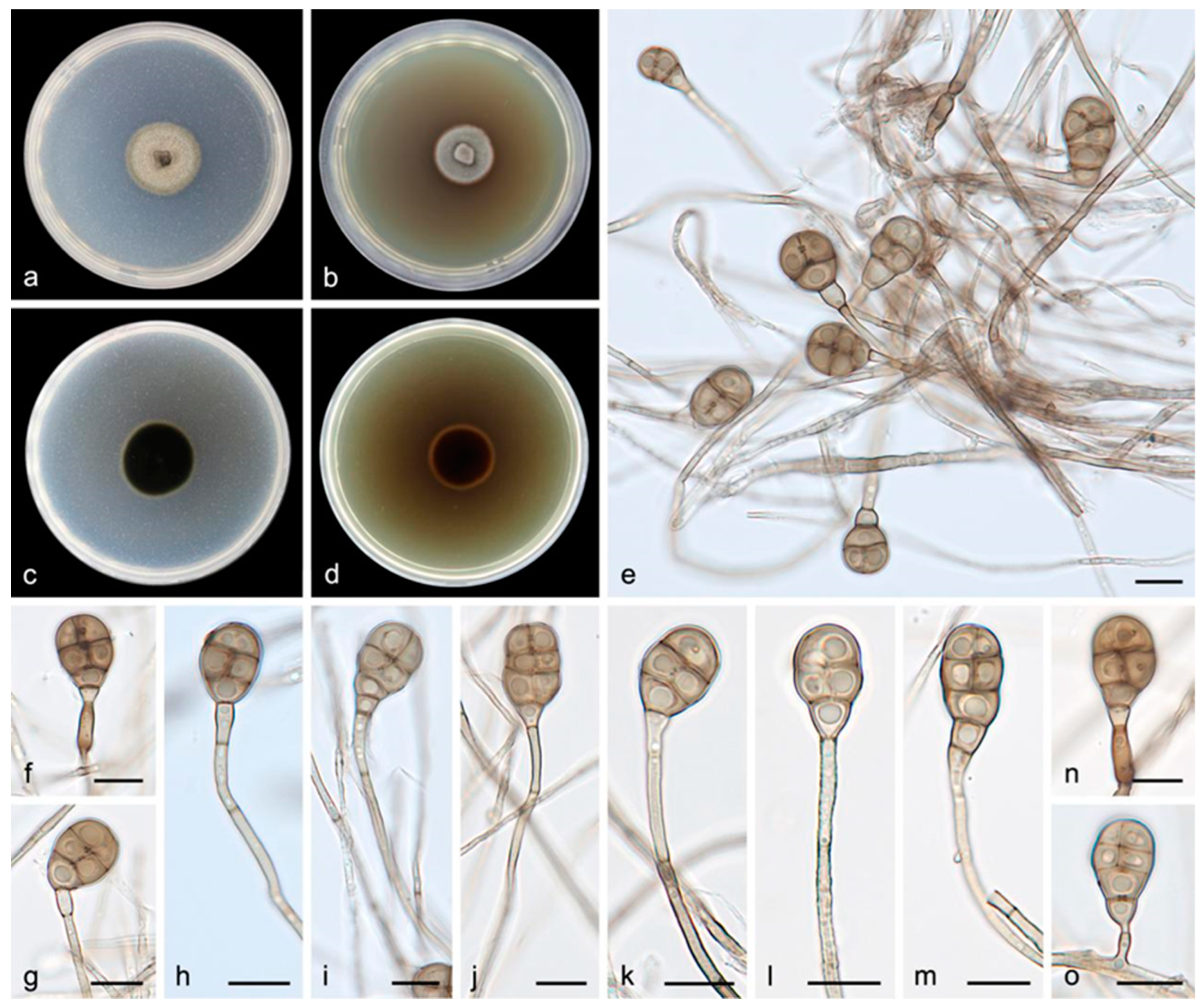
3.2.2. Acrodictys diaoluoshanensis S. Wang, J.W. Xia, X.Y. Liu, and X.G. Zhang, sp. nov.
3.2.3. Acrodictys ellisii S. Wang, J.W. Xia, X.Y. Liu, and X.G. Zhang, sp. nov.
3.2.4. Acrodictys pigmentosa S. Wang, J.W. Xia, X.Y. Liu, and X.G. Zhang, sp. nov.
3.3. Key to the Species of Acrodictys
| 1. Sexual morph known------------------------------------------------------------------------------------2 |
| 1’ Sexual morph unknown---------------------------------------------------------------------------------6 |
| 2. Maximum number of septa of ascospores > 3------------------------------------------------------3 |
| 2’ Maximum number of septa of ascospores ≤ 3------------------------------------------------------4 |
| 3. Conidia obovate---------------------------------------------------------------------------A. satwalekeri |
| 3’ Conidia pyriform---------------------------------------------------------------------------A. elaeidicola |
| 4. Conidia ellipsoid---------------------------------------------------------------------------------A. nigra |
| 4’ Conidia muriform-----------------------------------------------------------------------------------------5 |
| 5. Conidia size 23.0–34.0 × 18.0–22.0 μm----------------------------------------A. peruamazonensis |
| 5’ Conidia size 15.0–22.0 × 7.0–13.0 μm------------------------------------------------A. hainanensis |
| 6. Conidia with transverse septa only-------------------------------------------------------------------7 |
| 6’ Conidia with transverse and longitudinal septa-------------------------------------------------14 |
| 7. Conidia clavate--------------------------------------------------------------------------------------------8 |
| 7’ Conidia rounded-----------------------------------------------------------------------------------------10 |
| 8. Conidiogenous cells in groups--------------------------------------------------------------A. similis |
| 8’ Conidiogenous cells singly-----------------------------------------------------------------------------9 |
| 9. Conidiogenous cells size 70.0–100.0 × 4.0–6.0 μm------------------------------------A. aquatica |
| 9’ Conidiogenous cells size 98.0–142.0 × 4.0–6.0 μm---------------------------------A. fluminicola |
| 10. Conidiogenous cells branched---------------------------------------------------------A. caribensis |
| 10’ Conidiogenous cells unbranched------------------------------------------------------------------11 |
| 11. Conidiogenous cells septate-------------------------------------------------------------------------12 |
| 11’ Conidiogenous cells aseptate-----------------------------------------------------------------------13 |
| 12. Conidia spheroid---------------------------------------------------------------------------A. brooksiae |
| 12’ Conidia ellipsoidal--------------------------------------------------------------------------A. sacchari |
| 13. Conidia 2–3 transverse septa--------------------------------------------------------------A. elliptica |
| 13’ Conidia 4–9 transverse septa----------------------------------------------------------------A. liputii |
| 14. Conidiophores in groups-------------------------------------------------------------------A. furcata |
| 14’ Conidiophores singly---------------------------------------------------------------------------------15 |
| 15. Conidia maximum length > 100 μm--------------------------------------------A. septosporioides |
| 15’ Conidia maximum length < 100 μm---------------------------------------------------------------16 |
| 16. Conidiogenous cells lageniform--------------------------------------------------------------------17 |
| 16’ Conidiogenous cells cylindrical--------------------------------------------------------------------22 |
| 17. Conidiogenous cells determinate proliferations----------------------------------A. balladynae |
| 17’ Conidiogenous cells percurrent proliferations-------------------------------------------------18 |
| 18. Conidia subglobose or ellipsoidal-----------------------------------------------------------------19 |
| 18’ Conidia clavate or pyriform-------------------------------------------------------------------------20 |
| 19. Conidia size 12.0–22.0 × 8.0–16.0 μm------------------------------------------------A. irregularis |
| 19’ Conidia size 27.0–32.0 × 12.0–16.0 μm--------------------------------------------------A. oblonga |
| 20. Conidiogenous cells maximum length > 200 μm ------------------------------A. porosiseptata |
| 20’ Conidiogenous cells maximum length < 200 μm ----------------------------------------------21 |
| 21. Conidia size 20.0–30.0 × 13.0–16.0 μm---------------------------------------------A. bambusicola |
| 21’ Conidia size 17.0–27.0 × 11.0–15.0 μm----------------------------------------------A. atroapicula |
| 22. Conidiogenous cells with percurrent proliferations-------------------------------------------23 |
| 22’ Conidiogenous cells with determinate proliferations-----------------------------------------25 |
| 23. Conidiogenous cells maximum length > 60 μm------------------------------------A. micheliae |
| 23’ Conidiogenous cells maximum length < 60 μm------------------------------------------------24 |
| 24. Conidia size 28.0–32.0 × 8.0–12.0 μm---------------------------------------------------A. lignicola |
| 24’ Conidia size 16.0–20.0 × 12.0–15.0 μm-------------------------------------------------A. papillata |
| 25. Conidiogenous cells maximum length > 80 μm------------------------------------------------26 |
| 25’ Conidiogenous cells maximum length < 80 μm------------------------------------------------27 |
| 26. Conidia size 18.0–26.0 × 10.0–16.0 μm, exceeding 23 and 13 um in length and width, respectively----------------------------------------------------------------------------A. bawanglingensis |
| 26’ Conidia size 18.0–22.0 × 10.0–13.0 μm----------------------------------------------------A. ellisii |
| 27. Conidia size 18.0–22.0 × 10.0–13.0 μm------------------------------------A. diaoluoshanensis |
| 27’ Conidia size 12.0–24.0 × 7.0–12.0 μm----------------------------------------------A. pigmentosa |
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ellis, M.B. Dematiaceous Hyphomycetes II. Mycol. Pap. 1961, 79, 1–23. [Google Scholar]
- Baker, W.A.; Partridge, E.C.; Morgan-Jones, G. Notes on hyphomycetes LXXXI. Acrodictyella obovata, a new lignicolous, dematiaceous genus and species collected in Alabama. Mycotaxon 2001, 78, 29–35. [Google Scholar]
- Baker, W.A.; Partridge, E.C.; Morgan-Jones, G. Notes on hyphomycetes LXXXV. Junewangia, a genus in which to classify four Acrodictys species and a new taxon. Mycotaxon 2002, 81, 293–319. [Google Scholar]
- Baker, W.A.; Morgan-Jones, G. Notes on hyphomycetes. XCI. Pseudoacrodictys, a novel genus for seven taxa formerly placed in Acrodictys. Mycotaxon 2003, 85, 371–391. [Google Scholar]
- Baker, W.A.; Partridge, E.C.; Morgan-Jones, G. Notes on hyphomycetes LXXXVII. Rhexoacrodictys, a new segregate genus to accommodate four species previously classified in Acrodictys. Mycotaxon 2002, 82, 95–113. [Google Scholar]
- Zhao, G.Z.; Cao, A.X.; Zhang, T.Y.; Liu, X.Z. Acrodictys (Hyphomycetes) and related genera from China. Mycol. Prog. 2011, 10, 67–83. [Google Scholar] [CrossRef]
- Xia, J.W.; Ma, Y.R.; Li, Z.; Zhang, X.G. Acrodictys-like wood decay fungi from southern china, with two new families Acrodictyaceae and Junewangiaceae. Sci. Rep. 2017, 7, 7888. [Google Scholar] [CrossRef]
- Guo, L.D.; Hyde, K.D.; Liew, E.C.Y. Identification of endophytic fungi from Livistona chinensis based on morphology and rDNA sequences. New Phytol. 2000, 147, 617–630. [Google Scholar] [CrossRef]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for Bigger Datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef]
- Luo, Z.L.; Hyde, K.D.; Liu, J.K.; Maharachchikumbura, S.S.N.; Jeewon, R.; Bao, D.F.; Bhat, D.J.; Lin, C.G.; Li, W.L.; Yang, J.; et al. Freshwater Sordariomycetes. Fungal Divers. 2019, 99, 451–660. [Google Scholar] [CrossRef]
- Katoh, K.; Rozewicki, J.; Yamada, K.D. MAFFT online service: Multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 2017, 20, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Nylander, J.A.A. MrModelTest v. 2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Miller, M.A.; Pfeiffer, W.; Schwartz, T. The CIPRES science gateway: Enabling high-impact science for phylogenetics researchers with limited resources. In Proceedings of the 1st Conference of the Extreme Science and Engineering Discovery Environment. Bridging from the Extreme to the Campus and Beyond, Chicago, IL, USA, 16 July 2012; Association for Computing Machinery: San Diego, CA, USA, 2012; p. 8. [Google Scholar] [CrossRef]
- Stamatakis, A. RAxML Version 8: A tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 2014, 30, 1312–1313. [Google Scholar] [CrossRef] [PubMed]
- Huelsenbeck, J.P.; Ronquist, F. MRBAYES: Bayesian inference of phylogeny. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Huelsenbeck, J.P. MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 2003, 19, 1572–1574. [Google Scholar] [CrossRef] [PubMed]
- Ronquist, F.; Teslenko, M.; Van, D.M.P.; Ayres, D.L.; Darling, A.; Höhna, S.; Larget, B.; Liu, L.; Suchard, M.A.; Huelsenbeck, J.P. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [PubMed]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, F.J.R.M.; Lee, S.H.; Taylor, L.; Shawe-Taylor, J. Amplification and direct sequencing of fungal ribosomal rna genes for phylogenetics. In PCR Protocols: A Guide to Methods and Applications; Innis, M.A., Gelfand, D.H., Sninsky, J.J., Eds.; Academic Press Inc.: New York, NY, USA, 1990; pp. 315–322. [Google Scholar] [CrossRef]
- Rehner, S.A.; Samuels, G.J. Taxonomy and phylogeny of Gliocladium analysed from nuclear large subunit ribosomal DNA sequences. Mycol. Res. 1994, 98, 625–634. [Google Scholar] [CrossRef]
- Vilgalys, R.; Hester, M. Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 1990, 172, 4238–4246. [Google Scholar] [CrossRef]
- Glass, N.L.; Donaldson, G.C. Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 1995, 61, 1323–1330. [Google Scholar] [CrossRef]
- Liu, Y.J.; Whelen, S.; Hall, B.D. Phylogenetic Relationships among Ascomycetes: Evidence from an RNA polymerse II subunit. Mol. Biol. Evol. 1999, 16, 1799–1808. [Google Scholar] [CrossRef]
- Wu, W.P.; Zhuang, W. Sporidesmium, Endophragmiella and Related Genera from China; Fungal Diversity Research Series; Fungal Diversity Press: Chiang Rai, Thailand, 2005; Volume 15, p. 351. [Google Scholar]
- Jiang, N.; Voglmayr, H.; Bian, D.R.; Piao, C.G.; Wang, S.K.; Li, Y. Morphology and Phylogeny of Gnomoniopsis (Gnomoniaceae, Diaporthales) from Fagaceae Leaves in China. J. Fungi. 2021, 7, 792. [Google Scholar] [CrossRef]
- Xia, J.W.; Mu, T.C.; Zhang, Z.X.; Li, Z.; Zhang, X.G. Neoacrodictys elegans gen. & sp. nov. from Hainan Province, China. Mycotaxon 2022, 137, 63–71. [Google Scholar] [CrossRef]
- Zhang, Z.X.; Liu, R.Y.; Liu, S.B.; Mu, T.C.; Zhang, X.G.; Xia, J.W. Morphological and phylogenetic analyses reveal two new species of Sporocadaceae from Hainan, China. MycoKeys 2022, 88, 171–192. [Google Scholar] [CrossRef] [PubMed]


| Locus | Primer | Sequence (5′–3′) | PCR Program | References |
|---|---|---|---|---|
| ITS | ITS5 ITS4 | GGA AGT AAA AGT CGT AAC AAG G TCC TCC GCT TAT TGA TAT GC | (94 °C for 30 s, 55 °C for 30 s, 72 °C for 45 s) × 29 cycles | [18] |
| LSU | LR0R LR5 | GTA CCC GCT GAA CTT AAG C TCC TGA GGG AAA CTT CG | (94 °C for 30 s, 48 °C for 50 s, 72 °C for 90 s) × 35 cycles | [19,20] |
| SSU | NS1 NS4 | GTA GTC ATA TGC TTG TCT C CTT CCG TCA ATT CCT TTA AG | (94 °C for 30 s, 50 °C for 45 s, 72 °C for 90 s) × 35 cycles | [18] |
| tub2 | Bt-2a Bt-2b | GGT AAC CAA ATC GGT GCT GCT TTC ACC CTC AGT GTA GTG ACC CTT GGC | (95 °C for 30 s, 56 °C for 30 s, 72 °C for 60 s) × 35 cycles | [21] |
| rpb2 | fRPB2-5F fRPB2-7R | GGG GWG AYC AGA AGA AGG C CCC ATR GCT TGY TTR CCC AT | (95 °C for 50 s, 56 °C for 50 s, 72 °C for 120 s) × 37 cycles | [22] |
| Species | Strain | GenBank Accession Number | ||||
|---|---|---|---|---|---|---|
| ITS | LSU | SSU | rpb2 | tub2 | ||
| Acrodictys aquatica | MFLUCC 18-0356 * | MG835711 | MG835712 | – | – | – |
| A. bambusicola | CGMCC 3.18641 | KU999973 | KX033564 | KX033535 | – | KX036219 |
| A. bawanglingensis | SAUCC 1342 * | ON606324 | ON614219 | ON620164 | ON859853 | ON859845 |
| SAUCC 1343 | ON606325 | ON614220 | ON620165 | ON859852 | ON859844 | |
| A. diaoluoshanensis | SAUCC 1601 * | ON645265 | ON644407 | ON645269 | ON859847 | ON859839 |
| SAUCC 1602 | ON645266 | ON644408 | ON645270 | ON859846 | ON859838 | |
| A. ellisii | SAUCC 1471 * | ON645254 | ON644405 | ON645267 | ON859851 | ON859843 |
| SAUCC 1472 | ON645255 | ON644406 | ON645268 | ON859850 | ON859842 | |
| A. elaeidicola | CGMCC 3.18642 | KU999977 | KX033568 | KX033539 | – | – |
| CGMCC 3.18643 | KU999978 | KX033569 | KX033540 | – | – | |
| A. fluminicola | KUMCC 15-0240 * | MK828642 | MK849786 | – | – | – |
| A. globulosa | CGMCC 3.18644 | KU999970 | KX033562 | KX033532 | – | – |
| A. hainanensis | CGMCC 3.18645 * | KU999974 | KX033565 | KX033536 | – | – |
| A. liputii | CGMCC 3.18646 | KU999966 | KX033558 | KX033528 | – | – |
| CGMCC 3.18647 | KU999979 | KX033570 | KX033541 | – | – | |
| MFLUCC 18-0323 | MZ412512 | MZ412524 | MZ413269 | – | – | |
| A. malabarica | CGMCC 3.18648 | KU999968 | KX033560 | KX033530 | – | – |
| A. peruamazonensis | CGMCC 3.18649 | KU999969 | KX033561 | KX033531 | – | – |
| A. pigmentosum | SAUCC 1591 * | ON606326 | ON614221 | ON620166 | ON859849 | ON859841 |
| SAUCC 1592 | ON606327 | ON614222 | ON620167 | ON859850 | ON859842 | |
| A. porosiseptata | CGMCC 3.18650 | KU999967 | KX033559 | KX033529 | – | KX036220 |
| Fluminicola saprophytica | MFLUCC 15-0976 * | MF374358 | MF374367 | MF374375 | MF370954 | – |
| Species | Shape | Conidial Size (μm) | Septa |
|---|---|---|---|
| Acrodictys aquatica | Clavate | 20.0–27.0 × 10.0–17.0 | 3–4 transverse septa |
| A. bambusicola | Broadly clavate or pyriform | 17.0–36.0 × 12.0–18.0 | 2–5 transverse and 1 or more longitudinal septa |
| A. bawanglingensis | Obovoid to obpyriform | 18.0–26.0 × 10.0–16.0 | 3 transverse and 1–3 longitudinal septa |
| A. diaoluoshanensis | Obovoid to obpyriform | 18.0–22.0 × 10.0–13.0 | 3 transverse and 1–2 longitudinal septa |
| A. ellisii | Obovoid to obpyriform | 17.0–22.0 × 11.0–14.0 | 3 transverse and 1–3 longitudinal septa |
| A. elaeidicola | Turbinate, pyriform or clavate | 17.0–26.0 × 11.0–19.0 | 3 transverse and 1–3 longitudinal septa |
| A. fluminicola | Broadly clavate, obovoid to pyriform | 24.0–30.0 × 13.0–17.0 | 2–3 transverse and a few longitudinal septa |
| A. globulosa | Subglobose | 22.0–27.0 × 17.0–23.0 | 2 transverse and several longitudinal and oblique septa |
| A. hainanensis | Oblong to obovoid | 15.0–22.0 × 7.0–13.0 | 3–5 transverse and several longitudinal or oblique septa |
| A. liputii | Subglobose | 18.5–22.5 × 13.5–17.5 | 2–3 parallel transverse and 2 perpendicular longitudinal septa |
| A. malabarica | Gangliar, somewhat top-shaped | 16.0–21.0 × 14.0–17.0 | 3 transverse and 2–4 longitudinal septa |
| A. peruamazonensis | Ellipsoidal | 28.0–36.0 × 17.0–21.0 | 3–4 transverse and some longitudinal or oblique septa |
| A. pigmentosa | Obovoid to obpyriform | 12.0–24.0 × 7.0–12.0 | 1–4 transverse septa and 1–3 longitudinal septa |
| A. porosiseptata | Broadly clavate to pyriform | 25.0–30.0 × 13.5–16.5 | 4–5 transverse and 3 perpendicular longitudinal septa |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wang, S.; Liu, R.; Liu, S.; Zhang, Z.; Xia, J.; Li, D.; Liu, X.; Zhang, X. Morphological and Phylogenetic Analyses Reveal Four New Species of Acrodictys (Acrodictyaceae) in China. J. Fungi 2022, 8, 853. https://doi.org/10.3390/jof8080853
Wang S, Liu R, Liu S, Zhang Z, Xia J, Li D, Liu X, Zhang X. Morphological and Phylogenetic Analyses Reveal Four New Species of Acrodictys (Acrodictyaceae) in China. Journal of Fungi. 2022; 8(8):853. https://doi.org/10.3390/jof8080853
Chicago/Turabian StyleWang, Shi, Rongyu Liu, Shubin Liu, Zhaoxue Zhang, Jiwen Xia, Duhua Li, Xiaoyong Liu, and Xiuguo Zhang. 2022. "Morphological and Phylogenetic Analyses Reveal Four New Species of Acrodictys (Acrodictyaceae) in China" Journal of Fungi 8, no. 8: 853. https://doi.org/10.3390/jof8080853
APA StyleWang, S., Liu, R., Liu, S., Zhang, Z., Xia, J., Li, D., Liu, X., & Zhang, X. (2022). Morphological and Phylogenetic Analyses Reveal Four New Species of Acrodictys (Acrodictyaceae) in China. Journal of Fungi, 8(8), 853. https://doi.org/10.3390/jof8080853








