1. Introduction
Fusarium head blight (FHB) incited by
Fusarium graminearum (FG) fungus is ranked as the one of the prime annihilating fungal diseases of wheat (
Triticum aestivum L.) globally [
1]. The published literature indicated that this disease drastically reduces the crop yield, leading to huge economic losses [
2,
3,
4]. It has been noticed that yield losses are primarily linked with poor quality seed production. The contamination of infected seed grains with mycotoxins has been observed [
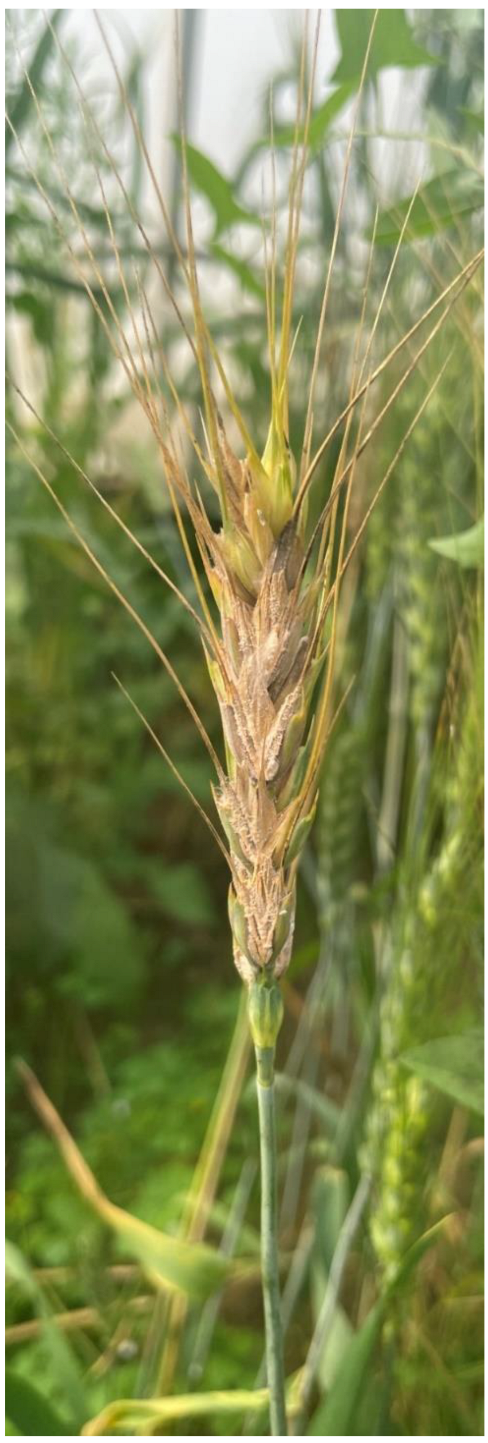
5]. In India, the typical symptoms produced by the fungus appear majorly on the glumes and rachis of wheat plants in the form of water-soaked lesions. Later on, the fungus spreads within the wheat ear heads, resulting in the partial bleaching to complete blighting of attacked ear heads (
Figure 1). Historically, the disease was first noticed in India in the Siang District of Arunachal Pradesh in the year 1974 [
6]. Later on, the disease has been reported by other workers from Wellington [
7] and Gurdaspur (Punjab, India) [
8]. Bagga et al. [
9] documented that the disease heavily attacked the wheat cultivar PDW 274 in Dera Baba Nanak region of Gurdaspur in Punjab district of India and resulted in noteworthy yield reduction. In addition to India, chronic appearance of the disease has been observed in different corners of the world and major regions including China, Brazil, USA, Canada, the former USSR, Eastern and Western Europe, Romania, etc. which account for more than 50% of global production [
10,
11]. In recent reports, it has been cautioned that
Fusarium head blight is liable to enhance under reduced tillage-based wheat cultivation and further aggravated with climate shift especially in the northern part of India, which is recognized as the main wheat basket of India [
4].
FG showed a broad host range, and it has the ability to infect different plants such as maize, sorghum, millets, rye, triticale, oats, etc. [
12]. Various research evidence indicated that FHB disease is highly prone to humid to semi-humid areas of the world, especially where heavy and frequent rainfall with a high level of moisture exists in the atmosphere throughout the wheat cultivation season [
13,
14]. Unfortunately, such type of weather is prevalent in the northern part of India, especially during and after the anthesis stage of wheat, which directly affects the crop yield [
4]. It has been observed that the fungus in the off-season survive in infected wheat straw in different grasses of the wheat family origin and in crop residues that remain in the soil after wheat harvest [
15,
16]. The incidence and severity of the FHB disease is determined by numerous factors such as quantity of airborne inocula (both inside and outside of the field), and the prevalence of humidity during and after the anthesis period [
14,
17]. Currently, fungicide (e.g., Tebuconazole, Triazoles, Prochloraz, etc.) sprays are important methods to preclude and conquer FHB disease in a short time and lessen
Fusarium toxins production, despite agro-ecological and resistance development problems [
2,
18,
19,
20]. Hence, the deployment of FHB-resistant varieties is a sustainable, cost-effective, and environmentally friendly approach to FHB management. Unfortunately, the majority of the popular varieties cultivated in India are prone to FHB disease [
4]. Moreover, the effectiveness of resistant varieties and implementation of different methods singly or in an integrated manner for the control of
Fusarium spp. still exist as a daunting task because of the complex nature of FG and fungicide-resistant development in natural FG populations [
21]. Therefore, an appraisal of genetic diversity and population structure of FG becomes mandatory to understand its evolutionary relationships with respect to environmental change, selection pressures, and other forces (e.g., mutation, genetic drift, gene flow, etc.) linked with evolutionary change [
22,
23]. Different research groups also advocated the essentiality of genetic structure analysis to decode the modes of recombination and distribution of isolates within and among populations [
24,
25]. Most importantly, FHB-resistant genotypes could be judiciously deployed if the pathogen population is known. At present, limited information is available regarding the distribution, genetic diversity and population structure of the FG population in India.
Molecular marker-driven technologies play a significant role in species identification because of their potential usage in exploring the population structure and genetic diversity within the fungal species and their isolates [
26,
27,
28,
29,
30,
31,
32,
33,
34]. It is worth noting that high-throughput sequencing of amplicon markers from conserved genomic regions has provided new opportunities to decipher
Fusarium diversity in agricultural crops in recent times. The molecular identification of fungi is primarily based on the internal transcribed spacer region (ITS) [
35,
36]. However, it has been observed that a number of species of
Fusarium genus comprise orthologous regions and as a consequence, ITS region-based identification resulted in erratic and unreliable results. In this context, genomic regions representing translocation elongation factor 1-α (TEF), β-tubulin (TUB), histones (HIS) and calmodulin (CAL) have been widely used by different group of researchers to distinguish
Fusarium spp. [
33,
37,
38,
39,
40,
41,
42,
43,
44,
45,
46]. Yli-Mattila et al. [
47] demonstrated the application if ITS, IGS, mtSSU, and TUB genomic region comparisons predicting the variation of
Fusarium spp. Similarly, TEF-1α, β-tubulin and histone 3 regions have been explored by Webb et al. [
48] and Taha et al. [
49] for dissecting the genetic variability among
F. oxysporum isolates. Recently, Fulton et al. [
50] employed two different housekeeping genes (BT and TEF) for analyzing the phylogenetic kinship of
F. oxysporum f. sp.
niveum isolates from the major watermelon-producing regions in north, central, and south Florida. A flooding of reports are available that highlight the significance of TUB, TEF and HIS in resolving boundaries of fungal species and further revealed species identification and discrimination based on the combined sequences of gene loci as a more valuable genomic resource than single gene loci [
51,
52]. Unfortunately, not a single report regarding the dissection of genetic variation and population structure of FG in the major wheat-growing belt of North India using combined housekeeping gene sequences is available.
The history of FG in wheat in India is not too old, and therefore, the northern plains, a major wheat-growing belt of India, offers a paragon site to pinpoint the prime evolutionary mechanism acting on a newly developing FG isolate as it dispersed topically and regionally within the Indo-gangetic plains of India. Therefore, the current research was planned with an intention to obtain the answers to the question of whether FG contains distinct evolutionary lineages in the northern plains of India. The other questions include: (i) Is the phylogenetic and evolutionary structure of FG indicative of recombination or mutation?; and (ii) Is there a biogeographic relationship with the evolutionary structure of FG? The answers to these questions are essential to improve the understanding of the ecology of the plant pathogen and devise a better management strategy for the management of FHB disease in wheat.
4. Discussion
FG is one of the emerging and economically important diseases affecting wheat in India [
4,
65]. Diversity analysis is essential to infer the population genetics of such an important disease for framing cost-effective management tactics. In this context, a comprehensive understanding of genetic variation of the Indian FG population becomes necessity. Conserved region-based DNA markers are reported as one of the potential tools for determining the genetic variation, speciation of fungal pathogens and inferring their ancestral background [
35,
66]. FG diversity analysis based on TEF, HIS and TUB genes has discovered the existence of different putative subspecies of
Fusarium in the Asian and sub-Saharan Africa terrains [
67]. As Indian FG populations have not been analyzed so far to determine their diversity and population structure, therefore, the current investigation presented for the first time effort to quantify genetic variability in FG in northern plains of India using three highly versatile molecular markers, i.e., TUB, TEF and HIS. In this study, attempts have been made to (i) determine the identity of FG infecting wheat in the northern plains of India; (ii) confirm the level of DNA polymorphism of TUB, TEF and HIS sequences; (iii) decode the haplotype diversity and their distribution; and (iv) work out the phylogenetic kinship between FG isolates of Indian origin.
Previous published literature indicated that TEF is highly recommended for species delineation and rejected ITS sequences in offering superior taxa resolution for several of the fungal genera (e.g.,
Trichoderma and
Fusarium) of the ascomycetes group [
31,
40]. From earlier published reports, it is cleared that when the distance between different
Fusarium spp. to the nearest neighbor fungal species was deduced on the basis of ITS regions, very low genetic distance values were obtained, which ultimately resulted in the inferior resolution and poor taxon placement in phylogenetic lineages [
68,
69]. It is important to mention that slow evolving genes, for instance, TEF and HIS, serve excellent options in inferring phylogeny-based relationships. Contrarily, more recent evolutionary and speciation events can be captured from the gene sequences (e.g., TUB) displaying high evolutionary rates. Therefore, all the three genes were satisfying all the basic requirements needed for the phylogenetic analysis, as the concatenated sequences were composed of both variable introns and conserved exons [
70] and hence selected for the present research investigation. According to the results of the current study and others [
40], it is crucial to ascertain the DNA polymorphism in the sequences to be employed for the identification of intraspecies kinships and inferring phylogenetic lineages because of their direct influence on the correct and precise identity with strong phylogenetic arrangement and signal strength. Additionally, its influence on genetic diversity indices such as haplotype diversity and their dispersion have been reported [
40]. The low level of genetic diversity noticed for wheat-associated FG populations in the northern plains of India based on concatenated gene sequences is not a rare event, as similar observations regarding the existence of a low level of phylogenetic diversity for FG populations in wheat has been documented by Castellá and Cabañes [
71].
The significance of evolutionary forces on the FG population has been studied by conducting neutrality tests, which provides evidence regarding the divergence of combined gene loci sequence data from neutrality statistics. Moreover, the significant positive Tajima’s D values highlighted that all the three gene loci are experiencing population bottlenecks in PB, HR and RJ, where the FG population appears to be uniform and only a few sequences are in a deciding role for the development of the nascent population. This indicates that FG biology and the colonization pattern could act as one important factor behind population bottlenecks. Contrarily, significant negative values of Tajima’s D and F statistics in case of the WB population showed a strong purifying selection. This observation has been matched with the earlier report of Zhao et al. [
72], where strong purifying selection in accordance with their important biological roles in cellular processes in fungi has been noticed.
Geography and climate are two of the most important epidemic linked factors and strongly influence the prevalence and establishment of
Fusarium spp. as pathogens in various types of plants [
71,
73]. Usually, in nature, a mixture of both old and new haplotypes in the form of living descendants in field populations exists. However, the relationship between the two types of haplotypes may be reticulate and non-bifurcated because of non-dichotomous historical events, etc. [
74]. Similarly, in the present study, a minimum of recombination events based on the combined three gene loci has been observed in the FG populations. This clearly reflects the mechanism of intragenic recombination behind the genetic variability in the pathogen population in the regions. However, it is important to mention here that Ma et al. [
75] also confirmed the significance of horizontal gene transfer in the evolution of
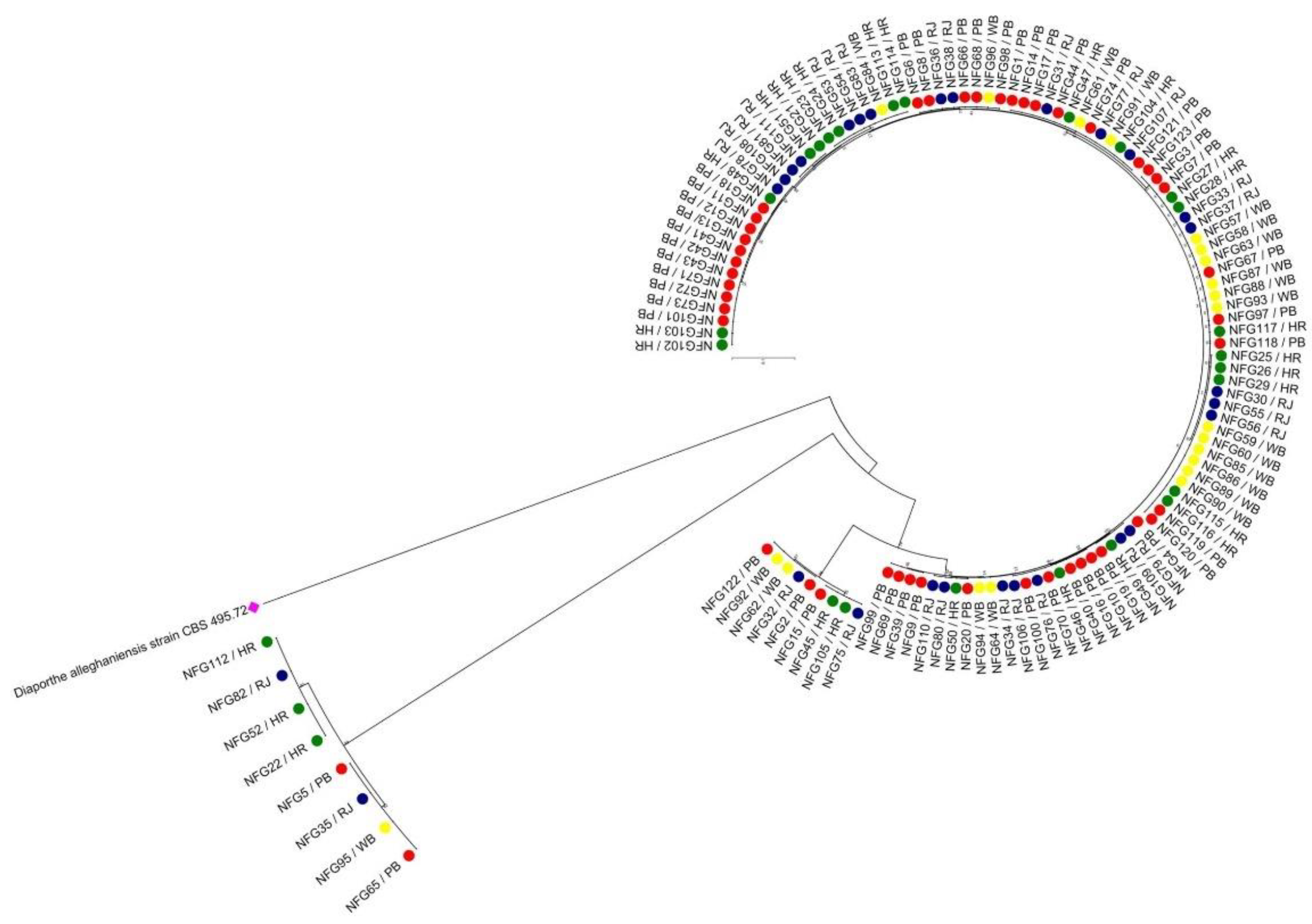
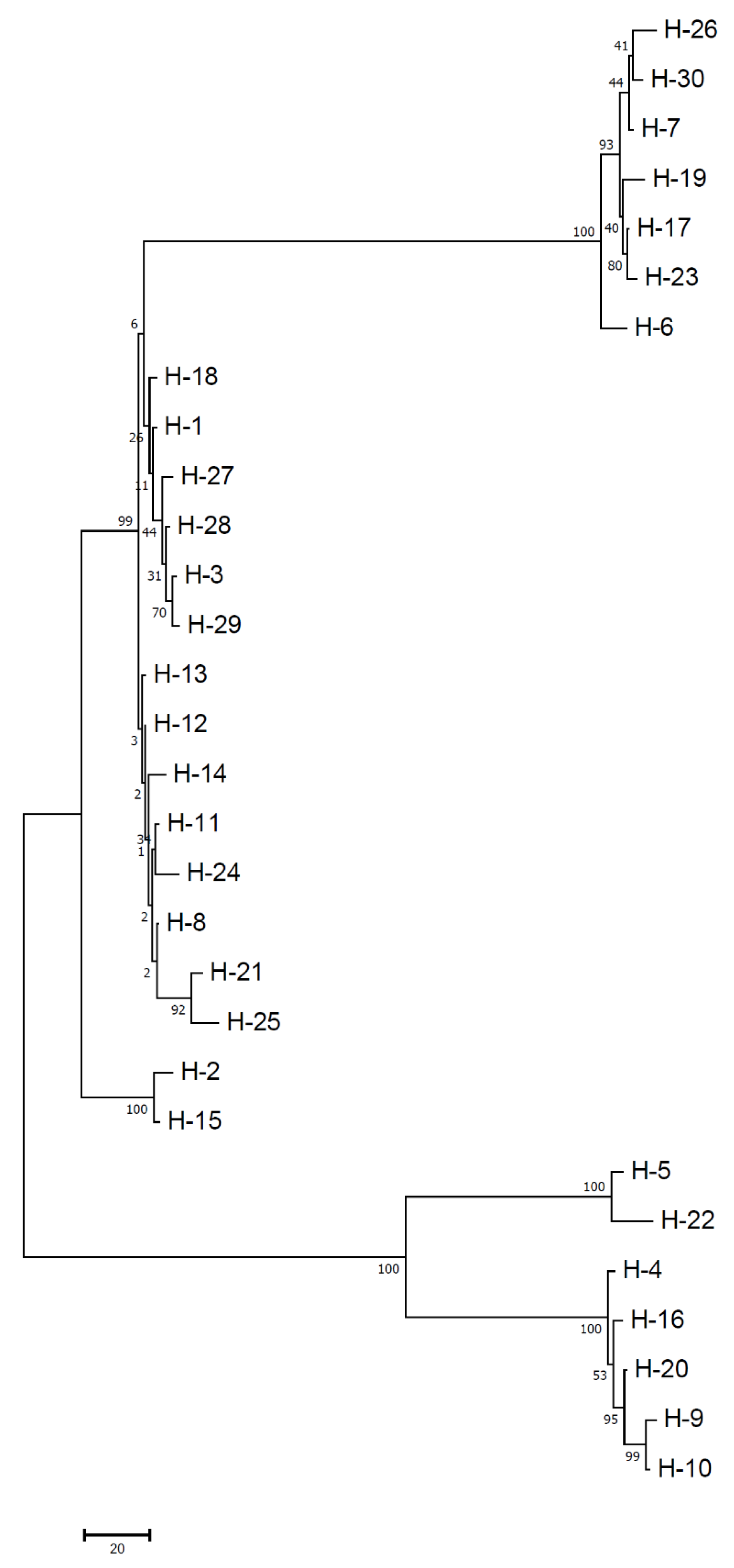
Fusarium genomes. Thus, high-frequency haplotypes (H1, H2 and H3) have been present in the population for a long time and detected from PB, RJ and WB population dominantly. Contrarily, other seem to be less frequent haplotypes and indicate toward more recent mutation events. Moreover, the terminal haplotypes (e.g., H2, H3, H5, H13–H15, H21, H22, H24–H26, H29, and H30) located at the network tips in the present study indicate a holocene pedigree rather than old ancestry (H1, H18 and H28) as placed in the interiors of the haplotype network. Carbone and Kohn [
76] reported that newer haplotypes generally show restricted geographical dispersion in comparison to ancestral haplotypes, which have a wide geographical distribution and show restricted gene flow. However, the current study indicated toward a reticulate network, where both ancestral and nascent haplotypes occupied an internal celestial point of the network. The pattern of haplotype network and haplotype distribution in the studied sites of the northern plains illustrate that the FG isolates were separated from other regional isolates by a series of mutational events rather than geographical location (e.g., PB, HR, RJ and WB). These points indicate toward the possibility of the operation of both direct and indirect factors in the distribution of FG fungus, which is seed-, soil- and air-borne in nature and further supported by the haplotype network. The possibility of human-driven transfer of fungi in contaminated plant material and the transfer of FG inocula belong to other regions such as PB (where the maximum number of haplotypes was observed) or vice versa, which is followed by host adaptation and panoramic spatiotemporal dispersion. It has been observed that a number of haplotypes displayed only a difference at one site in the present study when compared with their genetically closest haplotype. These findings clearly reflect the significance of mutation in developing haplotype diversity as observed by earlier researchers in case of other fungal crop plant pathogens [
30,
77,
78].
Haplotype analysis provides important information on the existence of different types of haplotypes (h), their diversity (Hd) and frequency. Usually, Hd values varied between 0 and 1, which reflects zero to high level of haplotype diversity, respectively [
79]. In the current investigation, Hd (0.962–0.981) values revealed high levels of genetic variability in all the states. PB reflected maximum diversity (i.e., 27 haplotypes from 47 FG isolates) followed by RJ (i.e., 21 haplotypes from 28 FG isolates). It is important to mention that several haplotypes have been shared among different populations that clearly hinted toward the significance of asexual reproduction and effective spore migration. Two other points may help to explain the observed level of high genetic variability among FG isolates. These include the possibility of the existence of multiple founder populations, which resulted in population admixture as well as dispersion due to the assemblage of different alleles in FG populations, as clearly extrapolated from mutation events (both cumulative and shared mutational events) recorded in the current research. This observation has been supported by the high level of population admixture noticed in network analysis in the current study. The results of the current study also pointed toward the mutation as an essential factor for the generation of diverse regional populations of FG that later resulted in the developments of a number of mutants from which new and virulent type isolates can arise. Moreover, sharing a number of haplotypes among FG populations supports multiple introductions of the identical haplotypes, which may be due to the asexual reproductive phase of FG. Overall, it seems that the analyzed FG populations are admixture ones and occupied haplotypes from different populations in the northern plains of India.
The population genetics analysis performed in the current study revealed that the FG population from RJ seems genetically similar with other populations of PB, HR and WB. This statement is clearly supported by the values of high gene flow (Nm = 11.89–88.51) and low genetic differentiation (Fst = 0.002−0.025). A similar observation of high gene flow among different subpopulations of
F. graminearum in Canada and the USA has been documented [
25,
80]. Furthermore, the ANOVA analysis conducted in present study supports the previous statement, as a very high level of genetic variation (100%) among individuals within the population has been recorded. These observations further indicate toward the greater possibility of sexual reproduction in the regions and are well supported with earlier research findings related to the population genetic structure of FG populations from wheat in Canada and the USA [
25]. There might be another plausible reason, i.e., infected wheat seed movement as a planting material among different regions as a prime cause of gene flow between isolates of distinct origins. However, the role of long-distance spore transfer of FG in determining the gene migration cannot be omitted because of the air-borne mode of dissemination of the fungus [
81].
A high variation in the aggressiveness of FG isolates has been observed in FG isolates collections from different wheat-growing states. These results are in agreement with earlier workers’ findings in both within-field populations and crossing populations [
82,
83,
84]. The study shows that highly aggressive isolates are widely distributed in the northern plains of India and reflects the robust genetic diversity among regional FG populations. However, an exhaustive understanding of localized FG populations and their potential to surmount disease management tactics is essential to secure the wheat farmers from this emerging disease threat in India.